The Synthetic Curcumin Analog HO-3867 Rescues Suppression of PLAC1 Expression in Ovarian Cancer Cells
Abstract
1. Introduction
2. Results
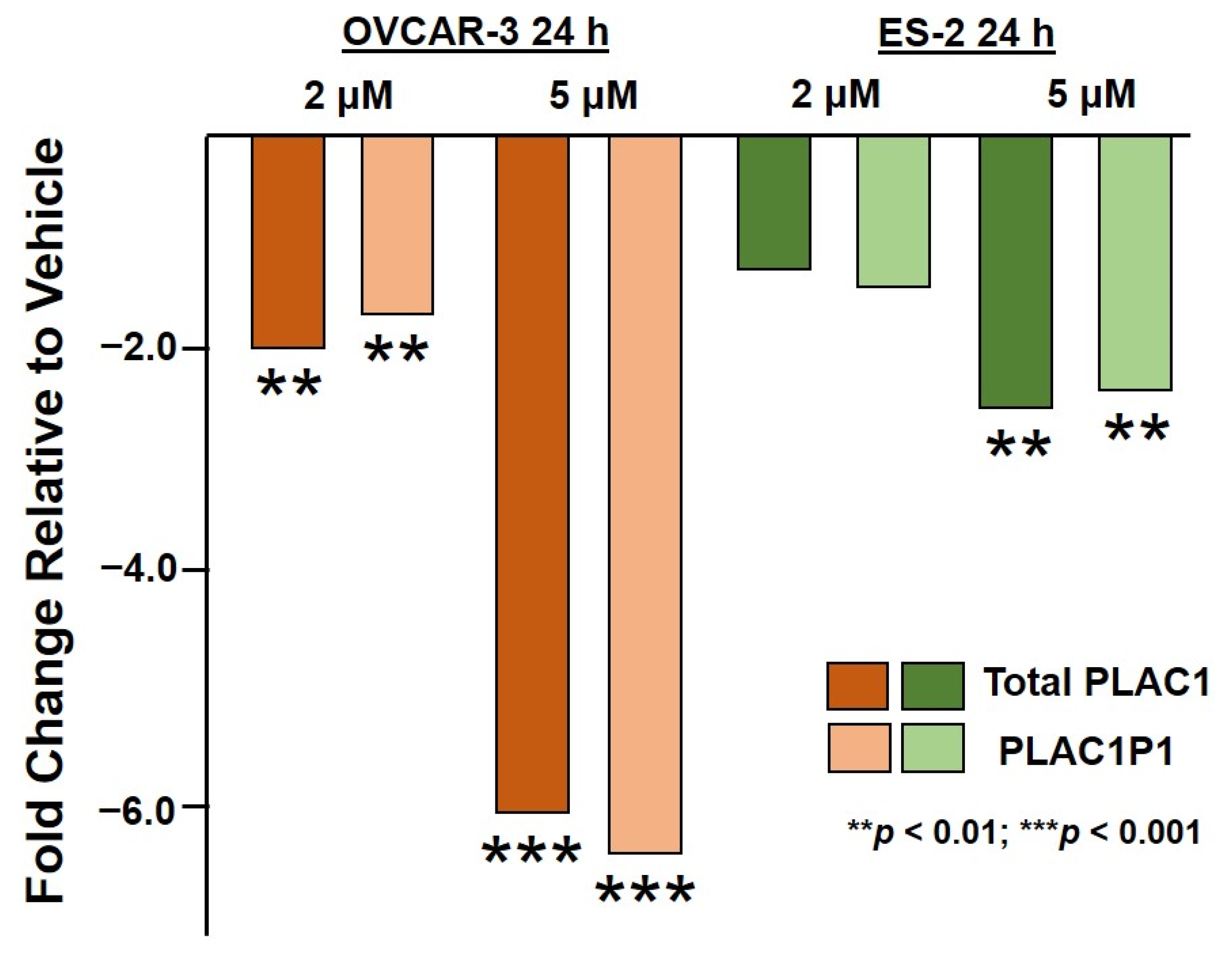
2.1. HO-3867 Treatment of Ovarian Cancer Cells Restores Suppression of PLAC1 Transcription
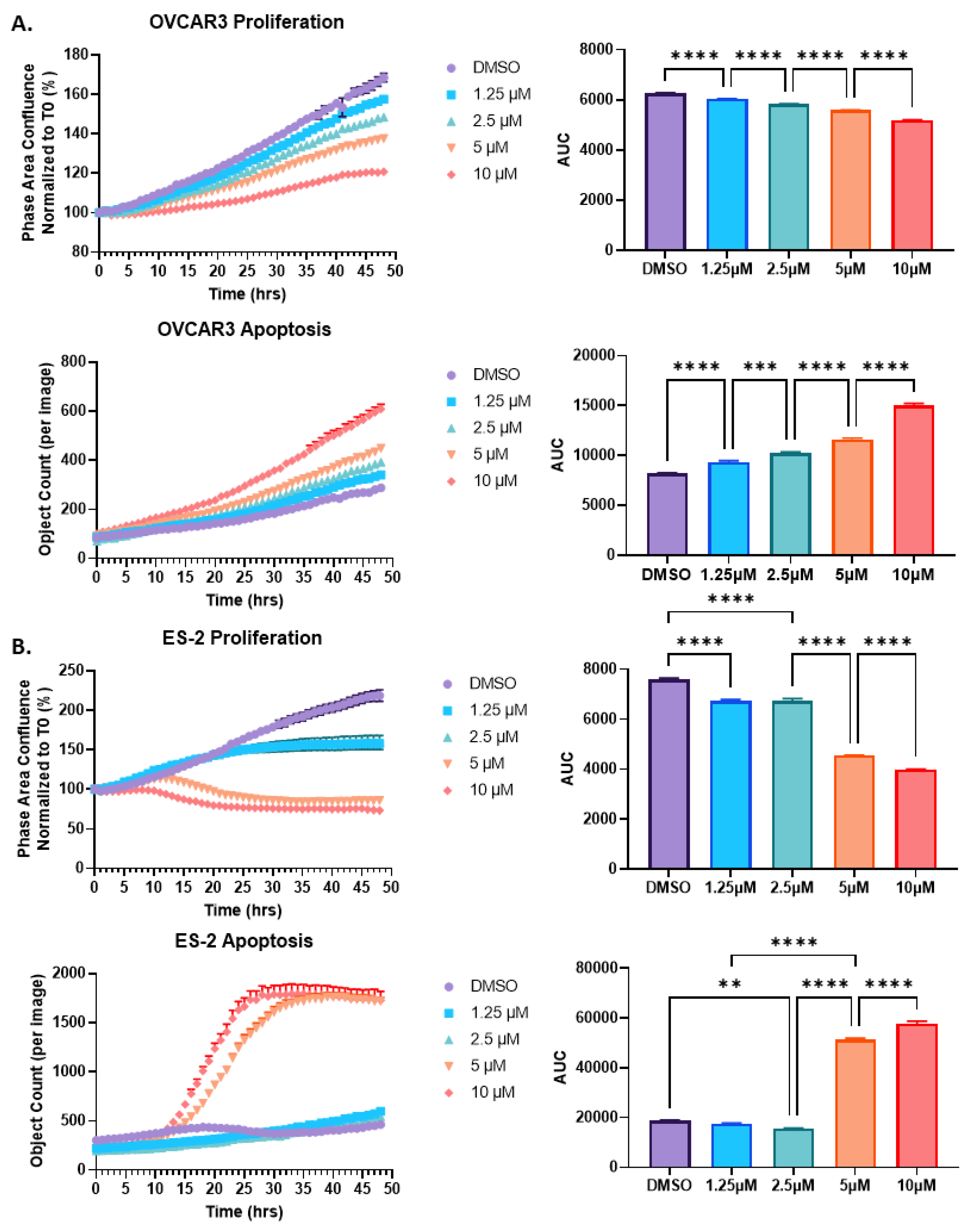
2.2. HO-3867 Treatment of Both OVCAR3 and ES-2 Ovarian Cancer Cells Results in a Reduction in Proliferation and an Increase in Apoptosis
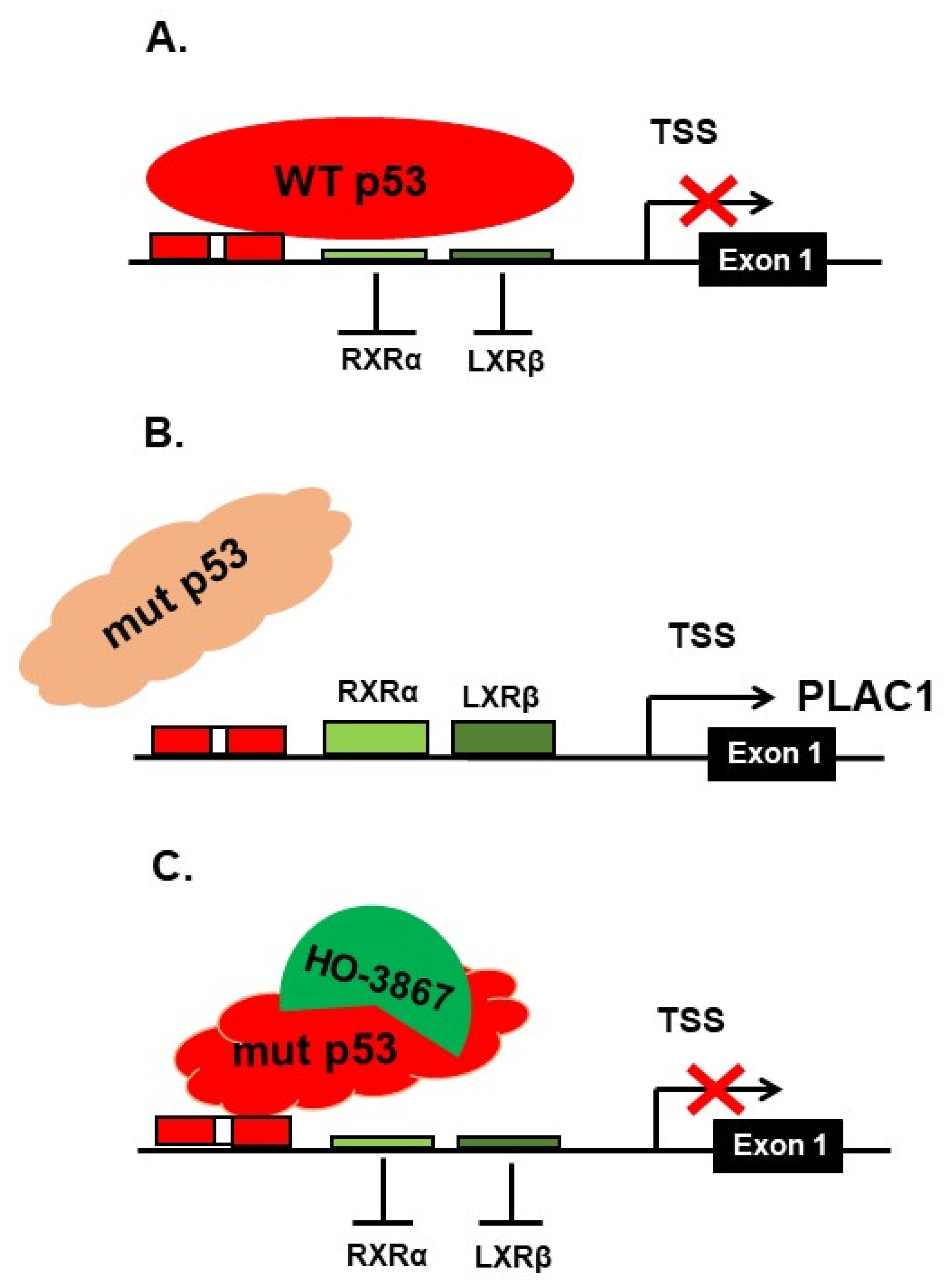
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. HO-3867 Treatment
4.3. RNA Purification and Quality Control
4.4. Real-Time PCR
4.5. Cell Proliferation and Apoptosis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cocchia, M.; Huber, R.; Pantano, S.; Chen, E.Y.; Ma, P.; Forabosco, A.; Ko, M.S.; Schlessinger, D. PLAC1, an Xq26 gene with placenta-specific expression. Genomics 2000, 68, 305–312. [Google Scholar] [CrossRef]
- Fant, M.; Barerra-Saldana, H.; Dubinsky, W.; Poindexter, B.; Bick, R. The PLAC1 protein localizes to membranous compartments in the apical region of the syncytiotrophoblast. Mol. Reprod. Dev. 2007, 74, 922–929. [Google Scholar] [CrossRef] [PubMed]
- Farina, A.; Concu, M.; Banzola, I.; Tempesta, A.; Vagnoni, S.; Gabrielli, S.; Mattioli, M.; Carinci, P.; Pilu, G.; Morano, D.; et al. PLAC1 mRNA in maternal blood correlates with Doppler waveform in uterine arteries in normal pregnancies at the second and third trimester. Ann. N. Y. Acad. Sci. 2006, 1075, 130–136. [Google Scholar] [CrossRef]
- Fujito, N.; Samura, O.; Miharu, N.; Tanigawa, M.; Hyodo, M.; Kudo, Y. Increased plasma mRNAs of placenta-specific 1 (PLAC1) and glial cells-missing 1 (GCM1) in mothers with pre-eclampsia. Hiroshima J. Med. Sci. 2006, 55, 9–15. [Google Scholar] [PubMed]
- Purwosunu, Y.; Sekizawa, A.; Farina, A.; Wibowo, N.; Okazaki, S.; Nakamura, M.; Samura, O.; Fujito, N.; Okai, T. Cell-free mRNA concentrations of CRH, PLAC1, and selectin-P are increased in the plasma of pregnant women with preeclampsia. Prenat. Diagn. 2007, 27, 772–777. [Google Scholar] [CrossRef]
- Kodama, M.; Miyoshi, H.; Fujito, N.; Samura, O.; Kudo, Y. Plasma mRNA concentrations of placenta-specific 1 (PLAC1) and pregnancy associated plasma protein A (PAPP-A) are higher in early-onset than late-onset pre-eclampsia. J. Obstet. Gynecol. Res. 2011, 37, 313–318. [Google Scholar] [CrossRef] [PubMed]
