1. Introduction
The collection and annotation of data on the appearance and occurrence of species are crucial pillars of biological research and practical nature conservation work focusing on biodiversity, climate change and species extinction [
1,
2]. The involvement of citizen communities is a cost effective approach to large scale data acquisition. Species observation datasets collected by the public have already been proven to improve data quality and to add significant value for understanding both basic and more applied aspects of mycology [
3,
4,
5,
6]. Citizen-science contributions provide more than 50% of all data accessible through the Global Biodiversity Information Facility [
7].
In citizen-science projects focusing on biodiversity, correct species identification is a challenge. Poor data quality is often quoted as a major concern about species data provided by untrained citizens [
8]. Some projects handle the issue by reducing the complexity of the species identification process, for example, by merging species into multitaxa indicator groups [
9], by focusing only on a subset of easily identifiable species or by involving human expert validators in the identification process. Other projects involve citizen-science communities in the data validation process. For instance, iNaturalist [
10] regards observations as research-grade labelled if three independent users have verified a suggested taxon based on an uploaded photo. Automatic image-based species identification can act both as a supplement or an alternative to these approaches.
We are interested in automating the process of fungi identification using machine learning. This has been made possible by the rapid progress of computer vision in the past decade, which was, to a great extent, facilitated by the existence of large-scale image collections. In the case of image recognition, the introduction of the ImageNet [
11] database and its use in the ILSVRC (The ImageNet Large Scale Visual Recognition Challenge) challenge [
12], together with PASCAL VOC [
13], helped start the CNN revolution. The same holds for the problem of fine-grained visual categorization (FGVC), where datasets and challenges like PlantCLEF [
14,
15,
16], iNaturalist [
17], CUB [
18], and Oxford Flowers [
19] have triggered the development and evaluation of novel approaches to fine-grained domain adaptation [
20], domain specific transfer learning [
21], image retrieval [
22,
23,
24], unsupervised visual representation [
25,
26], few-shot learning [
27], transfer learning [
21] and prior-shift [
28].
In this paper, we describe a system for AI-based fungi species recognition to help a citizen-science community — the Atlas of Danish Fungi. The system for fungi recognition “in the wild” achieved the best results in a Kaggle competition sponsored by the Danish Mycological Society, which was organized in conjunction with the Fine-Grained Categorization Workshop at CVPR 2018. The real-time identification tool (FungiVision) led to an increase in public interest in nature, quadrupling the number of citizens collecting data. It supports hands-on learning, much as children learn from their parents by asking direct and naïve questions that are answered on the spot. A supervised machine learning system with a human in the loop was created by linking the system to an existing mycological platform with an existing community-based validation process.
