Over 1000 nm Near-Infrared Multispectral Imaging System for Laparoscopic In Vivo Imaging
Abstract
1. Introduction
2. Materials and Methods
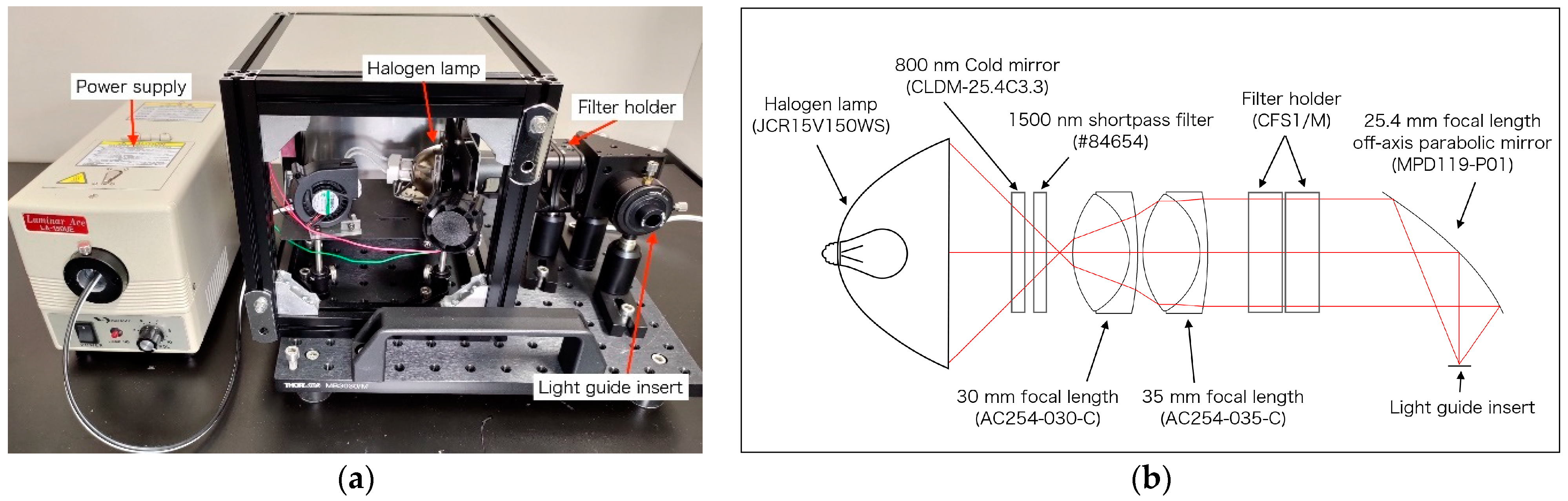
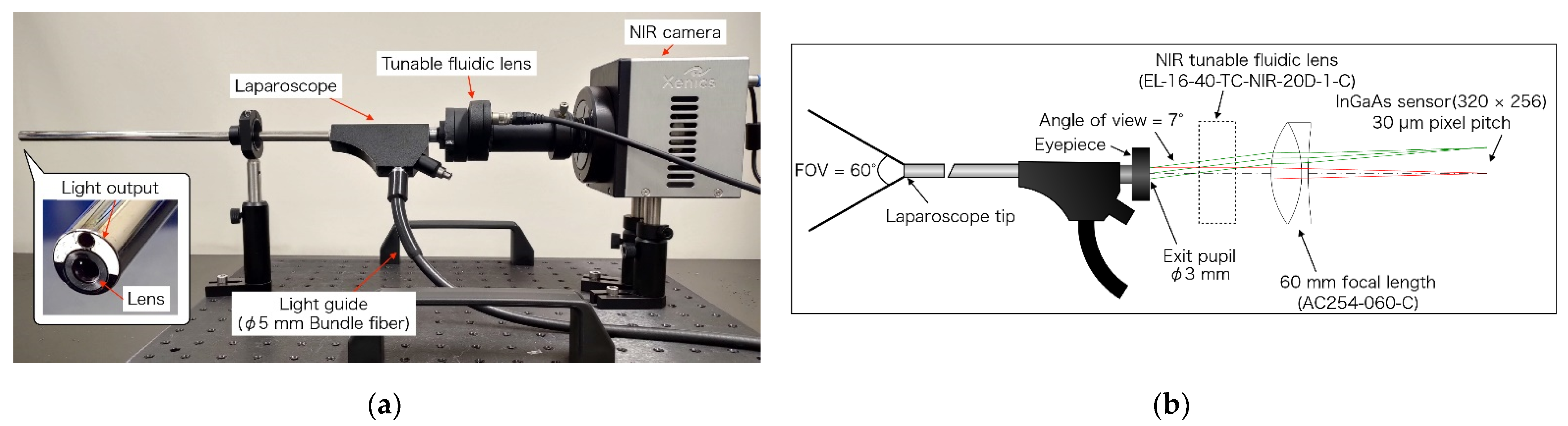
2.1. Light Source and Laparoscope for OTN-NIR Multispectral Imaging
2.2. Properties of the Laparoscope and Light Source
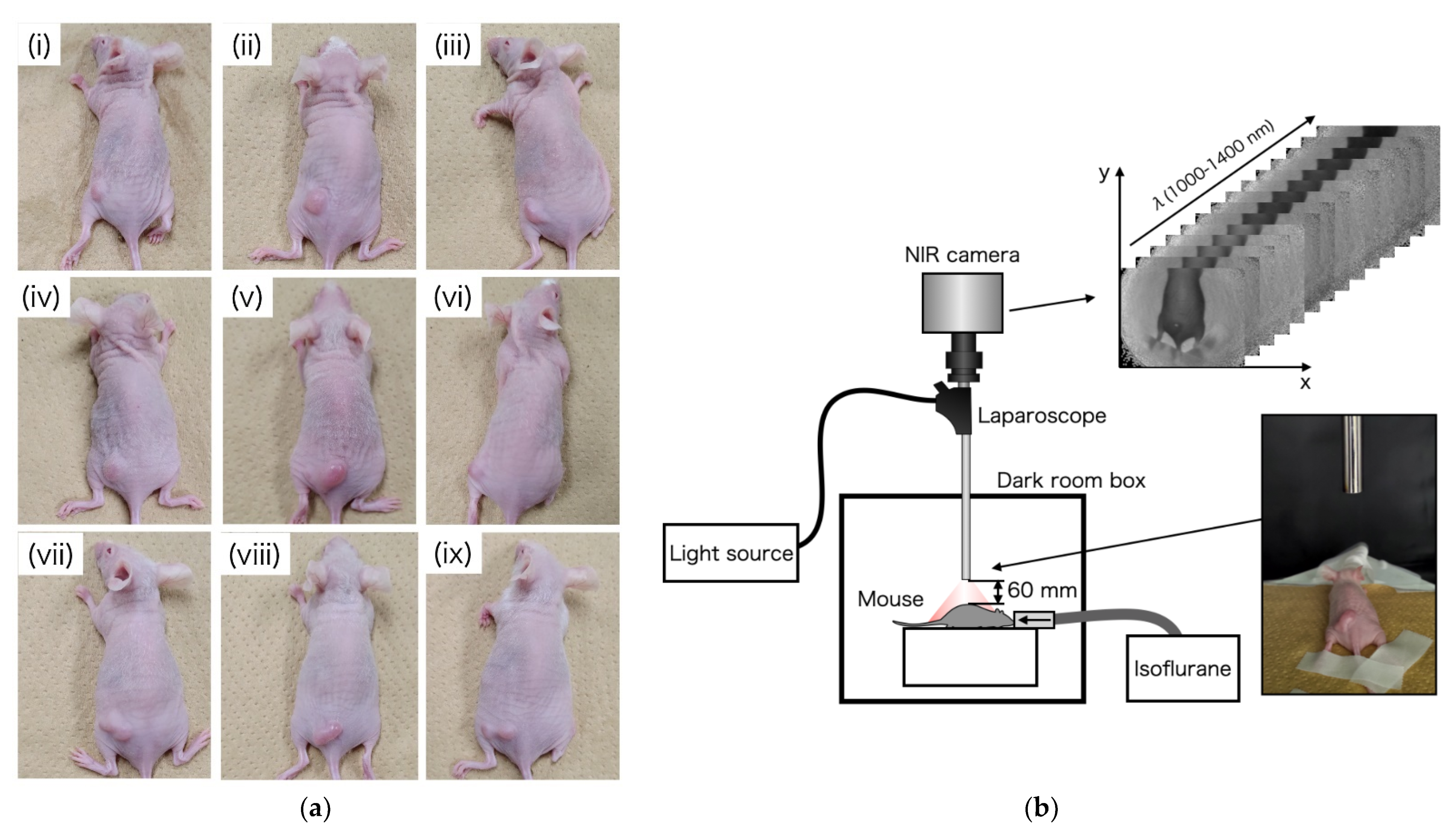
2.3. OTN-NIR Multispectral Imaging for Live Mouse
2.4. Data Processing
2.5. Classification Algorithm
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Hashimoto, D.A.; Rosman, G.; Rus, D.; Meireles, O.R. Artificial Intelligence in Surgery: Promises and Perils. Ann. Surg. 2018, 268, 70–76. [Google Scholar] [CrossRef] [PubMed]
- Kitaguchi, D.; Takeshita, N.; Matsuzaki, H.; Hasegawa, H.; Honda, R.; Teramura, K.; Oda, T.; Ito, M. Computer-assisted real-time automatic prostate segmentation during TaTME: A single-center feasibility study. Surg. Endosc. 2020, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.M.; Mancini, M.C.; Nie, S. Second window for in vivo imaging. Nat. Nanotechnol. 2009, 4, 710–711. [Google Scholar] [CrossRef] [PubMed]
- Zako, T.; Yoshimoto, M.; Hyodo, H.; Kishimoto, H.; Ito, M.; Kaneko, K.; Soga, K.; Maeda, M. Cancer-targeted near infrared imaging using rare earth ion-doped ceramic nanoparticles. Biomater. Sci. 2014, 3, 59–64. [Google Scholar] [CrossRef] [PubMed]
