1. Introduction
Motor imaging (MI) is the dynamic cognitive capability of generating mental movements without executing them. This mental process triggers the neurocognitive mechanisms that underlie voluntary movement planning, similar to how the action is performed realistically. MI has been proposed as a reliable tool in acquiring new motor skills to increase sports performance and physical therapy [
1,
2,
3,
4], in the development of professional motor skills learning [
5], and in improving balance and mobility outcomes in older adults and children with developmental coordination disorders [
6,
7], among others. There is sufficient experimental evidence that MI induces recovery and neuroplasticity in neurophysical regulation as the basis of motor learning [
8] and educational fields [
9]. Concerning this aspect, the Media and Information Literacy approach has been proposed by the United Nations Educational, Scientific, and Cultural Organization (UNESCO) to gather several vital human development capabilities. In practice, MI tasks are commonly solved from electroencephalography (EEG) records, which provide noninvasive measures and flexible portability at a relatively low-cost. However, EEG signals lack a suitable spatial resolution, not to mention the inter and intra-subject variability regarding the somatosensory cortex’s responses. Specifically, there is no consistency in the patterns among different subjects. Indeed, the variability arises within a session for the same subject because of a non-stationary, nonlinear, and low signal-to-noise ratio of EEG signals [
10]. Together with frequently used small sample datasets, all of these factors reduce MI systems’ performance based on EEG [
11,
12].
An enhanced approach to addressing this EEG data complexity is to conduct multiple training sessions to refine the modulation of sensorimotor rhythms (SMR). Nonetheless, the inter-subject variability, together with uncertain long-term effects and the apparent failure of some individuals to achieve self-regulation, makes a non-negligible portion of users (between
to
) develop insufficient coordination skills even after long training sessions. This inadequate performance of most brain-computer interface (BCI) systems (
BCI inefficiency) poses a challenge in MI research [
13]. To address this problem, the BCI performance model is enhanced in two directions: (
i) Developing guidelines in neural testing set-ups, practice, and instructions to ensure better performance of brain responses; and (
ii) Promoting evaluation tools to forecast the system performance may help identify the core issue of variability to incorporate compensating actions for the inefficiency when solving BCI-based tasks. In particular, a calibration strategy can be added, working hand in hand within the training stage. Therefore, it is possible to adapt the decoding scheme with an explicit brain pattern [
14], highlighting relevant BCI predictors to decrease training efforts and encourage user-centered MI [
15]. To date, several electrophysiological indicators have been reported to anticipate the MI inefficiency, like the direct assessment of the SMR, which extracts the power spectral density (PSD) from the resting wakefulness at motor cortex locations [
16]; a measure of the PSD uniformity of the resting-state data using spectral entropy [
17,
18]; and the PSD-based estimate to assess the dis/similarity (connectivity) of EEG signals at different locations in an attempt to understand the interdependency between functional and structural networks of corresponding cortical brain structures (like spectral coherence [
19,
20] or coherence-based correntropy spectral density [
21]), among others. To tackle the influence of artifacts and intertrial/inter-subject amplitude variability, phase-based relationships (phase synchronization) are more desirable as a functional connectivity (FC) measure of spatially distributed regions, dynamically interacting in accomplishing a mental task [
22]. It has been proved that the functional connectivity features measured by the phase lag index (or its weighted version—
wPLI and phase-locking value
PLV) can discriminate between different MI tasks [
23,
24].
Therefore, predicting motor performance from the resting motor-system functional connectivity can be determined as in [
16], showing that the efficient brain reconfiguration corresponds to a better MI performance [
25,
26]. Nevertheless, several conditions can affect their correct estimation and introduce spurious contributions, giving a potentially distorted measure of the real interactions (termed spurious connectivity) [
27]. Thus, FC estimation is highly time-dependent and fluctuates within multiple timescales, yielding inter-subject variations that remain a substantial problem [
28]. Specifically, the obstacles related to volume conduction and noise perturbations cause phase synchronization to incorporate thresholds applied to these FC measures to improve the connection sets’ discriminative ability. However, the threshold selection is generally far from being an automated procedure for big datasets [
29]. Undeterred by the promising evidence, there is a need to understand the learning mechanisms and the brain network reorganization, aiming to support the efficiency of BCI systems [
30].
