3D Deep Learning on Medical Images: A Review
Abstract
1. Introduction
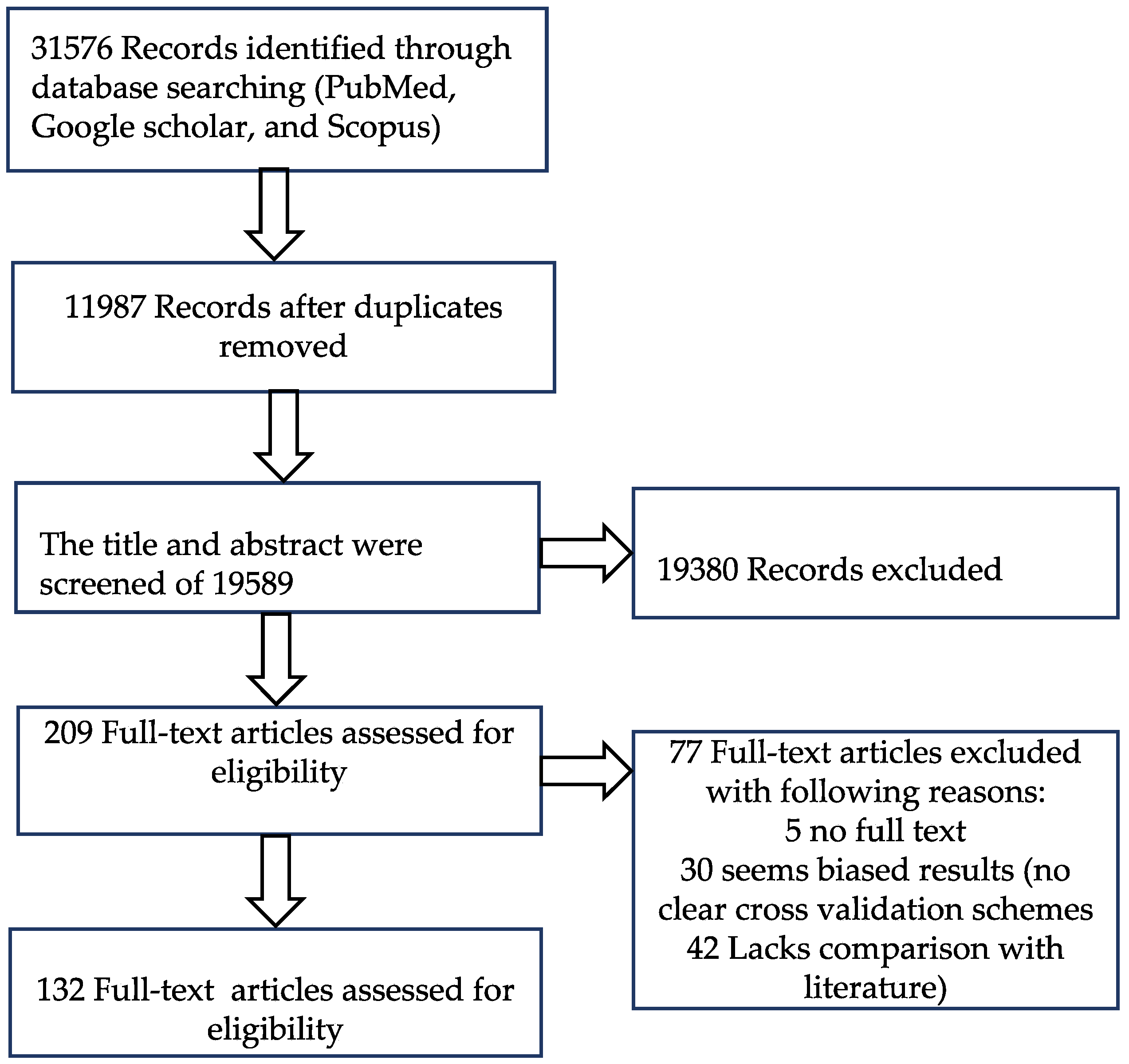
2. Materials and Methods
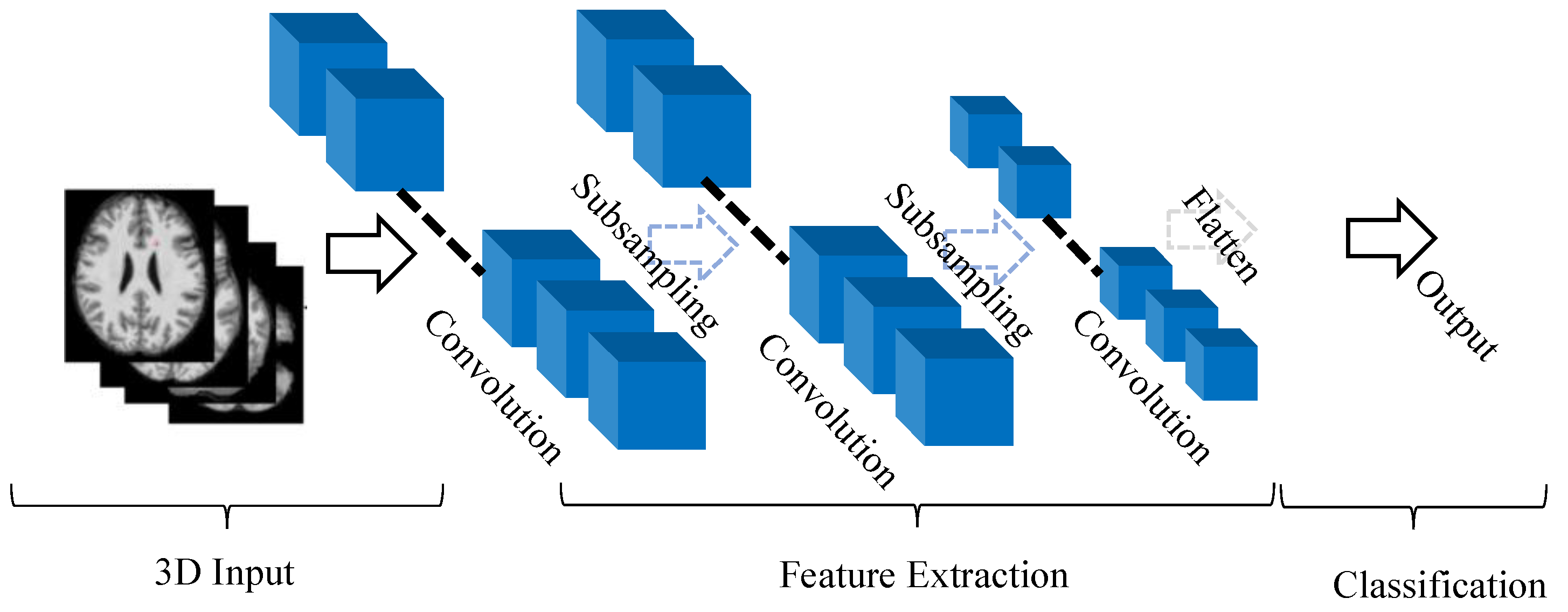
2.1. A Typical Architecture of 3D CNN
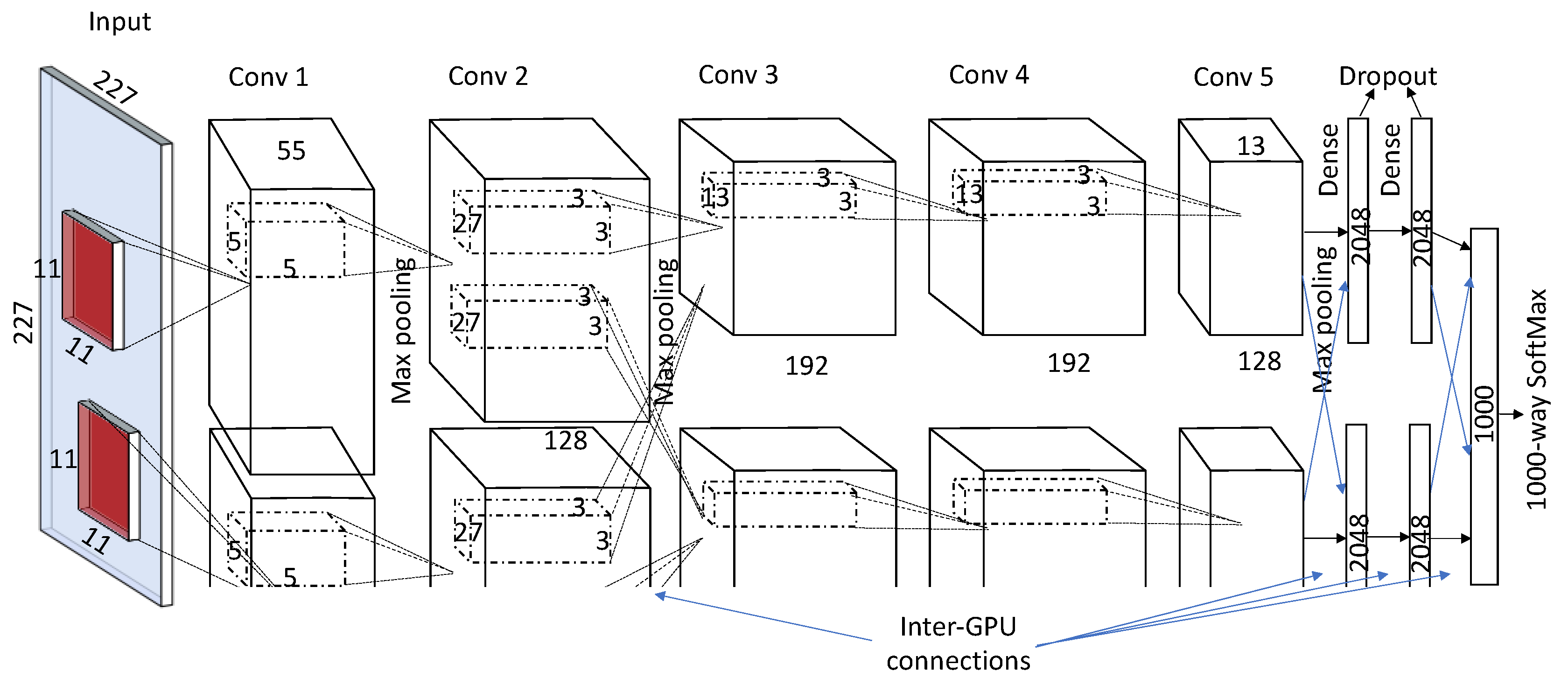
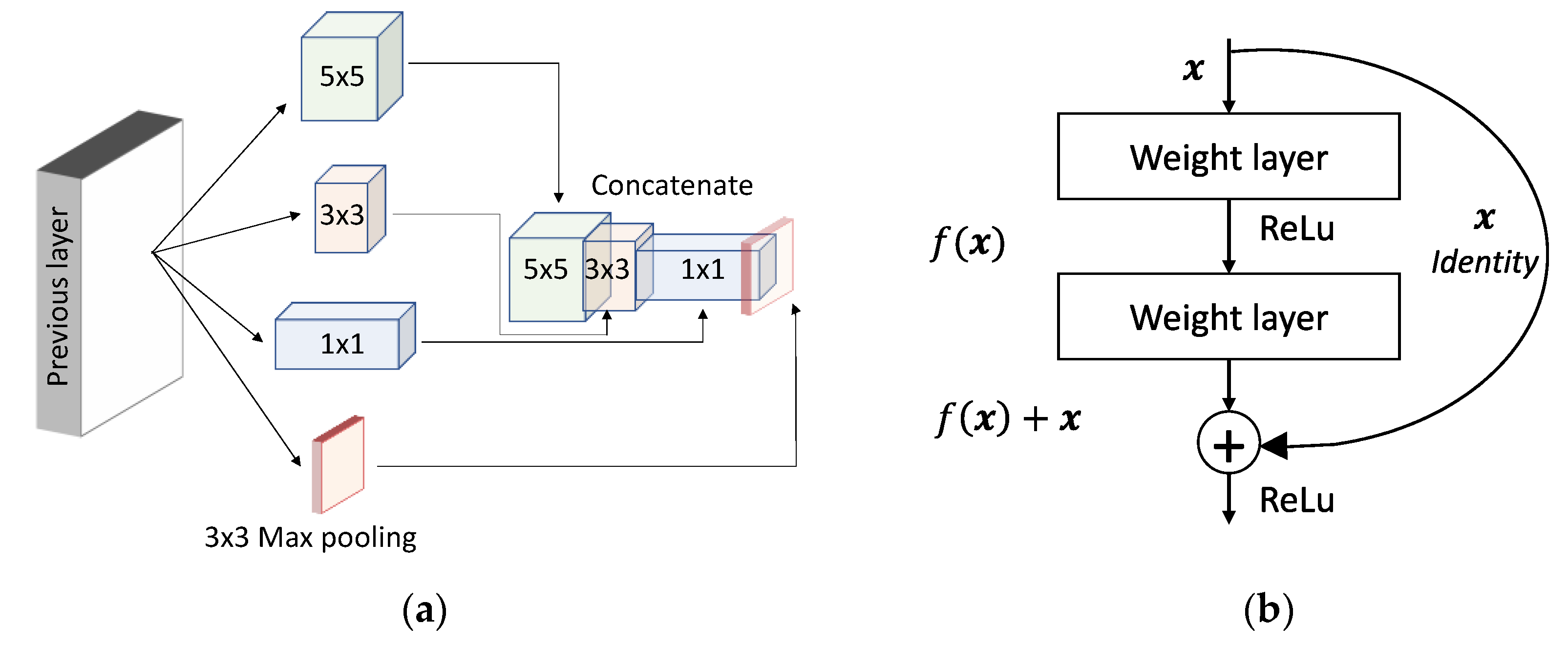
2.2. Breakthroughs in CNN Architectural Advances
3. 3D Medical Imaging Pre-Processing
4. Applications in 3D Medical Imaging
4.1. Segmentation
4.2. Classification
4.3. Detection and Localization
4.4. Registration
5. Challenges and Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Doi, K. Computer-Aided Diagnosis in Medical Imaging: Historical Review, Current Status and Future Potential. Comput. Med. Imaging Graph. 2007, 31, 198–211. [Google Scholar] [CrossRef]
- Miller, A.S.; Blott, B.H.; hames, T.K. Review of neural network applications in medical imaging and signal processing. Med. Biol. Eng. Comput. 1992, 30, 449–464. [Google Scholar] [CrossRef] [PubMed]
- Siedband, M.P. Medical imaging systems. Med. Instrum.-Appl. Des. 1998, 518–576. [Google Scholar]
- Prince, J.; Links, J. Medical Imaging Signals and Systems; Pearson: London, UK, 2006; pp. 315–379. [Google Scholar]
- Shapiro, R.S.; Wagreich, J.; Parsons, R.B.; Stancato-Pasik, A.; Yeh, H.C.; Lao, R. Tissue harmonic imaging sonography: Evaluation of image quality compared with conventional sonography. Am. J. Roentgenol. 1998, 171, 1203–1206. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, K.; Jinzaki, M.; Tanami, Y.; Ueno, A.; Yamada, M.; Kuribayashi, S. Virtual Monochromatic Spectral Imaging with Fast Kilovoltage Switching: Improved Image Quality as Compared with That Obtained with Conventional 120-kVp CT. Radiology 2011, 259, 257–262. [Google Scholar] [CrossRef] [PubMed]
- Thibault, J.-B.; Sauer, K.D.; Bouman, C.A.; Hsieh, J. A three-dimensional statistical approach to improved image quality for multislice helical CT. Med. Phys. 2007, 34, 4526–4544. [Google Scholar] [CrossRef]
- Marin, D.; Nelson, R.C.; Schindera, S.T.; Richard, S.; Youngblood, R.S.; Yoshizumi, T.T.; Samei, E. Low-Tube-Voltage, High-Tube-Current Multidetector Abdominal CT: Improved Image Quality and Decreased Radiation Dose with Adaptive Statistical Iterative Reconstruction Algorithm—Initial Clinical Experience. Radiology 2010, 254, 145–153. [Google Scholar] [CrossRef]
- Ker, J.; Singh, S.P.; Bai, Y.; Rao, J.; Lim, T.; Wang, L. Image Thresholding Improves 3-Dimensional Convolutional Neural Network Diagnosis of Different Acute Brain Hemorrhages on Computed Tomography Scans. Sensors 2019, 19, 2167. [Google Scholar] [CrossRef]
- Parmar, H.S.; Nutter, B.; Long, R.; Antani, S.; Mitra, S. Deep learning of volumetric 3D CNN for fMRI in Alzheimer’s disease classification. In Medical Imaging 2020: Biomedical Applications in Molecular, Structural, and Functional Imaging, Houston, TX, USA, 2020; Gimi, B.S., Krol, A., Eds.; SPIE: Bellingham, WA, USA, 2020; Volume 11317, p. 11. [Google Scholar]
