A Convolutional Neural Network-Based Method for Corn Stand Counting in the Field
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Setting
2.2. Image Preparation
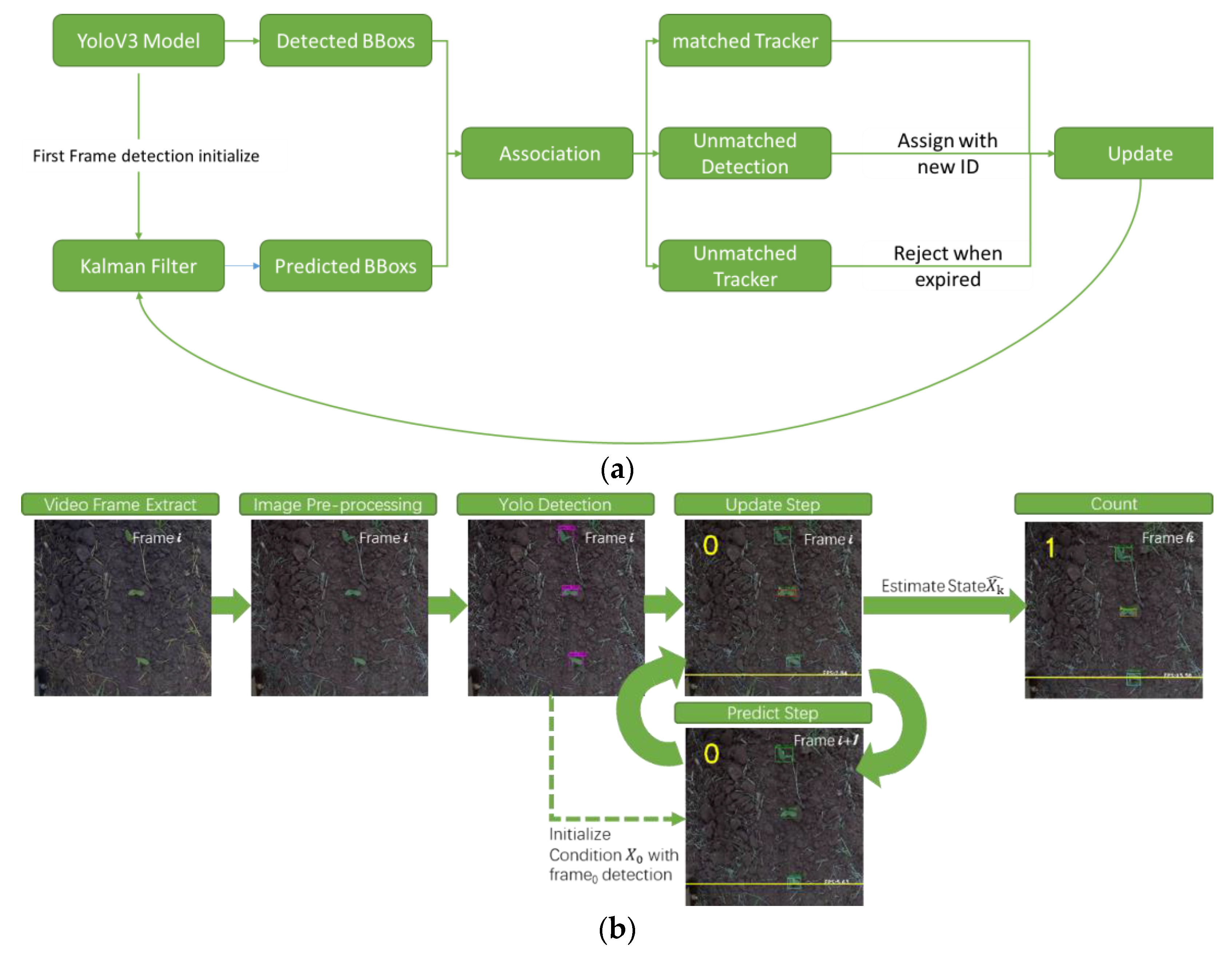
2.3. Corn Plant Stand Counting Pipeline
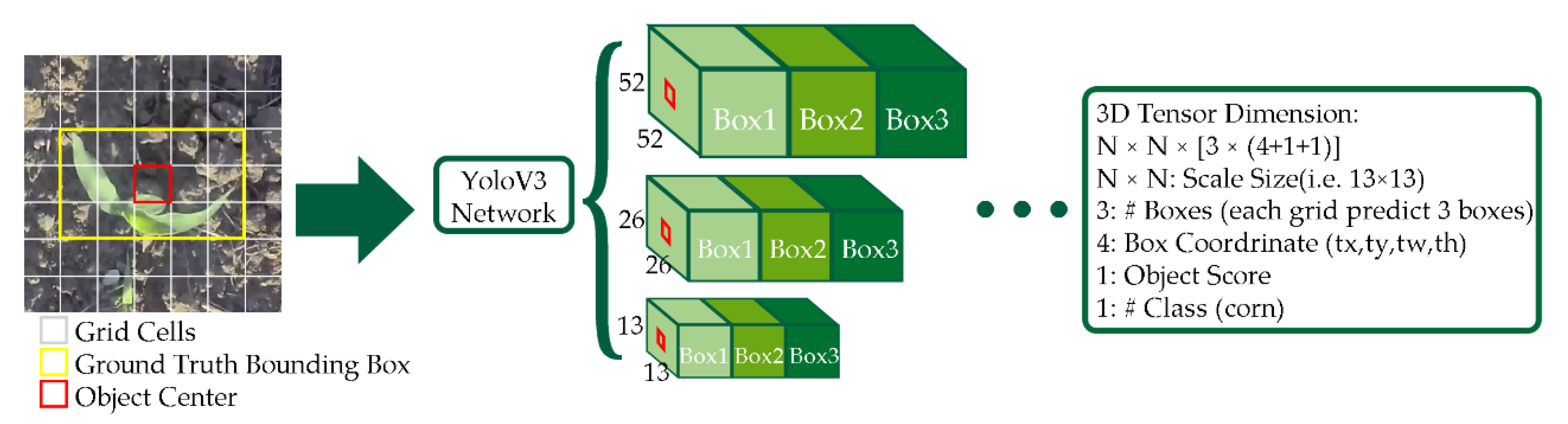
2.3.1. YoloV3 for Corn Plant Seedling Detection
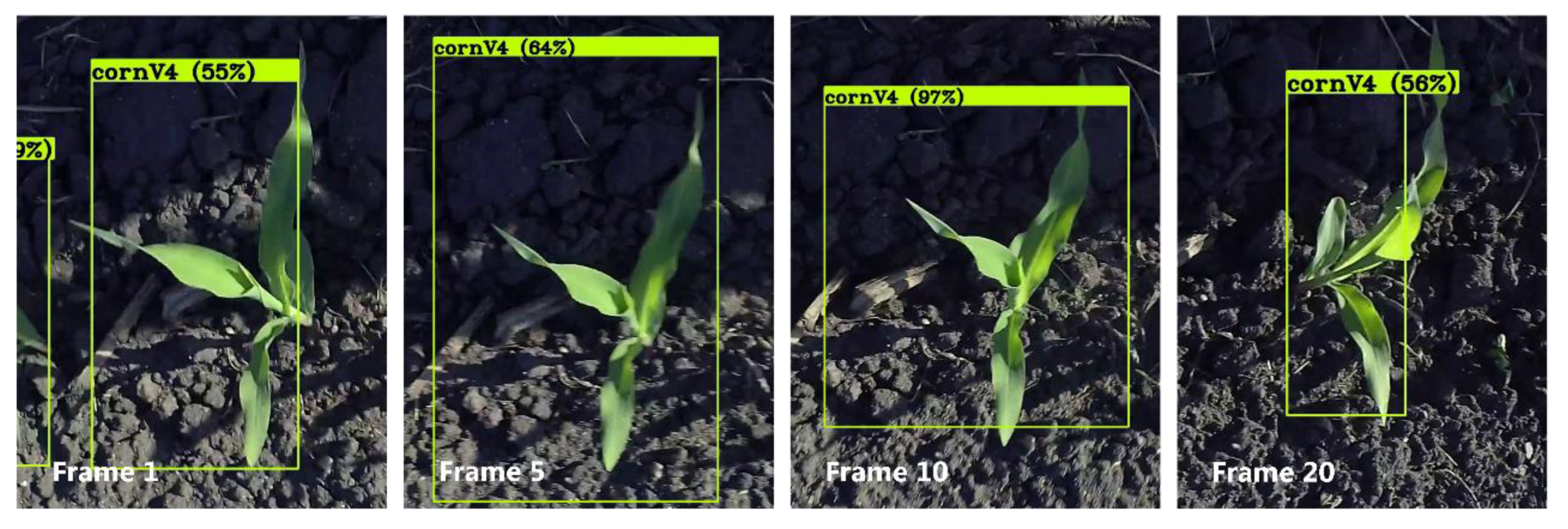
2.3.2. Corn Plant Seedling Tracking and Counting
3. Results
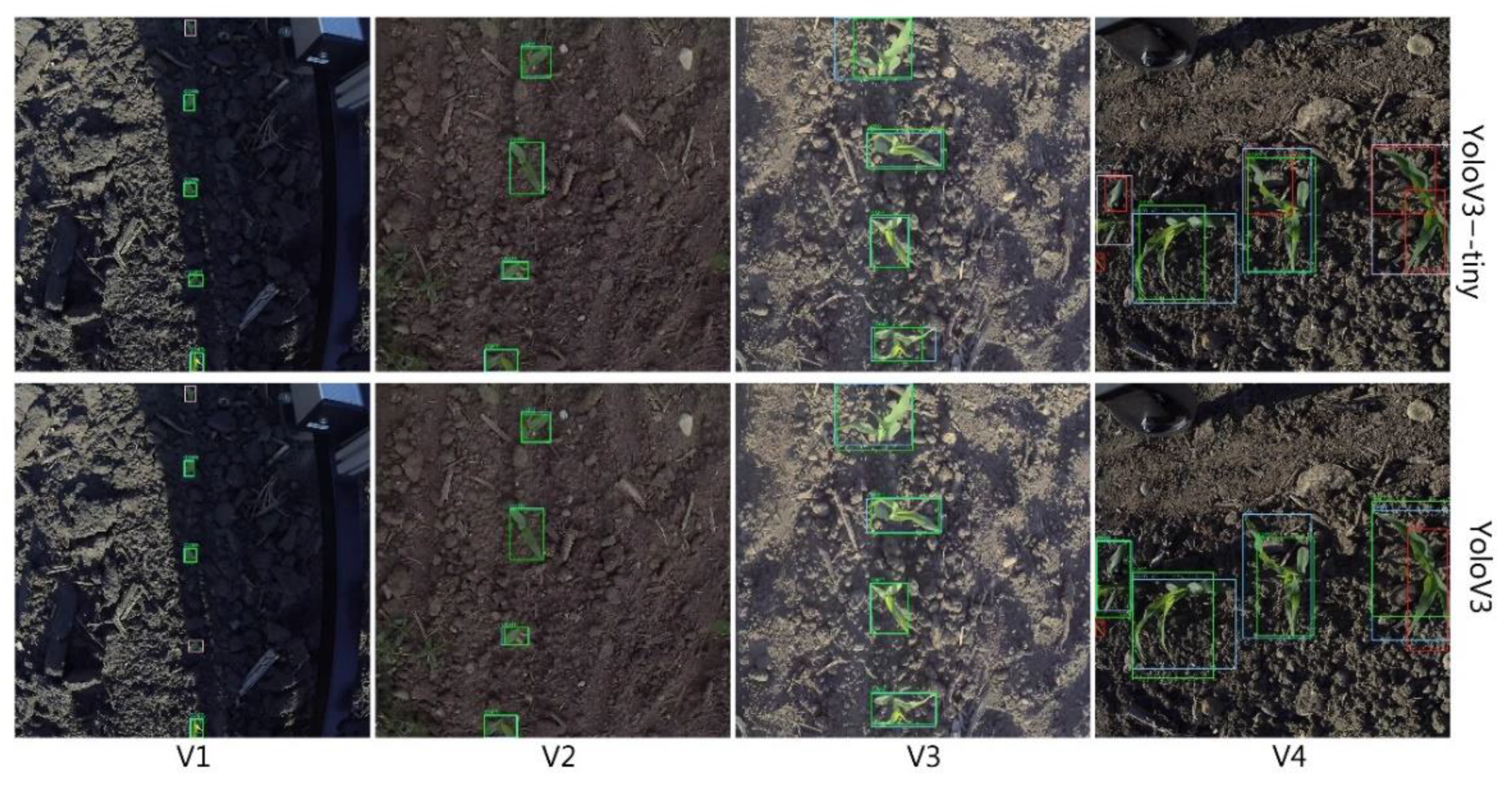
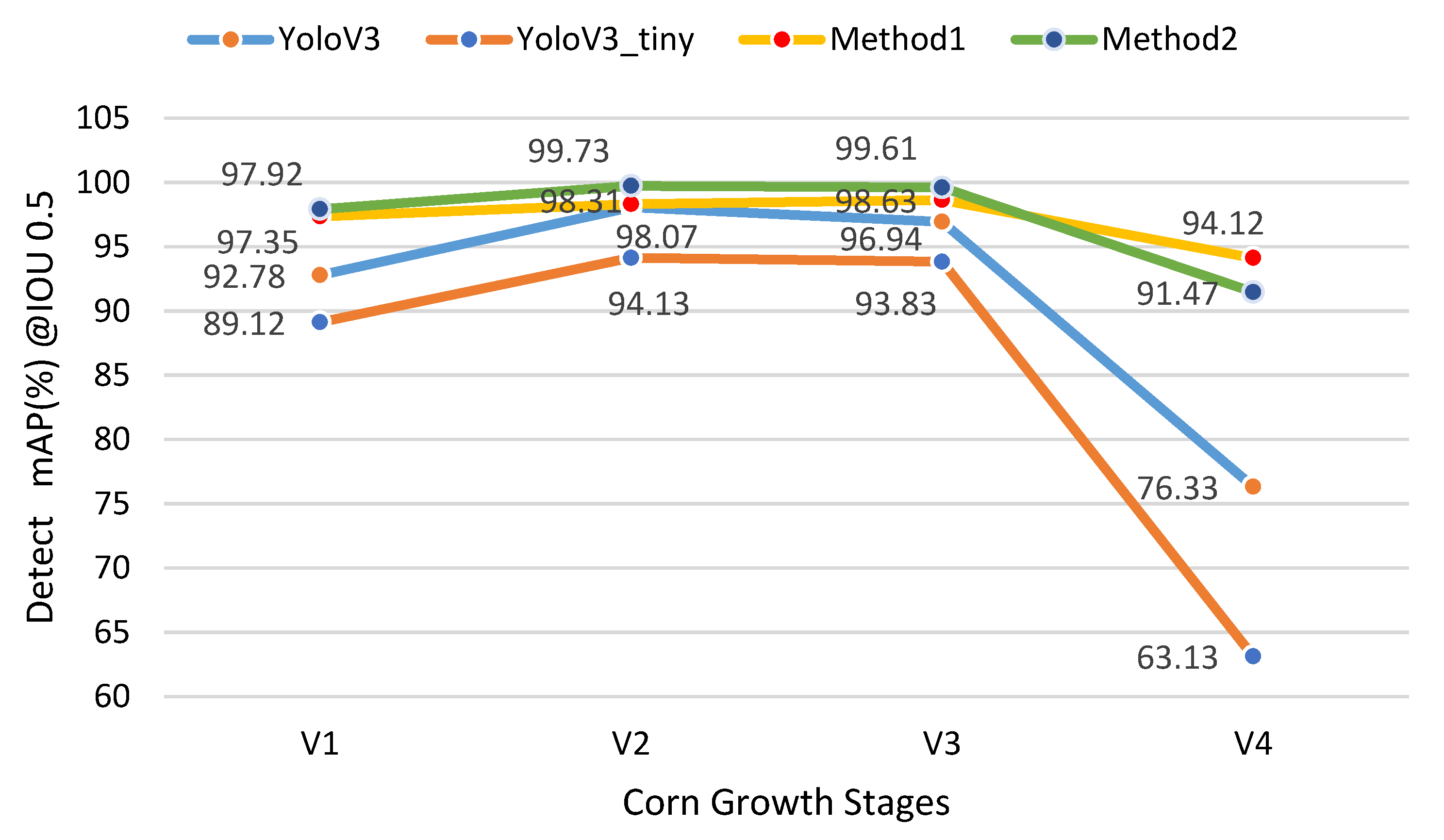
3.1. Detection
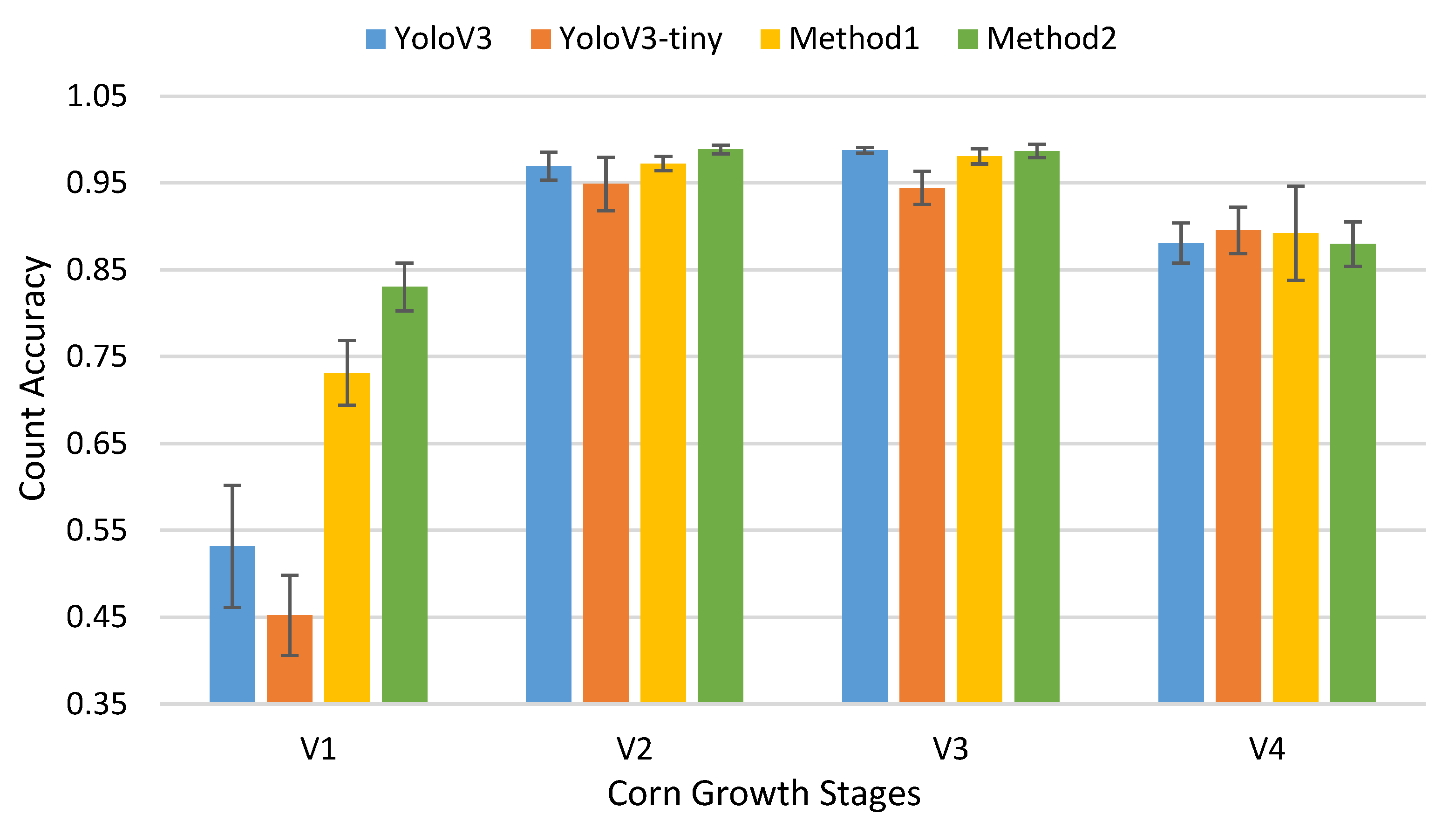
