Somatic Mutations Alter Interleukin Signaling Pathways in Grade II Invasive Breast Cancer Patients: An Egyptian Experience
Abstract
1. Introduction
2. Materials and Methods
2.1. Patient Samples
2.2. DNA Extraction, Library Preparation, and Data Processing
2.3. Identification and Classification of the Somatic Variants in Our Study
2.4. Pathway Analysis by Ingenuity Variant Analysis (IVA)
3. Results
3.1. Clinicopathological Characteristics of the Studied Sample Set
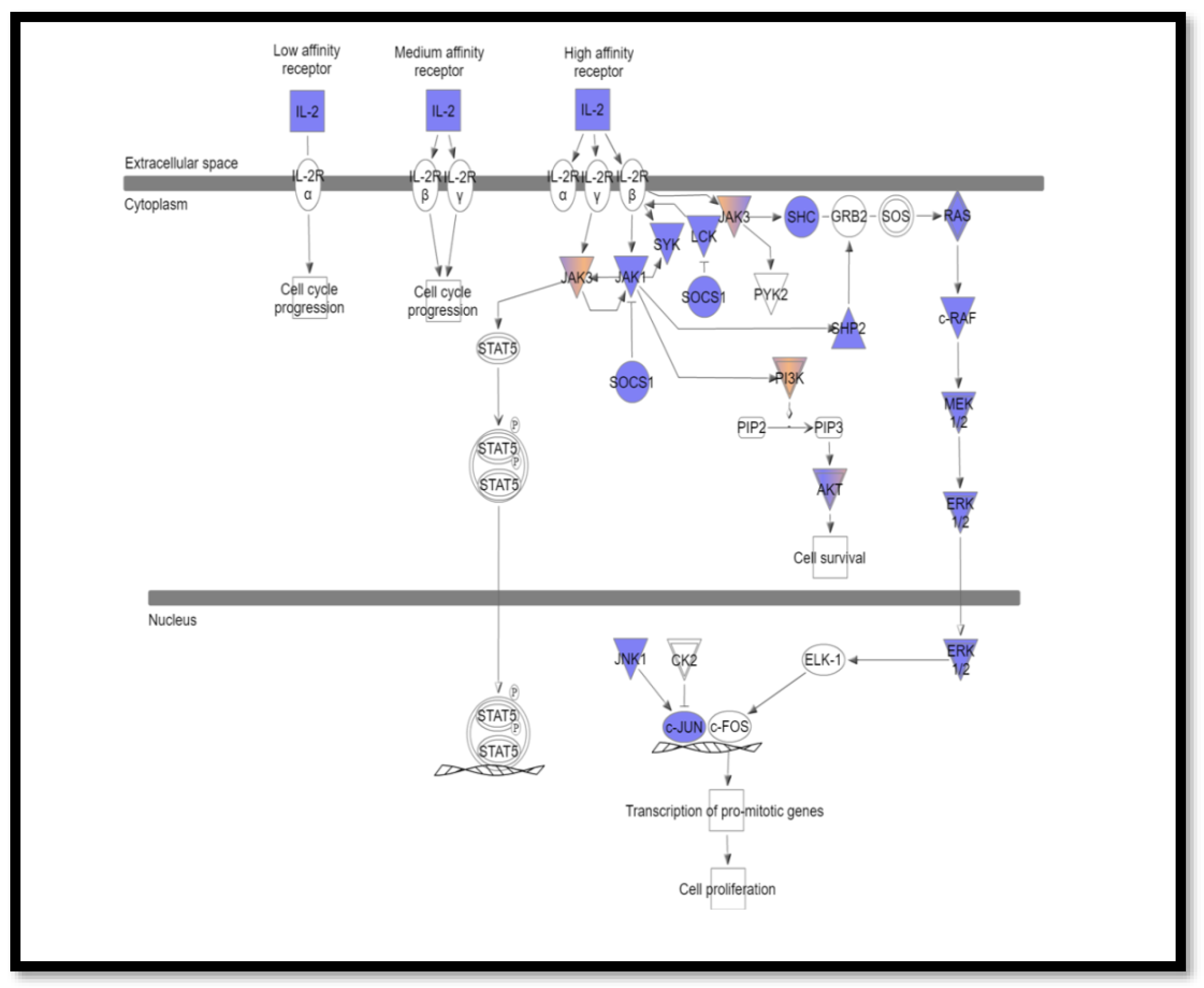
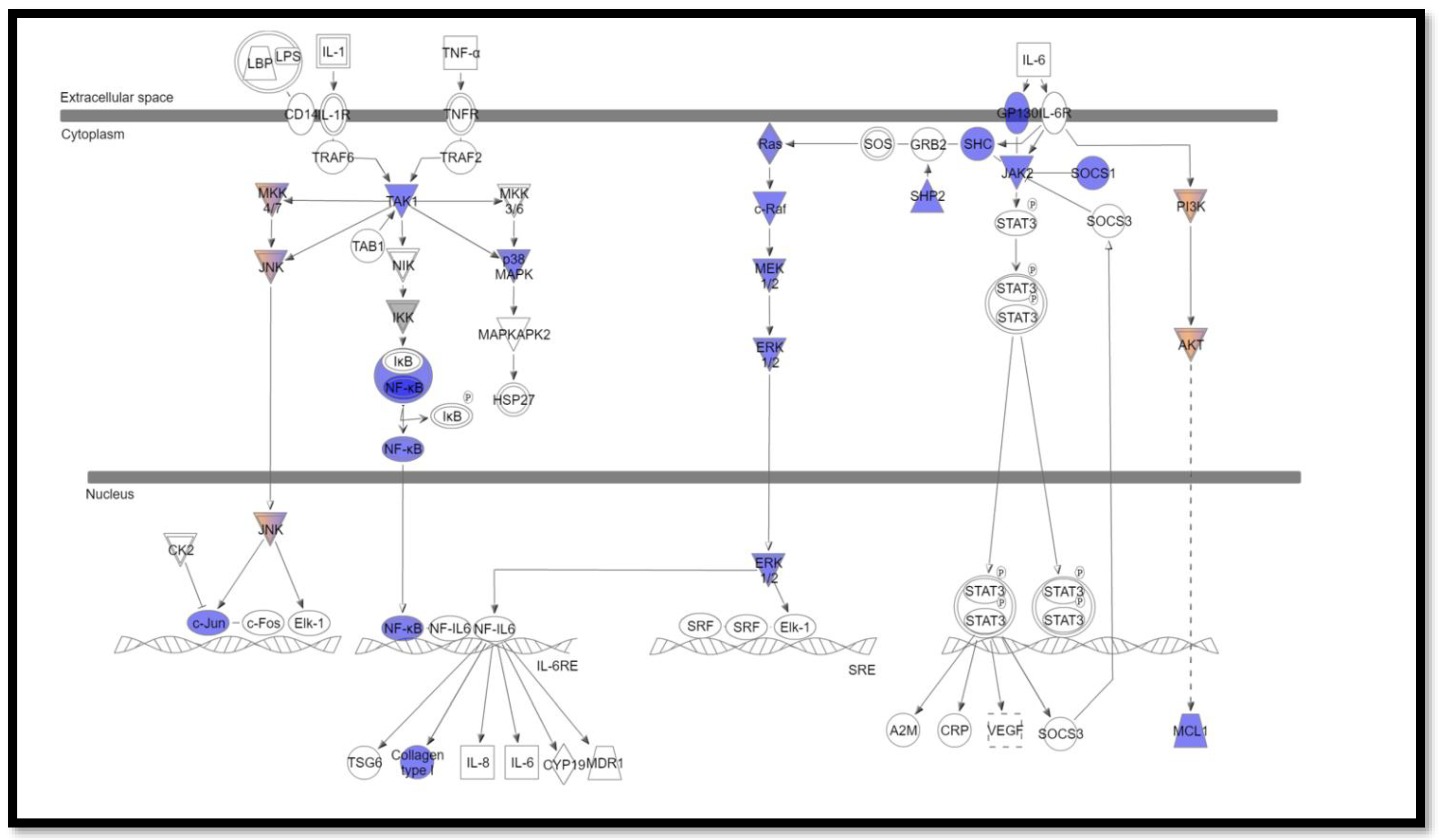
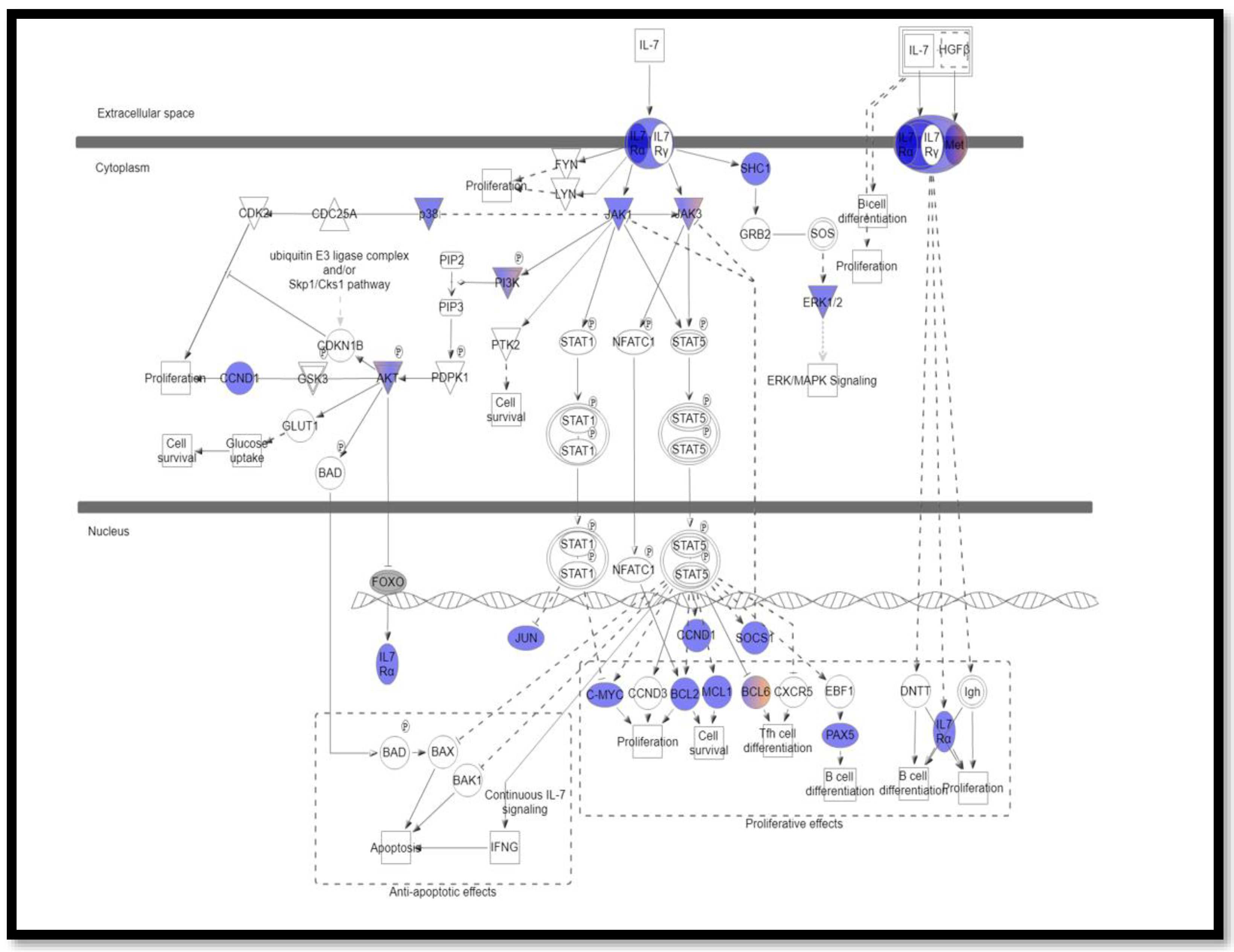
3.2. Somatic Mutations Involved in Interleukin Signaling Pathways
3.3. The Identified Likely Novel Mutations of Somatic Allele Origin
3.4. The Most Commonly Altered Pathways
4. Discussion
5. Conclusions
6. Recommendation
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- WHO. Breast Cancer Fact Sheet. 2021. Available online: https://www.who.int/news-room/fact-sheets/detail/breast-cancer (accessed on 7 July 2022).