As regards the prediction model, several regression methods are available for prognosticating MI accuracy from neurophysiological variables like simple and multiple linear regression [
31,
32], stepwise regression [
33], kernel regression [
34], and (kernel) support vector machine regression [
35], among others. Additionally, there is increasing use of regression approaches with neural networks that can be applied to the raw EEG data, simplifying BCI’s design pipelines by removing the need to extract features manually. However, several aspects degrade the prediction model performance, such as the fact that FC measures are prone to be influenced by outliers, which are to be removed before calculating correlations [
36]. Another drawback is the inter-trial variability of MI data (with a notable increase in subjects having low MI skills), which restricts prediction models with single-trial EEG data [
37]. One more issue influencing the regression model is the user’s categorization depending on their SMR activity (predictor) and classifier performance (target response) during the MI runs. Users are frequently adjusted to two partitions (skilled and non-skilled) divided by a single target value given in advance, as in [
38]. Still, as the number of subjects tested increases, the range of FC changes also rises. The partition-based method should also be sensitive in detecting predictor differences among subject clusters [
39,
40]. Therefore, the need for clustering into more partitions becomes more evident, as shown in [
41]. Lastly, the correlation coefficient (reflected in
r-squared) is often applied to assess the prediction shape, while the
p-value levels its statistical significance that can be implemented through several test procedures, as developed in [
42]. A common issue in neural network regression models, trained with small samples in MI studies, is their fitting to spurious residual variation (overfitting), apart from a controversial interpretation of
p-values [
43].
Here, to increase the prediction performance of the baseline linear regression models, we develop a deep network regression model devoted to prognosticating Motor Imagery Skills using EEG Functional Connectivity Indicators, appraising three procedures: leave-one-out cross-validation combined with Monte Carlo dropout layers, subject clustering of MI inefficiency, and transfer learning connecting neighboring runs. Our approach comprises functional connectivity predictors extracted from electroencephalographic signals to favor the data interpretability. To deal with the risk of overfitting prediction assessments because of the deep learning framework, we intend to preserve as much information as possible from the measured scalp potentials. Thus, to reach competitive values of prediction errors achieved by the leave-one-out cross-validation scheme, we introduce the following procedures: (i) Monte Carlo dropout layers to decrease the probability that the learned rules from specific training data cannot be generalized to new observations; (ii) Subject efficiency clustering to adapt the DNR estimator more effectively to complex EEG measurements inherent to BCI inefficiency subjects; (iii) For Prediction of Initial-training Synchronization, transfer learning of the weights inferred at the predecessor run to deal with the few-trial sets. The validation is performed in two MI databases (150 users) acquired in conditions close to real MI applications. Obtained results show how our approach can achieve a high prediction of pretraining desynchronization and initial training synchronization with adequate physiological interpretability. We further compared the DRN predictor prediction performance (on average, 0.8) with the results obtained by linear regression models that are reported, at least for DBI, in the baseline work [
44], presenting values of R-squared not exceeding 0.54.
The rest of the paper is organized as follows:
Section 2 briefly discusses the regression prediction model’s theoretical background.
Section 3 describes the experimental set-up, including both datasets evaluated.
Section 4 presents the assessment of Deep Regression Network performance and discusses the findings obtained to predict pretraining desynchronization and initial training synchronization. Lastly,
Section 5 concludes the paper.
3. Experimental Set-Up
The methodology for enhanced prediction of motor imagery skills using functional connectivity indicators is evaluated under a regression model to predict the bi-class accuracy response of subjects, embracing the following stages: (i) Predicting capability estimation of the pre-training desynchronization under a conventional linear regression model, testing different scenarios of input arrangements to improve the system performance; (ii) Prediction assessment of the pre-training desynchronization under the data-driven network regression model; (iii) Enhanced network prediction assessment using leave-one-out cross-validation combined with Monte Carlo dropout layers and clustering of subject inefficiency; (iv) Enhanced network regression prediction of initial-training synchronization with an additional transfer learning procedure.
