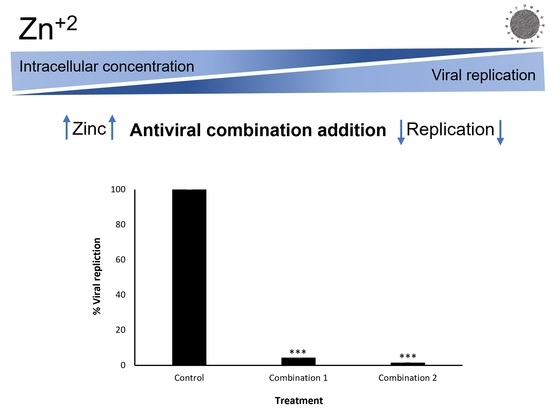
Inhibition of Respiratory RNA Viruses by a Composition of Ionophoric Polyphenols with Metal Ions
Abstract
:1. Introduction
2. Results and Discussion
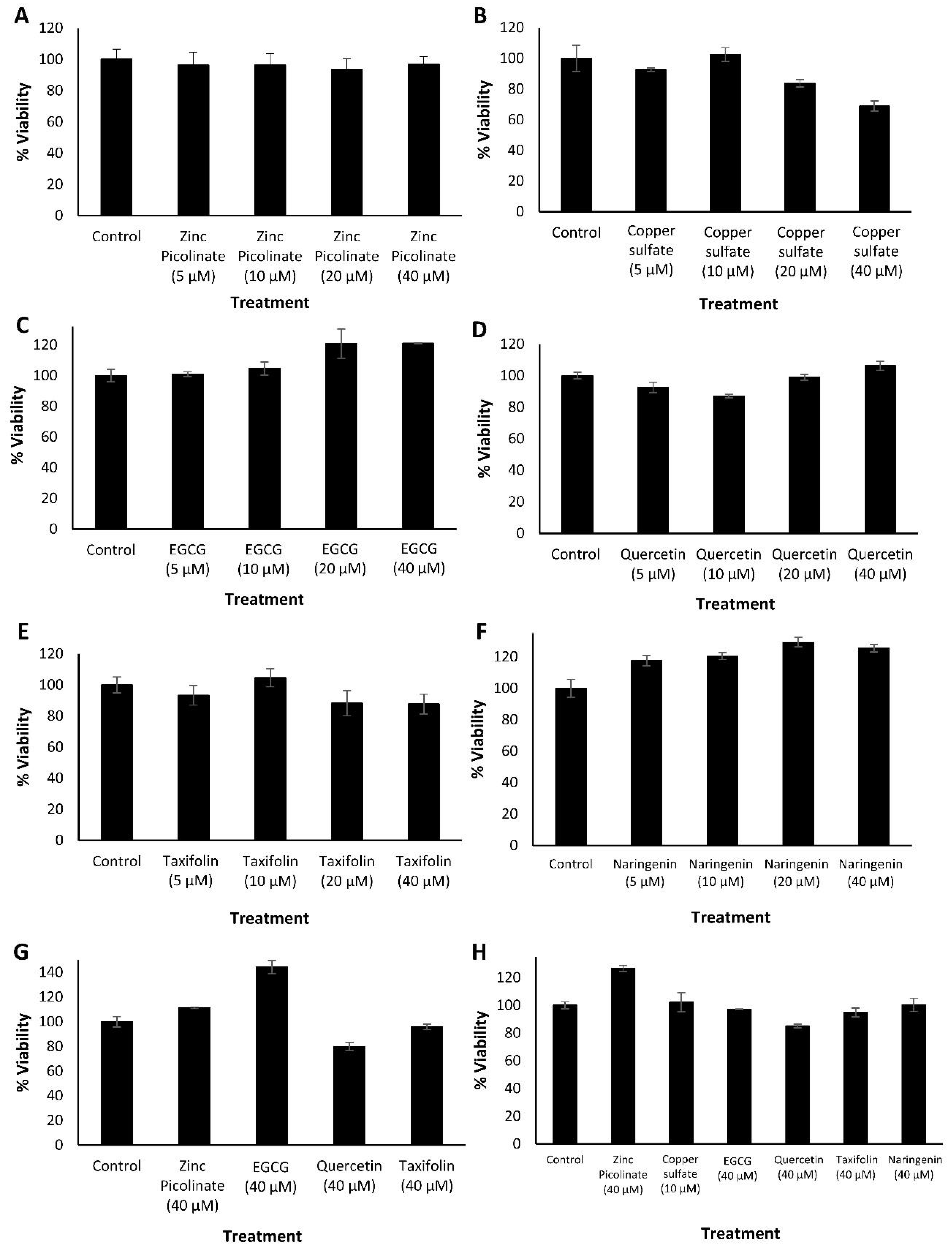
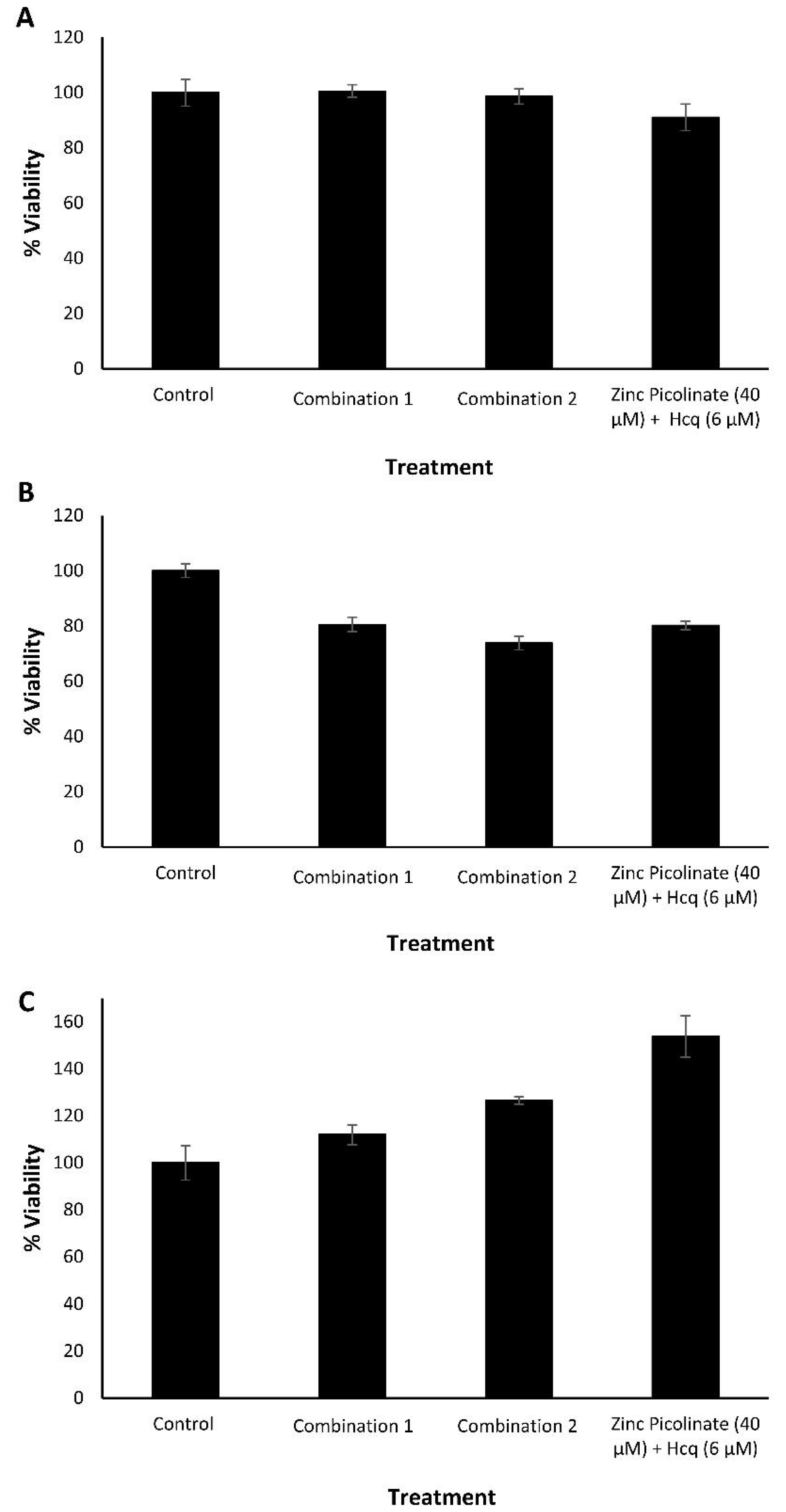
2.1. Biocompatibility of the Ingredients and Their Active Combinations
2.2. Inhibition of Intracellular Viral Replication by Natural Compound Combinations
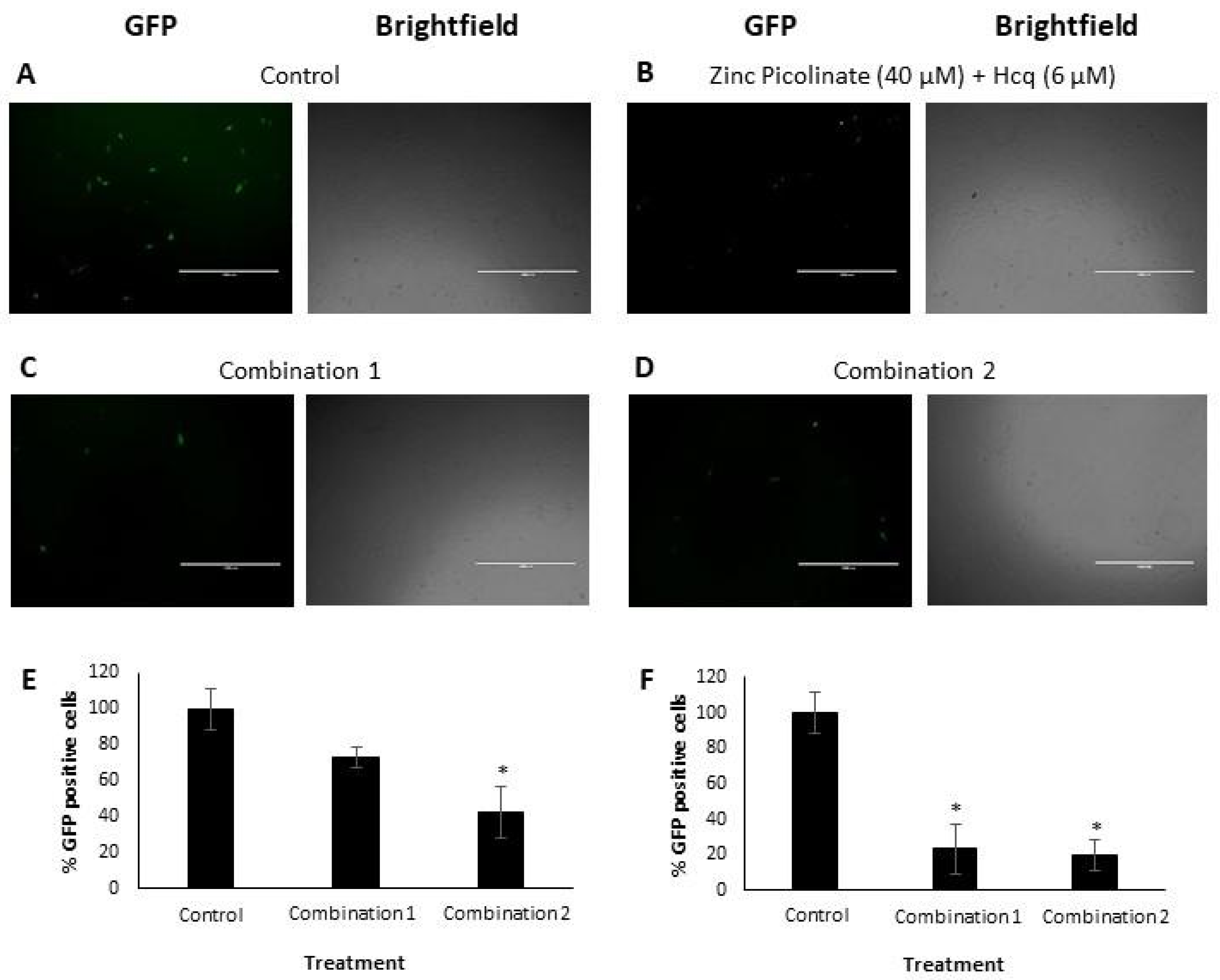
2.2.1. Flow Cytometry Analysis and Quantification
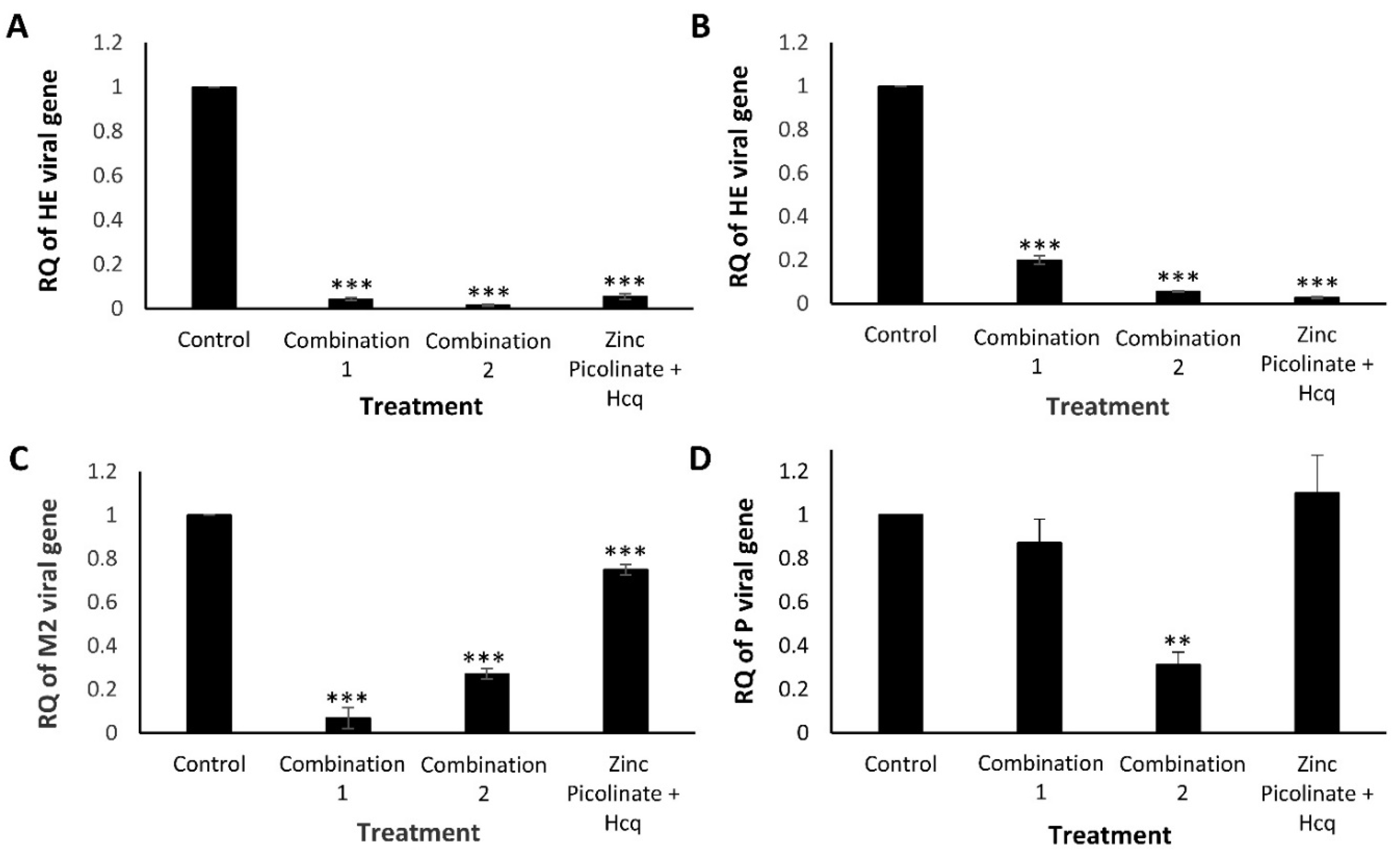
2.2.2. Quantitative Real-Time PCR Analysis
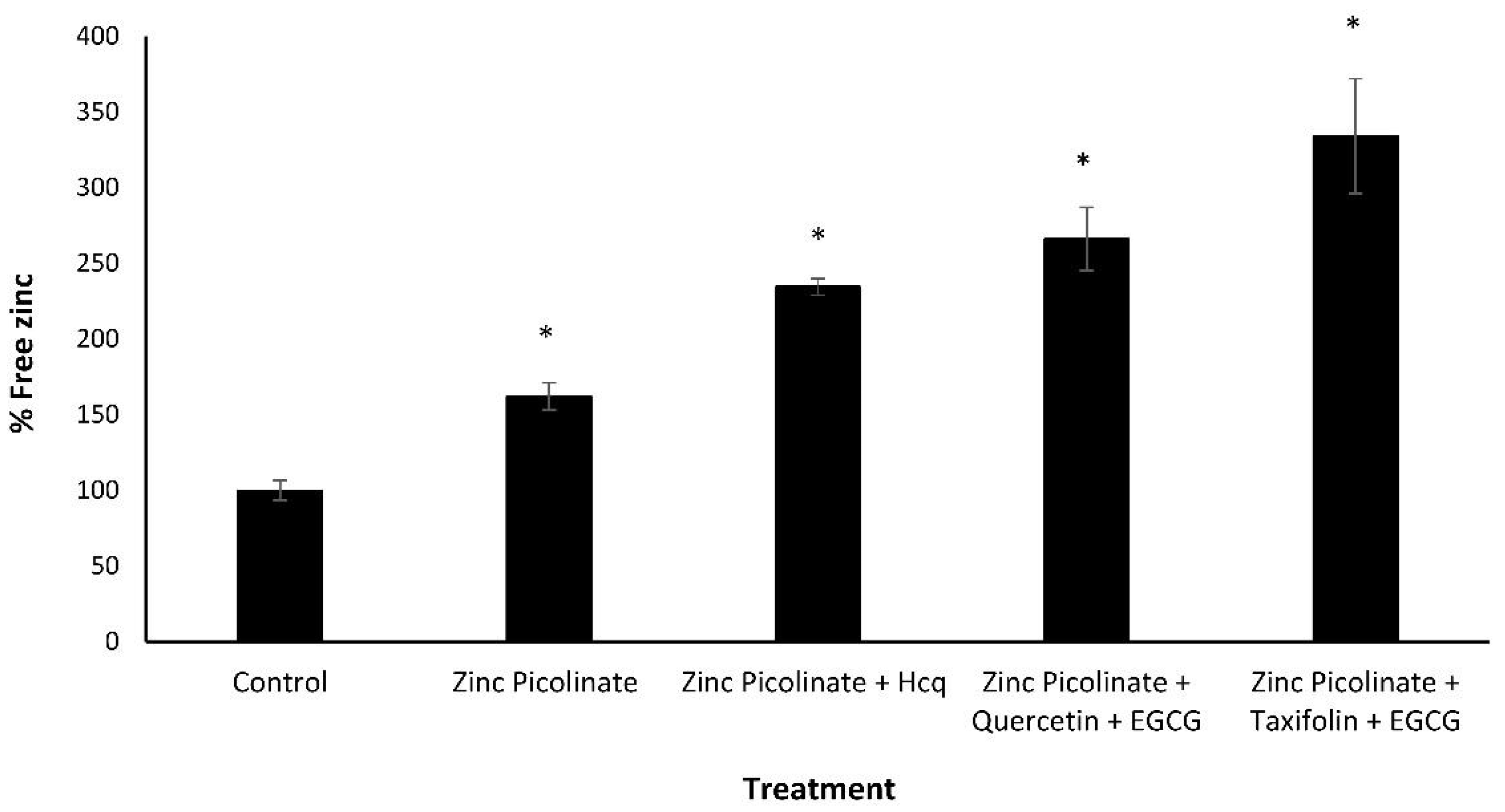
2.3. Increase of Intracellular Zinc Levels by the Natural Compound Combinations
3. Materials and Methods
3.1. Materials
- Reagents and Kits:
- Cell lines:
- Viral strains:
3.2. Experimental Methods
3.2.1. Cytotoxicity Evaluation Experiments
3.2.2. Viral Infection and Quantification by FACS
3.2.3. Viral Infection and Quantification by qRT-PCR
3.2.4. Free Zinc Quantification Assay
3.2.5. qRT-PCR Primers
| Virus/Cell Source | Primer Name | Sequence (5′-3′) |
|---|---|---|
| IAV | M2 Fw | CATGGAATGGCTAAAGACAAGACC |
| M2 Rev | CCATTAAGGGCATTTTGGACA | |
| hMPV | P Fw | GGCATGGGCAGACAACAGCGG |
| P Rev | GAATTCCTCTTCTTCAACAGG | |
| HCoV-OC43 | HE Fw | GGTTGTGACTATATCGTACC |
| HE Rev | GCGAATCAACAACCTGTACAG | |
| A549 and H1299 cells | Human GAPDH Fw | AGCCACATCGCTCAGACAC |
| Human GAPDH Rev | GCCCAATACGACCAAATCC | |
| Vero cells | Monkey GAPDH Fw | GAAGGCTGGGGCTCATTTGC |
| Monkey GAPDH Rev | ATGACGAACATGGGGGCGTC |
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Leung, N.H.L. Transmissibility and transmission of respiratory viruses. Nat. Rev. Microbiol. 2021, 19, 528–545. [Google Scholar] [CrossRef]
- Afewerky, H.K. Pathology, and pathogenicity of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Exp. Biol. Med. 2020, 245, 1299–1307. [Google Scholar] [CrossRef]
