A Feasibility Study on Deep Learning Based Brain Tumor Segmentation Using 2D Ellipse Box Areas
Abstract
:1. Introduction
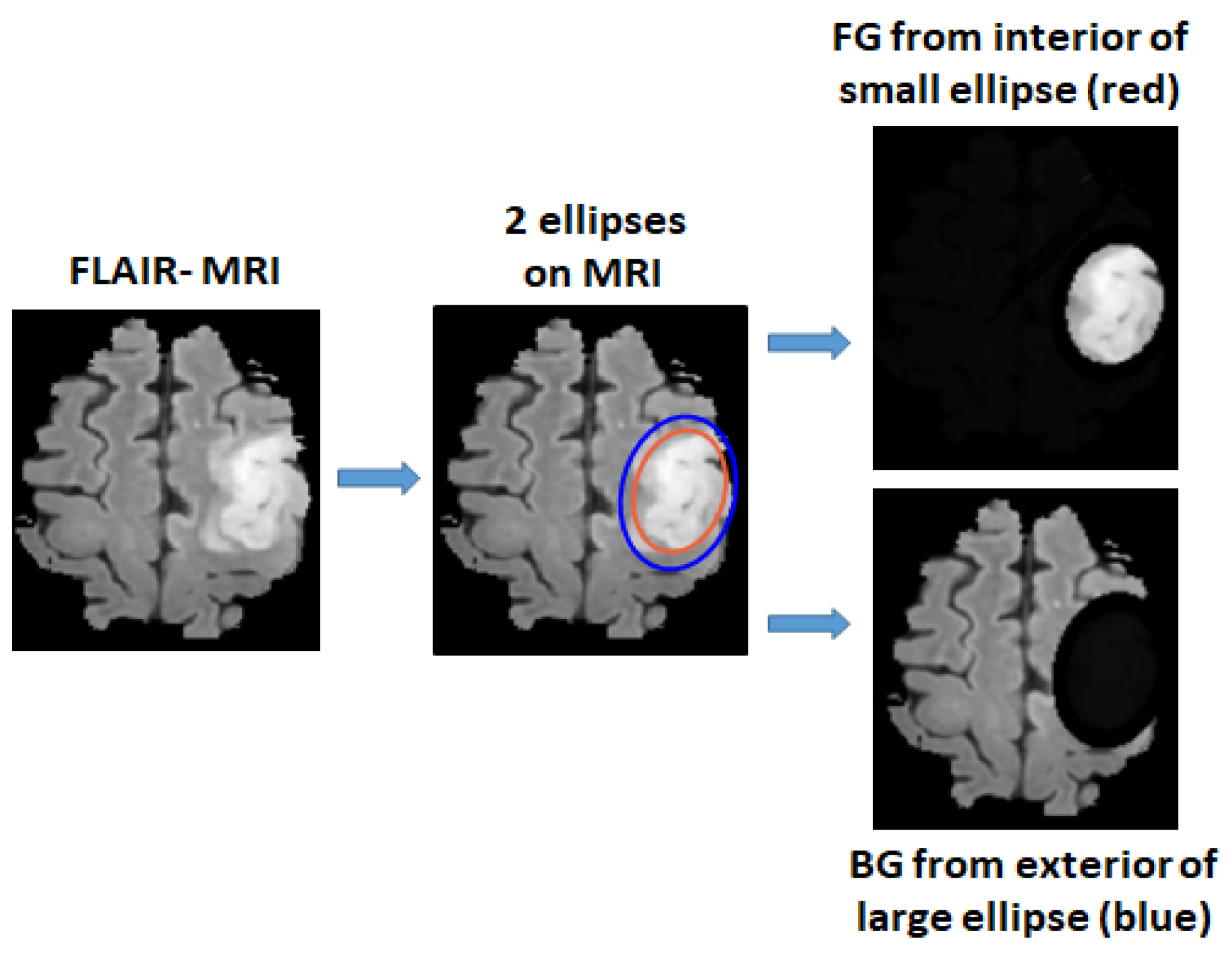
- We study the feasibility of the use of 2D ellipse box areas for training the deep network for brain tumor (glioma) segmentation plus a small number of annotated tumors.