The pre-training desynchronization assesses the relationship between the bi-class accuracy response and the electrophysiological indicators extracted from resting wakefulness data. We employ either resting-state or task-negative state before the cue-onset of the conventional MI trial timing for evaluation purposes. Besides, as the target response, we compute each subject’s classifier accuracy in distinguishing either MI class using the short-time sliding feature set extracted by the Common Spatial Patterns (CSP), which maximizes the class variance. To accurately extract the subject EEG dynamics over time, the sliding window is adjusted to 2 s, having an overlap of .
On the other hand, the pre-training desynchronization predictor relies on the fact that the change in neural activity, intentionally evoked by a mental imagery task, shows certain regularities through training runs or sessions. Accordingly, the pre-training indicator of neural desynchronization attempts to anticipate the MI responses evoked within every run’s wakefulness data.
3.1. MI Databases Description and Preprocessing
Giga-DBI: This MI dataset is publicly available at (
http://gigadb.org/dataset/100295, accessde on 30 January 2021). It gathers EEG records from fifty subjects (
), fixing the well-known
electrode configuration with
channels. The signal
comprises
s, at
Hz sample frequency. The MI protocol (see
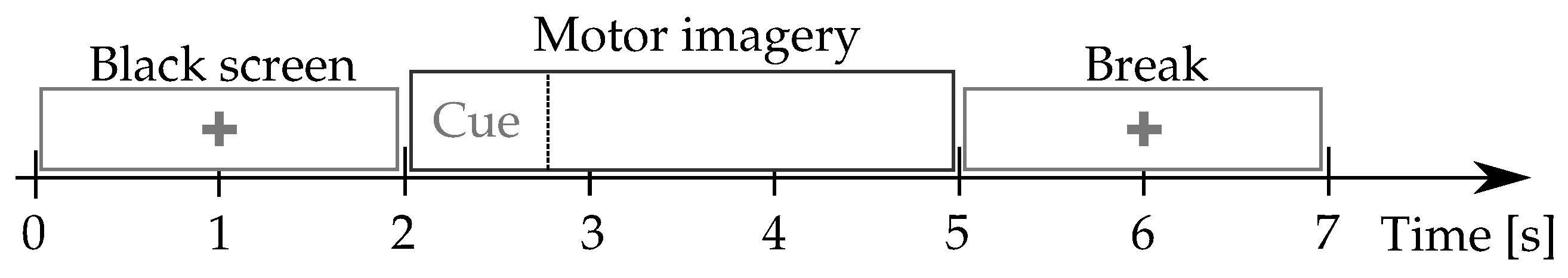
Figure 1) starts with a fixation cross shown on a black screen for 2 s. Further, a cue instruction is displayed depending on the MI instruction (label), which appears randomly within 3 s. For concrete testing, the cue asked to imagine moving his fingers, starting from the index finger and reaching the little one. Afterward, a blank screen is visible at the beginning of a break period (shown randomly between
and
s). Each MI run composes over 20 trials and a written cognitive quiz [
50]. Every subject performed five runs (on average) and a single-trial resting-state recording, lasting 60 s.