- Sanyaolu, A.; Okorie, C.; Marinkovic, A.; Patidar, R.; Younis, K.; Desai, P.; Hosein, Z.; Padda, I.; Mangat, J.; Altaf, M. Comorbidity and its Impact on Patients with COVID-19. SN Compr. Clin. Med. 2020, 2, 1069–1076. [Google Scholar] [CrossRef]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baj, J.; Karakuła-Juchnowicz, H.; Teresiński, G.; Buszewicz, G.; Ciesielka, M.; Sitarz, E.; Forma, A.; Karakuła, K.; Flieger, W.; Portincasa, P.; et al. COVID-19: Specific and non-specific clinical manifestations and symptoms: The current state of knowledge. J. Clin. Med. 2020, 9, 1753. [Google Scholar] [CrossRef]
- Liu, Y.C.; Kuo, R.L.; Shih, S.R. COVID-19: The first documented coronavirus pandemic in history. Biomed. J. 2020, 43, 328–333. [Google Scholar] [CrossRef]
- Lim, W.S. Pandemic flu: Clinical management of patients with an influenza-like illness during an influenza pandemic. Thorax 2007, 62, 1–46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wong, S.S.; Oshansky, C.M.; Guo, X.Z.J.; Ralston, J.; Wood, T.; Seeds, R.; Newbern, C.; Waite, B.; Reynolds, G.; Widdowson, M.A.; et al. Dela Severe Influenza Is Characterized by Prolonged Immune Activation: Results from the SHIVERS Cohort Study. J. Infect. Dis. 2018, 217, 245–256. [Google Scholar] [CrossRef]
- Huck, B.; Neumann-Haefelin, D.; Schmitt-Graeff, A.; Weckmann, M.; Mattes, J.; Ehl, S.; Falcone, V. Human metapneumovirus induces more severe disease and stronger innate immune response in BALB/c mice as compared with the respiratory syncytial virus. Respir. Res. 2007, 8, 6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, Y.; Liu, J.; Zhang, D.; Xu, Z.; Ji, J.; Wen, C. Cytokine Storm in COVID-19: The Current Evidence and Treatment Strategies. Front. Immunol. 2020, 11, 1708. [Google Scholar] [CrossRef]
- Mallah, S.I.; Ghorab, O.K.; Al-Salmi, S.; Abdellatif, O.S.; Tharmaratnam, T.; Iskandar, M.A.; Sefen, J.A.N.; Sidhu, P.; Atallah, B.; El-Lababidi, R.; et al. COVID-19: Breaking down a global health crisis. Ann. Clin. Microbiol. Antimicrob. 2021, 20, 1–36. [Google Scholar] [CrossRef] [PubMed]
- Salvatori, G.; Luberto, L.; Maffei, M.; Aurisicchio, L.; Aurisicchio, L.; Roscilli, G.; Roscilli, G.; Palombo, F.; Marra, E.; Marra, E. SARS-CoV-2 spike protein: An optimal immunological target for vaccines. J. Transl. Med. 2020, 18, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Jackson, C.B.; Farzan, M.; Chen, B.; Choe, H. Mechanisms of SARS-CoV-2 entry into cells. Nat. Rev. Mol. Cell Biol. 2021, 23, 3–20. [Google Scholar] [CrossRef] [PubMed]