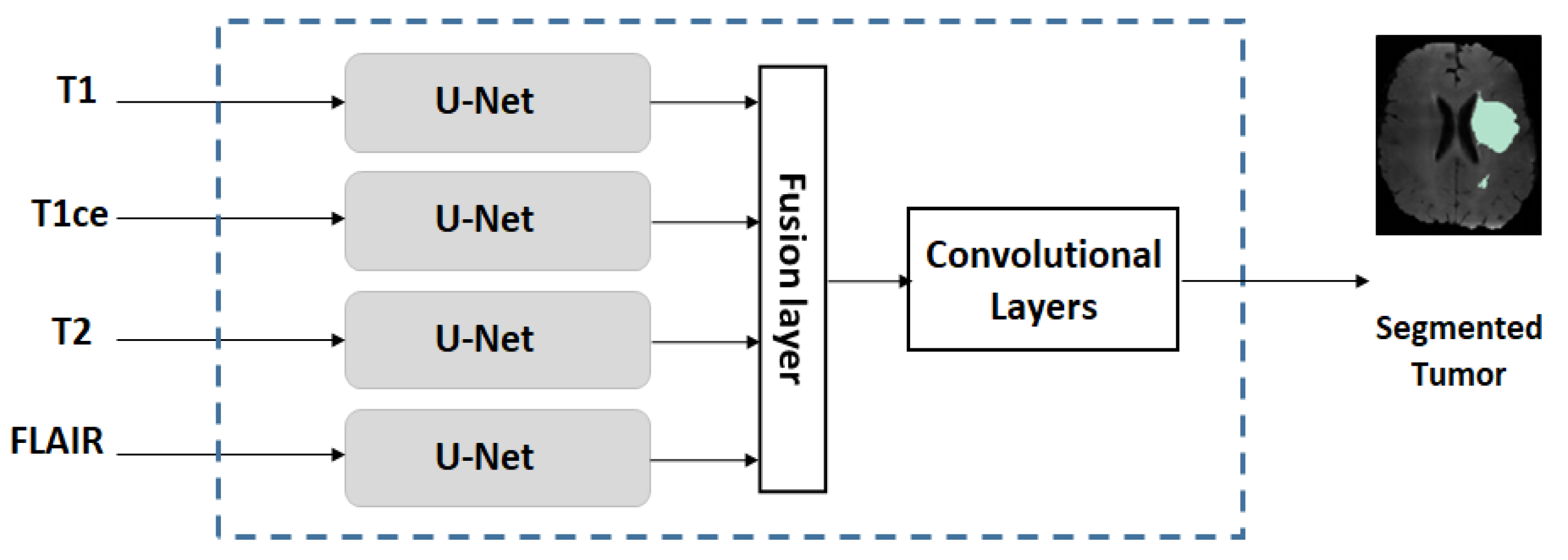
- We use a multi-stream U-Net for our experiments, which is an extended version of the conventional U-Net.
- We conduct studies on two scenarios: (a) if the training dataset is large/moderate, learning is conducted by pre-training on a large amount of FG and BG ellipse areas followed by refined-training on a small number of annotated tumor patients (<20); and (b) if the training dataset is small, learning is conducted in a fashion similar to the idea of transfer learning.
- We evaluate the performance of the proposed approach and compare the performance with the same network trained entirely by using annotated MRIs.
2. Proposed Method
2.1. Defining Foreground and Background Ellipse Box Areas Using Ellipses
2.2. Multi-Stream U-Net Used for Our Experiments
2.3. Training Strategies on Datasets with Different Sizes
2.3.1. Training on Dataset with Large/Moderate Size
2.3.2. Training on a Small Dataset
2.4. Other Issues
2.4.1. Strict Patient-Separated Splitting of Training/Validation/Testing Sets
2.4.2. Criteria for Performance Evaluation
Tumor Accuracy
Tumor Dice Score and Jaccard Index
3. Results and Performance Evaluation
3.1. Datasets, Setup, Pre-Processing
3.1.1. Datasets
3.1.2. Setup
3.1.3. Pre-Processing
3.2. Results, Comparison and Discussion
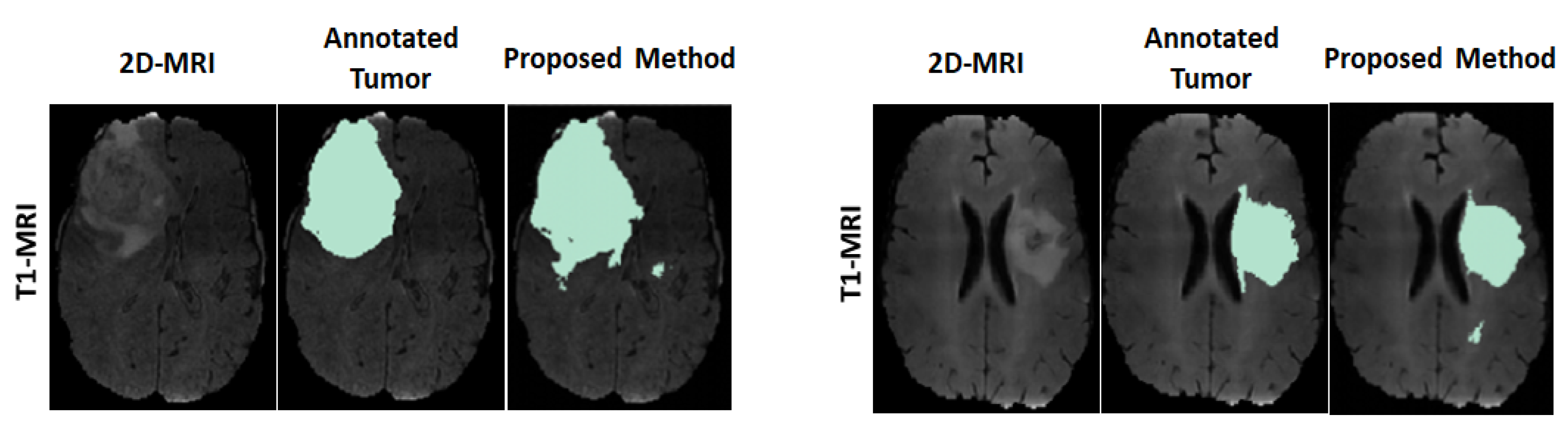
3.2.1. Results
3.2.2. Comparison
3.2.3. Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BG | Background |
| BN | Batch normalization |
| CNN | Convolutional Netural Network |
| FG | Foreground |
| FLAIR | Fluid-Attenuated Inversion Recovery |
| FN | False negative |
| GT | Ground truth |
| HGG | High Grade Glioma |
| LGG | Low Grade Glioma |
| MICCAI | Medical Image Computing Computer Assisted Intervention Society |
| MNI | Montreal Neurological Institute |
| MRI | Magentic Resonance Image |
| ReLU | Rectifier Linear Unit |
| T1 | T1 weighted |
| T2 | T2 weighted |
| T1ce | T1 weighted with Contrast Enhanced |
References
- Bø, H.K.; Solheim, O.; Jakola, A.S.; Kvistad, K.A.; Reinertsen, I.; Berntsen, E.M. Intra-rater variability in low-grade glioma segmentation. J. Neuro-Oncol. 2017, 131, 393–402. [Google Scholar] [CrossRef] [PubMed]
- White, D.R.; Houston, A.S.; Sampson, W.F.; Wilkins, G.P. Intra-and interoperator variations in region-of-interest drawing and their effect on the measurement of glomerular filtration rates. Clin. Nucl. Med. 1999, 24, 177–181. [Google Scholar] [CrossRef] [PubMed]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Munich, Germany, 5–9 October 2015; Springer: Berlin/Heidelberg, Germany, 2015; pp. 234–241. [Google Scholar]
- Diakogiannis, F.I.; Waldner, F.; Caccetta, P.; Wu, C. ResUNet-a: A deep learning framework for semantic segmentation of remotely sensed data. ISPRS J. Photogramm. Remote Sens. 2020, 162, 94–114. [Google Scholar] [CrossRef] [Green Version]