- Shen, D.; Wu, G.; Suk, H.-I. Deep Learning in Medical Image Analysis. Annu. Rev. Biomed. Eng. 2017, 19, 221–248. [Google Scholar] [CrossRef]
- Gruetzemacher, R.; Gupta, A.; Paradice, D. 3D deep learning for detecting pulmonary nodules in CT scans. J. Am. Med. Inform. Assoc. 2018, 25, 1301–1310. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.H.; Phillips, P.; Sui, Y.; Liu, B.; Yang, M.; Cheng, H. Classification of Alzheimer’s Disease Based on Eight-Layer Convolutional Neural Network with Leaky Rectified Linear Unit and Max Pooling. J. Med. Syst. 2018, 42, 85. [Google Scholar] [CrossRef] [PubMed]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet classification with deep convolutional neural networks. Commun. ACM 2017, 60, 84–90. [Google Scholar] [CrossRef]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015. [Google Scholar]
- Hoi, S.C.H.; Jin, R.; Zhu, J.; Lyu, M.R. Batch mode active learning and its application to medical image classification. In Proceedings of the 23rd International Conference on Machine Learning, New York, NY, USA, 25–29 June 2006; pp. 417–424. [Google Scholar]
- Rahman, M.M.; Bhattacharya, P.; Desai, B.C. A Framework for Medical Image Retrieval Using Machine Learning and Statistical Similarity Matching Techniques With Relevance Feedback. IEEE Trans. Inf. Technol. Biomed. 2007, 11, 58–69. [Google Scholar] [CrossRef]
- Wernick, M.; Yang, Y.; Brankov, J.; Yourganov, G.; Strother, S. Machine Learning in Medical Imaging. IEEE Signal. Process. Mag. 2010, 27, 25–38. [Google Scholar] [CrossRef] [PubMed]
- Criminisi, A.; Shotton, J.; Konukoglu, E. Decision forests: A unified framework for classification, regression, density estimation, manifold learning and semi-supervised learning. Found. Trends® Comput. Graph. Vis. 2012, 7, 81–227. [Google Scholar] [CrossRef]
- Singh, S.P.; Urooj, S. An Improved CAD System for Breast Cancer Diagnosis Based on Generalized Pseudo-Zernike Moment and Ada-DEWNN Classifier. J. Med. Syst. 2016, 40, 105. [Google Scholar] [CrossRef]
- Urooj, S.; Singh, S.P. Rotation invariant detection of benign and malignant masses using PHT. In Proceedings of the 2015 2nd International Conference on Computing for Sustainable Global Development (INDIACom), New Delhi, India, 11–13 March 2015. [Google Scholar]
- Ji, S.; Xu, W.; Yang, M.; Yu, K. 3D Convolutional Neural Networks for Human Action Recognition. IEEE Trans. Pattern Anal. Mach. Intell. 2013, 35, 221–231. [Google Scholar] [CrossRef]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; Altman, D.; Antes, G.; Atkins, D.; Barbour, V.; Barrowman, N.; Berlin, J.A.; et al. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. PLoS Med. 2009, 6, e1000097. [Google Scholar] [CrossRef]