- Wahl, A.; Gralinski, L.E.; Johnson, C.E.; Yao, W.; Kovarova, M.; Dinnon, K.H.; Liu, H.; Madden, V.J.; Krzystek, H.M.; De, C.; et al. SARS-CoV-2 infection is effectively treated and prevented by EIDD-2801. Nature 2021, 591, 451–457. [Google Scholar] [CrossRef] [PubMed]
- Ghoran, S.H.; El-Shazly, M.; Sekeroglu, N.; Kijjoa, A. Natural products from medicinal plants with anti-human coronavirus activities. Molecules 2021, 26, 1754. [Google Scholar] [CrossRef]
- Du, L.; Yang, Y.; Zhang, X. Neutralizing antibodies for the prevention and treatment of COVID-19. Cell. Mol. Immunol. 2021, 18, 2293–2306. [Google Scholar] [CrossRef]
- Cao, Y.; Wang, J.; Jian, F.; Xiao, T.; Song, W.; Yisimayi, A.; Huang, W.; Li, Q.; Wang, P.; An, R.; et al. Omicron escapes the majority of existing SARS-CoV-2 neutralizing antibodies. Nature 2021, 602, 657–663. [Google Scholar] [CrossRef]
- Hwang, Y.C.; Lu, R.M.; Su, S.C.; Chiang, P.Y.; Ko, S.H.; Ke, F.Y.; Liang, K.H.; Hsieh, T.Y.; Wu, H.C. Monoclonal antibodies for COVID-19 therapy and SARS-CoV-2 detection. J. Biomed. Sci. 2022, 29, 1–50. [Google Scholar] [CrossRef]
- Planas, D.; Saunders, N.; Maes, P.; Guivel-Benhassine, F.; Planchais, C.; Buchrieser, J.; Bolland, W.H.; Porrot, F.; Staropoli, I.; Lemoine, F.; et al. Considerable escape of SARS-CoV-2 Omicron to antibody neutralization. Nature 2021, 602, 671–675. [Google Scholar] [CrossRef]
- Wibmer, C.K.; Ayres, F.; Hermanus, T.; Madzivhandila, M.; Kgagudi, P.; Oosthuysen, B.; Lambson, B.E.; de Oliveira, T.; Vermeulen, M.; van der Berg, K.; et al. SARS-CoV-2 501Y.V2 escapes neutralization by South African COVID-19 donor plasma. Nat. Med. 2021, 27, 622–625. [Google Scholar] [CrossRef]
- Kistler, K.E.; Bedford, T. Evidence for adaptive evolution in the receptor-binding domain of seasonal coronaviruses OC43 and 229E. eLife 2021, 10, e64509. [Google Scholar] [CrossRef] [PubMed]