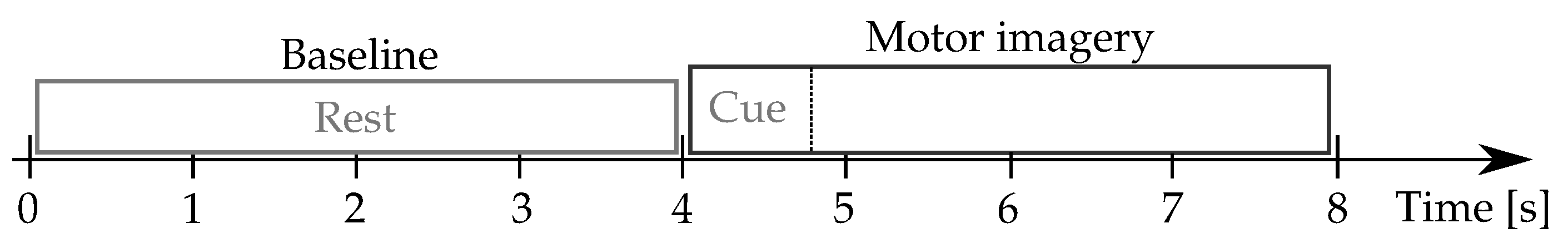
Physionet-DBII: This database, publicly available at (
https://physionet.org/content/eegmmidb/1.0.0/, accessde on 30 January 2021), holds
volunteers who properly performed the left and right-hand MI tasks, collecting a total average of
trials per subject. Besides, two one-minute baseline records are captured concerning a resting state trial (with eyes open and closed, respectively). The 64-channel EEG signals were recorded using the
international system, and sampled at
Hz.
Figure 2 describes the motor imagery timing.
Every raw EEG channel of either database was band-pass filtered in the frequency range
[4–40] Hz, covering the sensorimotor rhythms considered (
). Then, the band-passed EEG data are spatially filtered by a Laplacian filter centered on the selected electrode to improve the spatial resolution of EEG recordings, avoiding the influence of noise coming from neighboring channels and thus addressing the volume conduction problem (This filtering procedure was carried out using
Biosig Toolbox that is free available at
http://biosig.sourceforge.net, accessde on 30 January 2021). Further, the electrophysiological indicator set,
, based on phase synchronization is extracted using the
MNE package in Python, while the graph predictors are estimated using the Brain Connectivity Toolbox (brain-connectivity-toolbox.net).
3.2. Deep Network Regressor Set-Up and Performance Evaluation
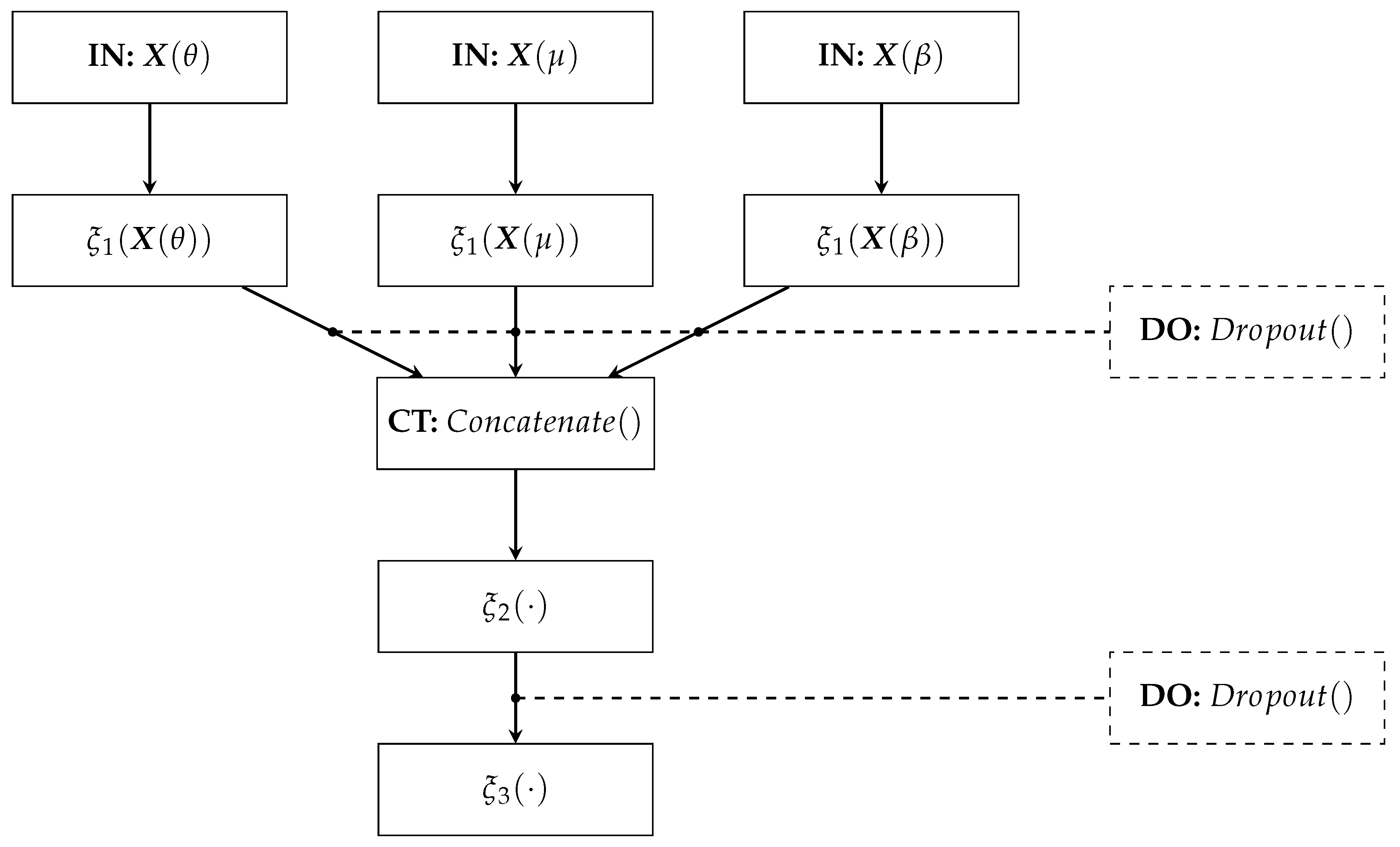
The proposed Deep Regression Network architecture comprises (see
Figure 3):
- –
IN: We consider two inputs-layer arrangements: multivariate indicator (Being C the electrode and links number when using graph indexes and FC, respectively).
- –
: The first dense layer codes the input relevant patterns from phase synchronization features. Here, we fix the number of neurons as neurons, where ⌜·⌝ stands for the ceiling operator. A tanh-based activation is employed to reveal non-linear relationships.
- –
CT: A concatenate layer is applied to append the resulting feature maps from the set of patterns extracted in . In particular, all phase synchronization-based features (coded as connectivity matrices) are stacked into a single block, sizing .
- –
: This fully-connected layer aims to preserve the predicted patterns assembled in the CT layer to fed a linear regressor. The number of neurons is fixed as . Again, the tanh is used as activation function.