- Kotto-Kome, A.C.; Silva, C.; Whiteman, V.; Kong, X.; Fant, M.E. Circulating anti-PLAC1 antibodies during pregnancy and in women with reproductive failure: A preliminary analysis. ISRN Immunol. 2011, 530491. [Google Scholar] [CrossRef]
- Jackman, S.M.; Kong, X.; Fant, M.E. Plac1 (placenta-specific 1) is essential for normal placental and embryonic development. Mol. Reprod. Dev. 2012, 79, 564–572. [Google Scholar] [CrossRef] [PubMed]
- Matteo, M.; Greco, P.; Levi Setti, P.E.; Morenghi, E.; De Rosario, F.; Massenzio, F.; Albani, E.; Totaro, P.; Liso, A. Preliminary evidence for high anti-PLAC1 antibody levels in infertile patients with repeated unexplained implantation failure. Placenta 2013, 34, 335–339. [Google Scholar] [CrossRef] [PubMed]
- Zanello, M.; Sekizawa, A.; Purwosunu, Y.; Curti, A.; Farina, A. Circulating mRNA for the PLAC1 gene as a second trimester marker (14–18 weeks’ gestation) in the screening for late preeclampsia. Fetal Diagn. Ther. 2014, 36, 196–201. [Google Scholar] [CrossRef] [PubMed]
- Ibanoglu, M.C.; Ozgu-Erdinc, A.S.; Uygur, D. Maternal Plac1 protein levels in early- and late-onset preeclampsia. Ginekol. Pol. 2018, 89, 147–152. [Google Scholar] [CrossRef] [PubMed]
- Ibanoglu, M.C.; Ozgu-Erdinc, A.S.; Kara, O.; Topcu, H.O.; Uygur, D. Association of Higher Maternal Serum Levels of Plac1 Protein with Intrauterine Growth Restriction. Z. Geburtshilfe Neonatol. 2019, 223, 285–288. [Google Scholar] [CrossRef]
- Wan, L.; Sun, D.; Xie, J.; Du, M.; Wang, P.; Wang, M.; Lei, Y.; Wang, H.; Wang, H.; Dong, M. Declined placental PLAC1 expression is involved in preeclampsia. Medicine. 2019, 98, e17676. [Google Scholar] [CrossRef] [PubMed]
- Levine, L.; Habertheuer, A.; Ram, C.; Korutla, L.; Schwartz, N.; Hu, R.W.; Reddy, S.; Freas, A.; Zielinski, P.D.; Harmon, J.; et al. Syncytiotrophoblast extracellular microvesicle profiles in maternal circulation for noninvasive diagnosis of preeclampsia. Sci. Rep. 2020, 10, 6398. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, N.; Timur, H.; Ugurlu, E.N.; Yilmaz, S.; Ozgu-Erdinc, A.S.; Erkilinc, S.; Inal, H.A. Placenta specific protein-1 in recurrent pregnancy loss and in In Vitro Fertilisation failure: A prospective observational case-control study. J. Obstet. Gynaecol. 2020, 40, 843–848. [Google Scholar] [CrossRef]
- Chen, J.; Pang, X.W.; Liu, F.F.; Dong, X.Y.; Wang, H.C.; Wang, S.; Zhang, Y.; Chen, W.F. [PLAC1/CP1 gene expression and autologous humoral immunity in gastric cancer patients]. Beijing Da Xue Xue Bao Yi Xue Ban 2006, 38, 124–127. (In Chinese) [Google Scholar]