- Ibrahim, A.S.; Khaled, H.M.; Mikhail, N.N.; Baraka, H.; Kamel, H. Cancer Incidence in Egypt: Results of the National Population-Based Cancer Registry Program. J. Cancer Epidemiol. 2014, 2014, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of Cancer: The Next Generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Balkwill, F.; Mantovani, A. Inflammation and cancer: Back to Virchow? Lancet 2001, 357, 539–545. [Google Scholar] [CrossRef] [PubMed]
- Bruni, D.; Angell, H.K.; Galon, J. The immune contexture and Immunoscore in cancer prognosis and therapeutic efficacy. Nat. Rev. Cancer 2020, 20, 662–680. [Google Scholar] [CrossRef] [PubMed]
- Manore, S.G.; Doheny, D.L.; Wong, G.L.; Lo, H.W. IL-6/JAK/STAT3 Signaling in Breast Cancer Metastasis: Biology and Treatment. Front. Oncol. 2022, 12, 866014. [Google Scholar] [CrossRef]
- Al-Rawi, M.A.; Rmali, K.; Watkins, G.; Mansel, R.E.; Jiang, W.G. Aberrant expression of interleukin-7 (IL-7) and its signalling complex in human breast cancer. Eur. J. Cancer 2004, 40, 494–502. [Google Scholar] [CrossRef]
- Vilsmaier, T.; Rack, B.; König, A.; Friese, K.; Janni, W.; Jeschke, U.; Acher, T.W. Influence of Circulating Tumour Cells on Production of IL-1α, IL-1β and IL-12 in Sera of Patients with Primary Diagnosis of Breast Cancer before Treatment. Anticancer Res. 2016, 36, 5227–5236. [Google Scholar] [CrossRef][Green Version]
- Kinder, M.; Chislock, E.; Bussard, K.M.; Shuman, L.; Mastro, A.M. Metastatic breast cancer induces an osteoblast inflammatory response. Exp. Cell Res. 2008, 314, 173–183. [Google Scholar] [CrossRef]
- Kamalakar, A.; Bendre, M.S.; Washam, C.L.; Fowler, T.W.; Carver, A.; Dilley, J.D.; Bracey, J.W.; Akel, N.S.; Margulies, A.G.; Skinner, R.A.; et al. Circulating interleukin-8 levels explain breast cancer osteolysis in mice and humans. Bone 2014, 61, 176–185. [Google Scholar] [CrossRef]
- Todorović-Raković, N.; Milovanović, J. Interleukin-8 in breast cancer progression. J. Interferon Cytokine Res. 2013, 33, 563–570. [Google Scholar] [CrossRef] [PubMed]
- Johnstone, C.N.; Chand, A.; Putoczki, T.L.; Ernst, M. Emerging roles for IL-11 signaling in cancer development and progression: Focus on breast cancer. Cytokine Growth Factor Rev. 2015, 26, 489–498. [Google Scholar] [CrossRef]
- Lang, G.-T.; Jiang, Y.-Z.; Shi, J.-X.; Yang, F.; Li, X.-G.; Pei, Y.-C.; Shao, Z.M. Characterization of the genomic landscape and actionable mutations in Chinese breast cancers by clinical sequencing. Nat. Commun. 2020, 11, 5679. [Google Scholar] [CrossRef] [PubMed]
- Fachal, L.; Aschard, H.; Beesley, J.; Barnes, D.R.; Allen, J.; Kar, S.; Jacubowksa, A. Fine-mapping of 150 breast cancer risk regions identifies 191 likely target genes. Nat. Genet. 2020, 52, 56–73. [Google Scholar] [CrossRef] [PubMed]
- Nassar, A.; Abouelhoda, M.; Mansour, O.; Loutfy, S.A.; Hafez, M.M.; Gomaa, M.; Bahnassy, A.; Youssef, A.S.E.-D.; Lotfy, M.M.; Ismail, H.; et al. Targeted next generation sequencing identifies somatic mutations in a cohort of Egyptian breast cancer patients. JAR 2020, 24, 149–157. [Google Scholar] [CrossRef] [PubMed]
- Nassar, A.; Lymona, A.; Lotfy, M.; Youssef, A.; Mohanad, M.; Manie, T.; Youssef, M.M.G.; Farahat, I.G.; Zekri, A.-R.N. Tumor Mutation Burden Prediction Model in Egyptian Breast Cancer patients based on Next Generation Sequencing. Asian Pac. J. Cancer Prev. 2021, 22, 2053–2059. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Henikoff, S.; Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 2009, 4, 1073–1081. [Google Scholar] [CrossRef] [PubMed]
