MindReader: Unsupervised Classification of Electroencephalographic Data
Abstract
1. Introduction
2. Materials and Methods
2.1. Dataset Analyzed
2.2. Software
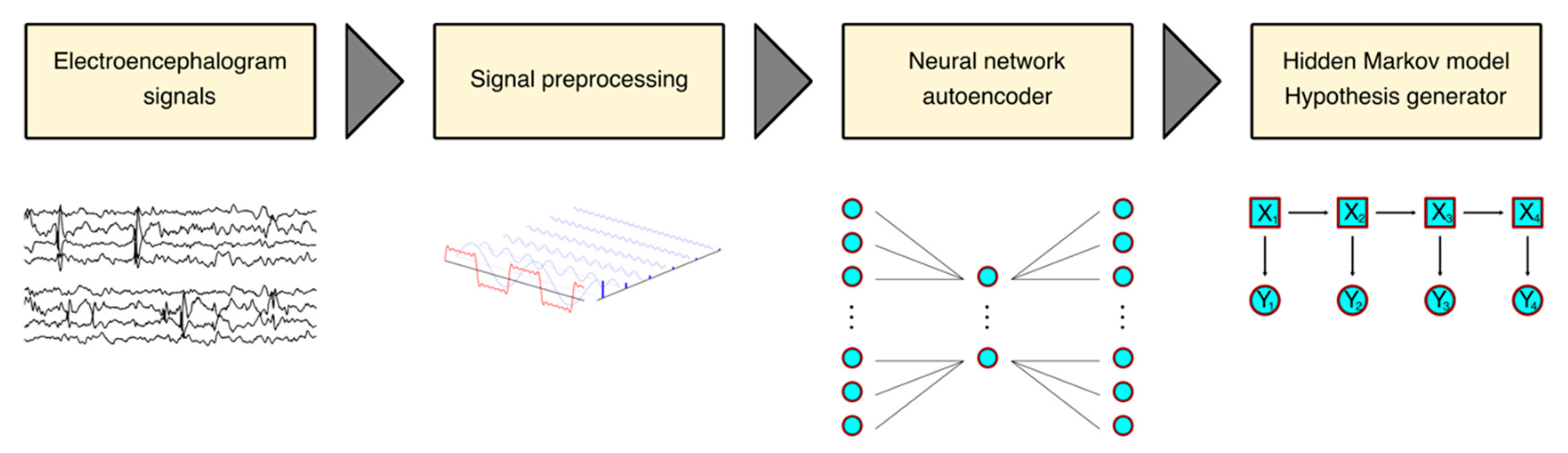
2.3. Architecture
- This is embarrassingly parallelizable for performance purposes;
- The computational complexity for a single channel is O(T), with T being the recording time, times O(N) with N denoting the number of channels. Even though memory consumption is low, it currently scales at O(T)*O(N), which can be further optimized, e.g., for deployment in embedded systems;
- MindReader is adaptable for different EEG montages, i.e., electrode placement;
- Identifying electrical anomalies independently allows for spatial localization per channel as well as hypothesizing the physiological relationship among different areas of the brain;
- Epileptogenic/irritative zones are potentially detectable and physically mappable. Notably, MindReader does not require specialized hardware and can be easily used after installation under any operating system: Linux, Windows, or OSX. Moreover, due to MindReader’s short run-time, it is potentially applicable in live interpretations.
3. Results
3.1. Physionet Dataset
3.2. MindReader Predictive Performance
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Olejniczak, P. Neurophysiologic basis of EEG. J. Clin. Neurophysiol. 2006, 23, 186–189. [Google Scholar] [CrossRef] [PubMed]
- Beghi, E.; Giussani, G.; Nichols, E.; Abd-Allah, F.; Abdela, J.; Abdelalim, A.; Abraha, H.N.; Adib, M.G.; Agrawal, S.; Alahdab, F.; et al. Global, regional, and national burden of epilepsy, 1990–2016: A systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 2019, 18, 357–375. [Google Scholar] [CrossRef]
- Baldin, E.; Hauser, W.A.; Buchhalter, J.R.; Hesdorffer, D.C.; Ottman, R. Yield of epileptiform electroencephalogram abnormalities in incident unprovoked seizures: A population-based study. Epilepsia 2014, 55, 1389–1398. [Google Scholar] [CrossRef]
- Schreiner, A.; Pohlmann-Eden, B. Value of the Early Electroencephalogram after a First Unprovoked Seizure. Clin. Electroencephalogr. 2003, 34, 140–144. [Google Scholar] [CrossRef] [PubMed]
- Tatum, W.O.; Rubboli, G.; Kaplan, P.W.; Mirsatari, S.M.; Radhakrishnan, K.; Gloss, D.; Caboclo, L.O.; Drislane, F.W.; Koutroumanidis, M.; Schomer, D.L.; et al. Clinical utility of EEG in diagnosing and monitoring epilepsy in adults. Clin. Neurophysiol. 2018, 129, 1056–1082. [Google Scholar] [CrossRef] [PubMed]
- Taran, S.; Ahmed, W.; Bui, E.; Prisco, L.; Hahn, C.D.; McCredie, V.A. Educational initiatives and implementation of electroencephalography into the acute care environment: A protocol of a systematic review. Syst. Rev. 2020, 9, 175. [Google Scholar] [CrossRef]
- Hannun, A.Y.; Rajpurkar, P.; Haghpanahi, M.; Tison, G.H.; Bourn, C.; Turakhia, M.P.; Ng, A.Y. Cardiologist-level arrhythmia detection and classification in ambulatory electrocardiograms using a deep neural network. Nat. Med. 2019, 25, 65–69. [Google Scholar] [CrossRef]
- Yang, X.; Zhang, X.; Yang, M.; Zhang, L. 12-Lead ECG arrhythmia classification using cascaded convolutional neural network and expert feature. J. Electrocardiol. 2021, 67, 56–62. [Google Scholar] [CrossRef]
- Allam, J.P.; Samantray, S.; Behara, C.; Kurkute, K.K.; Sinha, V.K. 8—Customized deep learning algorithm for drowsiness detection using single-channel EEG signal. In Artificial Intelligence-Based Brain-Computer Interface; Bajaj, V., Sinha, G.R., Eds.; Academic Press: Cambridge, MA, USA, 2022; pp. 189–201. [Google Scholar]