- Thorne, L.G.; Bouhaddou, M.; Reuschl, A.-K.; Zuliani-Alvarez, L.; Polacco, B.; Pelin, A.; Batra, J.; Whelan, M.V.X.; Ummadi, M.; Rojc, A.; et al. Evolution of enhanced innate immune evasion by the SARS-CoV-2 B.1.1.7 UK variant. Nature 2021, 602, 487–495. [Google Scholar] [CrossRef] [PubMed]
- Jones, N. Why easing COVID restrictions could prompt a fierce flu rebound. Nature 2021, 598, 395. [Google Scholar] [CrossRef] [PubMed]
- Lu, D. Children’s immunity at risk. New Sci. 2021, 250, 8–9. [Google Scholar] [CrossRef]
- Burrel, S.; Hausfater, P.; Dres, M.; Pourcher, V.; Luyt, C.E.; Teyssou, E.; Soulié, C.; Calvez, V.; Marcelin, A.G.; Boutolleau, D. Co-infection of SARS-CoV-2 with other respiratory viruses and performance of lower respiratory tract samples for the diagnosis of COVID-19. Int. J. Infect. Dis. 2021, 102, 10–13. [Google Scholar] [CrossRef]
- Jang, M.; Park, Y.I.; Cha, Y.E.; Park, R.; Namkoong, S.; Lee, J.I.; Park, J. Tea Polyphenols EGCG and Theaflavin Inhibit the Activity of SARS-CoV-2 3CL-Protease in Vitro. Evid.-Based Complement. Altern. Med. 2020, 2020, 5630838. [Google Scholar] [CrossRef]
- Hu, Z.; Lin, J.; Chen, J.; Cai, T.; Xia, L.; Liu, Y.; Song, X.; He, Z. Overview of Viral Pneumonia Associated With Influenza Virus, Respiratory Syncytial Virus, and Coronavirus, and Therapeutics Based on Natural Products of Medicinal Plants. Front. Pharmacol. 2021, 12, 630834. [Google Scholar] [CrossRef]
- Maison, N.; Peck, A.; Illi, S.; Meyer-Buehn, M.; Von Mutius, E.; Hübner, J.; Von Both, U. The rising of old foes: Impact of lockdown periods on “non-SARS-CoV-2” viral respiratory and gastrointestinal infections. Infection 2022, 1, 3. [Google Scholar] [CrossRef]
- Chiu, N.C.; Chi, H.; Tai, Y.L.; Peng, C.C.; Tseng, C.Y.; Chen, C.C.; Tan, B.F.; Lin, C.Y. Impact of Wearing Masks, Hand Hygiene, and Social Distancing on Influenza, Enterovirus, and All-Cause Pneumonia During the Coronavirus Pandemic: Retrospective National Epidemiological Surveillance Study. J. Med. Internet Res. 2020, 22, e21257. [Google Scholar] [CrossRef]