- Okubo, K.; Kitagawa, Y.; Hosokawa, N.; Umezawa, M.; Kamimura, M.; Kamiya, T.; Ohtani, N.; Soga, K. Visualization of quantitative lipid distribution in mouse liver through near-infrared hyperspectral imaging. Biomed. Opt. Express 2021, 12, 823–835. [Google Scholar] [CrossRef] [PubMed]
- Ehlen, L.; Zabarylo, U.J.; Speichinger, F.; Bogomolov, A.; Belikova, V.; Bibikova, O.; Artyushenko, V.; Minet, O.; Beyer, K.; Kreis, M.E.; et al. Synergy of Fluorescence and Near-Infrared Spectroscopy in Detection of Colorectal Cancer. J. Surg. Res. 2019, 242, 349–356. [Google Scholar] [CrossRef]
- Rey-Barroso, L.; Burgos-Fernández, F.J.; Delpueyo, X.; Ares, M.; Royo, S.; Malvehy, J.; Puig, S.; Vilaseca, M. Visible and Extended Near-Infrared Multispectral Imaging for Skin Cancer Diagnosis. Sensors 2018, 18, 1441. [Google Scholar] [CrossRef] [PubMed]
- Lu, G.; Fei, B. Medical hyperspectral imaging: A review. J. Biomed. Opt. 2014, 19, 010901. [Google Scholar] [CrossRef]
- Yoon, J.; Joseph, J.; Waterhouse, D.J.; Luthman, A.S.; Gordon, G.S.D.; Di Pietro, M.; Januszewicz, W.; Fitzgerald, R.C.; Bohndiek, S.E. A clinically translatable hyperspectral endoscopy (HySE) system for imaging the gastrointestinal tract. Nat. Commun. 2019, 10, 1902. [Google Scholar] [CrossRef] [PubMed]
- Akbari, H.; Uto, K.; Kosugi, Y.; Kojima, K.; Tanaka, N. Cancer detection using infrared hyperspectral imaging. Cancer Sci. 2011, 102, 852–857. [Google Scholar] [CrossRef]
