Liver Tumor Segmentation in CT Scans Using Modified SegNet
Abstract
1. Introduction
2. Basic Concepts
3. Materials and Method
3.1. Dataset
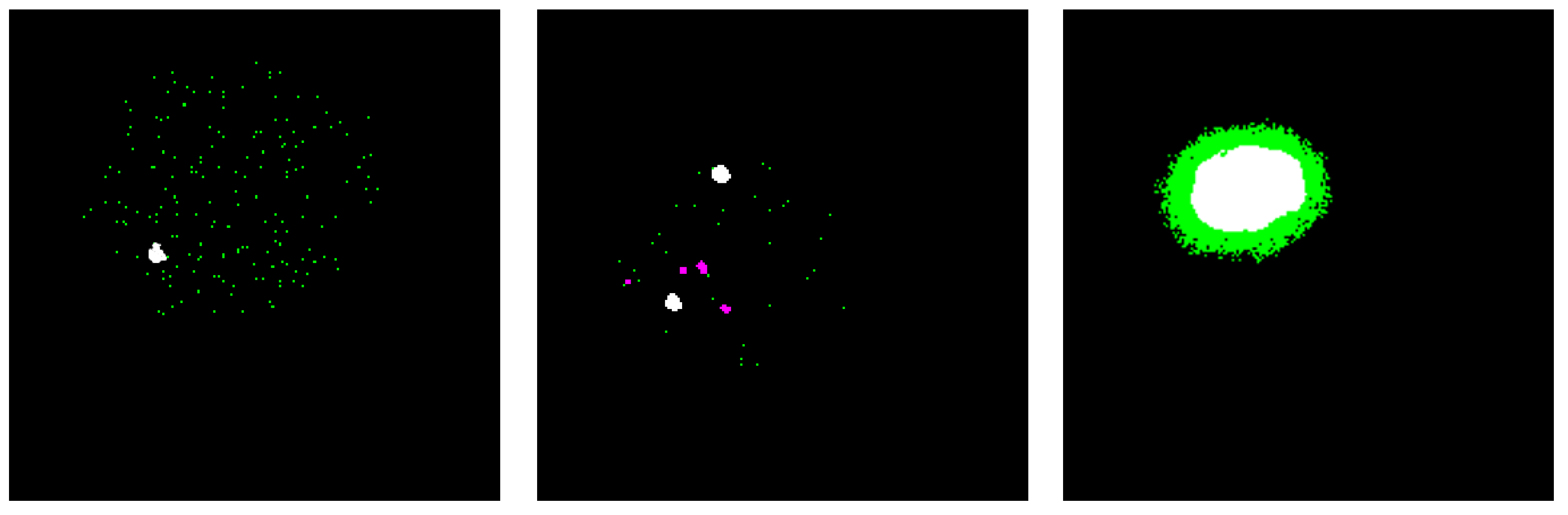
3.2. Image Preprocessing
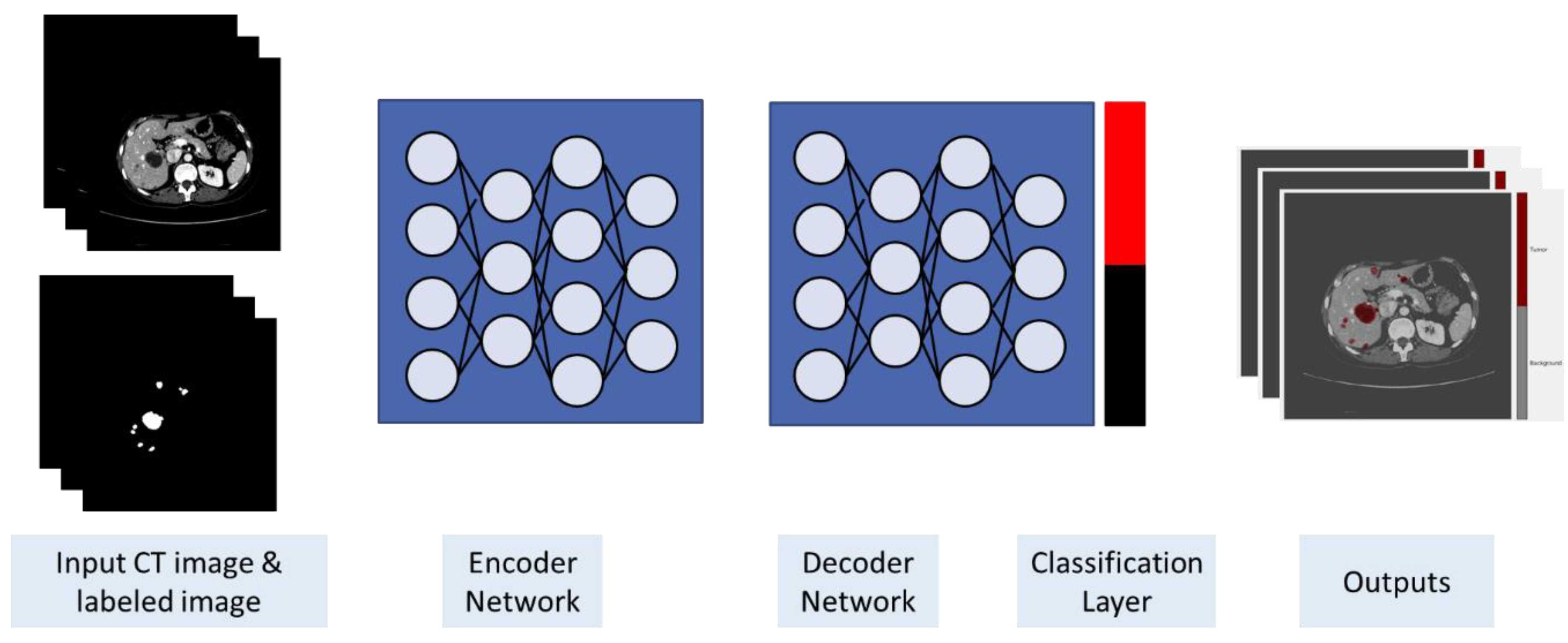
3.3. Training and Classification
4. Experimental Results
4.1. Evaluation Metrics
4.2. Data Set and Preprocessing
4.3. Training and Classification
4.4. Testing and Evaluation
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Al-Shaikhli, S.D.S.; Yang, M.Y.; Rosenhahn, B. Automatic 3d liver segmentation using sparse representation of global and local image information via level set formulation. arXiv, 2015; arXiv:abs/1508.01521. [Google Scholar]
- Whitcomb, E.; Choi, W.T.; Jerome, K.R.; Cook, L.; Landis, C.; Ahn, J.; Westerhoff, M. Biopsy Specimens From Allograft Liver Contain Histologic Features of Hepatitis C Virus Infection After Virus Eradication. Clin. Gastroenterol. Hepatol. 2017, 15, 1279–1285. [Google Scholar] [CrossRef] [PubMed]
- Woźniak, M.; Połap, D. Bio-Inspired methods modeled for respiratory disease detection from medical images. Swarm Evol. Comput. 2018, 41, 69–96. [Google Scholar] [CrossRef]