- te Velthuis, A.J.W.; van den Worml, S.H.E.; Sims, A.C.; Baric, R.S.; Snijder, E.J.; van Hemert, M.J. Zn2+ inhibits coronavirus and arterivirus RNA polymerase activity in vitro and zinc ionophores block the replication of these viruses in cell culture. PLoS Pathog. 2010, 6, e1001176. [Google Scholar] [CrossRef]
- Wessels, I.; Rolles, B.; Rink, L. The Potential Impact of Zinc Supplementation on COVID-19 Pathogenesis. Front. Immunol. 2020, 11, 1712. [Google Scholar] [CrossRef] [PubMed]
- Maywald, M.; Wessels, I.; Rink, L. Zinc signals and immunity. Int. J. Mol. Sci. 2017, 18, 2222. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gammoh, N.Z.; Rink, L. Zinc in infection and inflammation. Nutrients 2017, 9, 624. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Read, S.A.; Obeid, S.; Ahlenstiel, C.; Ahlenstiel, G. The Role of Zinc in Antiviral Immunity. Adv. Nutr. 2019, 10, 696–710. [Google Scholar] [CrossRef] [Green Version]
- Carlucci, P.M.; Ahuja, T.; Petrilli, C.; Rajagopalan, H.; Jones, S.; Rahimian, J. Zinc sulfate in combination with a zinc ionophore may improve outcomes in hospitalized COVID-19 patients. J. Med. Microbiol. 2020, 69, 1228–1234. [Google Scholar] [CrossRef]
- Warnes, S.L.; Little, Z.R.; Keevil, C.W. Human coronavirus 229E remains infectious on common touch surface materials. MBio 2015, 6, e01697-15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodriguez, K.; Saunier, F.; Rigaill, J.; Audoux, E.; Botelho-Nevers, E.; Prier, A.; Dickerscheit, Y.; Pillet, S.; Pozzetto, B.; Bourlet, T.; et al. Evaluation of in vitro activity of copper gluconate against SARS-CoV-2 using confocal microscopy-based high content screening. J. Trace Elem. Med. Biol. 2021, 68, 126818. [Google Scholar] [CrossRef]
- da Silva, A.P.G. Fighting coronaviruses with natural polyphenols. Biocatal. Agric. Biotechnol. 2021, 37, 102179. [Google Scholar] [CrossRef]
- Paraiso, I.L.; Revel, J.S.; Stevens, J.F. Potential use of polyphenols in the battle against COVID-19. Curr. Opin. Food Sci. 2020, 32, 149–155. [Google Scholar] [CrossRef]