- Liu, Z.; Wang, H.; Li, Q. Tongue Tumor Detection in Medical Hyperspectral Images. Sensors 2011, 12, 162–174. [Google Scholar] [CrossRef]
- Baltussen, E.J.M.; Sterenborg, H.J.C.M.; Ruers, T.J.M.; Dashtbozorg, B. Optimizing algorithm development for tissue classification in colorectal cancer based on diffuse reflectance spectra. Biomed. Opt. Express 2019, 10, 6096–6113. [Google Scholar] [CrossRef]
- Kho, E.; De Boer, L.L.; Van De Vijver, K.K.; Van Duijnhoven, F.; Peeters, M.-J.T.V.; Sterenborg, H.J.; Ruers, T.J. Hyperspectral Imaging for Resection Margin Assessment during Cancer Surgery. Clin. Cancer Res. 2019, 25, 3572–3580. [Google Scholar] [CrossRef]
- Kho, E.; Boer, L.L.; Post, A.L.; Van De Vijver, K.K.; Jóźwiak, K.; Sterenborg, H.J.; Ruers, T.J. Imaging depth variations in hyperspectral imaging: Development of a method to detect tumor up to the required tumor-free margin width. J. Biophotonics 2019, 12, e201900086. [Google Scholar] [CrossRef] [PubMed]
- De Koning, S.G.B.; Weijtmans, P.; Karakullukcu, M.B.; Shan, C.; Baltussen, E.J.M.; Smit, L.A.; Van Veen, R.L.P.; Hendriks, B.H.W.; Sterenborg, H.J.C.M.; Ruers, T.J.M. Toward assessment of resection margins using hyperspectral diffuse reflection imaging (400–1700 nm) during tongue cancer surgery. Lasers Surg. Med. 2020, 52, 496–502. [Google Scholar] [CrossRef]