- Mohamed, A.S.E.; Mohammed Salem, A.-M.; Hegazy, D.; Howida Shedeed, A. Improved Watershed Algorithm for CT Liver Segmentation Using Intraclass Variance Minimization. ICIST 2017. [Google Scholar] [CrossRef]
- Bellver, M.; Maninis, K.-K.; Pont-Tuset, J.; Nieto, X.G.; Torres, J.; Gool, L.V. Detection-Aided liver lesion segmentation using deep learning. Machine Learning 4 Health Workshop - NIPS. arXiv, 2017; arXiv:abs/1711.11069. [Google Scholar]
- Ben-Cohen, A.; Klang, E.; Amitai, M.M.; Goldberger, J.; Greenspan, H. Anatomical data augmentation for CNN based pixel-wise classification. In Proceedings of the IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), Washington, DC, USA, 4–7 April 2018; pp. 1096–1099. [Google Scholar]
- Bi, L.; Kim, J.; Kumar, A.; Feng, D. Automatic liver lesion detection using cascaded deep residual networks. arXiv, 2017; arXiv:abs/1704.02703. [Google Scholar]
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global cancer statistics. CA Cancer J. Clin. 2017, 61, 60–90. [Google Scholar]
- World Health Organization. Available online: http://www.emro.who.int/media/news/world-hepatitis-day-in-egypt-focuses-on-hepatitis-b-and-c-prevention (accessed on 24 February 2020).