3.2. Track and Count
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mandic, V.; Krnjaja, V.; Bijelic, Z.; Tomic, Z.; Simic, A.; Stanojkovic, A.; Petricevic, M.; Caro-Petrovic, V. The effect of crop density on yield of forage maize. Biotechnol. Anim. Husb. Biotehnol. Stoc. 2015, 31, 567–575. [Google Scholar] [CrossRef]
- Yu, X.; Zhang, Q.; Gao, J.; Wang, Z.; Borjigin, Q.; Hu, S.; Zhang, B.; Ma, D. Planting density tolerance of high-yielding maize and the mechanisms underlying yield improvement with subsoiling and increased planting density. Agronomy 2019, 9, 370. [Google Scholar] [CrossRef]
- Basiri, M.H.; Nadjafi, F. Effect of plant density on growth, yield and essential oil characteristics of Iranian Tarragon (Artemisia dracunculus L.) landraces. Sci. Hortic. 2019, 257, 108655. [Google Scholar] [CrossRef]
- Zhi, X.; Han, Y.; Li, Y.; Wang, G.; Du, W.; Li, X.; Mao, S.; Feng, L. Effects of plant density on cotton yield components and quality. J. Integr. Agric. 2016, 15, 1469–1479. [Google Scholar] [CrossRef]