- Frabasile, S.; Koishi, A.C.; Kuczera, D.; Silveira, G.F.; Verri, W.A.; Dos Santos, C.N.D.; Bordignon, J. The citrus flavanone naringenin impairs dengue virus replication in human cells. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Panche, A.N.; Diwan, A.D.; Chandra, S.R. Flavonoids: An overview. J. Nutr. Sci. 2016, 5. [Google Scholar] [CrossRef] [Green Version]
- Al-Karmalawy, A.A.; Farid, M.M.; Mostafa, A.; Ragheb, A.Y.; Mahmoud, S.H.; Shehata, M.; Abo Shama, N.M.; Gaballah, M.; Mostafa-Hedeab, G.; Marzouk, M.M. Naturally available flavonoid aglycones as potential antiviral drug candidates against SARS-CoV-2. Molecules 2021, 26, 6559. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.Y.; Pai, T.K.; Han, O. Effect of bioactive dietary polyphenols on zinc transport across the intestinal caco-2 cell monolayers. J. Agric. Food Chem. 2011, 59, 3606–3612. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clergeaud, G.; Dabbagh-Bazarbachi, H.; Ortiz, M.; Fernández-Larrea, J.B.; O’Sullivan, C.K. A simple liposome assay for the screening of zinc ionophore activity of polyphenols. Food Chem. 2016, 197, 916–923. [Google Scholar] [CrossRef]
- Bernatova, I.; Liskova, S. Mechanisms modified by (−)-epicatechin and taxifolin relevant for the treatment of hypertension and viral infection: Knowledge from preclinical studies. Antioxidants 2021, 10, 467. [Google Scholar] [CrossRef] [PubMed]
- Clementi, N.; Scagnolari, C.; D’Amore, A.; Palombi, F.; Criscuolo, E.; Frasca, F.; Pierangeli, A.; Mancini, N.; Antonelli, G.; Clementi, M.; et al. Naringenin is a powerful inhibitor of SARS-CoV-2 infection in vitro. Pharmacol. Res. 2021, 163, 105255. [Google Scholar] [CrossRef] [PubMed]
- Riva, A.; Ronchi, M.; Petrangolini, G.; Bosisio, S.; Allegrini, P. Improved Oral Absorption of Quercetin from Quercetin Phytosome®, a New Delivery System Based on Food Grade Lecithin. Eur. J. Drug Metab. Pharmacokinet. 2019, 44, 169–177. [Google Scholar] [CrossRef] [Green Version]
- Talmi-Frank, D.; Altboum, Z.; Solomonov, I.; Udi, Y.; Jaitin, D.A.; Klepfish, M.; David, E.; Zhuravlev, A.; Keren-Shaul, H.; Winter, D.R.; et al. Extracellular Matrix Proteolysis by MT1-MMP Contributes to Influenza-Related Tissue Damage and Mortality. Cell Host Microbe 2016, 20, 458–470. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Williams, J.V. Human metapneumovirus: An important cause of respiratory disease in children and adults. Curr. Infect. Dis. Rep. 2005, 7, 204–210. [Google Scholar] [CrossRef] [PubMed]