- World Health Organization Hepatitis. Available online: https://www.who.int/hepatitis/news-events/egypt-hepatitis-c-testing/en/ (accessed on 24 February 2020).
- Chlebus, G.; Meine, H.; Moltz, J.H.; Schenk, A. Neural network-Based automatic liver tumor segmentation with random forest-Based candidate filtering. arXiv, 2017; arXiv:abs/1706.00842. [Google Scholar]
- Christ, P.F.; Elshaer, M.E.A.; Ettlinger, F.; Tatavarty, S.; Bickel, M.; Bilic, P.; Rempfler, M.; Armbruster, M.; Hofmann, F.; D’Anastasi, M.; et al. Automatic liver and lesion segmentation in CT using cascaded fully convolutional neural networks and 3d conditional random fields. arXiv, 2016; arXiv:abs/1610.02177. [Google Scholar]
- Christ, P.F.; Ettlinger, F.; Grün, F.; Elshaer, M.E.A.; Lipkova, J.; Schlecht, S.; Ahmaddy, F.; Tatavarty, S.; Bickel, M.; Bilic, P.; et al. Automatic liver and tumor segmentation of CT and MRI volumes using cascaded fully convolutional neural networks. arXiv, 2017; arXiv:abs/1702.05970. [Google Scholar]
- Wei, W.; Zhou, B.; Połap, D.; Woźniak, M. A regional adaptive variational PDE model for computed tomography image reconstruction. Pattern Recognit. 2019, 92, 64–81. [Google Scholar] [CrossRef]
- Varma, D.R. Managing DICOM images: Tips and tricks for the radiologist. Indian J. Radiol. Imaging 2012, 22, 4–13. [Google Scholar] [CrossRef] [PubMed]
- The American Cancer Society medical and editorial content team. Liver Cancer Early Detection, Diagnosis, and Staging. Available online: https://www.cancer.org/content/dam/CRC/PDF/Public/8700.00.pdf (accessed on 22 July 2019).
- Wozniak, M.; Polap, D.; Capizzi, G.; Sciuto, G.L.; Kosmider, L.; Frankiewicz, K. Small lung nodules detection based on local variance analysis and probabilistic neural network. Comput. Methods Programs Biomed. 2018, 161, 173–180. [Google Scholar] [CrossRef]
- Mohamed, A.S.E.; Salem, M.A.-.M.; Hegazy, D.; Shedeed, H. Probablistic-Based framework for medical CT images segmentation. In Proceedings of the 2015 IEEE Seventh International Conference on Intelligent Computing and Information Systems (ICICIS), Cairo, Egypt, 12–14 December 2015; pp. 149–155. [Google Scholar] [CrossRef]
- Salem, M.A.; Meffert, B. Resolution mosaic EM algorithm for medical image segmentation. In Proceedings of the 2009 International Conference on High Performance Computing & Simulation, Leipzig, Germany, 21–24 June 2009; pp. 208–215. [Google Scholar] [CrossRef]
- El-Regaily, S.A.; Salem, M.A.M.; Abdel Aziz, M.H.; Roushdy, M.I. Multi-View convolutional neural network for lung nodule false positive reduction. Expert Syst. Appl. 2019. [Google Scholar] [CrossRef]
- Atteya, M.A.; Salem, M.A.-M.; Hegazy, D.; Roushdy, M.I. Image segmentation and particles classification using texture analysis method. Rev. Bras. de Eng. Biomed. 2016, 32, 243–252. [Google Scholar] [CrossRef][Green Version]
- Han, X. Automatic liver lesion segmentation using A deep convolutional neural network method. arXiv, 2017; arXiv:abs/1704.07239. [Google Scholar]
- National Institute of Biomedical Imaging and Bioengineering (NIBIB). Computed Tomography (CT), URL. Available online: https://www.nibib.nih.gov/science-education/science-topics/computed-tomography-ct (accessed on 18 May 2019).