- Zhang, D.; Sun, Z.; Feng, L.; Bai, W.; Yang, N.; Zhang, Z.; Du, G.; Feng, C.; Cai, Q.; Wang, Q.; et al. Maize plant density affects yield, growth and source-sink relationship of crops in maize/peanut intercropping. Field Crop. Res. 2020, 257, 107926. [Google Scholar] [CrossRef]
- Li, X.; Han, Y.; Wang, G.; Feng, L.; Wang, Z.; Yang, B.; Du, W.; Lei, Y.; Xiong, S.; Zhi, X.; et al. Response of cotton fruit growth, intraspecific competition and yield to plant density. Eur. J. Agron. 2020, 114, 125991. [Google Scholar] [CrossRef]
- Fischer, R.A.; Moreno Ramos, O.H.; Ortiz Monasterio, I.; Sayre, K.D. Yield response to plant density, row spacing and raised beds in low latitude spring wheat with ample soil resources: An update. Field Crop. Res. 2019, 232, 95–105. [Google Scholar] [CrossRef]
- Chen, R.; Chu, T.; Landivar, J.A.; Yang, C.; Maeda, M.M. Monitoring cotton (Gossypium hirsutum L.) germination using ultrahigh-resolution UAS images. Precis. Agric. 2018, 19, 161–177. [Google Scholar] [CrossRef]
- Fidelibus, M.W.; Mac Aller, R.T.F. Methods for Plant Sampling. Available online: http://www.sci.sdsu.edu/serg/techniques/mfps.html (accessed on 3 July 2020).