From the computer vision perspective, the application of the system to citizen-science data collection creates a valuable continuous stream of labelled examples for a challenging fine-grained visual classification task. Based on observations submitted to the Atlas of Danish Fungi, we introduce a novel fine-grained dataset and benchmark, the Danish Fungi 2020 (DF20). The dataset is unique in its taxonomy-accurate class labels, small number of errors, highly unbalanced long-tailed class distribution, rich observation metadata, and well-defined class hierarchy. DF20 has zero overlap with ImageNet, allowing unbiased comparison of models fine-tuned from publicly available ImageNet checkpoints. The proposed evaluation protocol enables testing the ability to improve classification using metadata — for example, precise geographic location, habitat and substrate, facilitates classifier calibration testing, and finally allows us to study the impact of the device settings on the classification performance.
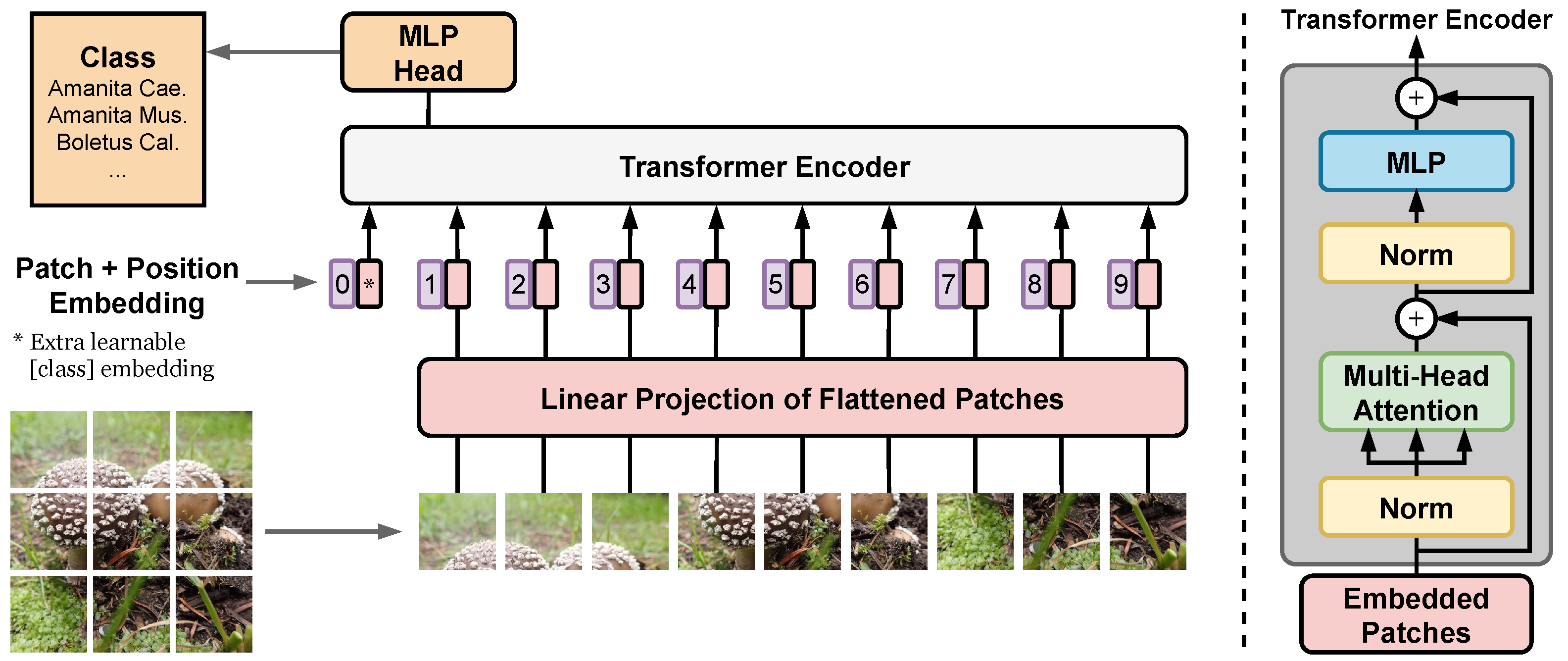
Finally, we present a substantial upgrade of the first version of the fungi recognition service by: (i) shifting from CNN towards Vision Transformers (ViT), we achieved state-of-the-art results in fine-grained classification; (ii) utilizing a simple procedure for including metadata in the decision process, improving the classification accuracy by more than 2.95 percentage points, reducing the error rate by 15%; (iii) increasing the amount of training data obtained with the help of the online identification tool. A new Vision Transformer architecture, which lowers the recognition error of the current system by 46.75%, is under review before deployment. By providing a stream of labeled data in one direction, and an improvement of the FungiVision in the other, the collaboration creates a virtuous cycle that helps both communities.
This paper is an extended version of our two papers published in WACV 2020 [
29] and WACV 2022 [
30].
3. Data
All experiments were based on datasets collected from the Atlas of Danish Fungi, which is described in
Section 3.1. The details of the particular dataset are presented in
Section 3.2,
Section 3.3 and
Section 3.4. Quantitative parameters of the used datasets are summarized in
Table 1. For reference, the table includes iNaturalist 2021, the richest (in the number of species) and largest (in the number of observations) publicly available fungi dataset not based on the Atlas of Danish Fungi. The species from the
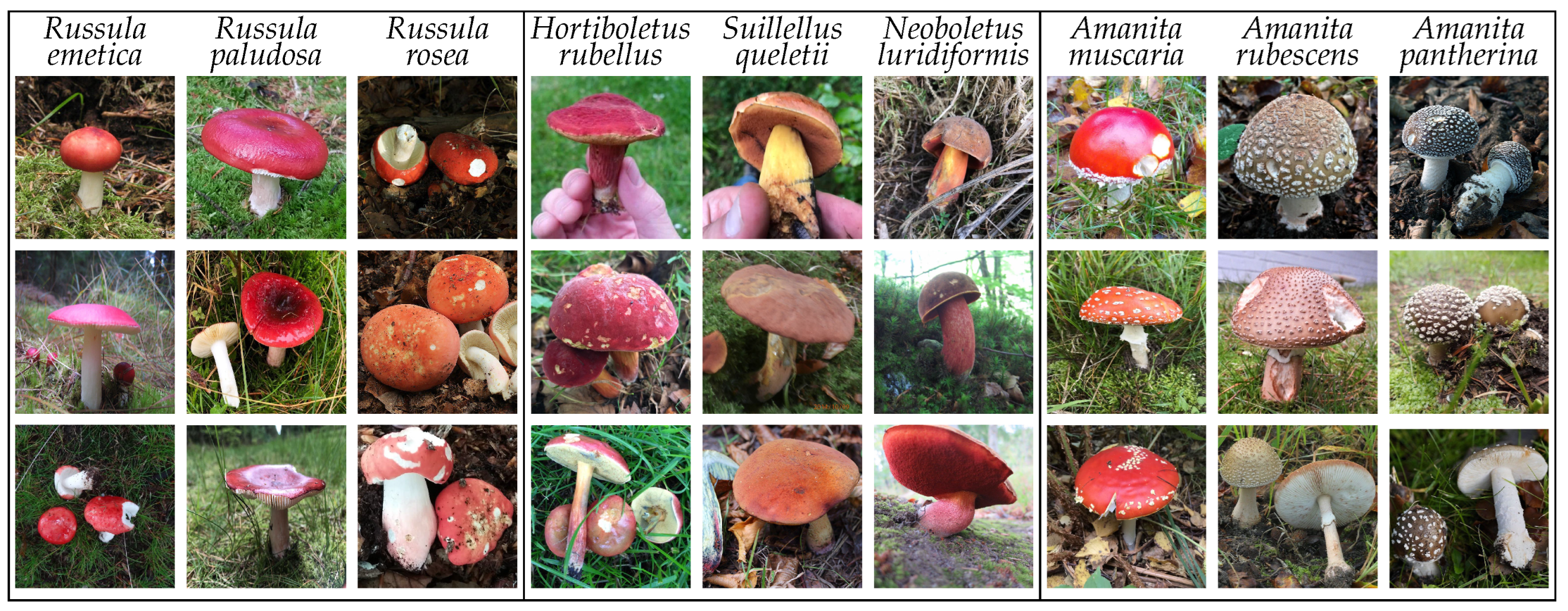
Fungi kingdom are, by nature, visually similar, thus introducing a challenging machine learning problem. The existing high intra- and inter-class similarities and differences present in the data are visualized in
Figure 1.
