1. Introduction
Traditional diagnosis of histological samples is based on manual examination of the morphological features of specimens by skilled pathologists. In recent years, the use of computer-aided technologies for aiding in this task is an emerging trend, with the main goal to reduce the intra and inter-observer variability [
1]. Such technologies are intended to improve the diagnosis of pathological slides to be more reproducible, increase objectivity, and save time in the routine examination of samples [
2,
3]. Although most of the research carried out in computational pathology has been in the context of RGB (red, green, and blue) image analysis [
4,
5,
6,
7], hyperspectral (HS) and multispectral imaging are shown as promising technologies to aid in the histopathological analysis of samples.
Hyperspectral imaging (HSI) is a technology capable of capturing both the spatial and spectral features of the materials that are imaged. Recently, this technology has proven to provide advantages in the diagnosis of different types of diseases [
8,
9,
10]. In the field of histopathological analysis, this technology has been used for different applications, such as the visualization of multiple biological markers within a single tissue specimen with inmunohistochemistry [
11,
12,
13], the digital staining of samples [
14,
15], or diagnosis.
The analysis of HS images is usually performed in combination with machine learning approaches [
16]. Traditionally, feature-based methods are used, such as supervised classifiers. Awan et al. performed automatic classification of colorectal tumor samples identifying four types of tissues: normal, tumor, hyperplastic polyp, and tubular adenoma with low-grade dysplasia. Using different types of feature extraction and band selection methods followed by support vector machines (SVM) classification, the authors found that the use of a higher number of spectral bands improved the classification accuracy [
17]. Wang et al. analyzed hematoxylin and eosin (H&E) skin samples to facilitate the diagnosis of melanomas. The authors proposed a customized spatial-spectral classification method, which provided an accurate identification of melanoma and melanocytes with high specificity and sensitivity [
18]. Ishikawa et al. presented a method for pancreatic tumor cell identification using HSI. They first proposed a method to remove the influence of the staining in the HS data, and then they applied SVM classification [
19].
Although these authors have proven the feasibility of the feature learning methods for the diagnosis of histopathological samples using HS information, the performance of these approaches may be improved by using deep learning (DL) schemes. DL approaches automatically learn from the data in which features are optimal for classification, potentially outperforming handcrafted features [