- Heimann, T.; Van Ginneken, B.; Styner, M. Sliver07 Grand Challenge. Available online: https://sliver07.grand-challenge.org/Home/ (accessed on 20 July 2019).
- Heimann, T.; van Ginneken, B.; Styner, M.A.; Arzhaeva, Y.; Aurich, V.; Bauer, C.; Beck, A.; Becker, C.; Beichel, R.; Bekes, G.; et al. Comparison and evaluation of methods for liver segmentation from ct datasets. IEEE Trans. Med. Imaging 2009, 28, 1251–1265. [Google Scholar] [CrossRef]
- Salem, M.A.-M.; Atef, A.; Salah, A.; Shams, M. Recent survey on medical image segmentation. Comput. Vis. Concepts Methodol. Tools Appl. 2018, 129–169. [Google Scholar] [CrossRef]
- Horn, Z.C.; Auret, L.; Mccoy, J.T.; Aldrich, C.; Herbst, B.M. Performance of convolutional neural networks for feature extraction in froth flotation sensing. IFAC Pap. Line 2017, 50, 13–18. [Google Scholar] [CrossRef]
- Li, W.; Jia, F.; Hu, Q. Automatic segmentation of liver tumor in CT images with deep convolutional neural networks. J. Comput. Commun. 2015, 3, 146–151. [Google Scholar] [CrossRef]
- Li, X.; Chen, H.; Qi, X.; Dou, Q.; Fu, C.; Heng, P. H-DenseUNet: Hybrid Densely Connected UNet for Liver and Tumor Segmentation From CT Volumes. IEEE Trans. Med. Imaging 2018, 37, 2663–2674. [Google Scholar] [CrossRef]
- Rajeshwaran, M.; Ahila, A. Segmentation of Liver Cancer Using SVM Techniques. Int. J. Comput. Commun. Inf. Syst. (IJCCIS) 2014, 6, 78–80. [Google Scholar]
- El-Regaily, S.A.; Salem, M.A.M.; Aziz, M.H.A.; Roushdy, M.I. Lung nodule segmentation and detection in computed tomography. In Proceedings of the 8th IEEE International Conference on Intelligent Computing and Information Systems (ICICIS), Cairo, Egypt, 5–7 December 2017; pp. 72–78. [Google Scholar] [CrossRef]
- Yuan, Y. Hierarchical convolutional-Deconvolutional neural networks for automatic liver and tumor segmentation. arXiv, 2017; arXiv:abs/1710.04540. [Google Scholar]
- Yang, D.; Xu, D.; Zhou, S.K.; Georgescu, B.; Chen, M.; Grbic, S.; Metaxas, D.N.; Comaniciu, D. Automatic liver segmentation using an adversarial image-To-Image network. arXiv, 2017; arXiv:abs/1707.08037. [Google Scholar]
- Shafaey, M.A.; Salem, M.A.-M.; Ebied, H.M.; Al-Berry, M.N.; Tolba, M.F. Deep Learning for Satellite Image Classification. In Proceedings of the 3rd International Conference on Intelligent Systems and Informatics (AISI2018), Cairo, Egypt, 1–3 September 2018. [Google Scholar]
- Sutskever, I.; Vinyals, O.; Le, Q.V. Sequence to sequence learning with neural networks. arXiv, 2014; arXiv:abs/1409.3215. [Google Scholar]
- Ke, Q.; Zhang, J.; Wei, W.; Damaševĭcius, R.; Wozniak, M. Adaptive Independent Subspace Analysis of Brain Magnetic Resonance Imaging Data. IEEE Access 2019, 7, 12252–12261. [Google Scholar] [CrossRef]
- Badrinarayanan, V.; Kendall, A.; Cipolla, R. SegNet: A Deep Convolutional Encoder-Decoder Architecture for Image Segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2481–2495. [Google Scholar] [CrossRef]