3.1. Atlas of Danish Fungi
The Atlas of Danish Fungi [
55,
56,
57] is supported by more than 4000 volunteers who have contributed more than 1 million content-checked observations of approximately 8300 fungi species, many with expert-validated class labels. The project has resulted in a vastly improved knowledge of fungi in Denmark [
57]. More than 180 species belonging to
Basidiomycota — a group of fungi that produces their sexual spores (basidiospores) on club-shaped spore-producing structures (basidia) supported by macroscopic fruit bodies including toadstools, puffbals, polypores and other types — have been added to the list of known Danish species in the first atlas period (2009–2013) alone [
57]. In addition, several species that were considered extinct were re-discovered [
63]. In the second project period (2015–2022) several improved search and assistance functions have been developed that present features relating to the individual species and their identification [
63], making it much easier to include an understanding of endangered species in nature management and decision-making.
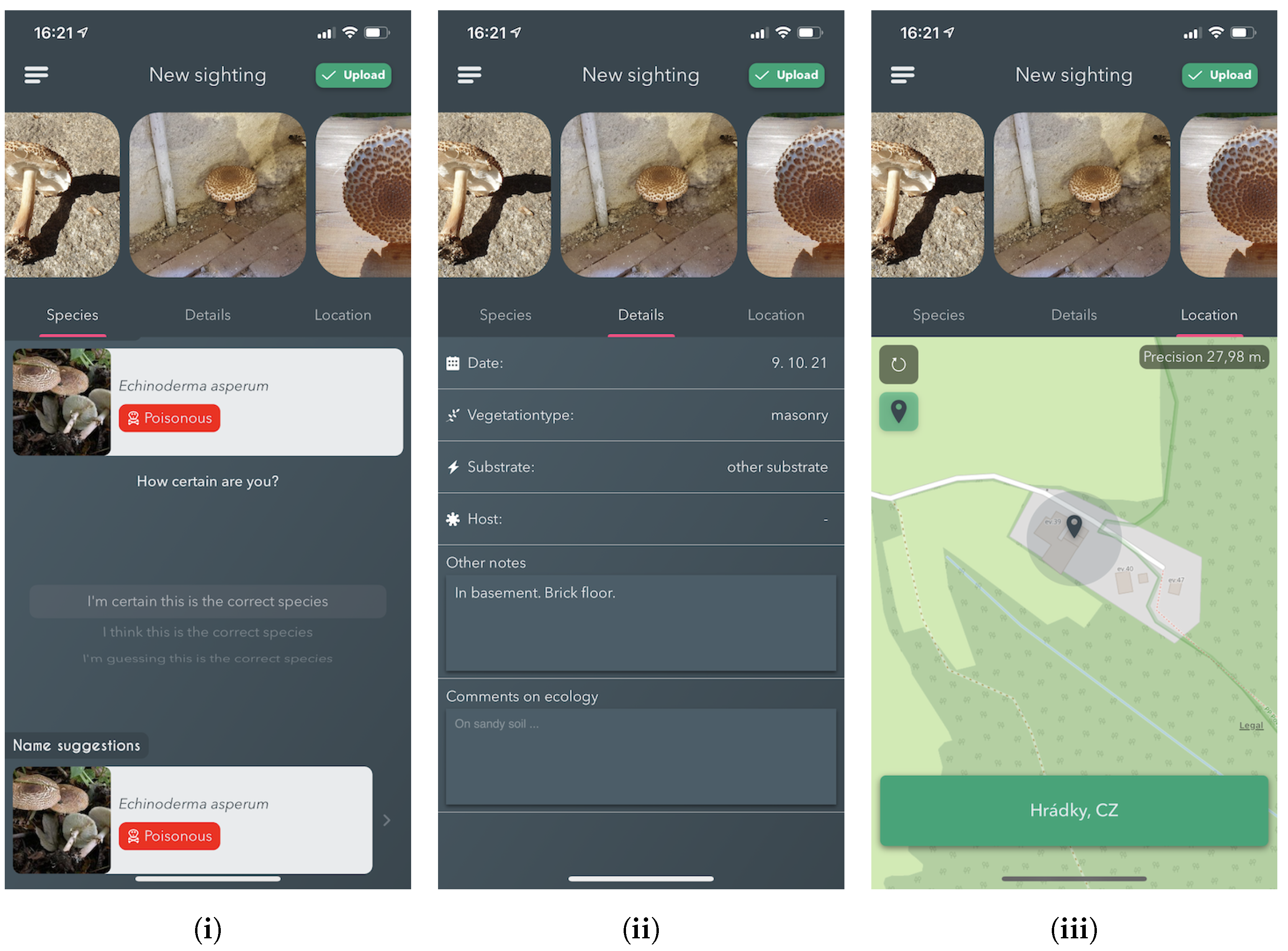
Annotation Process
Since 2017, the Atlas of Danish Fungi has had interactive labelling procedure for all submitted observations. When a user submits a fungal sighting (record) at species level, a “reliability score” (1–100) is calculated based on following factors:
Species rarity, that is, its relative frequency in the Atlas;
The geographical distribution of the species;
Phenology of the species, its seasonality;
User’s historical species-level proposal precision;
As above, within the proposal’s higher taxon rank.
Subsequently, other users may agree with the proposed species’ identity, increasing the identification score following the same principles, or proposing alternative identification for non-committal suggestions. Once the submission reaches a score of 80, the label (identification) is considered approved by community validation. Simultaneously, a small group of taxonomic experts (expert validators) monitor most of the observation on their own. Expert validators have the power to approve or reject species identifications regardless of the score in the interactive validation. Community-validated and expert-validated Svampeatlas records are published in the GBIF, weekly, since 2016. As of the end of October 2021, the data in GBIF included 955,392 occurrences with 504,165 images [
64]. Since 2019, the Atlas of Danish Fungi observation identification has been further streamlined thanks to an image recognition system [
29] — FungiVision.