- Keshavarz Valian, N.; Pourakbari, B.; Asna Ashari, K.; Hosseinpour Sadeghi, R.; Mahmoudi, S. Evaluation of human coronavirus OC43 and SARS-COV-2 in children with respiratory tract infection during the COVID-19 pandemic. J. Med. Virol. 2021, 94, 1450–1456. [Google Scholar] [CrossRef]
- Schirtzinger, E.E.; Kim, Y.; Davis, A.S. Improving human coronavirus OC43 (HCoV-OC43) research comparability in studies using HCoV-OC43 as a surrogate for SARS-CoV-2. J. Virol. Methods 2022, 299, 114317. [Google Scholar] [CrossRef] [PubMed]
- Shittu, M.O.; Afolami, O.I. Improving the efficacy of chloroquine and hydroxychloroquine against SARS-CoV-2 may require zinc additives-A better synergy for future COVID-19 clinical trials. Infez. Med. 2020, 28, 192–197. [Google Scholar] [PubMed]
- Mayor-Ibarguren, A.; Busca-Arenzana, C.; Robles-Marhuenda, Á. A Hypothesis for the Possible Role of Zinc in the Immunological Pathways Related to COVID-19 Infection. Front. Immunol. 2020. [Google Scholar] [CrossRef]
- Tecen-Yucel, K.; Aras-Atik, E.; Bayraktar-Ekincioglu, A. Does therapeutic drug monitoring of hydroxychloroquine improve treatment outcome in intensive care unit patients with COVID-19? Int. J. Clin. Pract. 2021, 75. [Google Scholar] [CrossRef] [PubMed]
- Sabo, Y.; Ehrlich, M.; Bacharach, E. The Conserved YAGL Motif in Human Metapneumovirus Is Required for Higher-Order Cellular Assemblies of the Matrix Protein and for Virion Production. J. Virol. 2011, 85, 6594–6609. [Google Scholar] [CrossRef] [Green Version]
- Bray, M. Highly pathogenic RNA viral infections: Challenges for antiviral research. Antivir. Res. 2008, 78, 1–8. [Google Scholar] [CrossRef]
- Terrier, O.; Si-Tahar, M.; Ducatez, M.; Chevalier, C.; Pizzorno, A.; Le Goffic, R.; Crepin, T.; Simon, G.; Naffakh, N. Influenza viruses and coronaviruses: Knowns, unknowns, and common research challenges. PLoS Pathog. 2021, 17, e1010106. [Google Scholar] [CrossRef]