20]. In the case of HS images, both the spatial and spectral features are exploited simultaneously. Recently, only a few researchers employed DL for the classification of HS images for histopathological applications. Malon et al. proposed the use of a convolutional neural network (CNN) for the detection of mitotic cells within breast cancer specimens [
21]. Haj-Hassan et al. also used CNNs for the classification of colorectal cancer tissue, showing performance improvements compared to traditional feature learning approaches [
22].
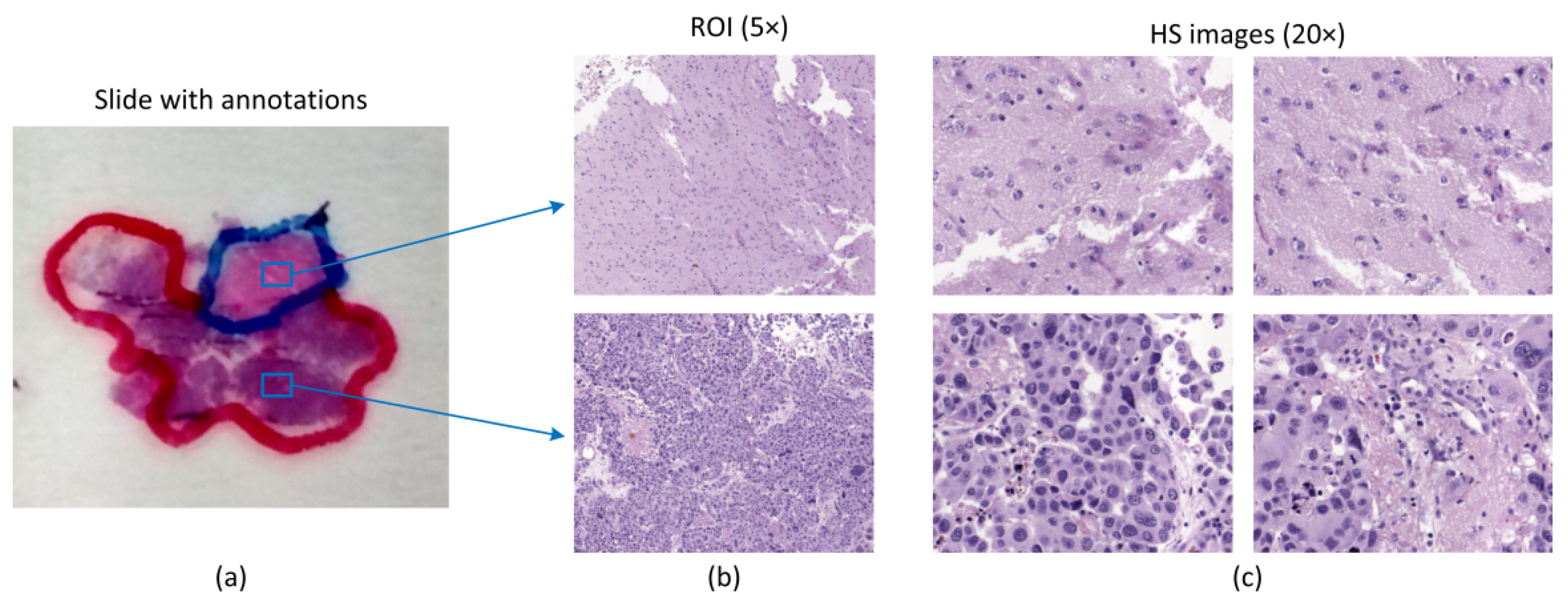
In this paper, we propose the use of CNNs for the classification of hematoxylin and eosin (H&E) stained brain tumor samples. Specifically, the main goal of this work was to differentiate between high-grade gliomas (i.e., glioblastoma (GB)) and non-tumor tissue. In a previous study, we presented a feature learning approach for this type of disease [
23]. Although such research was shown as a useful proof-of-concept on the possibilities of HS for histopathological analysis of GB, it presented some limitations, such as poor spatial and spectral resolution, and the lack of a rigorous experimental design. In this work, the image quality and spectral range have been significantly improved, resulting in a more appropriate experimental design for realistic clinical applications.
4. Discussion and Conclusion
In this research work, we present a hyperspectral microscopic imaging system and a deep learning approach for the classification of hyperspectral images of H&E pathological slides of brain tissue samples with glioblastoma of human patients.
As described in the introduction, this research is the continuation of a previous research that presented some drawbacks [
23]. Firstly, the total number of HS cubes used in our previous work was limited to 40, only 4 HS cubes per patient. Secondly, the instrumentation used in this previous work presented limitations in both the spectral and spatial information. Regarding the spectral information, the spectral range was restricted to 419–768 nm due to limitations of the microscope optical path. The spatial information was limited due to the use of a scanning platform unable to image the complete scene, so the analysis of the HS images was restricted to a low magnification (5×), which was not sufficient to image the morphological features of the sample. Additionally, the main goal of such previous work was to develop a preliminary proof-of-concept on the use of HSI for the differentiation of tumor and non-tumor samples, showing promising results.