- Landrum, M.J.; Lee, J.M.; Benson, M.; Brown, G.R.; Chao, C.; Chitipiralla, S.; Gu, B.; Hart, J.; Hoffman, D.; Jang, W.; et al. ClinVar: Improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 2018, 46, D1062–D1067. [Google Scholar] [CrossRef] [PubMed]
- Rentzsch, P.; Witten, D.; Cooper, G.M.; Shendure, J.; Kircher, M. CADD: Predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 2019, 47, D886–D894. [Google Scholar] [CrossRef]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef]
- Feng, Y.; Spezia, M.; Huang, S.; Yuan, C.; Zeng, Z.; Zhang, L.; Ji, X.; Liu, W.; Huang, B.; Luo, W.; et al. Breast cancer development and progression: Risk factors, cancer stem cells, signaling pathways, genomics, and molecular pathogenesis. Genes Dis. 2018, 5, 77–106. [Google Scholar] [CrossRef]
- Xu, X.; Dai, W.; Li, C. Interleukins in the treatment of melanoma. Chin. Med. J. 2022, 135, 393–399. [Google Scholar] [CrossRef] [PubMed]
- Briukhovetska, D.; Dörr, J.; Endres, S.; Libby, P.; Dinarello, C.A.; Kobold, S. Interleukins in cancer: From biology to therapy. Nat. Rev. Cancer 2021, 21, 481–499. [Google Scholar] [CrossRef] [PubMed]
- Song, D.; Cui, M.; Zhao, G.; Fan, Z.; Nolan, K.; Yang, Y.; Lee, P.; Ye, F.; Zhang, D.Y. Pathway-based analysis of breast cancer. Am. J. Transl. Res. 2014, 6, 302–311. [Google Scholar] [PubMed]
- Valle-Mendiola, A.; Gutiérrez-Hoya, A.; Lagunas-Cruz, M.d.C.; Weiss-Steider, B.; Soto-Cruz, I. Pleiotropic Effects of IL-2 on Cancer: Its Role in Cervical Cancer. Mediat. Inflamm. 2016, 2016, 1–11. [Google Scholar] [CrossRef]
- Johnston, J.A.; Kawamura, M.; Kirken, R.A.; Chen, Y.-Q.; Blake, T.B.; Shibuya, K.; Ortaldo, J.R.; McVicar, D.W.; O’Shea, J.J. Phosphorylation and activation of the Jak-3 Janus kinase in response to interleukin-2. Nature 1994, 370, 151–153. [Google Scholar] [CrossRef]
- Suzuki, K.; Nakajima, H.; Saito, Y.; Saito, T.; Leonard, W.J.; Iwamoto, I. Janus kinase 3 (Jak3) is essential for common cytokine receptor γ chain (γc)-dependent signaling: Comparative analysis of γc, Jak3, and γc and Jak3 double-deficient mice. Int. Immunol. 2000, 12, 123–132. [Google Scholar] [CrossRef]
- Miyazaki, T.; Kawahara, A.; Fujii, H.; Nakagawa, Y.; Minami, Y.; Liu, Z.-J.; Oishi, I.; Silvennoinen, O.; Witthuhn, B.A.; Ihle, J.N.; et al. Functional Activation of Jak1 and Jak3 by Selective Association with IL-2 Receptor Subunits. Science 1994, 266, 1045–1047. [Google Scholar] [CrossRef]
- O’Shea, J.J.; Kontzias, A.; Yamaoka, K.; Tanaka, Y.; Laurence, A. Janus kinase inhibitors in autoimmune diseases. Ann. Rheum. Dis. 2013, 72, ii111–ii115. [Google Scholar] [CrossRef]
- O’Shea, J.J.; Schwartz, D.M.; Villarino, A.V.; Gadina, M.; McInnes, I.B.; Laurence, A. The JAK-STAT Pathway: Impact on Human Disease and Therapeutic Intervention. Annu. Rev. Med. 2015, 66, 311–328. [Google Scholar] [CrossRef]
- Reif, K.; Buday, L.; Downward, J.; Cantrell, D.A. SH3 domains of the adapter molecule Grb2 complex with two proteins in T cells: The guanine nucleotide exchange protein Sos and a 75-kDa protein that is a substrate for T cell antigen receptor-activated tyrosine kinases. Genes Dis. 1994, 269, 14081–14087. [Google Scholar] [CrossRef]
- Reif, K.; Burgering, B.M.T.; Cantrell, D.A. Phosphatidylinositol 3-Kinase Links the Interleukin-2 Receptor to Protein Kinase B and p70 S6 Kinase. J. Biol. Chem. 1997, 272, 14426–14433. [Google Scholar] [CrossRef] [PubMed]