- Ahmed, A.H.; Salem, M.A. Mammogram-Based cancer detection using deep convolutional neural networks. In Proceedings of the 2018 13th International Conference on Computer Engineering and Systems (ICCES), Cairo, Egypt, 18–19 December 2018; pp. 694–699. [Google Scholar] [CrossRef]
- Lu, F.; Wu, F.; Hu, P.; Peng, Z.; Kong, D. Automatic 3d liver location and segmentation via convolutional neural networks and graph cut. Int. J. Comput. Assist Radiol. Surg. 2017, 12, 171–182. [Google Scholar] [CrossRef]
- Wang, P.; Qian, Y.; Soong, F.K.; He, L.; Zhao, H. Part-Of-Speech tagging with bidirectional long short-Term memory recurrent neural network. arXiv, 2015; arXiv:abs/1510.06168. [Google Scholar]
- Abdelhalim, A.; Salem, M.A. Intelligent Organization of Multiuser Photo Galleries Using Sub-Event Detection. In Proceedings of the 12th International Conference on Computer Engineering & Systems (ICCES), Cairo, Egypt, 19–20 December 2017; Available online: https://ieeexplore.ieee.org/document/347/ (accessed on 22 January 2020).
- Elsayed, O.A.; Marzouky, N.A.M.; Atefz, E.; Mohammed, A.-S. Abnormal Action detection in video surveillance. In Proceedings of the 9th IEEE International Conference on Intelligent Computing and Information Systems (ICICIS), Cairo, Egypt, 8–10 December 2019. [Google Scholar]
- El-Masry, M.; Fakhr, M.; Salem, M. Action Recognition by Discriminative EdgeBoxes. IET Comput. Vis. Dec. 2017. Available online: http://digital-library.theiet.org/content/journals/10.1049/iet-cvi.2017.0335 (accessed on 22 January 2020). [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for largescale image recognition. arXiv, 2014; arXiv:abs/1409.1556. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. arXiv, 2015; arXiv:abs/1505.04597. [Google Scholar]
- Ke, Q.; Zhang, J.; Wei, W.; Połap, D.; Woźniak, M.; Kośmider, L.; Damaševĭcius, R. A neuro-Heuristic approach for recognition of lung diseases from X-Ray images. Expert Syst. Appl. 2019, 126, 218–232. [Google Scholar] [CrossRef]
- Noh, H.; Hong, S.; Han, B. Learning Deconvolution Network for Semantic Segmentation. In Proceedings of the 2015 IEEE International Conference on Computer Vision (ICCV), Santiago, Chile, 7–13 December 2015; pp. 1520–1528. [Google Scholar]
- Vadera, S.; Jone, J. Couinaud classification of hepatic segments. Available online: https://radiopaedia.org/articles/couinaud-classification-of-hepatic-segments (accessed on 28 November 2019).
- Zayene, O.; Jouini, B.; Mahjoub, M.A. Automatic liver segmentation method in CT images. arXiv, 2012; arXiv:abs/1204.1634. [Google Scholar]
- IRCAD. Hôpitaux Universitaires. 1, place de l’Hôpital, 67091 Strasbourg Cedex, FRANCE. Available online: https://www.ircad.fr/contact/ (accessed on 2 February 2020).