- Koslowski, M.; Sahin, U.; Mitnacht-Kraus, R.; Seitz, G.; Huber, C.; Türeci, O. A placenta-specific gene ectopically activated in many human cancers is essentially involved in malignant cell processes. Cancer Res. 2007, 67, 9528–9534. [Google Scholar] [CrossRef]
- Silva, W.A., Jr.; Gnjatic, S.; Ritter, E.; Chua, R.; Cohen, T.; Hsu, M.; Jungbluth, A.A.; Altorki, N.K.; Chen, Y.T.; Old, L.J.; et al. PLAC1, a trophoblast-specific cell surface protein, is expressed in a range of human tumors and elicits spontaneous antibody responses. Cancer Immunol. 2007, 7, 18. [Google Scholar]
- Dong, X.Y.; Peng, J.R.; Ye, Y.J.; Chen, H.S.; Zhang, L.J.; Pang, X.W.; Li, Y.; Zhang, Y.; Wang, S.; Fant, M.E.; et al. Plac1 is a tumor-specific antigen capable of eliciting spontaneous antibody responses in human cancer patients. Int. J. Cancer 2008, 122, 2038–2043. [Google Scholar] [CrossRef]
- Liu, F.F.; Dong, X.Y.; Pang, X.W.; Xing, Q.; Wang, H.C.; Zhang, H.G.; Li, Y.; Yin, Y.H.; Fant, M.; Ye, Y.J.; et al. The specific immune response to tumor antigen CP1 and its correlation with improved survival in colon cancer patients. Gastroenterology 2008, 134, 998–1006. [Google Scholar] [CrossRef]
- Tchabo, N.E.; Mhawech-Fauceglia, P.; Caballero, O.L.; Villella, J.; Beck, A.F.; Miliotto, A.J.; Liao, J.; Andrews, C.; Lele, S.; Old, L.J.; et al. Expression and serum immunoreactivity of developmentally restricted differentiation antigens in epithelial ovarian cancer. Cancer Immunol. 2009, 9, 6. [Google Scholar]
- Koslowski, M.; Türeci, O.; Biesterfeld, S.; Seitz, G.; Huber, C.; Sahin, U. Selective activation of trophoblast-specific PLAC1 in breast cancer by CCAAT/enhancer-binding protein beta (C/EBPbeta) isoform 2. J. Biol. Chem. 2009, 284, 28607–28615. [Google Scholar] [CrossRef]
- Devor, E.J.; Leslie, K.K. The oncoplacental gene placenta-specific protein 1 is highly expressed in endometrial tumors and cell lines. Obstet. Gynecol. Int. 2013, 807849. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Lu, J.; Xiao, J.; Upadhyay, G.; Umans, R.; Kallakury, B.; Yin, Y.; Fant, M.E.; Kopelovich, L.; Glazer, R.I. PPARδ induces estrogen receptor-positive mammary neoplasia through an inflammatory and metabolic phenotype linked to mTOR activation. Cancer Res. 2013, 73, 4349–4361. [Google Scholar] [CrossRef]