- Ward, S.G.; Cantrell, D.A. Phosphoinositide 3-kinases in T lymphocyte activation. Curr. Opin. Immunol. 2001, 13, 332–338. [Google Scholar] [CrossRef]
- Gu, H.; Maeda, H.; Moon, J.J.; Lord, J.D.; Yoakim, M.; Nelson, B.H.; Neel, B.G. New Role for Shc in Activation of the Phosphatidylinositol 3-Kinase/Akt Pathway. Curr. Opin. Immunol. 2000, 20, 7109–7120. [Google Scholar] [CrossRef] [PubMed]
- Migone, T.-S.; Rodig, S.; Cacalano, N.A.; Berg, M.; Schreiber, R.D.; Leonard, W.J. Functional Cooperation of the Interleukin-2 Receptor β Chain and Jak1 in Phosphatidylinositol 3-Kinase Recruitment and Phosphorylation. Mol. Cell Biol. 1998, 18, 6416–6422. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bromberg, J.; Wang, T.C. Inflammation and Cancer: IL-6 and STAT3 Complete the Link. Cancer Cell 2009, 15, 79–80. [Google Scholar] [CrossRef]
- Hosseini, F.; Hassannia, H.; Mahdian-Shakib, A.; Jadidi-Niaragh, F.; Enderami, S.E.; Fattahi, M.; Anissian, A.; Mirshafiey, A.; Kokhaei, P. Targeting of crosstalk between tumor and tumor microenvironment by β-D mannuronic acid (M2000) in murine breast cancer model. Cancer Med. 2017, 6, 640–650. [Google Scholar] [CrossRef]
- Lokau, J.; Garbers, C. Activating mutations of the gp130/JAK/STAT pathway in human diseases. Adv. Protein Chem. Struct. Biol. 2019, 16, 283–309. [Google Scholar]
- Heinrich, P.C.; Behrmann, I.; Haan, S.; Hermanns, H.M.; Müller-Newen, G.; Schaper, F. Principles of interleukin (IL)-6-type cytokine signalling and its regulation. Biochem. J. 2003, 374, 1–20. [Google Scholar] [CrossRef]
- Bousoik, E.; Aliabadi, H.M. “Do We Know Jack” About JAK? A Closer Look at JAK/STAT Signaling Pathway. Front. Oncol. 2018, 8, 287. [Google Scholar] [CrossRef]
- Johnston, P.A.; Grandis, J.R. STAT3 SIGNALING: Anticancer Strategies and Challenges. Mol. Interv. 2011, 11, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Schaper, F.; Rose-John, S. Interleukin-6: Biology, signaling and strategies of blockade. Cytokine Growth Factor Rev. 2015, 26, 475–487. [Google Scholar] [CrossRef] [PubMed]
- Masjedi, A.; Hashemi, V.; Hojjat-Farsangi, M.; Ghalamfarsa, G.; Azizi, G.; Yousefi, M.; Jadidi-Niaragh, F. The significant role of interleukin-6 and its signaling pathway in the immunopathogenesis and treatment of breast cancer. Biomed. Pharmacother. 2018, 108, 1415–1424. [Google Scholar] [CrossRef]
- Motallebnezhad, M.; Jadidi-Niaragh, F.; Qamsari, E.S.; Bagheri, S.; Gharibi, T.; Yousefi, M. The immunobiology of myeloid-derived suppressor cells in cancer. Tumor Biol. 2015, 37, 1387–1406. [Google Scholar] [CrossRef]
- Erez, N.; Glanz, S.; Raz, Y.; Avivi, C.; Barshack, I. Cancer Associated Fibroblasts express pro-inflammatory factors in human breast and ovarian tumors. Biochem. Biophys. Res. Commun. 2013, 437, 397–402. [Google Scholar] [CrossRef] [PubMed]
- Degryse, S.; de Bock, C.E.; Demeyer, S.; Govaerts, I.; Bornschein, S.; Verbeke, D.; Jacobs, K.; Binos, S.; Skerrett-Byrne, D.A.; Murray, H.C.; et al. Mutant JAK3 phosphoproteomic profiling predicts synergism between JAK3 inhibitors and MEK/BCL2 inhibitors for the treatment of T-cell acute lymphoblastic leukemia. Leukemia 2018, 32, 788–800. [Google Scholar] [CrossRef]
- Pike, K.A.; Hatzihristidis, T.; Bussières-Marmen, S.; Robert, F.; Desai, N.; Miranda-Saavedra, D.; Pelletier, J.; Tremblay, M.L. TC-PTP regulates the IL-7 transcriptional response during murine early T cell development. Sci. Rep. 2017, 7, 13275. [Google Scholar] [CrossRef]