- Golan, R.; Jacob, C.; Denzinger, J. Lung nodule detection in CT images using deep convolutional neural networks. In Proceedings of the 2016 International Joint Conference on Neural Networks (IJCNN), Vancouver, BC, Canada, 24–29 July 2016; pp. 243–250. [Google Scholar]
- El-Regaily, S.A.; Salem, M.A.; Abdel Aziz, M.H.A.; Roushdy, M.I. Survey of Computer Aided Detection Systems for Lung Cancer in Computed Tomography. Current. Med. Imaging 2018, 14, 3–18. [Google Scholar] [CrossRef]
- Perez, L.; Wang, J. The Effectiveness of Data Augmentation in Image Classification Using Deep Learning. arXiv, 2017; arXiv:abs/1712.04621. [Google Scholar]
- Sun, C.; Guo, S.; Zhang, H.; Li, J.; Chen, M.; Ma, S.; Jin, L.; Liu, X.; Li, X.; Qian, X. Automatic segmentation of Liver tumors from multiphase contrast-Enhanced CT images based on FCN. Artificial Intell. Med. 2017, 83, 58–66. [Google Scholar] [CrossRef] [PubMed]
- Badura, P.; Wieclawe, W. Calibrating level set approach by granular computing in computed tomography abdominal organs segmentation. Appl. Soft Comput. 2016, 49, 887–900. [Google Scholar] [CrossRef]
- Shi, C.; Cheng, Y.; Liu, F.; Wang, Y.; Bai, J.; Tamura, S. A hierarchical local region-Based sparse shape composition for liver segmentation in CT scans. Pattern Recognit. 2015. [Google Scholar] [CrossRef]
- Hesamian, M.H.; Jia, W.; He, X.; Kennedy, P. Deep Learning Techniques for Medical Image Segmentation: Achievements and challenges. J. Digit. Imaging 2019. [Google Scholar] [CrossRef] [PubMed]
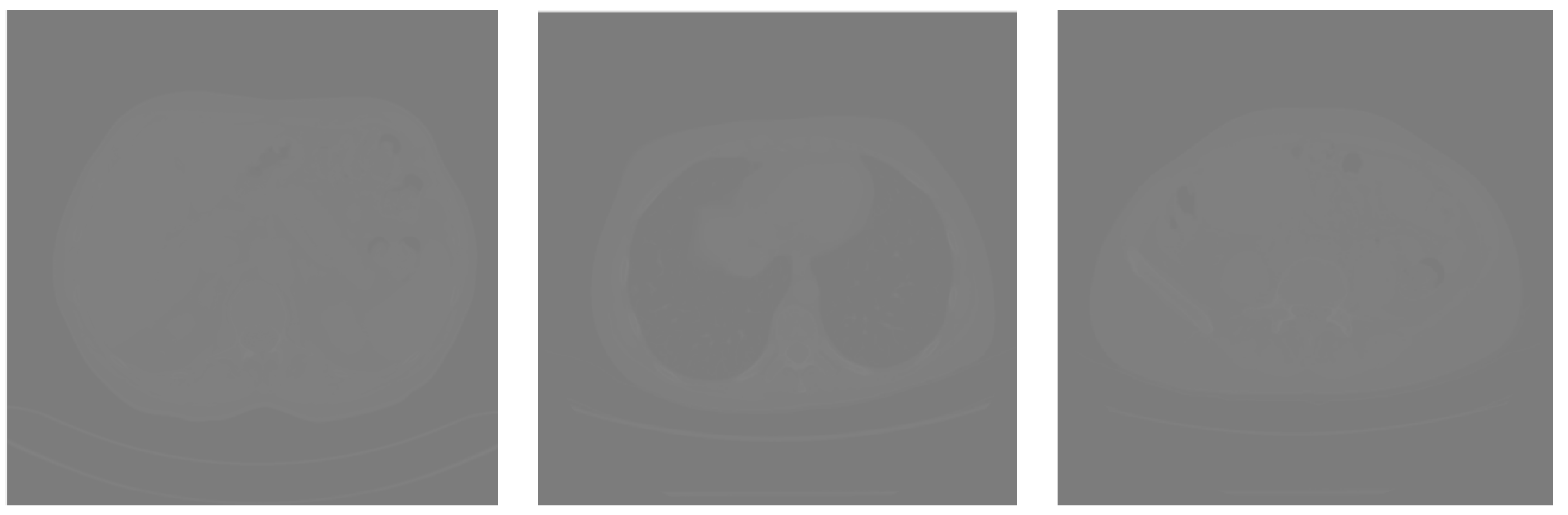





| Predicted | |||
|---|---|---|---|
| Positive | Negative | ||
| Actual | Positive | TP | FN |
| Negative | FP | TN | |
| # Training Images | Overall Ac. | Mean Ac. | Tumor Ac. (TPR) | Background Ac. (TNR) | Weighted IoU | F1-Score | |
|---|---|---|---|---|---|---|---|
| Case 1 | 70 | 91.27% | 93.43% | 95.58% | 91.27% | 91.26% | 21 91% |
| Case 2 | 124 | 93.36% | 95.18% | 97.34% | 93.03% | 92.99% | 46.58% |
| Case 3 | 260 | 94.57% | 97.26% | 99.99% | 94.52% | 93.72% | 62.16% |
| Case 1 | Predicted | ||
|---|---|---|---|
| Tumor | Others | ||