- Pezeshk, A.; Hamidian, S.; Petrick, N.; Sahiner, B. 3-D Convolutional Neural Networks for Automatic Detection of Pulmonary Nodules in Chest CT. IEEE J. Biomed. Heal. Inform. 2019, 23, 2080–2090. [Google Scholar] [CrossRef]
- Springenberg, J.T.; Dosovitskiy, A.; Brox, T.; Riedmiller, M. Striving for simplicity: The all convolutional net. In Proceedings of the 3rd International Conference on Learning Representations, ICLR 2015–Workshop Track Proceedings, San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- Kukačka, J.; Golkov, V.; Cremers, D. Regularization for Deep Learning: A Taxonomy. arXiv 2017, arXiv:1710.10686. [Google Scholar]
- Srivastava, N.; Hinton, G.; Krizhevsky, A.; Salakhutdinov, R. Dropout: A Simple Way to Prevent Neural Networks from Overfitting. J. Mach. Learn. Res. 2014, 15, 1929–1958. [Google Scholar]
- Ioffe, S.; Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. In Proceedings of the 32nd International Conference on Machine Learning, ICML 2015, International Machine Learning Society (IMLS), Lile, France, 6–11 July 2015; Volume 1, pp. 448–456. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. In Proceedings of the 3rd International Conference on Learning Representations, ICLR 2015, Conference Track Proceedings, San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the Inception Architecture for Computer Vision. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27 June–28 July 2016; Volume 1, pp. 2818–2826. [Google Scholar]
- Szegedy, C.; Ioffe, S.; Vanhoucke, V.; Alemi, A.A. Inception-v4, inception-ResNet and the impact of residual connections on learning. In Proceedings of the 31st AAAI Conference on Artificial Intelligence, AAAI 2017, San Francisco, CA, USA, 4–9 February 2017; pp. 4278–4284. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27 June–28 July 2016; Volume 1, pp. 770–778. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2015; Volume 9351, pp. 234–241. [Google Scholar]
- Çiçek, Ö.; Abdulkadir, A.; Lienkamp, S.S.; Brox, T.; Ronneberger, O. 3D U-net: Learning dense volumetric segmentation from sparse annotation. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2016; Volume 9901, pp. 424–432. [Google Scholar]
- Ker, J.; Wang, L.; Rao, J.; Lim, T. Deep Learning Applications in Medical Image Analysis. IEEE Access 2018, 6, 9375–9389. [Google Scholar] [CrossRef]
- Burt, J. Volumetric Quantification of Cardiovascular Structures from Medical Imaging. U.S. Patent 9,968,257, 15 May 2018. [Google Scholar]
- Esteban, O.; Markiewicz, C.J.; Blair, R.W.; Moodie, C.A.; Isik, A.I.; Erramuzpe, A.; Kent, J.D.; Goncalves, M.; DuPre, E.; Snyder, M.; et al. fmriprep: A Robust Preprocessing Pipeline for fMRI Data—Fmriprep version documentation. Nat. Methods 2019, 111–116. [Google Scholar] [CrossRef] [PubMed]