- –
: A one-neuron layer with linear activation is used to predict the MI skill value .
- –
DO: This Dropout layer randomly skips neurons according to drop rate. We fix the drop rate at 0.2 empirically.
For measuring the relationship between the response variable and the composite predictor, we build the set
, where
is computed using our Deep Learning Regressor following a leave-one-out cross-validation strategy along with the
M subjects. The quantity measures account for the influence to predict the acceptance rate on the electrodes performed by individuals, namely, for computation of value
, one individual is picked out as the training set and the remaining ones as the testing set. Then, the coefficient of determination (noted as
R) is computed. Besides, a
p-value is computed from a two-sided t-test whose null hypothesis is that the regression slope is zero [
44]. It is worth noting that such a hypothesis testing is used, as in state-of-the-art works [
38,
44], because our Deep Learning Regressor aims to code the no consistency in the brain patterns among different subjects to favor a linear dependency between
and
. Moreover, to provide a comparison with Neural Network-based regression strategies, the real-valued measures of
Mean Absolute Error (MAE), and
Root Squared Error (RMSE) are also assessed, as carried out in [
51,
52]:
where
stands for the variance operator.
5. Concluding Remarks
Here, we develop a methodology for predicting MI practicing’s neurophysiological inefficiency using EEG phase synchronization measures. A deep network regression evaluates over 150 subjects’ predicting capability in assessing the pre-training desynchronization and the initial training synchronization. The prediction estimates should help determine whether a specific user needs to undergo an additional calibration, supplying interpretation of subjects’ learning properties. Although our algorithm training can be time-consuming, growing considerably as the database set increases, such a training stage can be implemented offline. Once the Deep Network Regressor’s weights are learned, our predictor evaluation is as fast as baseline models, enabling real-time applications like the run-based prediction of initial-training synchronization.