- Sato, D.; Takamatsu, T.; Umezawa, M.; Kitagawa, Y.; Maeda, K.; Hosokawa, N.; Okubo, K.; Kamimura, M.; Kadota, T.; Akimoto, T.; et al. Distinction of surgically resected gastrointestinal stromal tumor by near-infrared hyperspectral imaging. Sci. Rep. 2020, 10, 21852. [Google Scholar] [CrossRef] [PubMed]
- Akimoto, K.; Ike, R.; Maeda, K.; Hosokawa, N.; Takamatsu, T.; Soga, K.; Yokota, H.; Sato, D.; Kuwata, T.; Ikematsu, H.; et al. Wavelength Bands Reduction Method in Near-Infrared Hyperspectral Image based on Deep Neural Network for Tumor Lesion Classification. Adv. Image Video Process. 2021, 9, 273–281. [Google Scholar] [CrossRef]
- Zuzak, K.J.; Naik, S.C.; Alexandrakis, G.; Hawkins, D.; Behbehani, K.; Livingston, E. Intraoperative bile duct visualization using near-infrared hyperspectral video imaging. Am. J. Surg. 2008, 195, 491–497. [Google Scholar] [CrossRef] [PubMed]
- Zuzak, K.J.; Naik, S.C.; Alexandrakis, G.; Hawkins, D.; Behbehani, K.; Livingston, E.H. Characterization of a Near-Infrared Laparoscopic Hyperspectral Imaging System for Minimally Invasive Surgery. Anal. Chem. 2007, 79, 4709–4715. [Google Scholar] [CrossRef]
- Clancy, N.T.; Jones, G.; Maier-Hein, L.; Elson, D.S.; Stoyanov, D. Surgical spectral imaging. Med. Image Anal. 2020, 63, 101699. [Google Scholar] [CrossRef] [PubMed]
- Zako, T.; Ito, M.; Hyodo, H.; Yoshimoto, M.; Watanabe, M.; Takemura, H.; Kishimoto, H.; Kaneko, K.; Soga, K.; Maeda, M. Extra-luminal detection of assumed colonic tumor site by near-infrared laparoscopy. Surg. Endosc. 2015, 30, 4153–4159. [Google Scholar] [CrossRef]