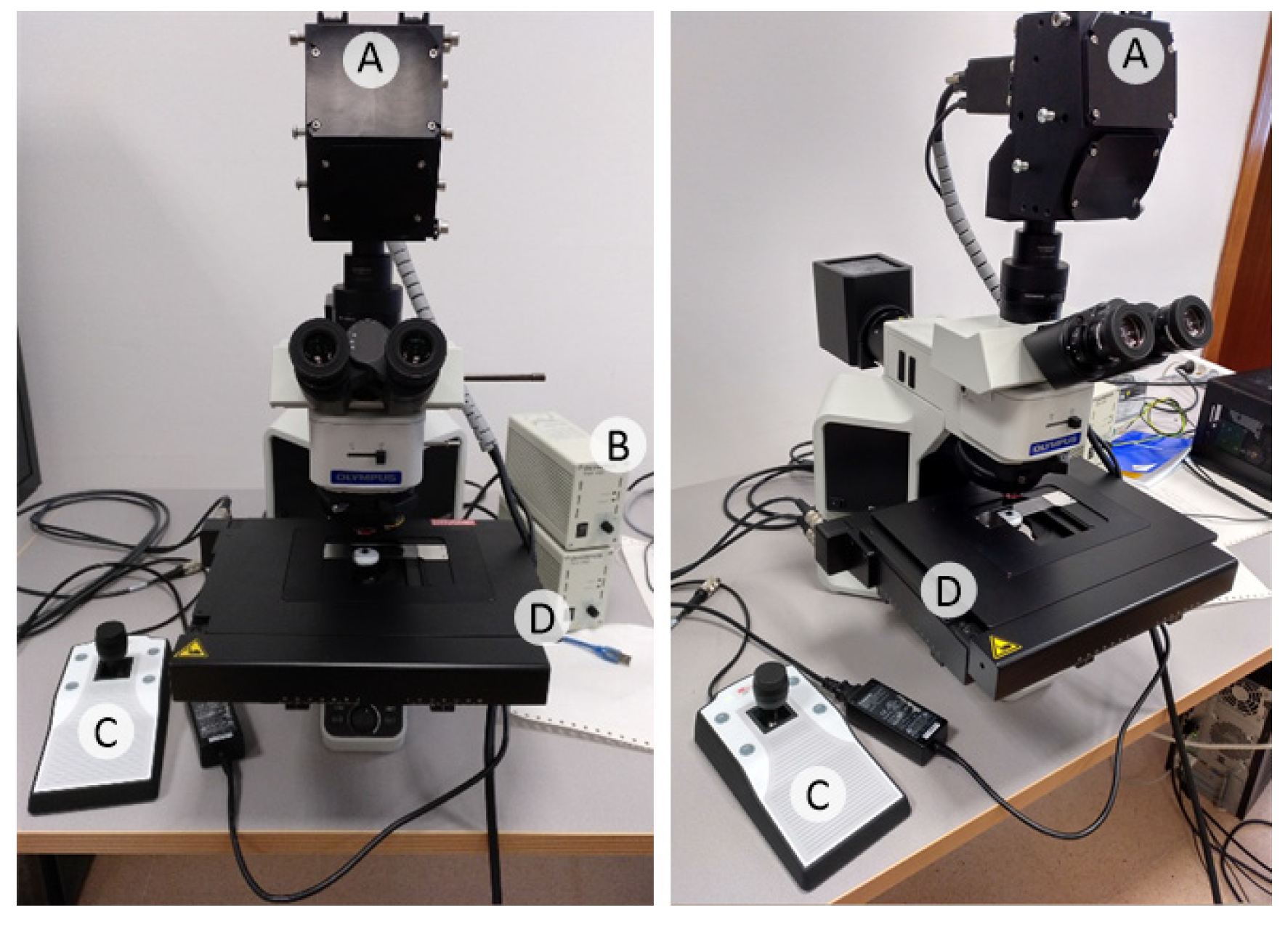
In this work, an improved acquisition system capable of capturing high-quality images in a higher magnification (20×), and with a higher spectral range (400–1000 nm) has been used to capture a total amount of 517 HS cubes. The use of 20× magnification allows the classifier to exploit both the spectral and the spatial differences of the samples to make a decision.