- Alansary, A.; Kamnitsas, K.; Davidson, A.; Khlebnikov, R.; Rajchl, M.; Malamateniou, C.; Rutherford, M.; Hajnal, J.V.; Glocker, B.; Rueckert, D.; et al. Fast Fully Automatic Segmentation of the Human Placenta from Motion Corrupted MRI. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Cham, Switzerland, 2016; pp. 589–597. [Google Scholar]
- Yang, C.; Rangarajan, A.; Ranka, S. Visual Explanations from Deep 3D Convolutional Neural Networks for Alzheimer’s Disease Classification. AMIA Annu. Symp. Proc. AMIA Symp. 2018, 2018, 1571–1580. [Google Scholar] [PubMed]
- Jones, D.K.; Griffin, L.D.; Alexander, D.C.; Catani, M.; Horsfield, M.A.; Howard, R.; Williams, S.C.R. Spatial Normalization and Averaging of Diffusion Tensor MRI Data Sets. Neuroimage 2002, 17, 592–617. [Google Scholar] [CrossRef] [PubMed]
- Jnawali, K.; Arbabshirani, M.; Rao, N. Deep 3D Convolution Neural Network for CT Brain Hemorrhage Classification. In Proceedings of the Medical Imaging 2018: Computer-Aided Diagnosis, International Society for Optics and Photonics, Houston, TX, USA, 10–15 February 2018; p. 105751C. [Google Scholar]
- Dubost, F.; Adams, H.; Bortsova, G.; Ikram, M. 3D Regression Neural Network for the Quantification of Enlarged Perivascular Spaces in Brain MRI. Med. Image Anal. 2019, 51, 89–100. [Google Scholar] [CrossRef]
- Lian, C.; Liu, M.; Zhang, J.; Zong, X.; Lin, W.; Shen, D. Automatic Segmentation of 3D Perivascular Spaces in 7T MR Images Using Multi-Channel Fully Convolutional Network. In Proceedings of the International Society for Magnetic Resonance in Medicine, Scientific Meeting and Exhibition, Paris, France, 16–21 June 2018; pp. 5–7. [Google Scholar]
- Pauli, R.; Bowring, A.; Reynolds, R.; Chen, G.; Nichols, T.E.; Maumet, C. Exploring fMRI Results Space: 31 Variants of an fMRI Analysis in AFNI, FSL, and SPM. Front. Neuroinform. 2016, 10. [Google Scholar] [CrossRef]
- Parker, D.; Liu, X.; Razlighi, Q.R. Optimal slice timing correction and its interaction with fMRI parameters and artifacts. Med. Image Anal. 2017, 35, 434–445. [Google Scholar] [CrossRef]
- Goebel, R. Brain Voyager—Past, present, future. Neuroimage 2012, 62, 748–756. [Google Scholar] [CrossRef]
- Maes, F.; Collignon, A.; Vandemeulen, D.; Marchal, G.; Suetens, P. Multimodality image registration by maximization of mutual information. IEEE Trans. Med. Imaging 1997, 16, 187–198. [Google Scholar] [CrossRef] [PubMed]
- Maintz, J.A.; Viergever, M.A. A survey of medical image registration. Med. Image Anal. 1998, 2, 1–36. [Google Scholar] [CrossRef]
- Pluim, J.P.W.; Maintz, J.B.A.; Viergever, M.A. Interpolation Artefacts in Mutual Information Based Image Registration. Comput. Vis. Image Underst. 2000, 77, 211–232. [Google Scholar] [CrossRef]