3.2. The FGVCx Fungi Dataset
The FGVCx Fungi Classification Challenge provided an image dataset covering 1394 fungal species and is split into a training set with 85,578 images, a validation set with 4182 images, and a competition test set of 9758 images without publicly available labels. There is a substantial change of categorical priors between the training set and the validation set: The distribution of images per class is highly unbalanced in the training set, while the validation set distribution is uniform.
3.3. The Danish Fungi 2020 Dataset
The
Danish Fungi 2020 (DF20) dataset contains image observations from the Atlas of Danish Fungi belonging to species with more than 30 images. The data are observations collected before the end of 2020. Note that this includes more than 15 months of data collection using our automatic fungal identification service described later in
Section 5. The dataset consists of 295,938 images represent 1,604 species mainly from the
Fungi kingdom with a visually similar species. Unlike most computer vision datasets, DF20 include rich metadata acquired by citizen-scientists in the field while recording the observations that opens promising research direction in combining visual data with metadata like timestamp, location at multiple scales, substrate, habitat, taxonomy labels and camera device settings.
The DF20 datasets were randomly split — with respect to the class distribution — into the provided training and (public) test sets, where the training set contains of images of each species.
3.4. Test Observations from 2021
To independently compare models trained on the FGVCx Fungi dataset and the DF20, we used all validated observations submitted to the Atlas of Danish Fungi between 1 January 2021 and 31 October 2021. Only submissions that used the FungiVision system [
29] were used; we choose only the first image for each observation. With this approach, we ended up with a test set of 14,391 images belonging to 999 species. In the following text, we will denote this dataset as