From the obtained results of validation, the following aspects are to be emphasized:
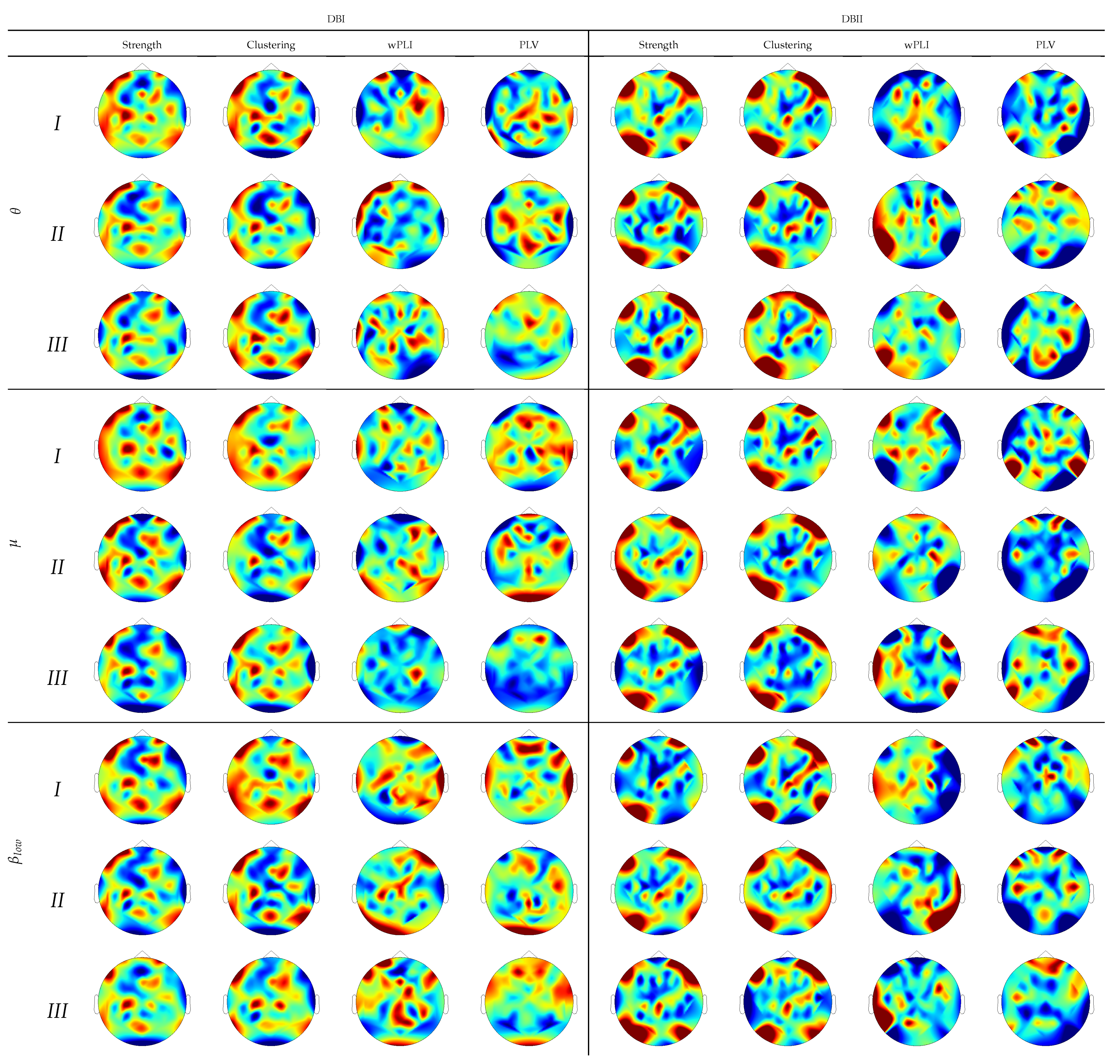
Electrophysiological predictors based on functional connectivity. We explore the Phase Locking Value and Weighted Phase Locking Index as connectivity measures together with their brain graph predictors (strength and clustering coefficient) to build a predictive regression model of BCI control. From the obtained results for the linear regression model (simple
Table 1 and multiple
Table 2), we can conclude that the FC predictors extracted from resting-state enable fair values of prediction performance (
R-squared below
) with notable variations, regardless of the input arrangement configuration employed. This behavior worsens in DBII that is an EEG collection with greater structure variableness.