- Huang, H.; Lin, L.; Tong, R.; Hu, H.; Zhang, Q.; Iwamoto, Y.; Han, X.; Chen, Y.W.; Wu, J. Unet 3+: A full-scale connected unet for medical image segmentation. In Proceedings of the ICASSP 2020—2020 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), Barcelona, Spain, 4–8 May 2020; pp. 1055–1059. [Google Scholar]
- Wang, F.; Jiang, R.; Zheng, L.; Meng, C.; Biswal, B. 3d u-net based brain tumor segmentation and survival days prediction. In Proceedings of the International MICCAI Brainlesion Workshop, Shenzhen, China, 17 October 2019; Springer: Berlin/Heidelberg, Germany, 2019; pp. 131–141. [Google Scholar]
- Kim, S.; Luna, M.; Chikontwe, P.; Park, S.H. Two-step U-Nets for brain tumor segmentation and random forest with radiomics for survival time prediction. In Proceedings of the International MICCAI Brainlesion Workshop, Shenzhen, China, 17 October 2019; Springer: Berlin/Heidelberg, Germany, 2019; pp. 200–209. [Google Scholar]
- Shi, W.; Pang, E.; Wu, Q.; Lin, F. Brain tumor segmentation using dense channels 2D U-Net and multiple feature extraction network. In Proceedings of the International MICCAI Brainlesion Workshop, Shenzhen, China, 17 October 2019; Springer: Berlin/Heidelberg, Germany, 2019; pp. 273–283. [Google Scholar]
- Dios, E.d.; Ali, M.B.; Gu, I.Y.H.; Vecchio, T.G.; Ge, C.; Jakola, A.S. Introduction to Deep Learning in Clinical Neuroscience. In Machine Learning in Clinical Neuroscience; Springer: Berlin/Heidelberg, Germany, 2022; pp. 79–89. [Google Scholar]
- Havaei, M.; Davy, A.; Warde-Farley, D.; Biard, A.; Courville, A.; Bengio, Y.; Pal, C.; Jodoin, P.M.; Larochelle, H. Brain tumor segmentation with deep neural networks. Med. Image Anal. 2017, 35, 18–31. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pereira, S.; Pinto, A.; Alves, V.; Silva, C.A. Brain tumor segmentation using convolutional neural networks in MRI images. IEEE Trans. Med. Imaging 2016, 35, 1240–1251. [Google Scholar] [CrossRef]
- Sun, Y.; Wang, C. A computation-efficient CNN system for high-quality brain tumor segmentation. Biomed. Signal Process. Control 2022, 74, 103475. [Google Scholar] [CrossRef]
- Das, S.; Swain, M.K.; Nayak, G.; Saxena, S. Brain tumor segmentation from 3D MRI slices using cascading convolutional neural network. In Advances in Electronics, Communication and Computing; Springer: Berlin/Heidelberg, Germany, 2021; pp. 119–126. [Google Scholar]
- Shan, C.; Li, Q.; Wang, C.H. Brain Tumor Segmentation using Automatic 3D Multi-channel Feature Selection Convolutional Neural Network. J. Imaging Sci. Technol. 2022, 1–9. [Google Scholar] [CrossRef]