| Actual | Tumor | 45.82% | 54.28% |
| Others | 0.31% | 99.69% | |
| Case 1 | Predicted | ||
|---|---|---|---|
| Tumor | Others | ||
| Actual | Tumor | 67.13% | 32.87% |
| Others | 3.04% | 96.95% | |
| Case 1 | Predicted | ||
|---|---|---|---|
| Tumor | Others | ||
| Actual | Tumor | 99.99% | 0% |
| Others | 5.48% | 94.52% | |
| Author (s) | Application | Method | Accuracy |
|---|---|---|---|
| Chlebus [11] | Liver tumor candidate classification | Random Forest | 90% |
| Christ [12] | Automatic liver and tumor segmentation of CT and MRI | Cascaded fully convolutional neural networks (CFCNs) with dense 3D conditional random fields (CRFs) | 94% |
| Yang [33] | Liver segmentation of CT volumes | A deep image-to-image network (DI2IN) | 95% |
| Bi [7] | Liver segmentation | Deep residual network (Cascaded ResNet) | 95.9% |
| Li [46] | Liver and tumor segmentation from CT Volumes | H-DenseUNet | 96.5% |
| Yuan [32] | Automatic liver and tumor segmentation | Hierarchical Convolutional—Deconvolutional Neural Networks | 96.7% |
| Wen Li et al. [28] | Patch-based liver tumor classification | Conventional Convolutional Neural Network (CNN) | 80.6% |
| Our method | Liver tumor in CT Scans segmentation | Modified SegNet | 98.8% |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Almotairi, S.; Kareem, G.; Aouf, M.; Almutairi, B.; Salem, M.A.-M. Liver Tumor Segmentation in CT Scans Using Modified SegNet. Sensors 2020, 20, 1516. https://doi.org/10.3390/s20051516
Almotairi S, Kareem G, Aouf M, Almutairi B, Salem MA-M. Liver Tumor Segmentation in CT Scans Using Modified SegNet. Sensors. 2020; 20(5):1516. https://doi.org/10.3390/s20051516
Chicago/Turabian StyleAlmotairi, Sultan, Ghada Kareem, Mohamed Aouf, Badr Almutairi, and Mohammed A.-M. Salem. 2020. "Liver Tumor Segmentation in CT Scans Using Modified SegNet" Sensors 20, no. 5: 1516. https://doi.org/10.3390/s20051516
APA StyleAlmotairi, S., Kareem, G., Aouf, M., Almutairi, B., & Salem, M. A.-M. (2020). Liver Tumor Segmentation in CT Scans Using Modified SegNet. Sensors, 20(5), 1516. https://doi.org/10.3390/s20051516