With regard to DNR, all considered FC predictors present similar and even performance, reaching more competitive prediction values (see
Table 3,
Table 4 and
Table 5). In terms of providing interpretation, the DNR weights mostly supporting the prediction performance are comparable in
wPLI and
PLV predictors (see
Figure 4). One more consideration is the limited effectiveness of the thresholding method, usually performed to remove false connections and noise. The thresholding performance may be jeopardized by the high intrasubject variability, demanding the application of subject-related tuning algorithms. Therefore, the network regression models ease the need for elaborate feature extraction procedures based on functional connectivity analysis. It is worth noting that the network regression estimator benefits from all considered rhythms (i.e.,
), though each contributes differently.
Quality of network regression models. While widely-common procedures can appraise linear regression models’ statistical significance, assessing and enhancing network regression models’ prediction quality is a much more challenging task because of the risk of overfitting [
56]. Here, we propose the leave-one-out cross-validation that includes Monte Carlo dropout layers (holding neurons with a probability of being ignored during training and validation) for decreasing the probability that the learned rules from specific training data cannot be generalized to new observations. As a result, the DNR prediction errors of
MAE and
RMSE fall by nearly half (see
Table 4). Furthermore, including the Monte Carlo dropout layers allows selecting a reduced set of FC links enhancing the prediction performance (see
Figure 4). Consequently, this aspect improves the physiological interpretability of network regression models.
In practice, assessment of resting-state activation is frequently performed with a reduced number of electrodes to reduce computational complexity and the set-up time. To this end, we evaluate the DRN performance for the predictor sets extracted over the sensorimotor area, showing that the channel selection strategy underperforms the whole electrode set’s inclusion. This issue becomes more manifest in subjects with a more prominent EEG variability (that is, high BCI inefficiency). As suggested in [