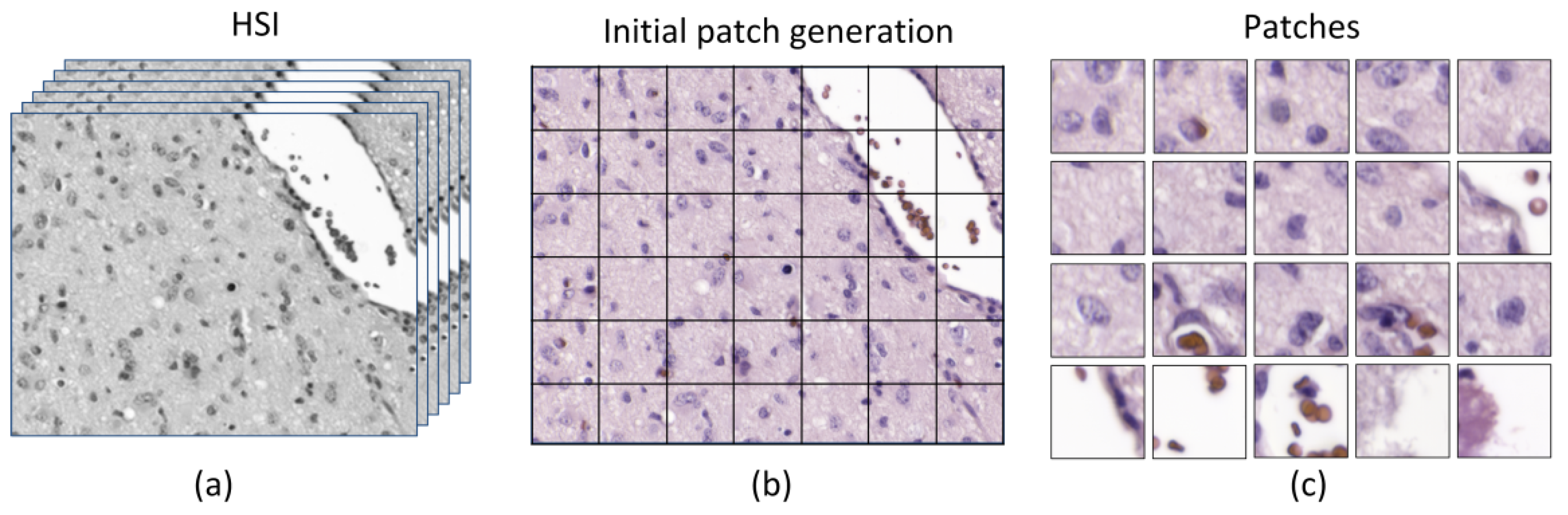
Such dataset was then used to train a CNN and to perform the classification between non-tumor and tumor tissue. Due to a limited number of patients involved in this study and with the aim to provide a data partition scheme with minimum bias, we decided to split the dataset in four different folds where the training, validation, and testing data belonged to different patients. Each fold was trained with 9 patients, where only 5 of them presented both types of samples, i.e., tumor and non-tumor tissue.
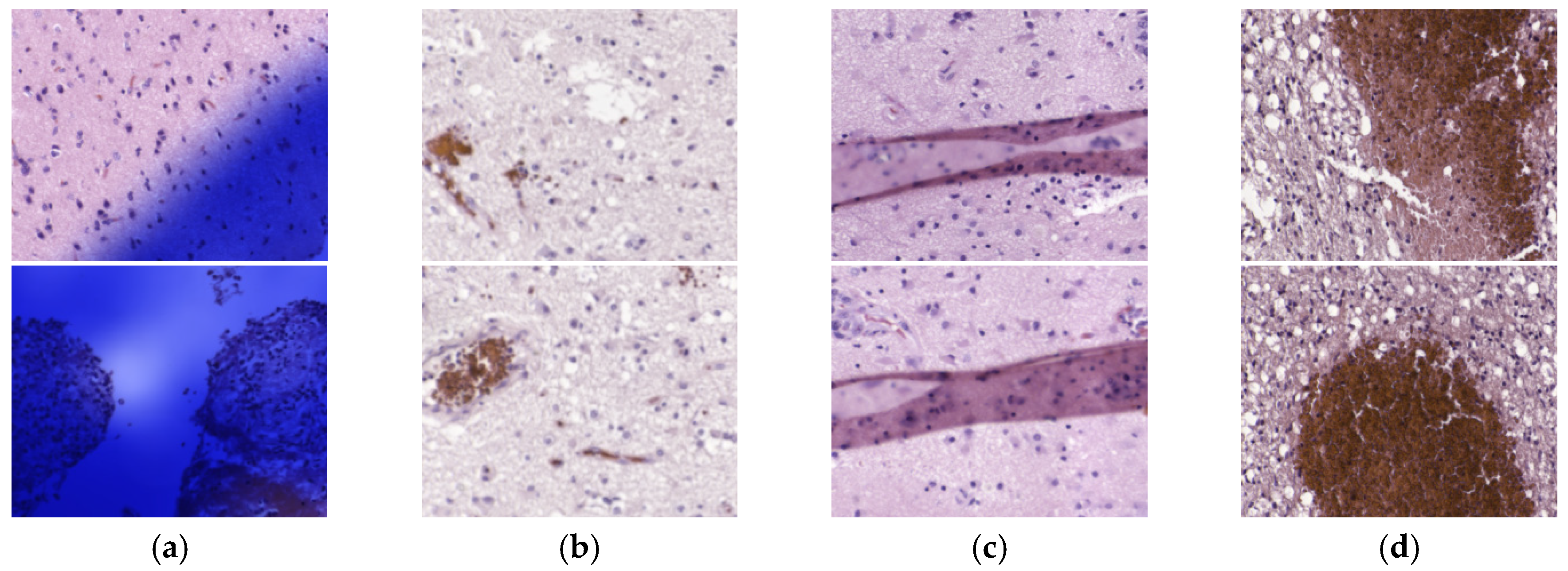
After selecting models with high AUC and balanced accuracy, sensitivity and specificity in the validation phase, some results on the test set were not accurate at all. For this reason, we carefully inspected the heat maps generated by the classifiers for each patient in order to find a rationale about the inaccurate results. After