- Devor, E.J.; Reyes, H.D.; Santillan, D.A.; Santillan, M.K.; Onukwugha, C.; Goodheart, M.J.; Leslie, K.K. Placenta-specific protein 1: A potential key to many oncofetal-placental OB/GYN research questions. Obstet. Gynecol. Int. 2014, 678984. [Google Scholar] [CrossRef] [PubMed]
- Ghods, R.; Ghahremani, M.H.; Madjd, Z.; Asgari, M.; Abolhasani, M.; Tavasoli, S.; Mahmoudi, A.R.; Darzi, M.; Pasalar, P.; Jeddi-Tehrani, M.; et al. High placenta-specific 1/low prostate-specific antigen expression pattern in high-grade prostate adenocarcinoma. Cancer Immunol. Immunother. 2014, 63, 1319–1327. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Zhang, H.; Shen, D.; Wang, S.; Ye, Y.; Chen, H.; Pang, X.; Song, Q.; He, P. Identification of two new HLA-A*0201-restricted cytotoxic T lymphocyte epitopes from colorectal carcinoma-associated antigen PLAC1/CP1. J. Gastroenterol. 2014, 49, 419–426. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Shen, D.; Kang, X.; Zhang, C.; Song, Q. New tumour antigen PLAC1/CP1, a potentially useful prognostic marker and immunotherapy target for gastric adenocarcinoma. J. Clin. Pathol. 2015, 68, 913–916. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Lin, X.; Di, X.; Chen, Y.; Zhao, H.; Wang, X. Oncogenic function of Plac1 on the proliferation and metastasis in hepatocellular carcinoma cells. Oncol. Rep. 2017, 37, 465–473. [Google Scholar] [CrossRef]
- Guo, L.; Xu, D.; Lu, Y.; Peng, J.; Jiang, L. Detection of circulating tumor cells by reverse transcription-quantitative polymerase chain reaction and magnetic activated cell sorting in the peripheral blood of patients with hepatocellular carcinoma. Mol. Med. Rep. 2017, 16, 5894–5900. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yin, Y.; Zhu, X.; Huang, S.; Zheng, J.; Zhang, M.; Kong, W.; Chen, Q.; Zhang, Y.; Chen, X.; Lin, K.; et al. Expression and clinical significance of placenta-specific 1 in pancreatic ductal adenocarcinoma. Tumour Biol. 2017, 39, 1–8. [Google Scholar] [CrossRef]
- Devor, E.J.; Gonzalez-Bosquet, J.; Warrier, A.; Reyes, H.D.; Ibik, N.V.; Schickling, B.M.; Newtson, A.; Goodheart, M.J.; Leslie, K.K. p53 mutation status is a primary determinant of placenta-specific protein 1 expression in serous ovarian cancers. Int. J. Oncol. 2017, 50, 1721–1728. [Google Scholar] [CrossRef] [PubMed]
- Devor, E.J.; Reyes, H.D.; Gonzalez-Bosquet, J.; Warrier, A.; Kenzie, S.A.; Ibik, N.V.; Miller, M.D.; Schickling, B.M.; Goodheart, M.J.; Thiel, K.W.; et al. Placenta-specific protein 1 expression in human papillomavirus 16/18-positive cervical cancers is associated with tumor histology. Int. J. Gynecol. Cancer 2017, 27, 784–790. [Google Scholar] [CrossRef]