- Ranjbarzadeh, R.; Bagherian Kasgari, A.; Jafarzadeh Ghoushchi, S.; Anari, S.; Naseri, M.; Bendechache, M. Brain tumor segmentation based on deep learning and an attention mechanism using MRI multi-modalities brain images. Sci. Rep. 2021, 11, 10930. [Google Scholar] [CrossRef]
- Dai, J.; He, K.; Sun, J. Boxsup: Exploiting bounding boxes to supervise convolutional networks for semantic segmentation. In Proceedings of the Proceedings of the IEEE International Conference on Computer Vision, Santiago, Chile, 7–13 December 2015; pp. 1635–1643.
- de Carvalho, O.L.F.; de Carvalho Júnior, O.A.; de Albuquerque, A.O.; Santana, N.C.; Guimarães, R.F.; Gomes, R.A.T.; Borges, D.L. Bounding Box-Free Instance Segmentation Using Semi-Supervised Iterative Learning for Vehicle Detection. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2022, 15, 3403–3420. [Google Scholar] [CrossRef]
- Zhan, D.; Liang, D.; Jin, H.; Wu, X. MBBOS-GCN: Minimum bounding box over-segmentation—Graph convolution 3D point cloud deep learning model. J. Appl. Remote Sens. 2022, 16, 016502. [Google Scholar] [CrossRef]
- Zhang, D.; Song, K.; Xu, J.; Dong, H.; Yan, Y. An image-level weakly supervised segmentation method for No-service rail surface defect with size prior. Mech. Syst. Signal Process. 2022, 165, 108334. [Google Scholar] [CrossRef]
- Zhou, X.; Girdhar, R.; Joulin, A.; Krähenbühl, P.; Misra, I. Detecting twenty-thousand classes using image-level supervision. arXiv 2022, arXiv:2201.02605. [Google Scholar]
- Cheng, B.; Parkhi, O.; Kirillov, A. Pointly-supervised instance segmentation. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, New Orleans, LA, USA, 19–24 June 2022; pp. 2617–2626. [Google Scholar]
- Khan, Z.H.; Gu, I.Y.H. Online domain-shift learning and object tracking based on nonlinear dynamic models and particle filters on Riemannian manifolds. Comput. Vis. Image Underst. 2014, 125, 97–114. [Google Scholar] [CrossRef]
- Yun, Y.; Gu, I.Y.H. Human fall detection in videos via boosting and fusing statistical features of appearance, shape and motion dynamics on Riemannian manifolds with applications to assisted living. Comput. Vis. Image Underst. 2016, 148, 111–122. [Google Scholar] [CrossRef]
- Zhang, Y.; Liao, Q.; Jiao, R.; Zhang, J. Uncertainty-Guided Mutual Consistency Learning for Semi-Supervised Medical Image Segmentation. arXiv 2021, arXiv:2112.02508. [Google Scholar] [CrossRef]
- Luo, X.; Chen, J.; Song, T.; Wang, G. Semi-supervised medical image segmentation through dual-task consistency. In Proceedings of the AAAI Conference on Artificial Intelligence, Virtual. 2–9 February 2021; Volume 35, pp. 8801–8809. [Google Scholar]