this, we detected four types of problems in the images that could worsened the results, namely the presence of ink or artifacts in the images, unfocused images, or excess of red blood cells. We reported both results, before and after cleaning wrong HS images, for a fair experimental design. We consider that the test results after removing such defective HS images are not biased because the rationale of removing the images from the test set is justifiable and transparent. These corrupted images were part of the training set, but it is unknown if the training process of the CNN was affected.
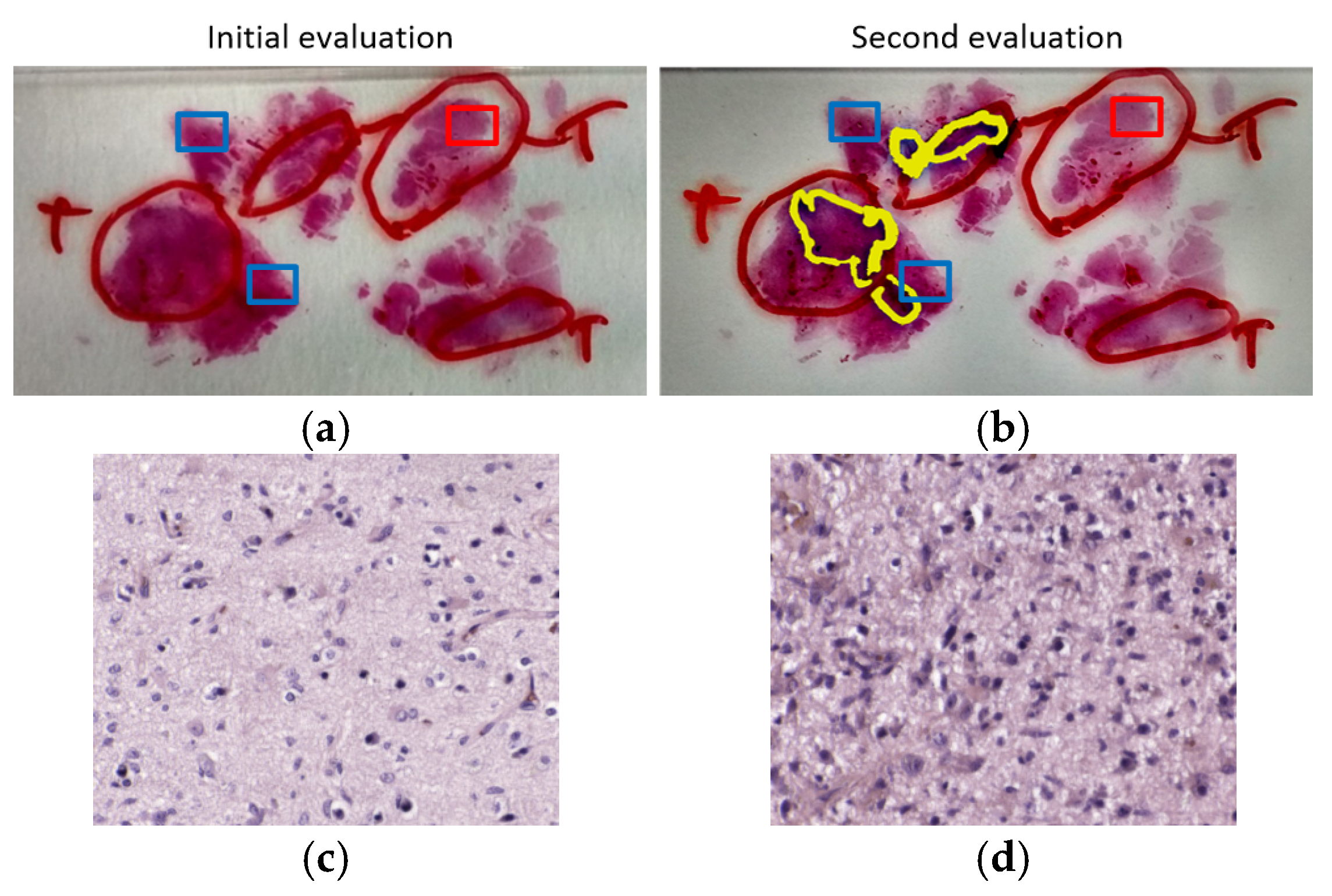
We also found a patient, P6, where the results were really inaccurate. For this reason, the regions of interest that were analyzed by HS were re-examined by the pathologists. After examining the sample, an atypical subtype of GB was found, and examination revealed that the ROIs selected as normal samples were close to the tumor area, which cannot be considered as non-tumor. Although the classification results were not valid for this patient, by using the outcomes of the CNN, we were able to identify a problem with the prior examination and ROI selection within the sample. Additionally, although this patient was used both as part of the training and as a patient used for validation, the results are not significantly affected by this fact. These results highlight the robustness of the CNN for tumor classification. Firstly, although the validation results of fold 1 were good when evaluating patient P6, the model from this fold was also capable of accurately classifying patients P1 and P11. Secondly, although patient P6 was used as training data for fold 3 and fold 4, the outcomes of these models were not proven to be significantly affected by contaminated training data.