- Turner, R.B.; Cetnarowski, W.E. Effect of treatment with zinc gluconate or zinc acetate on experimental and natural colds. Clin. Infect. Dis. 2000, 31, 1202–1208. [Google Scholar] [CrossRef]
- Prasad, A.S.; Beck, F.W.J.; Bao, B.; Snell, D.; Fitzgerald, J.T. Duration and severity of symptoms and levels of plasma interleukin-1 receptor antagonist, soluble tumor necrosis factor receptor, and adhesion molecules in patients with common cold treated with zinc acetate. J. Infect. Dis. 2008, 197, 795–802. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Song, J.; Liu, A.; Xiao, B.; Li, S.; Wen, Z.; Lu, Y.; Du, G. Research Progress of the Antiviral Bioactivities of Natural Flavonoids. Nat. Prod. Bioprospect. 2020, 10, 271–283. [Google Scholar] [CrossRef]
- Zakaryan, H.; Arabyan, E.; Oo, A.; Zandi, K. Flavonoids: Promising natural compounds against viral infections. Arch. Virol. 2017, 162, 2539–2551. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kreiser, T.; Zaguri, D.; Sachdeva, S.; Zamostiano, R.; Mograbi, J.; Segal, D.; Bacharach, E.; Gazit, E. Inhibition of Respiratory RNA Viruses by a Composition of Ionophoric Polyphenols with Metal Ions. Pharmaceuticals 2022, 15, 377. https://doi.org/10.3390/ph15030377
Kreiser T, Zaguri D, Sachdeva S, Zamostiano R, Mograbi J, Segal D, Bacharach E, Gazit E. Inhibition of Respiratory RNA Viruses by a Composition of Ionophoric Polyphenols with Metal Ions. Pharmaceuticals. 2022; 15(3):377. https://doi.org/10.3390/ph15030377
Chicago/Turabian StyleKreiser, Topaz, Dor Zaguri, Shreya Sachdeva, Rachel Zamostiano, Josef Mograbi, Daniel Segal, Eran Bacharach, and Ehud Gazit. 2022. "Inhibition of Respiratory RNA Viruses by a Composition of Ionophoric Polyphenols with Metal Ions" Pharmaceuticals 15, no. 3: 377. https://doi.org/10.3390/ph15030377
APA StyleKreiser, T., Zaguri, D., Sachdeva, S., Zamostiano, R., Mograbi, J., Segal, D., Bacharach, E., & Gazit, E. (2022). Inhibition of Respiratory RNA Viruses by a Composition of Ionophoric Polyphenols with Metal Ions. Pharmaceuticals, 15(3), 377. https://doi.org/10.3390/ph15030377