- Volpi, D.; Tullis, I.D.C.; Barber, P.R.; Augustyniak, E.M.; Smart, S.C.; Vallis, K.A.; Vojnovic, B. Electrically tunable fluidic lens imaging system for laparoscopic fluorescence-guided surgery. Biomed. Opt. Express 2017, 8, 3232–3247. [Google Scholar] [CrossRef] [PubMed]
- Kang, H.G.; Song, S.H.; Han, Y.B.; Lee, H.-Y.; Kim, K.M.; Hong, S.J. Proof-of-concept of a multimodal laparoscope for simultaneous NIR/gamma/visible imaging using wavelength division multiplexing. Opt. Express 2018, 26, 8325–8339. [Google Scholar] [CrossRef] [PubMed]
- Svozil, D.; Kvasnicka, V.; Pospichal, J. Introduction to multi-layer feed-forward neural networks. Chemom. Intell. Lab. Syst. 1997, 39, 43–62. [Google Scholar] [CrossRef]
- Udelhoven, T.; Schütt, B. Capability of feed-forward neural networks for a chemical evaluation of sediments with diffuse reflectance spectroscopy. Chemom. Intell. Lab. Syst. 2000, 51, 9–22. [Google Scholar] [CrossRef]
- Stelzer, E.H. Contrast, resolution, pixelation, dynamic range and signal-to-noise ratio: Fundamental limits to resolution in fluorescence light microscopy. J. Microsc. 1998, 189, 15–24. [Google Scholar] [CrossRef]
- Kasai, M.; Yasuda, Y.; Takemura, H.; Mizoguchi, H.; Soga, K.; Kaneko, K. In vivo tumor spatial classification using PCA and K-means with NIR-hyperspectral data. J. Biomed. Eng. Med. Imaging 2016, 3, 45. [Google Scholar] [CrossRef]