- Li, Y.; Chu, J.; Li, J.; Feng, W.; Yang, F.; Wang, Y.; Zhang, Y.; Sun, C.; Yang, M.; Vasilatos, S.N.; et al. Cancer/testis antigen-Plac1 promotes invasion and metastasis of breast cancer through Furin/NICD/PTEN signaling pathway. Mol. Oncol. 2018, 12, 1233–1248. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Zha, T.Q.; He, X.; Chen, L.; Zhu, Q.; Wu, W.B.; Nie, F.Q.; Wang, Q.; Zang, C.S.; Zhang, M.L. Placenta-specific protein 1 promotes cell proliferation and invasion in non-small cell lung cancer. Oncol. Rep. 2018, 39, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Wang, X.; Shi, C.; Jin, L.; Hu, J.; Zhang, A.; Li, J.; Vijayendra, N.; Doodala, V.; Weiss, S.; et al. Plac1 is a key regulator of the inflammatory response and immune tolerance in mammary tumorigenesis. Sci. Rep. 2018, 8, 5717. [Google Scholar] [CrossRef]
- Lin, C.; Qian, P.; Zhang, Y.; Liu, Z.; Dai, K.; Sun, D. Plac1 promotes nasopharyngeal carcinoma cells proliferation, migration and invasion via Furin/NICD/PTEN pathway. Tissue Cell Ress. 2021, 69, 101480. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Li, L.; Du, J.; Pan, C.; Zhang, C.; Chen, Y. Placenta-specific protein 1 enhances liver metastatic potential and is associated with the PI3K/AKT/NF-κB signaling pathway in colorectal cancer. Eur. J. Cancer Prev. 2021, 30, 220–231. [Google Scholar] [CrossRef]
- Chen, Y.; Moradin, A.; Schlessinger, D.; Nagaraja, R. RXRα and LXR activate two promoters in placenta- and tumor-specific expression of PLAC1. Placenta 2011, 32, 877–884. [Google Scholar] [CrossRef] [PubMed]
- Devor, E.J. Placenta-specific protein 1 (PLAC1) is a unique onco-fetal-placental protein and an underappreciated therapeutic target in cancer. Integr. Cancer Sci. Ther. 2016, 3, 479–483. [Google Scholar] [CrossRef]
- Chen, Y.; Schlessinger, D.; Nagaraja, R. T antigen transformation reveals TP53/RB-dependent route to PLAC1 transcription activation in primary fibroblasts. Oncogenesis 2013, 2, e67. [Google Scholar] [CrossRef]
- Madan, E.; Parker, T.M.; Bauer, M.R.; Dhiman, A.; Pelham, C.J.; Nagane, M.; Kuppusamy, M.L.; Holmes, M.; Holmes, T.R.; Shaik, K.; et al. The curcumin analog HO-3867 selectively kills cancer cells by converting mutant p53 protein to transcriptionally active wildtype p53. J. Biol. Chem. 2018, 293, 4262–4276. [Google Scholar] [CrossRef] [PubMed]