- Wu, J.; Yang, G.; Yang, X.; Xu, B.; Han, L.; Zhu, Y. Automatic Counting of in situ Rice Seedlings from UAV Images Based on a Deep Fully Convolutional Neural Network. Remote Sens. 2019, 11, 691. [Google Scholar] [CrossRef]
- Feng, A.; Sudduth, K.; Vories, E.; Zhou, J. Evaluation of cotton stand count using UAV-based hyperspectral imagery. In Proceedings of the 2019 ASABE Annual International Meeting, Boston, MA, USA, 7–10 July 2019. [Google Scholar] [CrossRef]
- Zhao, B.; Zhang, J.; Yang, C.; Zhou, G.; Ding, Y.; Shi, Y.; Zhang, D.; Xie, J.; Liao, Q. Rapeseed seedling stand counting and seeding performance evaluation at two early growth stages based on unmanned aerial vehicle imagery. Front. Plant Sci. 2018, 9, 1–17. [Google Scholar] [CrossRef]
- Varela, S.; Dhodda, P.R.; Hsu, W.H.; Prasad, P.V.V.; Assefa, Y.; Peralta, N.R.; Griffin, T.; Sharda, A.; Ferguson, A.; Ciampitti, I.A. Early-season stand count determination in Corn via integration of imagery from unmanned aerial systems (UAS) and supervised learning techniques. Remote Sens. 2018, 10, 343. [Google Scholar] [CrossRef]
- Burud, I.; Lange, G.; Lillemo, M.; Bleken, E.; Grimstad, L.; Johan From, P. Exploring Robots and UAVs as Phenotyping Tools in Plant Breeding. IFAC-PapersOnLine 2017, 50, 11479–11484. [Google Scholar] [CrossRef]
- Sankaran, S.; Khot, L.R.; Carter, A.H. Field-based crop phenotyping: Multispectral aerial imaging for evaluation of winter wheat emergence and spring stand. Comput. Electron. Agric. 2015, 118, 372–379. [Google Scholar] [CrossRef]
- Castro, W.; Junior, J.M.; Polidoro, C.; Osco, L.P.; Gonçalves, W.; Rodrigues, L.; Santos, M.; Jank, L.; Barrios, S.; Valle, C.; et al. Deep learning applied to phenotyping of biomass in forages with uav-based rgb imagery. Sensors 2020, 20, 4802. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Li, C.; Paterson, A.H.; Robertson, J.S. DeepSeedling: Deep convolutional network and Kalman filter for plant seedling detection and counting in the field. Plant Methods 2019, 15, 1–19. [Google Scholar] [CrossRef]
- Shrestha, D.S.; Steward, B.L. Automatic corn plant population measurement using machine vision. Trans. Am. Soc. Agric. Eng. 2003, 46, 559–565. [Google Scholar] [CrossRef]
- Kayacan, E.; Zhang, Z.; Chowdhary, G. Embedded High Precision Control and Corn Stand Counting Algorithms for an Ultra-Compact 3D Printed Field Robot. Robotics 2018. [Google Scholar] [CrossRef]
- Li, Z.; Guo, R.; Li, M.; Chen, Y.; Li, G. A review of computer vision technologies for plant phenotyping. Comput. Electron. Agric. 2020, 176, 105672. [Google Scholar] [CrossRef]
- Ristorto, G.; Gallo, R.; Gasparetto, A.; Scalera, L.; Vidoni, R.; Mazzetto, F. A mobile laboratory for orchard health status monitoring in precision farming. Chem. Eng. Trans. 2017, 58, 661–666. [Google Scholar] [CrossRef]
- de Oliveira, L.T.; de Carvalho, L.M.T.; Ferreira, M.Z.; de Andrade Oliveira, T.C.; Júnior, F.W.A. Application of LIDAR to forest inventory for tree count in stands of Eucalyptus sp. Cerne 2012, 18, 175–184. [Google Scholar] [CrossRef]
- Li, B.; Xu, X.; Han, J.; Zhang, L.; Bian, C.; Jin, L.; Liu, J. The estimation of crop emergence in potatoes by UAV RGB imagery. Plant Methods 2019, 15, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Praveen Kumar, J.; Domnic, S. Image based leaf segmentation and counting in rosette plants. Inf. Process. Agric. 2019, 6, 233–246. [Google Scholar] [CrossRef]
- Gavrilescu, R.; Fo, C.; Zet, C.; Cotovanu, D. Faster R-CNN: An Approach to Real-Time Object Detection. In Proceedings of the 2018 International Conference and Exposition on Electrical And Power Engineering (EPE), Iasi, Romania, 18–19 October 2018; pp. 2018–2021. [Google Scholar]
- He, K.; Gkioxari, G.; Dollár, P.; Girshick, R. Mask R-CNN. IEEE Trans. Pattern Anal. Mach. Intell. 2020, 42, 386–397. [Google Scholar] [CrossRef] [PubMed]
- Redmon, J.; Divvala, S.; Girshick, R.; Farhadi, A. You only look once: Unified, real-time object detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 26 June–1 July 2016; pp. 779–788. [Google Scholar]