- Youssef, A.S.E.-D.; Abdel-Fattah, M.A.; Lotfy, M.M.; Nassar, A.; Abouelhoda, M.; Touny, A.O.; Hassan, Z.K.; Eldin, M.M.; Bahnassy, A.A.; Khaled, H.; et al. Multigene Panel Sequencing Reveals Cancer-Specific and Common Somatic Mutations in Colorectal Cancer Patients: An Egyptian Experience. Curr. Issues Mol. Biol. 2022, 44, 1332–1352. [Google Scholar] [CrossRef]
| Patients Characteristics | Total (n = 55) | Percentage (%) |
|---|---|---|
| Age: Median (Range) | 52.6 (32–73) | |
| <50 | 18 | 32.7 |
| ≥50 | 37 | 67.3 |
| Menopausal status | ||
| Premenopausal | 8 | 14.5 |
| Postmenopausal | 47 | 85.5 |
| Lymph Node involvement | ||
| 0 | 12 | 21.8 |
| 1 to 3 | 22 | 40 |
| >3 | 21 | 38.2 |
| Estrogen (ER) | ||
| Negative | 18 | 32.7 |
| Positive | 37 | 67.3 |
| Progesterone (PR) | ||
| Negative | 19 | 34.5 |
| Positive | 36 | 65.4 |
| HER-2 | ||
| Negative | 43 | 78.2 |
| Positive | 12 | 21.8 |
| Ki-67 | ||
| <14% | 19 | 34.5 |
| ≥14% | 36 | 65.4 |
| Gene | Position | Cosmic ID | Type | IVA Classification | Inferred Activity | SIFT/or PolyPhen-2 Prediction | CADD Phred Score | Affected Transcript | HGVS.c | HGVS.p | Cases (n = 55) | Occurrence in Cosmic Database |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AKT3 | Chr1:243675627 | COSM7264038 * | Deletion | PV | Loss | - | 32 | NM_001370074.1 | c.1353delA | p.K451fs*? | 9 (16.3%) | Large intestine = 1 |
| AKT3 | Chr1:243708813 | COSM8568034 * | Deletion | PV | Loss | - | - | NM_001370074.1 | c.1250del | p.K417Sfs*10 | 3 (5.4%) | Stomach = 1 |
| BCL2 | Chr18:60985665 | COSM7343104 * | Deletion | PV | Loss | - | - | NM_000633.2 | c.234_235del | p.G79Rfs*73 | 1 (1.8%) | Large intestine = 1 |
| BCL2 | Chr18:60985776 | COSM9113142 * | Deletion | PV | Loss | - | - | NM_000633.2 | c.124delG | p.A42fs*54 | 7 (12.7%) | Large intestine = 1 |
| IL6ST | Chr5:55247869 | COSM269367 * | Deletion | PV | Loss | - | 32 | NM_175767.3 | c.1587delA | p.V530* | 14 (25.4%) | Large intestine = 12 and Stomach = 1 |
| IL7R | Chr5:35867541 | COSM2687237 * | Deletion | PV | Loss | - | 20.2 | NM_002185.5 | c.361delA | p.I121* | 7 (12.7%) | Large intestine = 7 and Stomach = 4 |
| IL7R | Chr5:35873646 | COSM7342389 * | SNV | PV | Loss | Damaging | 25.3 | NM_002185.5 | c.602A > G | p.Y201C | 3 (5.4%) | Large intestine = 1 and Thyroid = 1 |
| JAK1 | Chr1:65325833 | COSM1560531 | Deletion | PV | Loss | - | 33 | NM_001321853.2 | c.1289delC | p.P386fs*2 | 13 (23.6%) | Large intestine = 8 and Breast = 3 |
| JAK1 | Chr1:65330630 | COSM1343915 * | Deletion | PV | Loss | - | 27.9 | NM_001321853.2 | c.884delA | p.N339fs*3 | 8 (14.5%) | Endometrium = 2 and Large intestine = 1 |
| JAK1 | Chr1:65339129 | COSM6912578 * | Deletion | PV | Loss | - | 32 | NM_001321853.2 | c.407delA | p.N136fs*32 | 5 (9%) | Prostate = 2 |
| JAK1 | Chr1:65339111 | COSM1639943 | Deletion | PV | Loss | - | 25.6 | NM_001321853.2 | c.425delA | p.K142fs*26 | 3 (5.4%) | Breast = 4 and Large intestine = 2 |
| JAK2 | Chr9:5078342 | COSM7286153 * | Deletion | PV | Loss | - | 33 | NM_001322199.1 | c.1586delA | p.N678fs*53 | 10 (18%) | Large intestine = 1 |
| JAK3 | Chr19:17954215 | COSM34216 * | SNV | PV | Gain | Possibly damaging | 19.01 | NM_000215.4 | c.394C>A | p.P132T | 3 (5.4%) | Lung = 1 and Hematopoietic and lymphoid = 1 |
| MCL1 | Chr1:150551952 | COSM5156964 * | Deletion | PV | Loss | - | 22.7 | NM_001197320.1 | c.55delG | p.A19fs*30 | 11 (20%) | Upper aerodigestive tract = 1 |