- Mullany, L.K.; Wong, K.-K.; Marciano, D.C.; Katsonis, P.; King-Crane, E.R.; Ren, Y.A.; Lichtarge, O.; Richards, J.S. Specific TP53 mutants overrepresented in ovarian cancer impact CNV, TP53 activity, responses to Nutlin-3a, and cell survival. Neoplasia 2015, 17, 789–803. [Google Scholar] [CrossRef]
- Joerger, A.C.; Fersht, A.R. Structure-function-rescue: The diverse nature of common p53 cancer mutants. Oncogene 2007, 26, 2226–2242. [Google Scholar] [CrossRef]
- Emerling, B.M.; Hurov, J.B.; Poulogiannis, G.; Tsukazawa, K.S.; Choo-Wing, R.; Wulf, G.M.; Bell, E.L.; Shim, H.-S.; Lamia, K.A.; Rameh, L.E.; et al. Depletion of a putatively druggable class of phosphatidylinositol kinases inhibits growth of p53-null tumors. Cell 2013, 155, 844–857. [Google Scholar] [CrossRef] [PubMed]
- Parrales, A.; Iwakuma, T. Targeting oncogenic mutant p53 for cancer therapy. Front. Oncol. 2015, 5, 288. [Google Scholar] [CrossRef] [PubMed]
- Bykov, V.J.N.; Issaeva, N.; Shilov, A.; Hultcrantz, M.; Pugacheva, E.; Chumakov, P.; Bergman, J.; Wiman, K.; Selivanova, G. Restoration of the tumor suppressor function to mutant p53 by a low molecular-weight compound. Nat. Med. 2002, 8, 282–288. [Google Scholar] [CrossRef] [PubMed]
- Kastan, M.B. Wild-type p53: Tumors can’t stand it. Cell 2007, 128, 837–840. [Google Scholar] [CrossRef]
- Vinyals, A.; Peinado, M.A.; Gonzalez-Garrigues, M.; Monzo, M.; Binfil, R.D.; Fabra, A. Failure of wild-type p53 gene therapy in human cancer cells expressing a mutant p53 protein. Gene Ther. 1999, 6, 22–33. [Google Scholar] [CrossRef]
- Watanabe, T.; Sullenger, B.A. Introduction of wild-type p53 activity in human cancer cells by ribozymes that repair mutant p53 transcripts. Proc. Natl. Acad. Sci. USA 2000, 97, 8490–8494. [Google Scholar] [CrossRef] [PubMed]
- Duffy, M.J.; Synnott, N.C.; Crown, J. Mutant p53 as a target for cancer treatment. Eur. J. Cancer 2017, 83, 258–265. [Google Scholar] [CrossRef]
- Vareki, S.M.; Salim, K.Y.; Danter, W.R.; Koropatnick, J. Novel anti-cancer drug COTI-2 synergizes with therapeutic agents and does not induce resistance or exhibit cross-resistance in human cancer cell lines. PLoS ONE 2018, 13, e0191766. [Google Scholar] [CrossRef] [PubMed]