DanishFungi 2021 or
DF21 in short.
6. Results
6.1. Machine Learning for Fungi Recognition in 2018
The FGVCx Fungi’18 competition test dataset on Kaggle was divided into two parts — public and private. Public results were calculated with approximately 70% of the test data, which were visible to all participants. The rest of the data were used for final competition evaluation to avoid bias towards the test images’ performance.
We chose our best performing system, that is, the ensemble of the six fine-tuned CNNs with 14 crops per test image and with predictions adjusted to new class priors, for the final submission to Kaggle. The accumulation of predictions was done by the mode from Top1 species per prediction as it had better preliminary scores on the public part of the FGVCx Fungi’18 test set.
Our submission to the challenge achieved the best scores in Top3 accuracy for both public and private leaderboards. The results of the top five teams are listed in
Table 5.
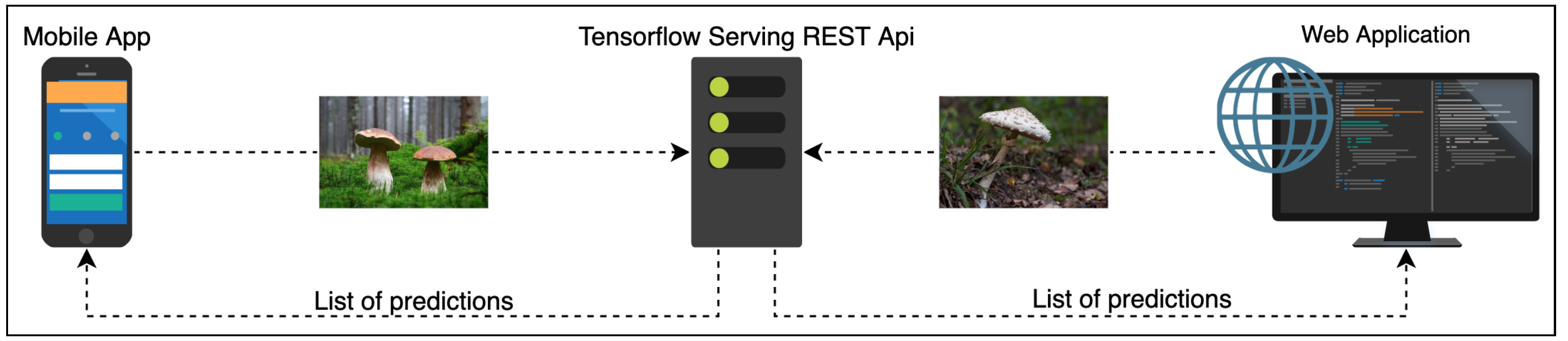
6.2. Online Classification Service
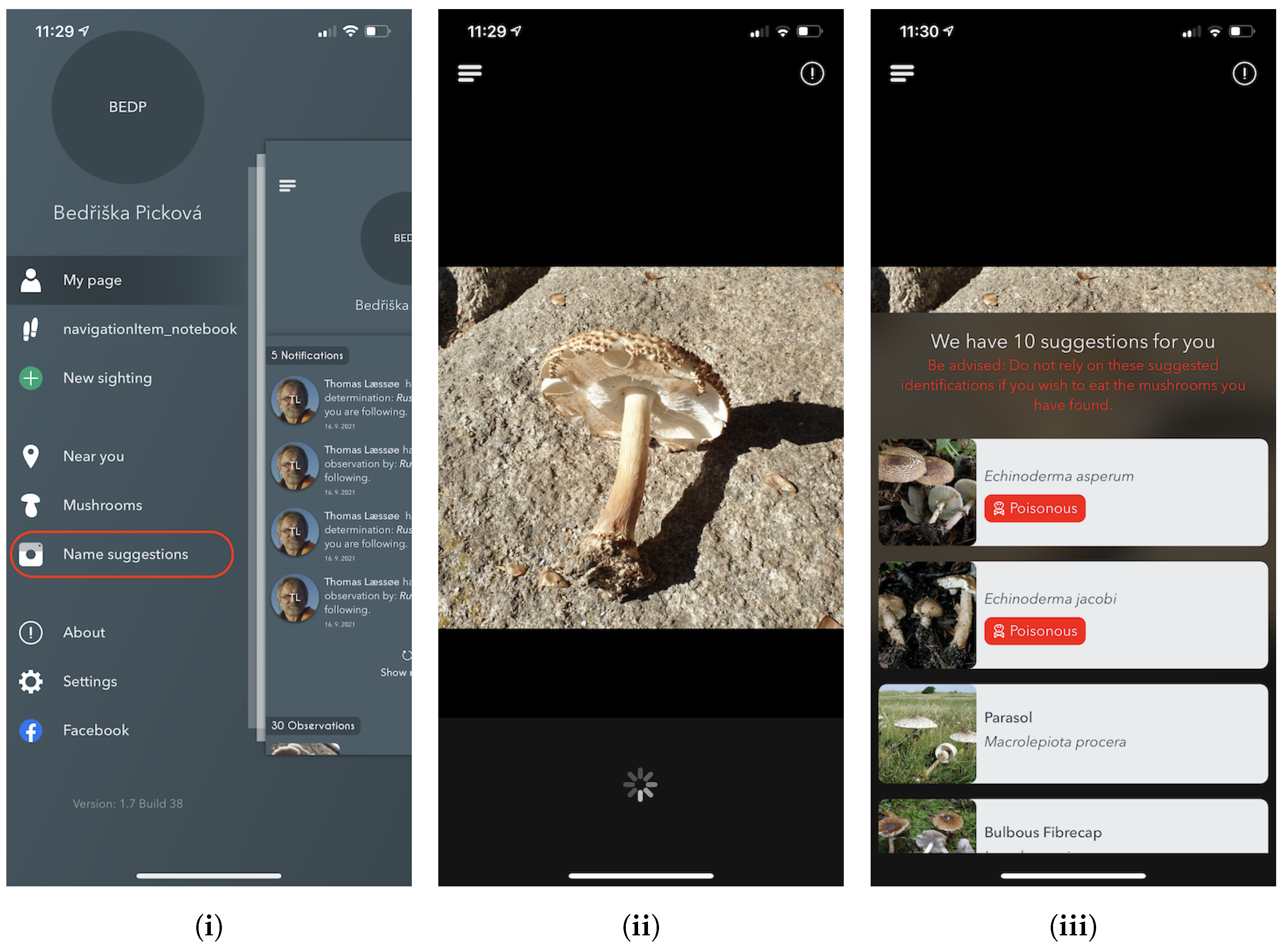
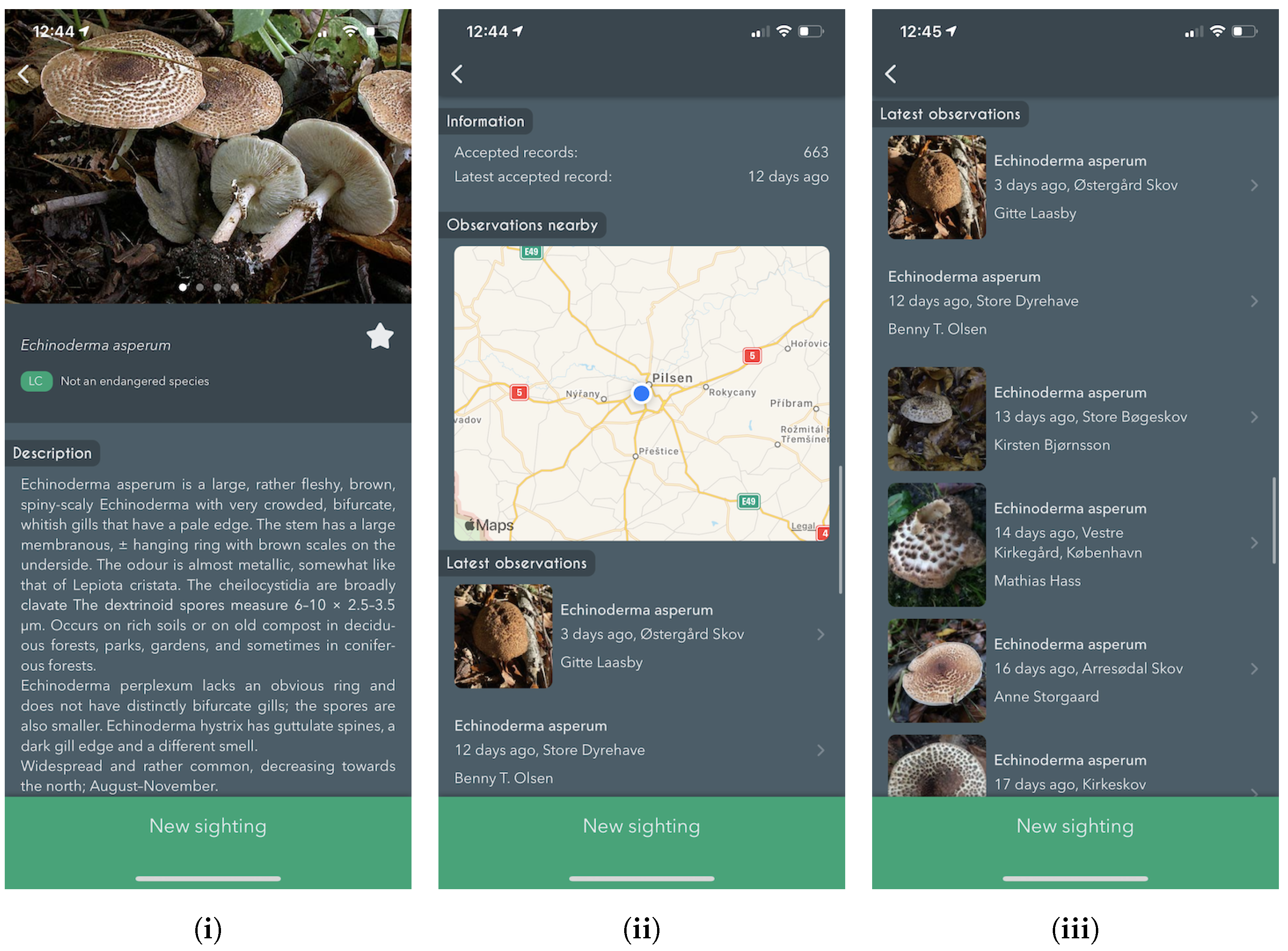
The experts behind the Atlas of Danish Fungi have been highly impressed by the performance of the system; in the application, the results of the system are referred to as an AI suggested species. A data evaluation on the DanishFungi 2021 — data that have been submitted for automatic recognition — has shown that only 7.18% were not approved by the community or expert validation, thus revealing a far better performance than most non-expert users in the system. Almost two thirds (69.28%) of the approved species identifications were based on the highest-ranking AI suggesting species ID. In contrast, another 12.28% were based on the second-highest-ranking AI suggested species ID and another 10.54% were based on the top 3–5 suggestions. In other words, the AI system achieved the Top1 accuracy of 69.28% and the Top1 accuracy of 92.82% in combination with citizen scientists.