- Redmon, J.; Farhadi, A. YOLOv3: An Incremental Improvement. arXiv 2018, arXiv:1804.02767. [Google Scholar]
- Redmon, J.; Farhadi, A. YOLO9000: Better, faster, stronger. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 6517–6525. [Google Scholar] [CrossRef]
- Khaki, S.; Pham, H.; Han, Y.; Kuhl, A.; Kent, W.; Wang, L. DeepCorn: A Semi-Supervised Deep Learning Method for High-Throughput Image-Based Corn Kernel Counting and Yield Estimation. arXiv 2020, arXiv:2007.10521. [Google Scholar]
- Zhu, X.; Zhu, M.; Ren, H. Method of plant leaf recognition based on improved deep convolutional neural network. Cogn. Syst. Res. 2018, 52, 223–233. [Google Scholar] [CrossRef]
- Quan, L.; Feng, H.; Lv, Y.; Wang, Q.; Zhang, C.; Liu, J.; Yuan, Z. Maize seedling detection under different growth stages and complex field environments based on an improved Faster R–CNN. Biosyst. Eng. 2019, 184, 1–23. [Google Scholar] [CrossRef]
- Fu, L.; Feng, Y.; Majeed, Y.; Zhang, X.; Zhang, J.; Karkee, M.; Zhang, Q. Kiwifruit detection in field images using Faster R-CNN with ZFNet. IFAC-PapersOnLine 2018, 51, 45–50. [Google Scholar] [CrossRef]
- Santos, T.T.; de Souza, L.L.; dos Santos, A.A.; Avila, S. Grape detection, segmentation, and tracking using deep neural networks and three-dimensional association. Comput. Electron. Agric. 2020, 170, 105247. [Google Scholar] [CrossRef]
- Fu, L.; Majeed, Y.; Zhang, X.; Karkee, M.; Zhang, Q. Faster R–CNN–based apple detection in dense-foliage fruiting-wall trees using RGB and depth features for robotic harvesting. Biosyst. Eng. 2020, 197, 245–256. [Google Scholar] [CrossRef]
- Shi, R.; Li, T.; Yamaguchi, Y. An attribution-based pruning method for real-time mango detection with YOLO network. Comput. Electron. Agric. 2020, 169, 105214. [Google Scholar] [CrossRef]
- Reis, M.J.C.S.; Morais, R.; Peres, E.; Pereira, C.; Contente, O.; Soares, S.; Valente, A.; Baptista, J.; Ferreira, P.J.S.G.; Bulas Cruz, J. Automatic detection of bunches of grapes in natural environment from color images. J. Appl. Log. 2012, 10, 285–290. [Google Scholar] [CrossRef]
- Liu, G.; Nouaze, J.C.; Mbouembe, P.L.T.; Kim, J.H. YOLO-tomato: A robust algorithm for tomato detection based on YOLOv3. Sensors 2020, 20, 2145. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Yang, G.; Wang, Z.; Wang, H.; Li, E.; Liang, Z. Apple detection during different growth stages in orchards using the improved YOLO-V3 model. Comput. Electron. Agric. 2019, 157, 417–426. [Google Scholar] [CrossRef]
- Fu, L.; Feng, Y.; Wu, J.; Liu, Z.; Gao, F.; Majeed, Y.; Al-Mallahi, A.; Zhang, Q.; Li, R.; Cui, Y. Fast and accurate detection of kiwifruit in orchard using improved YOLOv3-tiny model. Precis. Agric. 2020. [Google Scholar] [CrossRef]
- Rawla, P.; Sunkara, T.; Gaduputi, V.; Jue, T.L.; Sharaf, R.N.; Appalaneni, V.; Anderson, M.A.; Ben-Menachem, T.; Decker, G.A.; Fanelli, R.D.; et al. Plant-stand Count and Weed Identification Mapping Using Unmanned Aerial Vehicle Images. Gastrointest. Endosc. 2018, 10, 279–288. [Google Scholar] [CrossRef]
- Jin, X.; Liu, S.; Baret, F.; Hemerlé, M.; Comar, A. Estimates of plant density of wheat crops at emergence from very low altitude UAV imagery. Remote Sens. Environ. 2017, 198, 105–114. [Google Scholar] [CrossRef]
- Liu, S.; Baret, F.; Allard, D.; Jin, X.; Andrieu, B.; Burger, P.; Hemmerlé, M.; Comar, A. A method to estimate plant density and plant spacing heterogeneity: Application to wheat crops. Plant Methods 2017, 13, 1–11. [Google Scholar] [CrossRef]
- Abendroth, L.J.; Elomere, R.W.; Boyer, M.J.; Marlay, S.K. Corn Growth and Development; Iowa State Univ.: Ames, IA, USA, 2011. [Google Scholar]
- Torralba, A.; Russell, B.C.; Yuen, J. LabelMe: Online image annotation and applications. Proc. IEEE 2010, 98, 1467–1484. [Google Scholar] [CrossRef]
- Kalman, R.E. A new approach to linear filtering and prediction problems. J. Fluids Eng. Trans. ASME 1960, 82, 35–45. [Google Scholar] [CrossRef]
- Redmon, J. Darknet. Available online: http://pjreddie.com/darknet/ (accessed on 3 May 2020).