| MCL1 | Chr1:150551935 | COSM5734360 * | Deletion | PV | Loss | - | 15.72 | NM_001197320.1 | c.61_72delTTGGGGGCCGGC | p.L21_G24del | 1 (1.8%) | Large intestine = 1 and Stomach = 1 |
| MCL1 | Chr1:150551327 | COSM6923093 * | SNV | NV | Loss | Tolerated | 22.8 | NM_001197320.1 | c.680C>T | p.A227V | 1 (1.8%) | Thyroid = 1 and Hematopoietic and lymphoid = 1 |
| MTOR | Chr1:11199444 | COSM6716875 * | SNV | PV | Loss | Damaging | 32 | NM_004958.4 | c.5047C>T | p.R1683W | 1 (1.8%) | Large intestine = 1 |
| NRAS | Chr1:115252309 | COSM7177458 * | SNV | PV | Loss | Possibly Damaging | 23.4 | NM_002524.5 | c.331A>G | p.M111V | 1 (1.8%) | Urinary tract = 1 and Stomach = 1 |
| PIK3CA | Chr3:178936082 | COSM29329 | SNV | PV | Loss | Damaging | 31 | NM_006218.4 | c.1624G>A | p.E542K | 11 (20%) | Breast = 239 and Large intestine = 66 |
| SOCS1 | Chr16:11348846 | COSM19320 * | Deletion | PV | Loss | - | 24.3 | NM_003745.1 | c.490delG | p.A164fs*41 | 10 (18%) | Hematopoietic and lymphoid = 3 and Large intestine = 1 |
| SOCS1 | Chr16:11348998 | COSM6964419 * | Deletion | PV | Loss | - | 32 | NM_003745.1 | c.338delT | p.F113fs*5 | 5 (9%) | Large intestine = 1 |
| Gene | Position | Cosmic ID | Accession No. | Type | IVA Classification | Inferred Activity | Affected Transcript | HGVS.c | HGVS.p | Cases (n = 55) |
|---|---|---|---|---|---|---|---|---|---|---|
| SHC1 | Chr1:154947252 | Novel | SCV002574988 | Deletion | PV | Loss | NM_001130041.2 | c.33delA | p.K11fs*18 | 2 (3.6%) |
| SHC1 | Chr1:154947270 | Novel | SCV002574989 | Deletion | PV | Loss | NM_001130041.2 | c.51delT | p.F17fs*12 | 2 (3.6%) |
| IL2 | Chr4:123372936 | Novel | SCV002574990 | Deletion | PV | Loss | NM_000586.4 | c.433delT | p.C145fs*8 | 10 (18%) |
| IL2 | Chr4:123377462 | Novel | SCV002574991 | Deletion | PV | Loss | NM_000586.4 | c.134delT | p.L45* | 3 (5.3%) |
| IL7R | Chr5:35871245 | Novel | SCV002574992 | Deletion | PV | Loss | NM_002185.5 | c.470delA | p.K157fs*4 | 5 (9%) |
| JUN | Chr1:59248084 | Novel | SCV002574993 | Deletion | PV | Loss | NM_002228.4 | c.659delC | p.P220fs*6 | 7 (12.7%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nassar, A.; Zekri, A.R.N.; Elberry, M.H.; Lymona, A.M.; Lotfy, M.M.; Abouelhoda, M.; Youssef, A.S.E.-D. Somatic Mutations Alter Interleukin Signaling Pathways in Grade II Invasive Breast Cancer Patients: An Egyptian Experience. Curr. Issues Mol. Biol. 2022, 44, 5890-5901. https://doi.org/10.3390/cimb44120401
Nassar A, Zekri ARN, Elberry MH, Lymona AM, Lotfy MM, Abouelhoda M, Youssef ASE-D. Somatic Mutations Alter Interleukin Signaling Pathways in Grade II Invasive Breast Cancer Patients: An Egyptian Experience. Current Issues in Molecular Biology. 2022; 44(12):5890-5901. https://doi.org/10.3390/cimb44120401
Chicago/Turabian StyleNassar, Auhood, Abdel Rahman N. Zekri, Mostafa H. Elberry, Ahmed M. Lymona, Mai M. Lotfy, Mohamed Abouelhoda, and Amira Salah El-Din Youssef. 2022. "Somatic Mutations Alter Interleukin Signaling Pathways in Grade II Invasive Breast Cancer Patients: An Egyptian Experience" Current Issues in Molecular Biology 44, no. 12: 5890-5901. https://doi.org/10.3390/cimb44120401
APA StyleNassar, A., Zekri, A. R. N., Elberry, M. H., Lymona, A. M., Lotfy, M. M., Abouelhoda, M., & Youssef, A. S. E.-D. (2022). Somatic Mutations Alter Interleukin Signaling Pathways in Grade II Invasive Breast Cancer Patients: An Egyptian Experience. Current Issues in Molecular Biology, 44(12), 5890-5901. https://doi.org/10.3390/cimb44120401