57], the learned network weights depend on the variability resulting from the channel selection used, making the prediction performance vary notably from one subject partition to another.
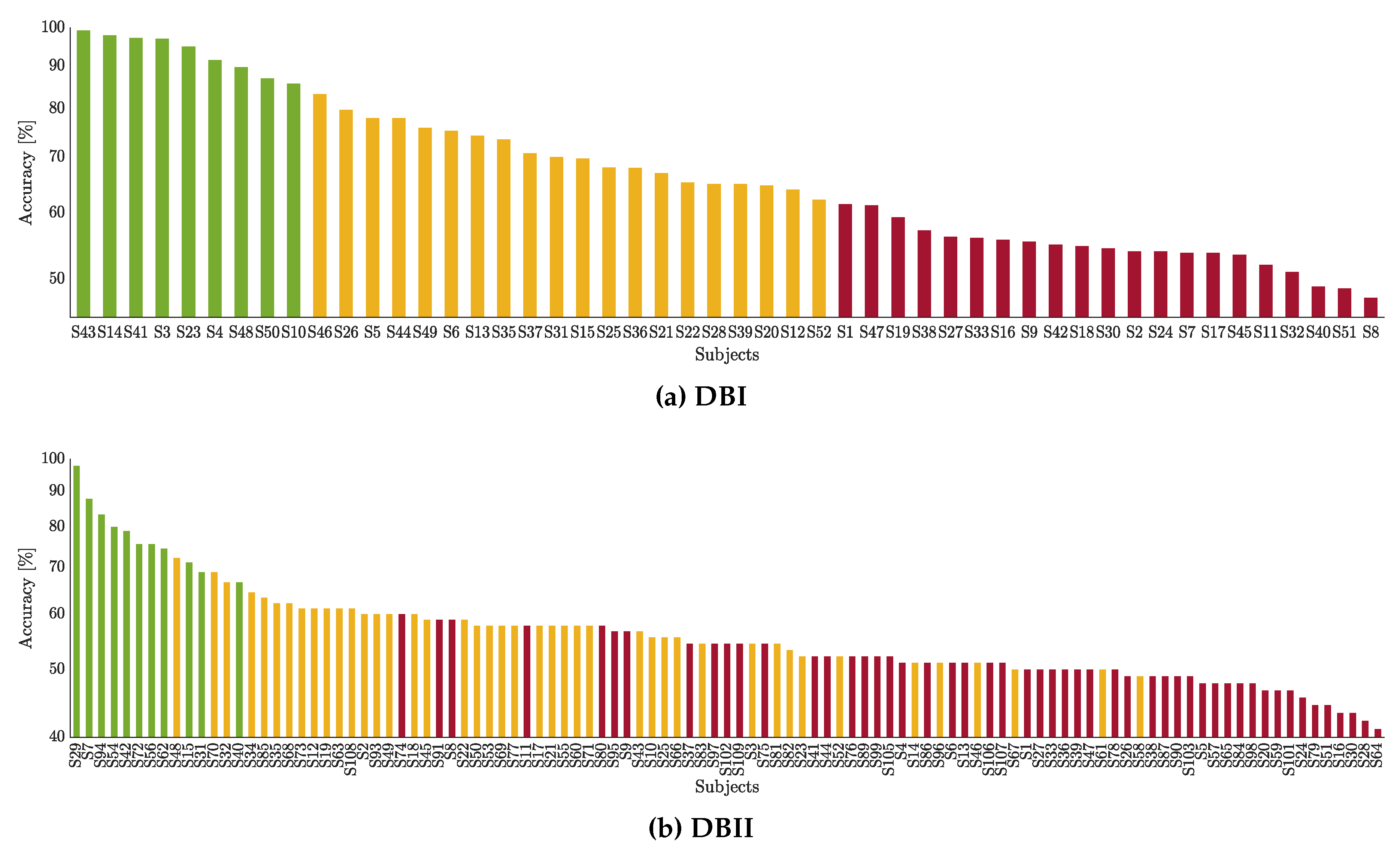
Regression assessments using subject clustering of BCI inefficiency. One more issue impacting the regression prediction is the user’s categorization depending on their SMR activity and classifier performance during the MI runs. The obtained results show that the prediction performance improves (the values of R-squared increase while the errors decrease) regardless of the subject group under consideration. Therefore, we hypothesize that the regression analysis using partitions may be more effective in databases with complex EEG measurements. Consequently, clustering combined with DNR models enhances understanding of the factors influencing subjects’ accuracy performance with significant BCI inefficiency.
DNR prediction with transfer learning. We also assess the DNR performance of predicting the MI accuracy at each run using the single-trial PLV predictor of wakefulness data. However, we associate the values learned from each run’s MI data with the weights inferred at the predecessor run to deal with the few-trials sets. Thus, compared to the all set performance, the initial-training synchronization prediction increases in each group of individuals.
For future work, the authors plan to enhance FC predictors’ feature extraction, providing a better understanding of their impact and interaction on BCI-related tasks to identify potential non-learners.
Profiting from MI-based BCI learning progression, dynamic network regression models must be developed to capture the sequence regression’s latent trends. In this line of analysis, the cluster-based enhancing procedure and the vector accuracy response should also account for FC predictors’ dynamic behavior. Intending to improve the DNR prediction, an extended panel of standardized and validated psychological questionnaires are to be included within the network estimator, accounting for user’s specific characteristics like daily motor activity and age.
One more aspect to explore is to adjust the DNR pipeline to learn the weights for supporting prediction in a broader clinical application class, relying on the ability of deep learning architectures to extract complex random structures from EEG data.