- Russakovsky, O.; Deng, J.; Su, H.; Krause, J.; Satheesh, S.; Ma, S.; Huang, Z.; Karpathy, A.; Khosla, A.; Bernstein, M.; et al. ImageNet Large Scale Visual Recognition Challenge. Int. J. Comput. Vis. 2015, 115, 211–252. [Google Scholar] [CrossRef]
- Li, X.; Wang, K.; Wang, W.; Li, Y. A multiple object tracking method using Kalman filter. In Proceedings of the 2010 IEEE International Conference on Information and Automation, Harbin, China, 20–23 June 2010; pp. 1862–1866. [Google Scholar] [CrossRef]
- Patel, H.A.; Thakore, D.G. Moving Object Tracking Using Kalman Filter. Int. J. Comput. Sci. Mob. Comput. 2013, 2, 326–332. [Google Scholar]
- Wojke, N.; Bewley, A.; Paulus, D. Simple online and realtime tracking with a deep association metric. In Proceedings of the 2017 IEEE International Conference on Image Processing (ICIP), Beijing, China, 17–20 September 2017; pp. 3645–3649. [Google Scholar] [CrossRef]
- Davies, D.; Palmer, P.L.; Mirmehdi, M. Detection and tracking of very small low-contrast objects. In Proceedings of the British Machine Vision Conference 1998, BMVC 1998, Southampton, UK, January 1998. [Google Scholar] [CrossRef]
- Blostein, S.; Huang, T. Detecting small, moving objects in image sequences using sequen- tial hypothesis testing. IEEE Trans. Signal Process. 1991, 39, 1611–1629. [Google Scholar] [CrossRef]
- Ffrench, P.A.; Zeidler, J.R.; Ku, W.H. Enhanced detectability ofsmall objects in correlated clutter using an improved 2-d adaptive lattice algorithm. IEEE Trans. IP 1997, 6, 383–397. [Google Scholar]





| Camera/ Platform | Study Object | Camera Height | Resolution | Detect /Classify | Count Performance | Reference |
|---|---|---|---|---|---|---|
| Hyperspectral /UAV | Cotton stand | 50 m | 1088 × 2048 | Segment according NDVI | Accuracy = 98% MAPE = 9% | Feng et al. (2019) [11] |
| RGB/UAV | Potato emergence | 30 m | 20 megapixel | Edge detection | R2 = 0.96 | Li et al. (2019) [23] |
| RGB/UAV | Rice seedling | 20 m | 5427 × 3648 | Simplified VGG-16 | Accuracy > 93% R2 = 0.94 | Wu et al., (2019) [10] |
| RGB/UAV | Winter wheat | 3~7 m | 6024 × 4024 | SVM | R2 = 0.73~0.91 RMSE = 21.66~80.48 | Jin et al. (2017) [42] |
| RGB/UAV | Rapeseed seeding stand | 20 m | 7360 × 4912 | Shape feature | R2 = 0.845 and 0.867 MAE = 9.79% and 5.11% | Zhao et al. (2018) [12] |
| RGB/UAS | Cotton seeding | 15~20 m | 4608 × 3456 | Supervised Maximum Likelihood Classifier | Average Accuracy = 88.6% | Chen et al. (2018) [8] |
| RGB/UAS | Corn early-season stand | 10 m | 6000 × 4000 | Supervised Learning | Classification Accuracy = 0.96 | Varela et al. [13] |
| Multispectral /UAV | Corn stand | 50 feet | 0.05 m ground spatial | “Search-Hands” Method | Accuracy > 99% | Rawla et al. (2018) [41] |
| RGB/Ground level (handheld, cart, and tractor) | Cotton seeding | 0.5 m | 1920 × 1080 | Faster RCNN Model | Accuracy = 93% R2 = 0.98 | Jiang et al. (2019) [17] |
| RGB/moving cart | Wheat density estimation | 1.5 m | 4608 × 3072 | Gamma Count Model | R2 = 0.83 RMSE = 48.27 | Liu et al. (2019) [43] |
| RGB/Hand-held camera | On-ear corn kernels | / | 1024 × 768 | Semi-Supervised Deep Learning | MAE = 44.91 RMSE = 65.92 | Khaki et al. (2020) [30] |
| mAP(%)@IoU0.5 | V1 | V2 | V3 | V4 | Average FPS |
|---|---|---|---|---|---|
| YoloV3 | 92.78 | 98.07 | 96.94 | 76.33 | 12.21 |
| YoloV3-tiny | 89.12 | 94.13 | 93.83 | 63.13 | 76.27 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, L.; Xiang, L.; Tang, L.; Jiang, H. A Convolutional Neural Network-Based Method for Corn Stand Counting in the Field. Sensors 2021, 21, 507. https://doi.org/10.3390/s21020507
Wang L, Xiang L, Tang L, Jiang H. A Convolutional Neural Network-Based Method for Corn Stand Counting in the Field. Sensors. 2021; 21(2):507. https://doi.org/10.3390/s21020507
Chicago/Turabian StyleWang, Le, Lirong Xiang, Lie Tang, and Huanyu Jiang. 2021. "A Convolutional Neural Network-Based Method for Corn Stand Counting in the Field" Sensors 21, no. 2: 507. https://doi.org/10.3390/s21020507
APA StyleWang, L., Xiang, L., Tang, L., & Jiang, H. (2021). A Convolutional Neural Network-Based Method for Corn Stand Counting in the Field. Sensors, 21(2), 507. https://doi.org/10.3390/s21020507