- Li, Q.; Wang, Y.; Liu, H.; Sun, Z. AOTF based hyperspectral tongue imaging system and its applications in computer-aided tongue disease diagnosis. In Proceedings of the 2010 3rd International Conference on Biomedical Engineering and Informatics, Yantai, China, 16–18 October 2010; Volume 4, pp. 1424–1427. [Google Scholar]
- Shao, H.; Chen, Y.; Yang, Z.; Jiang, C.; Li, W.; Wu, H.; Wang, S.; Yang, F.; Chen, J.; Puttonen, E.; et al. Feasibility Study on Hyperspectral LiDAR for Ancient Huizhou-Style Architecture Preservation. Remote Sens. 2019, 12, 88. [Google Scholar] [CrossRef]
- Soga, K.; Umezawa, M.; Okubo, K. Transparency in Biology: Making Invisible Visible; Springer: Singapore, 2021; p. 263. [Google Scholar] [CrossRef]
- Wirkert, S.J. Multispectral Image Analysis in Laparoscopy—A Machine Learning Approach to Live Perfusion Monitoring. Ph.D. Dissertation, Karlsruher Institut für Technologie (KIT), Karlsruhe, Germany, 2018. [Google Scholar] [CrossRef]
- Köhler, H.; Kulcke, A.; Maktabi, M.; Moulla, Y.; Jansen-Winkeln, B.; Barberio, M.; Diana, M.; Gockel, I.; Neumuth, T.; Chalopin, C. Laparoscopic system for simultaneous high-resolution video and rapid hyperspectral imaging in the visible and near-infrared spectral range. J. Biomed. Opt. 2020, 25, 086004. [Google Scholar] [CrossRef] [PubMed]


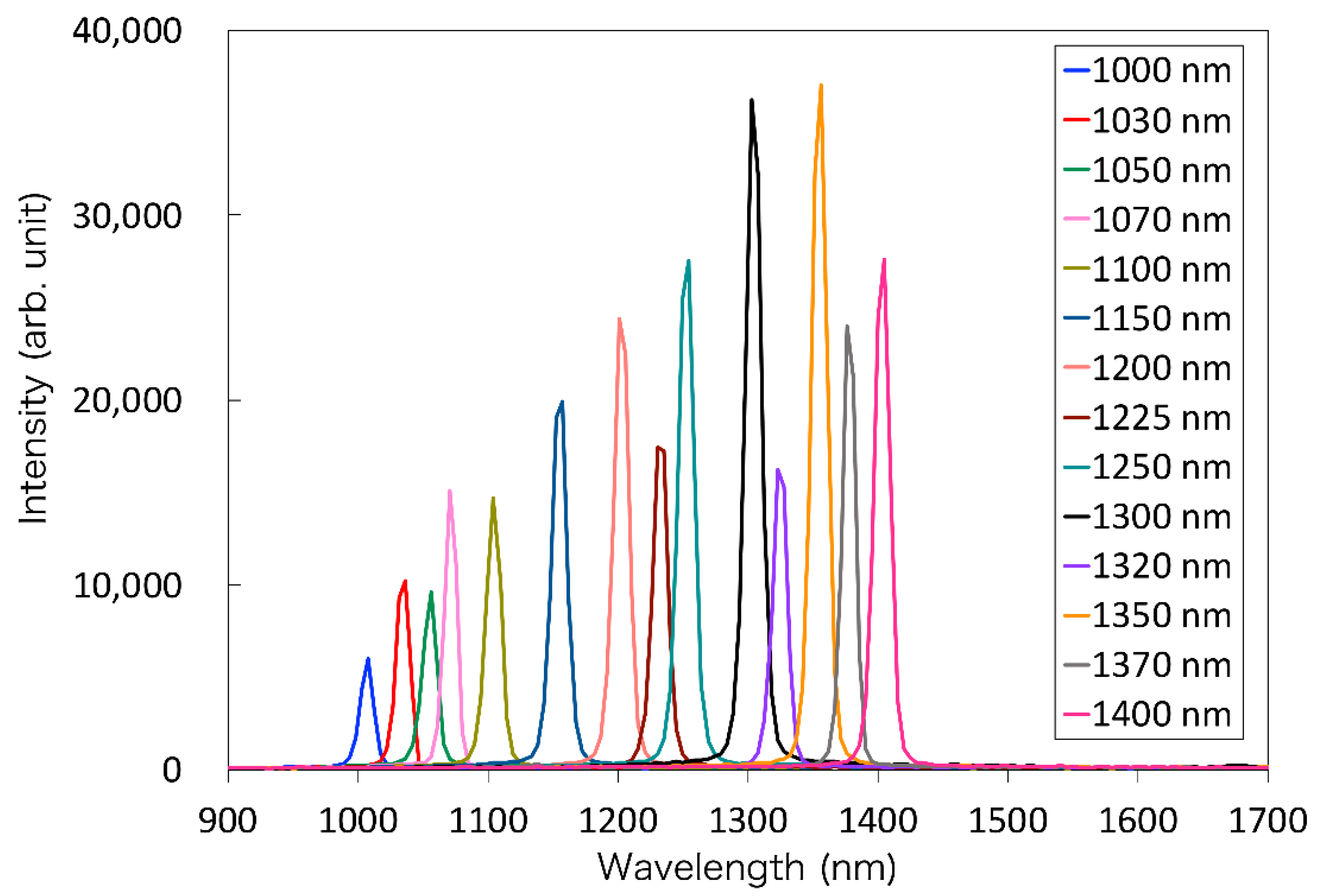