- Locharla, G.R.; Pogiri, R.; Allam, J.P. 9—EEG-based deep learning neural net for apnea detection. In Artificial Intelligence-Based Brain-Computer Interface; Bajaj, V., Sinha, G.R., Eds.; Academic Press: Cambridge, MA, USA, 2022; pp. 203–215. [Google Scholar]
- Beniczky, S.; Aurlien, H.; Brøgger, J.C.; Fuglsang-Frederiksen, A.; Martins-da-Silva, A.; Trinka, E.; Visser, G.; Rubboli, G.; Hjalgrim, H.; Stefan, H.; et al. Standardized computer-based organized reporting of EEG: SCORE. Epilepsia 2013, 54, 1112–1124. [Google Scholar] [CrossRef]
- Halford, J.J.; Pressly, W.B.; Benbadis, S.R.; Tatum, W.O., 4th; Turner, R.P.; Arain, A.; Pritchard, P.B.; Edwards, J.C.; Dean, B.C. Web-based collection of expert opinion on routine scalp EEG: Software development and interrater reliability. J. Clin. Neurophysiol. 2011, 28, 178–184. [Google Scholar] [CrossRef]
- Wüstenhagen, S.; Terney, D.; Gardella, E.; Meritam Larsen, P.; Rømer, C.; Aurlien, H.; Beniczky, S. EEG normal variants: A prospective study using the SCORE system. Clin. Neurophysiol. Pract. 2022, 7, 183–200. [Google Scholar] [CrossRef]
- Benbadis, S.R.; Tatum, W.O. Overintepretation of EEGs and misdiagnosis of epilepsy. J. Clin. Neurophysiol. 2003, 20, 42–44. [Google Scholar] [CrossRef]
- Kang, J.Y.; Krauss, G.L. Normal Variants Are Commonly Overread as Interictal Epileptiform Abnormalities. J. Clin. Neurophysiol. 2019, 36, 257–263. [Google Scholar] [CrossRef] [PubMed]
- Krauss, G.L.; Abdallah, A.; Lesser, R.; Thompson, R.E.; Niedermeyer, E. Clinical and EEG features of patients with EEG wicket rhythms misdiagnosed with epilepsy. Neurology 2005, 64, 1879–1883. [Google Scholar] [CrossRef]
- Santoshkumar, B.; Chong, J.J.; Blume, W.T.; McLachlan, R.S.; Young, G.B.; Diosy, D.C.; Burneo, J.G.; Mirsattari, S.M. Prevalence of benign epileptiform variants. Clin. Neurophysiol. 2009, 120, 856–861. [Google Scholar] [CrossRef]
- Fowle, A.J.; Binnie, C.D. Uses and abuses of the EEG in epilepsy. Epilepsia 2000, 41 (Suppl. 3), S10–S18. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, I.; Wang, X.; Zhu, M.; Wang, C.; Pi, Y.; Khan, J.A.; Khan, S.; Samuel, O.W.; Chen, S.; Li, G. EEG-Based Epileptic Seizure Detection via Machine/Deep Learning Approaches: A Systematic Review. Comput. Intell. Neurosci. 2022, 2022, 6486570. [Google Scholar] [CrossRef] [PubMed]
- Rathod, P.; Naik, S. Review on Epilepsy Detection with Explainable Artificial Intelligence. In Proceedings of the 2022 10th International Conference on Emerging Trends in Engineering and Technology—Signal and Information Processing (ICETET-SIP-22), Nagpur, India, 29–30 April 2022; IEEE: New York City, NY, USA, 2022. [Google Scholar]
- Goldberger, A.L.; Amaral, L.A.N.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.-K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet. Circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef]
- Innes, M. Flux: Elegant machine learning with Julia. J. Open Source Softw. 2018, 3, 602. [Google Scholar] [CrossRef]
- Kemp, B.; Olivan, J. European data format ‘plus’ (EDF+), an EDF alike standard format for the exchange of physiological data. Clin. Neurophysiol. 2003, 114, 1755–1761. [Google Scholar] [CrossRef] [PubMed]
- Kramer, M.A. Nonlinear principal component analysis using autoassociative neural networks. AIChE J. 1991, 37, 233–243. [Google Scholar] [CrossRef]
- Hinton, G.E.; Salakhutdinov, R.R. Reducing the Dimensionality of Data with Neural Networks. Science 2006, 313, 504–507. [Google Scholar] [CrossRef]