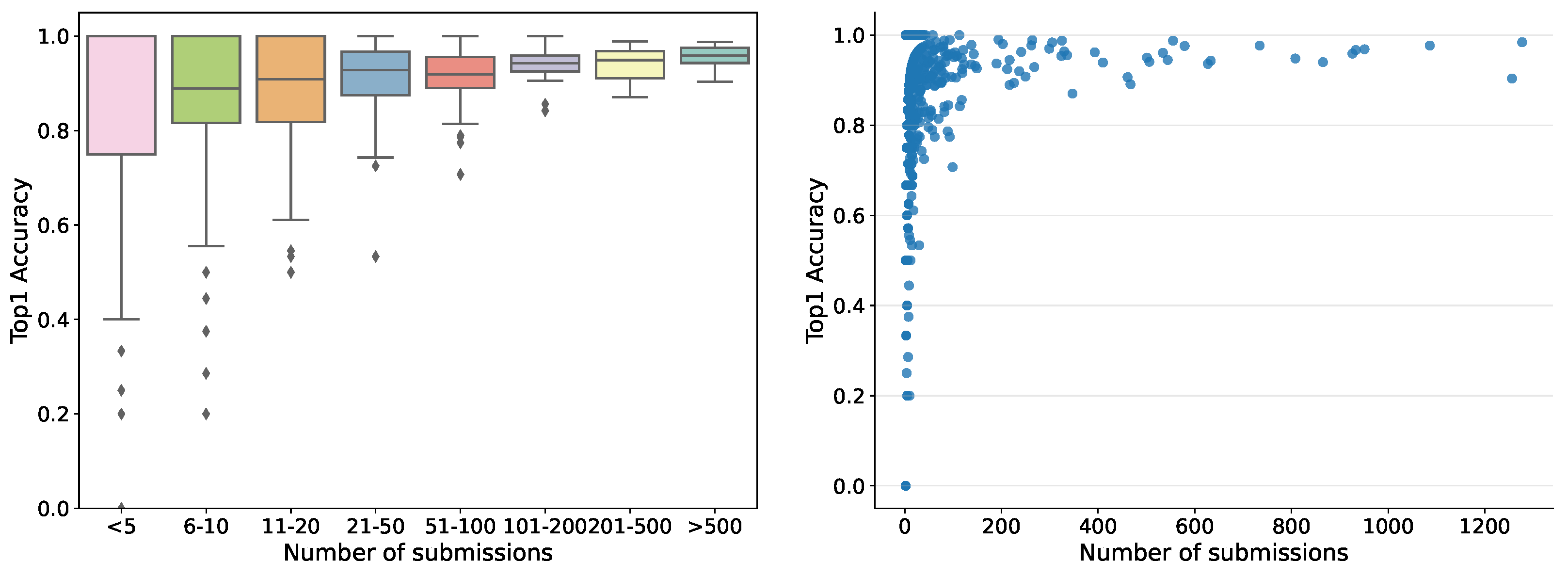
So far, the automatic recognition system has been tested by 1769 users — each submitting between one and 1277 records — who contributed 35,018 fungi sightings over the past 2 years. For users submitting more than ten records, the accuracy in terms of correct identifications guided by the system varied from 30% to 100%, pointing to quite considerable differences in how well different users have been able to identify the correct species using the system. Hence, the tool is not fully reliable but helps non-expert users to gain better identification skills. The accuracy was variable among the fungal morphogroups defined in the fungal atlas, varying from 24% to 100% for groups with more than ten records. The accuracy was tightly correlated with the obtained morphogroup user score based on the algorithms deployed in the Atlas of Danish Fungi to support community validation. Within the first month the server ran, more than 20,000 images were submitted for recognition. The dependence of human in the loop performance on the number of submissions, for example, recognition experience, is shown in
Figure 7.
6.3. Convolutional Neural Networks vs. Vision Transformers
In this section, we compare the performance of the well known CNN based models and ViT models in terms of Top1 and Top3 accuracy on the DF20 and the FGVCx Fungi’18 datasets and two different resolutions — 224 × 224 and 384 × 384.