- Lambert, J.M.R.; Gorzov, P.; Veprintsev, D.B.; Soderqvist, M.; Segerback, D.; Bergman, J.; Fersht, A.R.; Hainault, P.; Wiman, K.G.; Bykov, V.J.N. PRIMA-1 reactivates mutant p53 by covalently binding to the core domain. Cancer Cell 2019, 15, 376–388. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Bykov, V.J.N.; Wiman, K.; Zawacka-Pankua, J. APR-246 reactivates mutant p53 by targeting cysteines 124 and 277. Cell Death Dis. 2018, 9, 439. [Google Scholar] [CrossRef] [PubMed]
- Brachova, P.; Mueting, S.R.; Devor, E.J.; Leslie, K.K. Oncomorphic TP53 mutations in gynecologic cancers lose the normal protein:protein interactions with the microRNA microporcessing complex. J. Cancer Ther. 2014, 5, 506–516. [Google Scholar] [CrossRef]
- Fischer, M. Census and evaluation of p53 target genes. Oncogene 2017, 36, 3943–3956. [Google Scholar] [CrossRef]
- Nguyen, T.-A.T.; Grimm, S.A.; Bushel, P.R.; Li, J.; Li, Y.; Bennett, B.D.; Lavender, C.A.; Ward, J.M.; Fargo, D.C.; Anderson, C.W.; et al. Revealing a human p53 universe. Nucleic Acids Res. 2018, 46, 8153–8167. [Google Scholar] [CrossRef]
- Selvendiran, K.; Tong, L.; Bratasz, A.; Kuppusamy, M.L.; Ahmed, S.; Ravi, Y.; Trigg, N.J.; Rivera, B.K.; Kalai, T.; Hideg, K.; et al. Anticancer efficacy of a difluorodiarylidenyl piperidone (HO-3867) in human ovarian cancer cells and tumor xenographs. Mol. Cancer Ther. 2010, 9, 1169–1179. [Google Scholar] [CrossRef]
- Selvendiran, K.; Ahmed, S.; Dayton, A.; Kuppusamy, M.L.; Rivera, B.K.; Kalai, T.; Hideg, K.; Kuppusamy, P. HO-3867, a curcumin analog, sensitizes cisplatin-resistant ovarian carcinoma, leading to therapeutic synergy through STAT3 inhibition. Cancer Biol. Ther. 2011, 12, 837–845. [Google Scholar] [CrossRef]
- Rath, K.S.; Naidu, S.K.; Lata, P.; Bid, H.K.; Rivera, B.K.; McCann, G.A.; Tierney, B.J.; ElNaggar, A.C.; Bravo, V.; Leone, G.; et al. HO-3867, a safe STAT3 inhibitor, is selectively cytotoxic to ovarian cancer. Cancer Res. 2014, 74, 2316–2327. [Google Scholar] [CrossRef]
- Bixel, K.; Saini, U.; Bid, H.K.; Fowler, J.; Riley, M.; Wanner, R.; Dorayappan, K.D.P.; Rajendran, S.; Konishi, I.; Matsumura, N.; et al. Targeting STAT3 by HO-3867 induces apoptosis in ovarian clear cell carcinoma. Int. J. Cancer 2017, 141, 1856–1866. [Google Scholar] [CrossRef]
- Lindeboom, R.G.H.; Supek, F.; Lehner, B. The rules and impact of nonsense-mediated mRNA decay in human cancers. Nat. Genet. 2017, 48, 1112–1118. [Google Scholar] [CrossRef]
- Hamilton, T.C.; Young, R.C.; McKoy, W.M.; Grotzinger, K.R.; Green, J.A.; Chu, E.W.; Whang-Peng, J.; Rogan, A.M.; Green, W.R.; Ozols, R.F. Characterization of a human ovarian carcinoma cell line (NIH:OVCAR3) with androgen and estrogen receptors. Cancer Res. 1983, 43, 5379–5398. [Google Scholar] [PubMed]