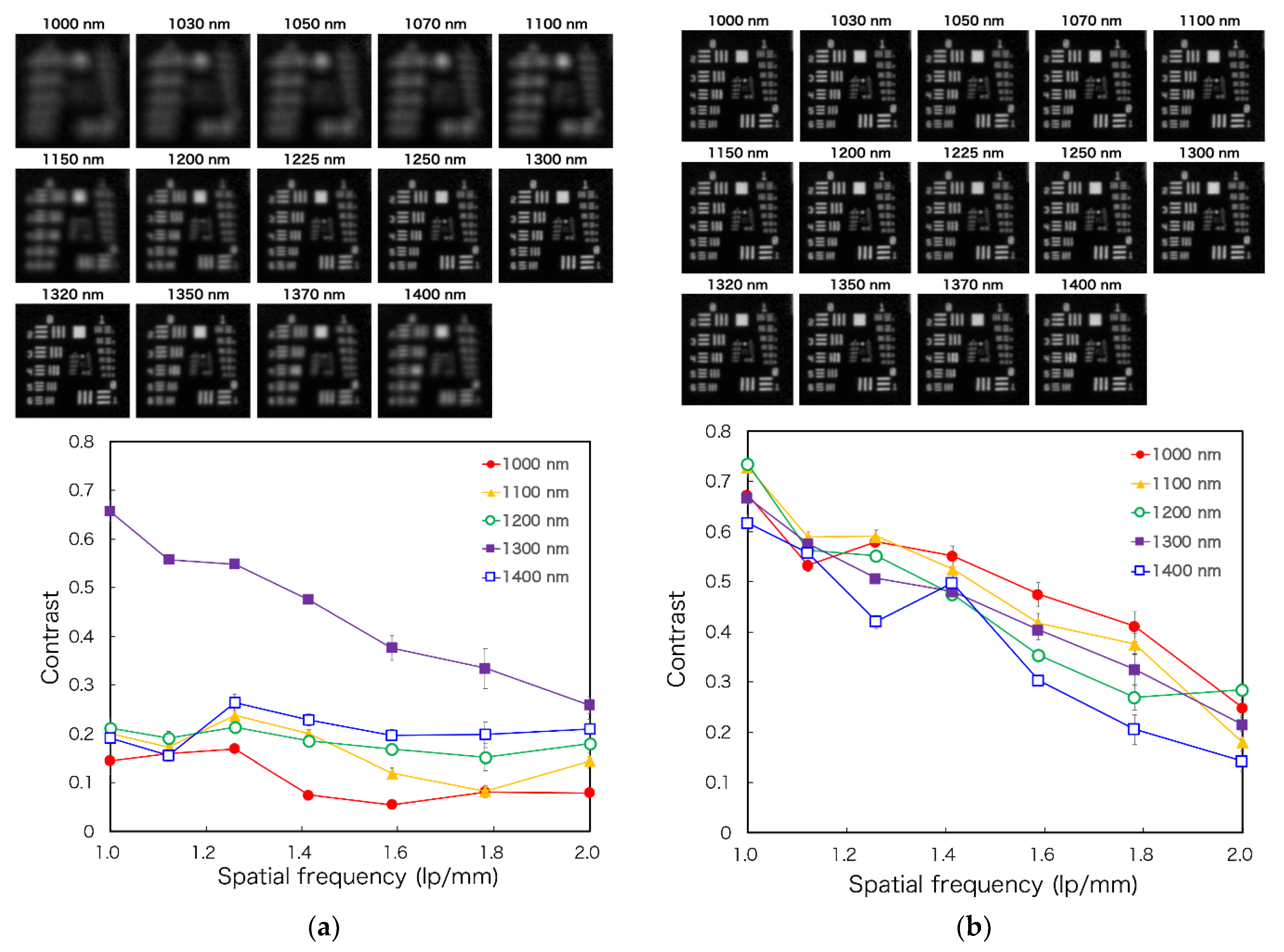

| Wavelength (nm) | Optical Power (dpt) | FWHM (nm) | Vendor | Model Number |
|---|---|---|---|---|
| 1000 | −2.03 | 10 | Edmund Optics | 65-782 |
| 1030 | −1.87 | 10 | Thorlabs, Inc | FLH1030-10 |
| 1050 | −1.55 | 10 | Edmund Optics | 65-783 |
| 1070 | −1.61 | 10 | Thorlabs, Inc | FBH1070-10 |
| 1100 | −1.42 | 10 | Edmund Optics | 65-784 |
| 1150 | −0.85 | 10 | Edmund Optics | 65-785 |
| 1200 | −0.69 | 10 | Edmund Optics | 65-786 |
| 1225 | −0.29 | 10 | IR System Co., Ltd. | NB-1225-010 nm |
| 1250 | −0.22 | 10 | Edmund Optics | 65-787 |
| 1300 | 0.29 | 12 | Edmund Optics | 65-788 |
| 1320 | 0.59 | 12 | Thorlabs, Inc | FB1320-12 |
| 1350 | 0.67 | 12 | Edmund Optics | 65-789 |
| 1370 | 0.94 | 10 | IR System Co., Ltd. | NB-1370-010 nm |
| 1400 | 1.18 | 12 | Edmund Optics | 65-790 |
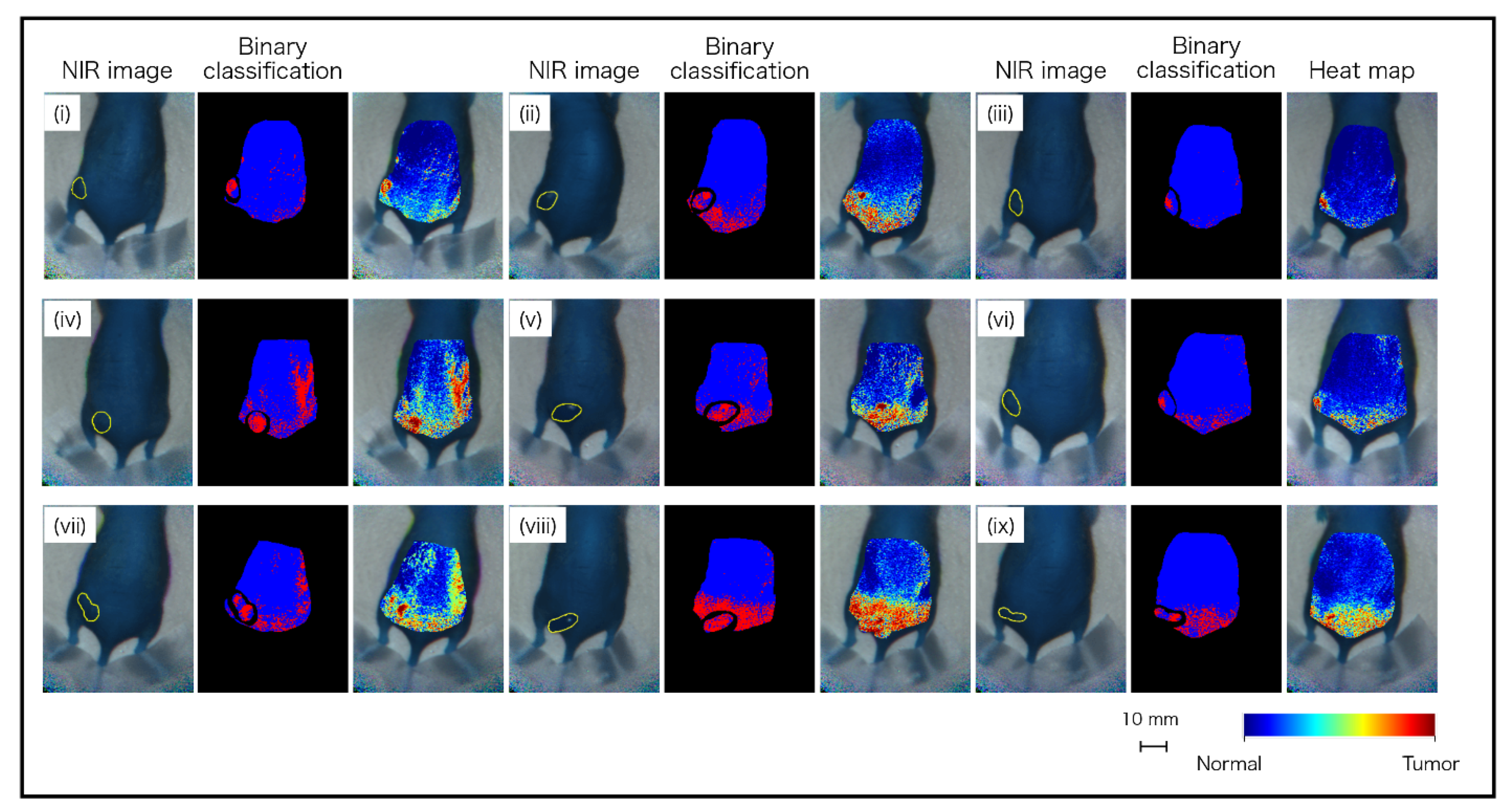
| No. | Tumor Volume (mm3) | Tumor (px) | Normal (px) | Specificity (%) | Sensitivity (%) | Accuracy (%) |
|---|---|---|---|---|---|---|
| i | 174.0 | 360 | 11,015 | 96.3 | 54.7 | 95.0 |
| ii | 526.0 | 507 | 11,527 | 89.4 | 53.5 | 87.9 |
| iii | 355.2 | 451 | 10,923 | 99.2 | 26.6 | 96.3 |
| iv | 416.1 | 565 | 10,322 | 80.9 | 72.9 | 80.5 |
| v | 821.9 | 735 | 8698 | 88.0 | 61.1 | 85.9 |
| vi | 276.5 | 558 | 12,472 | 94.6 | 21.3 | 91.5 |
| vii | 696.2 | 648 | 9453 | 87.1 | 52.3 | 84.9 |
| viii | 419.5 | 684 | 10,457 | 77.8 | 74.1 | 77.6 |