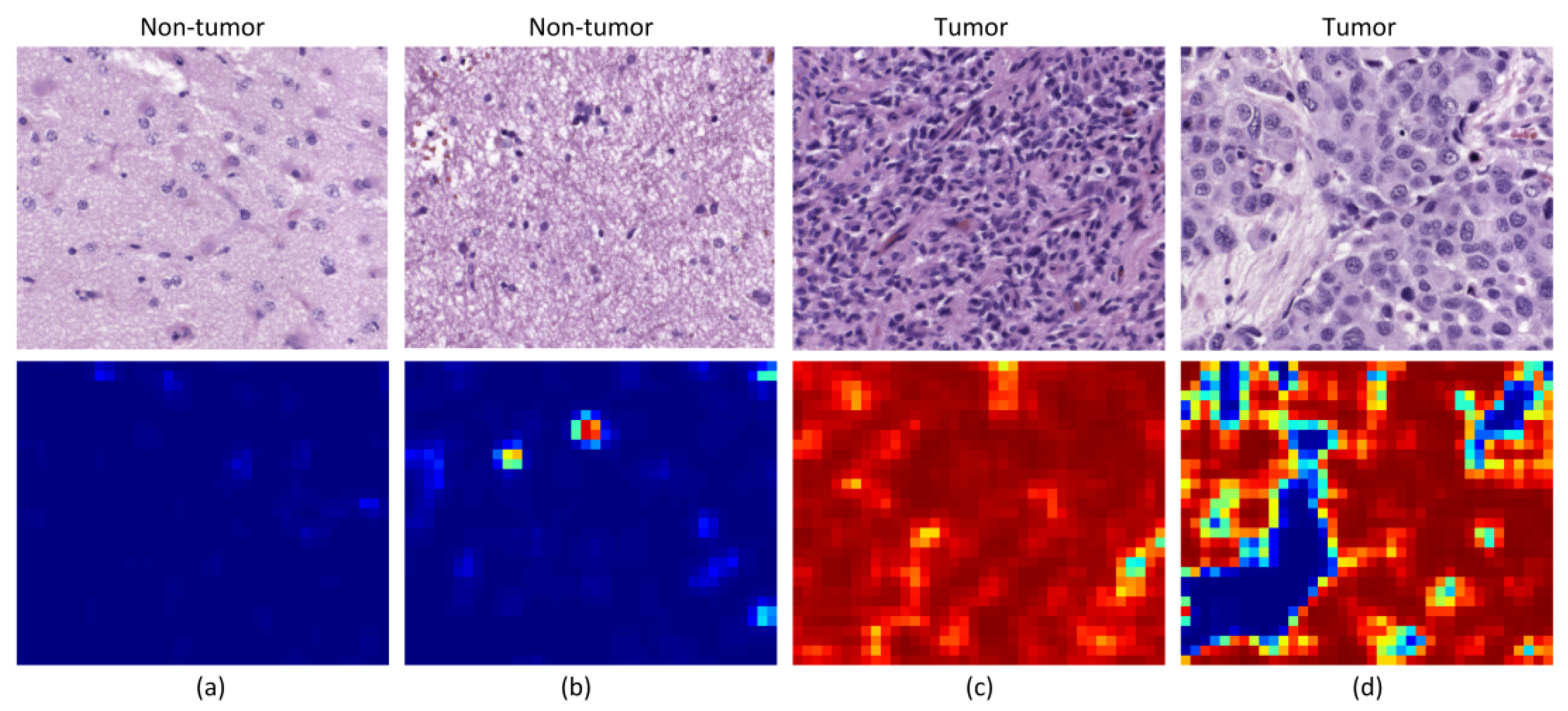
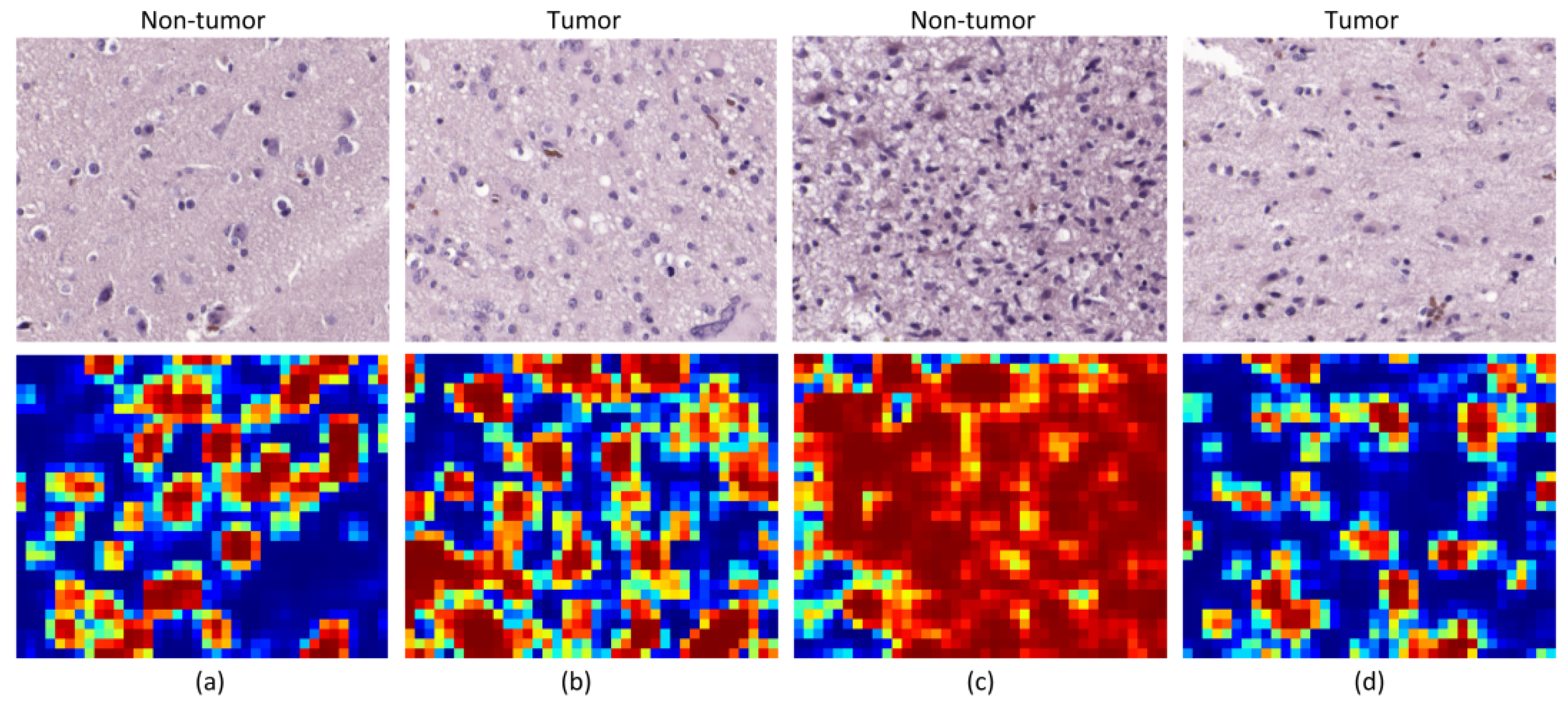
Although the results are not accurate in every patient, after excluding incorrectly labeled and contaminated HS images, nine patients showed accurate classification results (P1 to P3 and P8 to P13). Two patients provided acceptable results (P5 and P7), and only a single patient presented results that were slightly better than random guessing (P4). Nevertheless, these results can be considered promising for two main reasons. First, a limited number of samples were used for training, especially for the non-tumor class, which was limited to only five training patients for each CNN. Second, the high inter-patient variability shows significant differences between tumor samples among the different patients. As can be observed in the analysis of heat maps (
Figure 7 and
Figure 8), there is a significant heterogeneity in cellular morphology in different patients’ specimens, which makes GB detection an especially challenging application. To handle these challenges, the number of patients should be increased in future works, and to deal with the high inter-patient variability, HS data from more than a single patient should be used to validate the models.
Finally, we found that HSI data perform slightly better than RGB images for the classification. Such improvement is more evident when the classification is performed on challenging patients (e.g., P5 or P7) or in patients with only tumor samples. Furthermore, the classification results of HSI are shown to provide more balanced sensitivity and specificity, which is the goal for clinical applications, improving the average sensitivity and specificity by 7% and 9% with respect to the RGB imaging results, respectively. Nevertheless, more research should be performed to definitively demonstrate the superiority of HSI over conventional RGB imagery.