- An, J.; Cho, S. Variational Autoencoder based Anomaly Detection using Reconstruction Probability. Spec. Lect. IE 2015, 2, 1–18. [Google Scholar]
- Zamani, N.; Russell, P.; Lantz, H.; Hoeppner, M.P.; Meadows, J.R.; Vijay, N.; Mauceli, E.; Di Palma, F.; Lindblad-Toh, K.; Jern, P.; et al. Unsupervised genome-wide recognition of local relationship patterns. BMC Genom. 2013, 14, 347. [Google Scholar] [CrossRef]
- Ramantani, G.; Maillard, L.; Koessler, L. Correlation of invasive EEG and scalp EEG. Seizure 2016, 41, 196–200. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.K.; Kumar, K.; Seelamantula, C.S.; Singh Thakur, C. A Portable Ultrasound Imaging System Utilizing Deep Generative Learning-Based Compressive Sensing on Pre-Beamformed RF Signals. In Proceedings of the 2019 41st Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC), Berlin, Germany, 23–27 July 2019; pp. 2740–2743. [Google Scholar]
- You, S.; Hwan Cho, B.; Shon, Y.M.; Seo, D.W.; Kim, I.Y. Semi-supervised automatic seizure detection using personalized anomaly detecting variational autoencoder with behind-the-ear EEG. Comput. Methods Programs Biomed. 2022, 213, 106542. [Google Scholar] [CrossRef] [PubMed]
- Pisa, I.; Morell, A.; Vicario, J.L.; Vilanova, R. Denoising Autoencoders and LSTM-Based Artificial Neural Networks Data Processing for Its Application to Internal Model Control in Industrial Environments-The Wastewater Treatment Plant Control Case. Sensors 2020, 20, 3743. [Google Scholar] [CrossRef]
- Graves, A.; Schmidhuber, J. Framewise phoneme classification with bidirectional LSTM and other neural network architectures. Neural Netw. 2005, 18, 602–610. [Google Scholar] [CrossRef]
- Zhao, R.; Yan, R.; Wang, J.; Mao, K. Learning to Monitor Machine Health with Convolutional Bi-Directional LSTM Networks. Sensors 2017, 17, 273. [Google Scholar] [CrossRef]
- Luong, M.-T.; Pham, H.; Manning, C.D. Effective Approaches to Attention-based Neural Machine Translation. arXiv 2015, arXiv:1508.04025. [Google Scholar]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, L.; Polosukhin, I. Attention Is All You Need. In Proceedings of the Advances in Neural Information Processing Systems 30: Annual Conference on Neural Information Processing Systems 2017, Long Beach, CA, USA, 4–9 December 2017. [Google Scholar]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rivas-Carrillo, S.D.; Akkuratov, E.E.; Valdez Ruvalcaba, H.; Vargas-Sanchez, A.; Komorowski, J.; San-Juan, D.; Grabherr, M.G. MindReader: Unsupervised Classification of Electroencephalographic Data. Sensors 2023, 23, 2971. https://doi.org/10.3390/s23062971
Rivas-Carrillo SD, Akkuratov EE, Valdez Ruvalcaba H, Vargas-Sanchez A, Komorowski J, San-Juan D, Grabherr MG. MindReader: Unsupervised Classification of Electroencephalographic Data. Sensors. 2023; 23(6):2971. https://doi.org/10.3390/s23062971
Chicago/Turabian StyleRivas-Carrillo, Salvador Daniel, Evgeny E. Akkuratov, Hector Valdez Ruvalcaba, Angel Vargas-Sanchez, Jan Komorowski, Daniel San-Juan, and Manfred G. Grabherr. 2023. "MindReader: Unsupervised Classification of Electroencephalographic Data" Sensors 23, no. 6: 2971. https://doi.org/10.3390/s23062971
APA StyleRivas-Carrillo, S. D., Akkuratov, E. E., Valdez Ruvalcaba, H., Vargas-Sanchez, A., Komorowski, J., San-Juan, D., & Grabherr, M. G. (2023). MindReader: Unsupervised Classification of Electroencephalographic Data. Sensors, 23(6), 2971. https://doi.org/10.3390/s23062971