- Penney, G.P.; Weese, J.; Little, J.A.; Desmedt, P.; Hill, D.L.G.; Hawkes, D.J. A comparison of similarity measures for use in 2-D-3-D medical image registration. Med. Imaging IEEE Trans. 1998, 17, 586–595. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.N.; Yamany, S.M.; Mohamed, N.; Farag, A.A.; Moriarty, T. A modified fuzzy c-means algorithm for bias field estimation and segmentation of MRI data. IEEE Trans. Med. Imaging 2002, 21, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Xu, C.; Anderson, A.W.; Gore, J.C. MRI tissue classification and bias field estimation based on coherent local intensity clustering: A unified energy minimization framework. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2009; Volume 5636, pp. 288–299. [Google Scholar]
- Kamnitsas, K.; Ferrante, E.; Parisot, S.; Ledig, C.; Nori, A.V.; Criminisi, A.; Rueckert, D.; Glocker, B. DeepMedic for Brain Tumor Segmentation. In Proceedings of the International Workshop on Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries, Athens, Greece, 17 October 2016; pp. 138–149. [Google Scholar]
- Kamnitsas, K.; Ledig, C.; Newcombe, V.F.J.; Simpson, J.P.; Kane, A.D.; Menon, D.K.; Rueckert, D.; Glocker, B. Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med. Image Anal. 2017, 36, 61–78. [Google Scholar] [CrossRef]
- Casamitjana, A.; Puch, S.; Aduriz, A.; Vilaplana, V. 3D convolutional neural networks for brain tumor segmentation: A comparison of multi-resolution architectures. In Proceedings of the Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Athens, Greece, 2016; pp. 150–161. [Google Scholar]
- Zhou, C.; Ding, C.; Wang, X.; Lu, Z.; Tao, D. One-pass Multi-task Networks with Cross-task Guided Attention for Brain Tumor Segmentation. IEEE Trans. Image Process. 2020, 29, 4516–4529. [Google Scholar] [CrossRef]
- Chen, W.; Liu, B.; Peng, S.; Sun, J.; Qiao, X. S3D-UNET: Separable 3D U-Net for brain tumor segmentation. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2019; Volume 11384, pp. 358–368. [Google Scholar]
- Kayalibay, B.; Jensen, G.; van der Smagt, P. CNN-based Segmentation of Medical Imaging Data. arXiv 2017, arXiv:1701.03056. [Google Scholar]
- Isensee, F.; Kickingereder, P.; Wick, W.; Bendszus, M.; Maier-Hein, K.H. Brain tumor segmentation and radiomics survival prediction: Contribution to the BRATS 2017 challenge. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2018; Volume 10670, pp. 287–297. [Google Scholar]
- Peng, S.; Chen, W.; Sun, J.; Liu, B. Multi-Scale 3D U-Nets: An approach to automatic segmentation of brain tumor. Int. J. Imaging Syst. Technol. 2019, 30, 5–17. [Google Scholar] [CrossRef]
- Milletari, F.; Ahmadi, S.A.; Kroll, C.; Plate, A.; Rozanski, V.; Maiostre, J.; Levin, J.; Dietrich, O.; Ertl-Wagner, B.; Bötzel, K.; et al. Hough-CNN: Deep Learning for Segmentation of Deep Brain Regions in MRI and Ultrasound. Comput. Vis. Image Underst. 2017, 164, 92–102. [Google Scholar] [CrossRef]