- Lau, D.H.; Lewis, A.D.; Ehsan, M.N.; Sikic, B.I. Multifactorial mechanisms associated with broad cross-resistance of ovarian carcinoma cells selected by cyanomorpholino doxorubicin. Cancer Res. 1991, 51, 5181–5187. [Google Scholar] [PubMed]
- Schroeder, A.; Mueller, O.; Stocker, S.; Salowsky, R.; Leiber, M.; Gassmann, M.; Lightfoot, S.; Menzel, W.; Granzow, M.; Ragg, T. The RIN: An RNA integrity number for assigning integrity values to RNA measurements. BMC Mol. Biol. 2006, 7, 3. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta, C.(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative Ct method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]
- Snedecor, G.W.; Cochran, W.G. Statistical Methods, 8th ed.; State University Press: Ames, IA, USA, 1989. [Google Scholar]

| Locus | Sequence | Amplicon | Tm (°C) * |
|---|---|---|---|
| PLAC1 | 5′-CACCAGTGAGCACAAAGCCACATT-3 | 232 bp | 60.3 |
| 5′-CCATGAACCAGTCTATGGAG-3′ | 52.3 | ||
| PLAC1P1 | 5′-AAACTTACACGAGGAGTCTGTC-3′ | 371 bp # | 57.2 |
| 5′-CTGTGACCATGAACCAGTCTAT-3′ | 285 bp # | 54.2 | |
| 18S rRNA | 5′-AACTTTCGATGGTAGTCGCCG-3′ | 104 bp | 57.2 |
| 5′-CCTTGGATGTGGTAGCCGTTT-3′ | 54.2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Devor, E.J.; Schickling, B.M.; Lapierre, J.R.; Bender, D.P.; Gonzalez-Bosquet, J.; Leslie, K.K. The Synthetic Curcumin Analog HO-3867 Rescues Suppression of PLAC1 Expression in Ovarian Cancer Cells. Pharmaceuticals 2021, 14, 942. https://doi.org/10.3390/ph14090942
Devor EJ, Schickling BM, Lapierre JR, Bender DP, Gonzalez-Bosquet J, Leslie KK. The Synthetic Curcumin Analog HO-3867 Rescues Suppression of PLAC1 Expression in Ovarian Cancer Cells. Pharmaceuticals. 2021; 14(9):942. https://doi.org/10.3390/ph14090942
Chicago/Turabian StyleDevor, Eric J., Brandon M. Schickling, Jace R. Lapierre, David P. Bender, Jesus Gonzalez-Bosquet, and Kimberly K. Leslie. 2021. "The Synthetic Curcumin Analog HO-3867 Rescues Suppression of PLAC1 Expression in Ovarian Cancer Cells" Pharmaceuticals 14, no. 9: 942. https://doi.org/10.3390/ph14090942
APA StyleDevor, E. J., Schickling, B. M., Lapierre, J. R., Bender, D. P., Gonzalez-Bosquet, J., & Leslie, K. K. (2021). The Synthetic Curcumin Analog HO-3867 Rescues Suppression of PLAC1 Expression in Ovarian Cancer Cells. Pharmaceuticals, 14(9), 942. https://doi.org/10.3390/ph14090942