- Ali, M.B.; Gu, I.Y.H.; Berger, M.S.; Pallud, J.; Southwell, D.; Widhalm, G.; Roux, A.; Vecchio, T.G.; Jakola, A.S. Domain Mapping and Deep Learning from Multiple MRI Clinical Datasets for Prediction of Molecular Subtypes in Low Grade Gliomas. Brain Sci. 2020, 10, 463. [Google Scholar] [CrossRef]
- Ali, M.B.; Gu, I.Y.H.; Lidemar, A.; Berger, M.S.; Widhalm, G.; Jakola, A.S. Prediction of glioma-subtypes: Comparison of performance on a DL classifier using bounding box areas versus annotated tumors. BMC Biomed. Eng. 2022, 4, 4. [Google Scholar] [CrossRef]
- Pavlov, S.; Artemov, A.; Sharaev, M.; Bernstein, A.; Burnaev, E. Weakly supervised fine tuning approach for brain tumor segmentation problem. In Proceedings of the 2019 18th IEEE International Conference On Machine Learning And Applications (ICMLA), Boca Raton, FL, USA, 16–19 December 2019; pp. 1600–1605. [Google Scholar]
- Zhu, X.; Chen, J.; Zeng, X.; Liang, J.; Li, C.; Liu, S.; Behpour, S.; Xu, M. Weakly supervised 3d semantic segmentation using cross-image consensus and inter-voxel affinity relations. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, BC, Canada, 11–17 October 2021; pp. 2814–2824. [Google Scholar] [CrossRef]
- Xu, Y.; Gong, M.; Chen, J.; Chen, Z.; Batmanghelich, K. 3d-boxsup: Positive-unlabeled learning of brain tumor segmentation networks from 3d bounding boxes. Front. Neurosci. 2020, 14, 350. [Google Scholar] [CrossRef]
- Menze, B.H.; Jakab, A.; Bauer, S.; Kalpathy-Cramer, J.; Farahani, K.; Kirby, J.; Burren, Y.; Porz, N.; Slotboom, J.; Wiest, R.; et al. The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans. Med. Imaging 2014, 34, 1993–2024. [Google Scholar] [CrossRef]
- Bakas, S.; Akbari, H.; Sotiras, A.; Bilello, M.; Rozycki, M.; Kirby, J.S.; Freymann, J.B.; Farahani, K.; Davatzikos, C. Advancing the cancer genome atlas glioma MRI collections with expert segmentation labels and radiomic features. Sci. Data 2017, 4, 170117. [Google Scholar] [CrossRef] [Green Version]
- Bakas, S.; Reyes, M.; Jakab, A.; Bauer, S.; Rempfler, M.; Crimi, A.; Shinohara, R.T.; Berger, C.; Ha, S.M.; Rozycki, M.; et al. Identifying the best machine learning algorithms for brain tumor segmentation, progression assessment, and overall survival prediction in the BRATS challenge. arXiv 2018, arXiv:1811.02629. [Google Scholar]
- Jenkinson, M.; Beckmann, C.F.; Behrens, T.E.; Woolrich, M.W.; Smith, S.M. Fsl. Neuroimage 2012, 62, 782–790. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Avants, B.B.; Tustison, N.J.; Song, G.; Cook, P.A.; Klein, A.; Gee, J.C. A reproducible evaluation of ANTs similarity metric performance in brain image registration. Neuroimage 2011, 54, 2033–2044. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Block No. | No. of Units |
|---|---|
| Downstream-path | Conv2D, stride, BN, Max-pooling |
| 1 | [3 × 3, 32, ReLU] × 2, 2, BN, 2 × 2 |