Comparing well known CNN architectures on the DF20 dataset, we can see a similar behaviour to that on other datasets [
11,
17,
18]. The best performing model on the DF20 and input resolution of 384 × 384 was SE-ResNeXt-101 with a Top1 score of 78.72%. EfficientNetV2-L achieved a slightly lower accuracy of 77.83%. On a smaller input resolution (224 × 224), the best performing model was the EfficientNetV2-L, while achieving a better performance by 1.22% than SE-ResNeXt-101.
Comparing two ViT architectures — ViT-Base/16 and ViT-Large/16 — against the well-performing CNN models — EfficientNetV2-L and SE-ResNeXt-101 — on a DF20 dataset, we see a difference from the performance evaluation on ImageNet [
74,
75]. In our experiments, ViTs outperform state-of-the-art CNNs by a large margin in a 384 × 384 scenario. The best performing ViT model achieved an impressive Top1 accuracy of 80.45% while outperforming the SE-ResNeXt-101 by a significant margin of 1.73% on the images with 384 × 384 input size. In a 224 × 224 scenario, netiher CNNs nor ViT showed a superior performance. A wider performance comparison is shown in
Table 6.
6.4. Importance of the Metadata
Inspired by the common practice in mycology, we set up an experiment to show the importance of metadata for
Fungus species identification. Using the approach described in
Section 4.3, we improved performance in all measured metrics by a significant margin. We measured the performance improvement with all metadata types and their combinations. Overall, the habitat was most efficient in improving the performance. With the combination of habitat, substrate and month, we improved the ViT-Large/16 model’s performance on DF20 by 2.95% and 1.92% in Top1 and Top3, respectively, and the performance of the ViT-Base/16 model by 3.81% and 2.84% in Top1 and Top3. A detailed evaluation of the performance gain using different observation metadata and their combinations is shown in
Table 7.
6.5. Impact on Mycology–Atlas of Danish Fungi
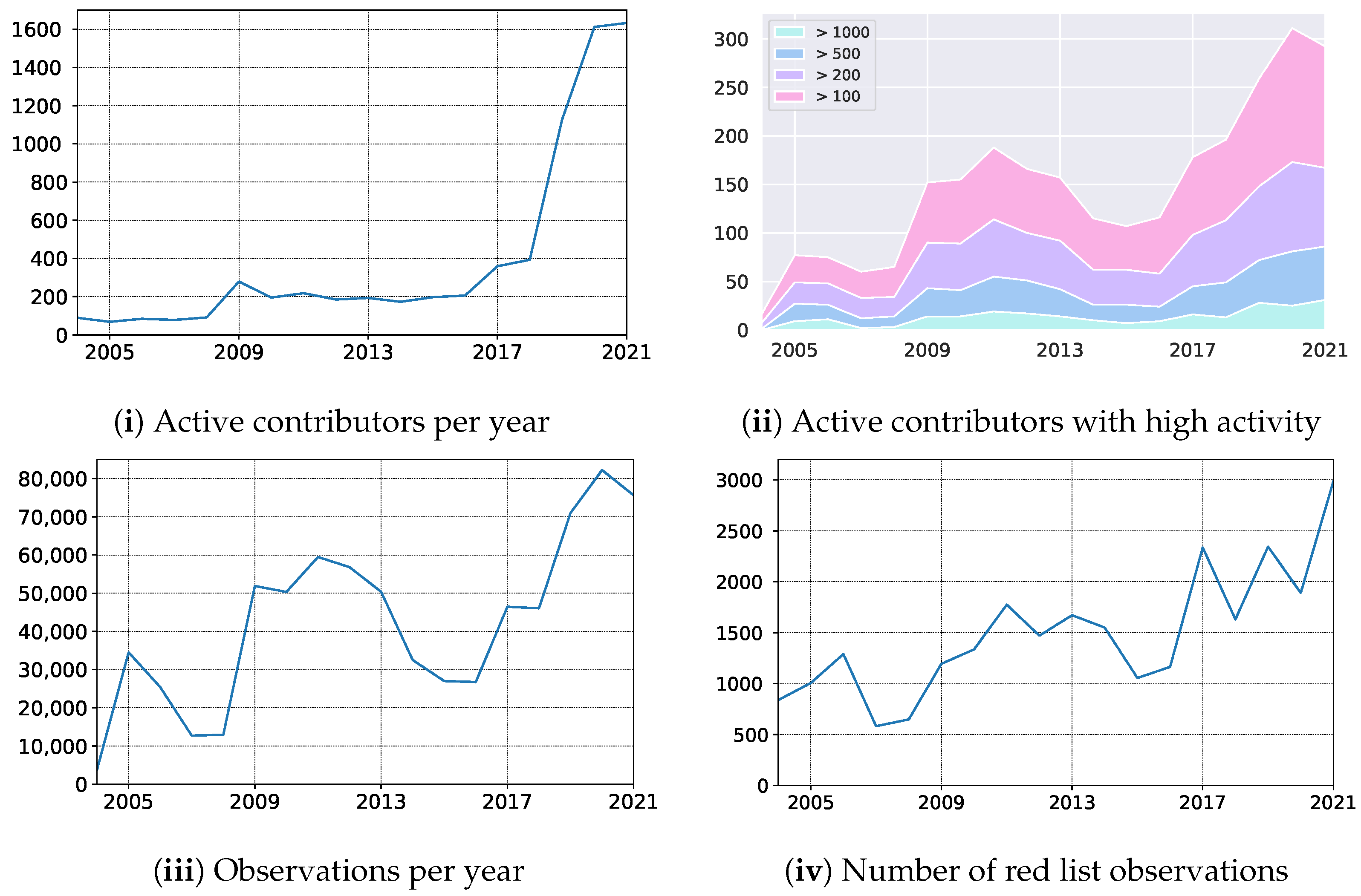
In October 2019, we launched the mobile application empowering Atlas of Danish Fungi with users with an image-based recognition tool for fungi species identification. The launch received good press coverage, including an appearance in the evening news on Danish National television — TV2. The launch led to an immediate increase in the user base, increasing the number of weekly contributors from 150 to 400. Besides, the number of yearly contributors quadrupled from 2018 to 2020, resulting in a 79% increase in submitted records (
Figure 8 (i) — the number of active contributors, (iii) yearly records). In parallel, the average number of records submitted per contributor dropped by 49% (from 117 to 51), indicating a substantial increase in less dedicated fungal recorders but with much broader geographical coverage. However, even the most active user groups with more than 100, 200, 500, and 1000 records a year also increased their size by 55%, 44%, 66%, and 92%, respectively (
Figure 8 (ii) — number of contributors with a various number of min records).