- Dolz, J.; Desrosiers, C. 3D fully convolutional networks for subcortical segmentation in MRI: A large-scale study. Neuroimage 2018, 170, 456–470. [Google Scholar] [CrossRef] [PubMed]
- Sato, D.; Hanaoka, S.; Nomura, Y.; Takenaga, T.; Miki, S.; Yoshikawa, T.; Hayashi, N.; Abe, O. A primitive study on unsupervised anomaly detection with an autoencoder in emergency head CT volumes. In Proceedings of the Medical Imaging 2018: Computer-Aided Diagnosis, Houston, TX, USA, 10–15 February 2018; p. 60. [Google Scholar]
- Dou, Q.; Yu, L.; Chen, H.; Jin, Y.; Yang, X.; Qin, J.; Heng, P.A. 3D deeply supervised network for automated segmentation of volumetric medical images. Med. Image Anal. 2017, 41, 40–54. [Google Scholar] [CrossRef] [PubMed]
- Zeng, G.; Yang, X.; Li, J.; Yu, L.; Heng, P.A.; Zheng, G. 3D U-net with multi-level deep supervision: Fully automatic segmentation of proximal femur in 3D MR images. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2017; Volume 10541, pp. 274–282. [Google Scholar]
- Zhu, Z.; Xia, Y.; Shen, W.; Fishman, E.; Yuille, A. A 3D coarse-to-fine framework for volumetric medical image segmentation. In Proceedings of the 2018 International Conference on 3D Vision, Verona, Italy, 5–8 September 2018; pp. 682–690. [Google Scholar]
- Yang, X.; Bian, C.; Yu, L.; Ni, D.; Heng, P.A. Hybrid loss guided convolutional networks for whole heart parsing. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2018; Volume 10663, pp. 215–223. [Google Scholar]
- Roth, H.R.; Oda, H.; Zhou, X.; Shimizu, N.; Yang, Y. Computerized Medical Imaging and Graphics An application of cascaded 3D fully convolutional networks for medical image segmentation. Comput. Med. Imaging Graph. 2018, 66, 90–99. [Google Scholar] [CrossRef]
- Yu, L.; Yang, X.; Qin, J.; Heng, P.A. 3D FractalNet: Dense volumetric segmentation for cardiovascular MRI volumes. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2017; Volume 10129, pp. 103–110. [Google Scholar]
- Li, X.; Chen, H.; Qi, X.; Dou, Q.; Fu, C.-W.; Heng, P.A. H-DenseUNet: Hybrid Densely Connected UNet for Liver and Liver Tumor Segmentation from CT Volumes. IEEE Trans. Med. Imaging 2018, 37, 2663–2674. [Google Scholar] [CrossRef] [PubMed]
- Ambellan, F.; Tack, A.; Ehlke, M.; Zachow, S. Automated segmentation of knee bone and cartilage combining statistical shape knowledge and convolutional neural networks: Data from the Osteoarthritis Initiative. Med. Image Anal. 2019, 52, 109–118. [Google Scholar] [CrossRef]
- Chen, L.; Shen, C.; Li, S.; Albuquerque, K.; Folkert, M.R.; Wang, J.; Maquilan, G. Automatic PET cervical tumor segmentation by deep learning with prior information. In Proceedings of the Physics in Medicine and Biology, Houston, TX, USA, 10–15 February 2018; p. 111. [Google Scholar]