| 2 | [3 × 3, 64, ReLU] × 2, 2, BN, 2 × 2 |
| 3 | [3 × 3, 128, ReLU] × 2, 2, BN, 2 × 2 |
| 4 | [3 × 3, 256, ReLU] × 2, 2, BN, 2 × 2 |
| 5 | [3 × 3, 512, ReLU] × 2, 2, BN, - |
| Upstream-path | Conv2DTranspose, concat, BN, Conv2D |
| 6 | [2 × 2, 128], concat, -, [3 × 3, 256, ReLU] × 2 |
| 7 | [2 × 2, 128], concat, BN, [3 × 3, 128, ReLU] × 2 |
| 8 | [2 × 2, 128], concat, BN, [3 × 3, 64, ReLU] × 2 |
| 9 | [2 × 2, 128], concat, BN, [3 × 3, 32, ReLU] × 2 |
| Dataset | T1/T1ce/T2/FLAIR | #2D | #2D */3D | #2D */3D | #2D */3D |
|---|---|---|---|---|---|
| Slices | (Training) | (Validation) | (Testing) | ||
| MICCAI | 285/285/285/285 | 9 | 1539/171 | 513/57 | 513/57 |
| US | 0/75/0/75 | 18 | 270/15 | - | 1080/60 |
| (a) MICCAI dataset | ||
| Predicted\True | Tumor () | Non-tumor () |
| Tumor | 83.88 (±0.08) | 1.54 (±0.09) |
| Non-tumor | 16.12 (±0.08) | 98.46 (±0.09) |
| (b) US dataset | ||
| Predicted\True | Tumor () | Non-tumor () |
| Tumor | 88.47 (±0.34) | 0.42 (±0.27) |
| Non-tumor | 11.53 (±0.34) | 99.58 (±0.27) |
| (a) | |||
|---|---|---|---|
| Dataset | Tumor Accuracy ()% | Dice Score () | Jaccard Index () |
| MICCAI | 83.88 (±0.08) | 0.8407 (±0.0006) | 0.7233 (±0.0028) |
| US | 88.47 (±0.34) | 0.9104 (±0.0021) | 0.8355 (±0.0029) |
| (b) | |||
| Dataset | Sensitivity ()% | Specificity ()% | False Positive ()% |
| MICCAI | 83.88 (±0.08) | 98.46(±0.09) | 1.54 (±0.09) |
| US | 88.47 (±0.34) | 99.58 (±0.27) | 0.42 (±0.27) |
| Datasets | Method | Tumor Accuracy % | Dice Score |
|---|---|---|---|
| MICCAI | Proposed | 83.88 (±0.08) | 0.8407 (±0.0006) |
| Conventional | 92.66 (±0.15) | 0.9001 (±0.0018) | |
| Degradation | |||
| US | Proposed | 88.47 () | 0.9104 () |
| Conventional | 91.08 () | 0.9263 () | |
| Degradation |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ali, M.B.; Bai, X.; Gu, I.Y.-H.; Berger, M.S.; Jakola, A.S. A Feasibility Study on Deep Learning Based Brain Tumor Segmentation Using 2D Ellipse Box Areas. Sensors 2022, 22, 5292. https://doi.org/10.3390/s22145292
Ali MB, Bai X, Gu IY-H, Berger MS, Jakola AS. A Feasibility Study on Deep Learning Based Brain Tumor Segmentation Using 2D Ellipse Box Areas. Sensors. 2022; 22(14):5292. https://doi.org/10.3390/s22145292
Chicago/Turabian StyleAli, Muhaddisa Barat, Xiaohan Bai, Irene Yu-Hua Gu, Mitchel S. Berger, and Asgeir Store Jakola. 2022. "A Feasibility Study on Deep Learning Based Brain Tumor Segmentation Using 2D Ellipse Box Areas" Sensors 22, no. 14: 5292. https://doi.org/10.3390/s22145292
APA StyleAli, M. B., Bai, X., Gu, I. Y.-H., Berger, M. S., & Jakola, A. S. (2022). A Feasibility Study on Deep Learning Based Brain Tumor Segmentation Using 2D Ellipse Box Areas. Sensors, 22(14), 5292. https://doi.org/10.3390/s22145292