Over a longer period, including the first very active recording period from 2009–2013, the shift from a relatively small but dedicated user group to a much larger group including more and less active contributors is even more evident.
Table 8 shows the comparison of our original FungiVision system and a newly proposed system, comprising a Vision Transformer trained on DF20 and utilizing the available metadata. With the new system, the Top-1 error in was reduced by 48.2%.
Building on the terminology suggested by Ceccaroni et al. [
87], the application of AI in the Atlas of Danish Fungi has mainly contributed to influencing human behaviour, that is, attracting many new contributors, who earlier tended to find fungi too challenging to identify. Anyway, based on our yearly evaluation — see
Table 9 — both the higher number of users and submissions containing more challenging species for identification did not affect the overall user performance. So far, we have not explored the educational and social benefits for new contributors in detail, but from casual oral and written responses from new contributors, the effects seem to be considerable. The large influx of new contributors has been a challenge for already associated expert users and professional experts associated with the project. The system is designed to be interactive and involving, requiring new users to be trained to submit high-quality records and contribute actively to the validation process. The development of automatic response options, for example, addressing common issues related to the poor quality of submitted photos or inadequate meta-data, could solve some of these issues in the future, potentially using AI to replace the time-consuming human evaluation of records.
6.6. Impact on AI–Fungi Recognition in 2021
We introduce a novel fine-grained dataset and benchmark based on the symbiotic relationship between Machine Learning and Mycology, the Danish Fungi 2020 (DF20). The dataset, constructed from observations submitted to the Atlas of Danish Fungi, is unique in its taxonomy-accurate class labels, small number of errors, highly unbalanced long-tailed class distribution, rich observation metadata, and well-defined class hierarchy. DF20 has zero overlap with ImageNet, allowing unbiased comparison of models fine-tuned from publicly available ImageNet checkpoints. The proposed evaluation protocol enables testing the ability to improve classification using metadata — for example, location, habitat, and substrate, facilitates classifier calibration testing and finally allows us to study the impact of the device settings on the classification performance.
Experimental comparison of selected CNN and ViT architectures shows that DF20 presents a challenging task. Interestingly, ViT achieves results superior to CNN baselines with 80.45% accuracy, reducing the CNN error by 9%.
A simple procedure for including metadata into the decision process improves the classification accuracy by more than 2.95 and 0.65 percentage points, reducing the error rate by 15% and 6.5% on Danish Fungi 2020 and Danish Fungi 2021, respectively.
In
Table 10, we present the comparison on the FGVCx Fungi’18 test set between our novel approach where we utilize the ViT architecture and metadata, and the single model developed back in 2018. We can see a significant increase in performance by 6.59% in terms of Top1 Accuracy. Evaluated via Kaggle using 80% central crop.
7. Conclusions
A machine learning system for automatic fungi recognition, a winner of a computer vision Kaggle challenge, was deployed as an online recognition service to help a community of citizen scientists identify the species of an observed specimen.
The development of the machine learning system for the Kaggle competition in
Section 4.1 showed the effect of calibrating outputs to new a priori probabilities, test-time data augmentation and ensembles: together, these “tricks” increased the recognition accuracy by almost 12% and helped us to score 1st in the FGVCx Fungi Classification competition hosted on Kaggle, achieving a Top3 Accuracy of 73%. The availability of the identification service helped to increase the activity and contributions of citizen scientists to the Atlas of Danish Fungi. Integration of the image recognition system into the Atlas of Danish Fungi has made community-based fungi observation identification easier: 92.82% of submissions labeled by users with the help of the FungiVision system were identified correctly.
The collected data allowed the creation of a novel fine-grained classification dataset — the Danish Fungi 2020 (DF20) — which has zero overlap with ImageNet, allowing unbiased comparison of models fine-tuned from publicly available ImageNet checkpoints. With the precise annotation and rich metadata coming with the DF20 dataset, we would like to encourage research in other areas of computer vision and machine learning, beyond fine-grained visual categorization. The datasets may serve as a benchmark for classifier calibration, loss functions, validation metrics, taxonomy and hierarchical learning, device dependency or time series based species prediction. For example, the standard loss function focusing on recognition accuracy ignores the practically important cost of predicting a species with high toxicity. The quantitative and qualitative analysis of CNNs and ViTs showed superior performance of the ViT in fine-grained classification. We present the baselines for processing the habitat, substrate and time (month) metadata. We show that — even with the simple method from
Section 4.3 — utilizing the metadata increases the classification performance significantly. A new Vision Transformer architecture, trained on DF20 and exploiting available metadata, with a recognition error 46.75% lower than that of the current system.
Cross science efforts, such as the collaboration described here, can develop tools for citizen-scientists that improve their skills and the quality of the data they generate. Along with data generated by DNA sequencing, this may help by lowering the taxonomic bias in the biodiversity information data available in the future. By providing a stream of labeled data in one direction and an accuracy increase in the other, the collaboration creates a virtuous cycle, helping both communities.