- Heinrich, M.P.; Oktay, O.; Bouteldja, N. OBELISK-Net: Fewer layers to solve 3D multi-organ segmentation with sparse deformable convolutions. Med. Image Anal. 2019, 54, 1–9. [Google Scholar] [CrossRef]
- Chen, S.; Zhong, X.; Hu, S.; Dorn, S.; Kachelrieß, M.; Lell, M.; Maier, A. Automatic multi-organ segmentation in dual-energy CT (DECT) with dedicated 3D fully convolutional DECT networks. Med. Phys. 2020, 47, 552–562. [Google Scholar] [CrossRef]
- Kruthika, K.R.; Rajeswari; Maheshappa, H.D. CBIR system using Capsule Networks and 3D CNN for Alzheimer’s disease diagnosis. Inform. Med. Unlocked 2019, 14, 59–68. [Google Scholar] [CrossRef]
- Feng, C.; Elazab, A.; Yang, P.; Wang, T.; Zhou, F.; Hu, H.; Xiao, X.; Lei, B. Deep Learning Framework for Alzheimer’s Disease Diagnosis via 3D-CNN and FSBi-LSTM. IEEE Access 2019, 7, 63605–63618. [Google Scholar] [CrossRef]
- Wegmayr, V.; Aitharaju, S.; Buhmann, J. Classification of brain MRI with big data and deep 3D convolutional neural networks. In Proceedings of the Medical Imaging 2018: Computer-Aided Diagnosis, Houston, TX, USA, 10–15 February 2018. [Google Scholar] [CrossRef]
- Gao, X.; Hui, R.; Biomedicine, Z.T. Classification of CT brain images based on deep learning networks. Omput. Methods Prog. Biomed. Elsevier 2017, 138, 49–56. [Google Scholar] [CrossRef]
- Nie, D.; Zhang, H.; Adeli, E.; Liu, L.; Shen, D. 3D deep learning for multi-modal imaging-guided survival time prediction of brain tumor patients. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Athens, Greece, 17–21 October 2016; Springer: Quebec City, QC, Canada, 2016; pp. 212–220. [Google Scholar]
- Zhou, J.; Luo, L.; Dou, Q.; Chen, H.; Chen, C.; Li, G.; Jiang, Z.; Heng, P. Weakly supervised 3D deep learning for breast cancer classification and localization of the lesions in MR images. J. Magn. Reson. Imaging 2019, 50, 1144–1151. [Google Scholar] [CrossRef] [PubMed]
- Ha, R.; Chang, P.; Mema, E.; Mutasa, S.; Karcich, J.; Wynn, R.T.; Liu, M.Z.; Jambawalikar, S. Fully Automated Convolutional Neural Network Method for Quantification of Breast MRI Fibroglandular Tissue and Background Parenchymal Enhancement. J. Digit. Imaging 2019, 32, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Zhou, Z.; Sher, D.; Zhang, Q.; Shah, J.; Pham, N.-L.; Jiang, S.B.; Wang, J. Combining many-objective radiomics and 3-dimensional convolutional neural network through evidential reasoning to predict lymph node metastasis in head and neck cancer. Phys. Med. Biol. 2019, 64, 075011. [Google Scholar] [CrossRef]
- Shaish, H.; Mutasa, S.; Makkar, J.; Chang, P.; Schwartz, L.; Ahmed, F. Prediction of lymph node maximum standardized uptake value in patients with cancer using a 3D convolutional neural network: A proof-of-concept study. Am. J. Roentgenol. 2019, 212, 238–244. [Google Scholar] [CrossRef] [PubMed]