| ix | 341.9 | 408 | 12,637 | 90.0 | 52.7 | 88.8 |
| total | - | 4916 | 97,504 | 89.5 | 53.5 | 87.8 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Takamatsu, T.; Kitagawa, Y.; Akimoto, K.; Iwanami, R.; Endo, Y.; Takashima, K.; Okubo, K.; Umezawa, M.; Kuwata, T.; Sato, D.; et al. Over 1000 nm Near-Infrared Multispectral Imaging System for Laparoscopic In Vivo Imaging. Sensors 2021, 21, 2649. https://doi.org/10.3390/s21082649
Takamatsu T, Kitagawa Y, Akimoto K, Iwanami R, Endo Y, Takashima K, Okubo K, Umezawa M, Kuwata T, Sato D, et al. Over 1000 nm Near-Infrared Multispectral Imaging System for Laparoscopic In Vivo Imaging. Sensors. 2021; 21(8):2649. https://doi.org/10.3390/s21082649
Chicago/Turabian StyleTakamatsu, Toshihiro, Yuichi Kitagawa, Kohei Akimoto, Ren Iwanami, Yuto Endo, Kenji Takashima, Kyohei Okubo, Masakazu Umezawa, Takeshi Kuwata, Daiki Sato, and et al. 2021. "Over 1000 nm Near-Infrared Multispectral Imaging System for Laparoscopic In Vivo Imaging" Sensors 21, no. 8: 2649. https://doi.org/10.3390/s21082649
APA StyleTakamatsu, T., Kitagawa, Y., Akimoto, K., Iwanami, R., Endo, Y., Takashima, K., Okubo, K., Umezawa, M., Kuwata, T., Sato, D., Kadota, T., Mitsui, T., Ikematsu, H., Yokota, H., Soga, K., & Takemura, H. (2021). Over 1000 nm Near-Infrared Multispectral Imaging System for Laparoscopic In Vivo Imaging. Sensors, 21(8), 2649. https://doi.org/10.3390/s21082649