- Oh, K.; Chung, Y.C.; Kim, K.W.; Kim, W.S.; Oh, I.S. Classification and Visualization of Alzheimer’s Disease using Volumetric Convolutional Neural Network and Transfer Learning. Sci. Rep. 2019, 9, 1–16. [Google Scholar] [CrossRef]
- Amidi, A.; Amidi, S.; Vlachakis, D.; Megalooikonomou, V.; Paragios, N.; Zacharaki, E. EnzyNet: Enzyme classification using 3D convolutional neural networks on spatial representation. PeerJ 2018, 6, e4750. [Google Scholar] [CrossRef]
- Dou, Q.; Chen, H.; Yu, L.; Zhao, L.; Qin, J.; Wang, D.; Mok, V.C.; Shi, L.; Heng, P.A. Automatic detection of cerebral microbleeds from MR images via 3D convolutional neural networks. IEEE Trans. Med. Imaging 2016, 35, 1182–1195. [Google Scholar] [CrossRef]
- Standvoss, K.; Goerke, L.; Crijns, T.; van Niedek, T.; Alfonso Burgos, N.; Janssen, D.; van Vugt, J.; Gerritse, E.; Mol, J.; van de Vooren, D.; et al. Cerebral microbleed detection in traumatic brain injury patients using 3D convolutional neural networks. In Proceedings of the Medical Imaging 2018: Computer-Aided Diagnosis, Houston, TX, USA, 10–15 February 2018; p. 48. [Google Scholar]
- Wolterink, J.M.; van Hamersvelt, R.W.; Viergever, M.A.; Leiner, T.; Išgum, I. Coronary Artery Centerline Extraction in Cardiac CT Angiography. Med. Image Anal. 2019, 51, 46–60. [Google Scholar] [CrossRef]
- Anirudh, R.; Thiagarajan, J.J.; Bremer, T.; Kim, H. Lung nodule detection using 3D convolutional neural networks trained on weakly labeled data. In Proceedings of the Medical Imaging 2016: Computer-Aided Diagnosis, San Diego, CA, USA, 27 February–3 March 2016; Volume 9785, p. 978532. [Google Scholar]
- Dou, Q.; Chen, H.; Yu, L.; Qin, J.; Heng, P.A. Multilevel Contextual 3-D CNNs for False Positive Reduction in Pulmonary Nodule Detection. IEEE Trans. Biomed. Eng. 2017, 64, 1558–1567. [Google Scholar] [CrossRef]
- Huang, X.; Shan, J.; Vaidya, V. Lung nodule detection in CT using 3D convolutional neural networks. In Proceedings of the International Symposium on Biomedical Imaging, Melbourne, Australia, 18–21 April 2017; pp. 379–383. [Google Scholar]
- Gu, Y.; Lu, X.; Yang, L.; Zhang, B.; Yu, D.; Zhao, Y.; Gao, L.; Wu, L.; Zhou, T. Automatic lung nodule detection using a 3D deep convolutional neural network combined with a multi-scale prediction strategy in chest CTs. Comput. Biol. Med. 2018, 103, 220–231. [Google Scholar] [CrossRef]
- Winkels, M.; Cohen, T.S. Pulmonary nodule detection in CT scans with equivariant CNNs. Med. Image Anal. 2019, 55, 15–26. [Google Scholar] [CrossRef] [PubMed]
- Gong, L.; Jiang, S.; Yang, Z.; Zhang, G.; Wang, L. Automated pulmonary nodule detection in CT images using 3D deep squeeze-and-excitation networks. Int. J. Comput. Assist. Radiol. Surg. 2019, 14, 1969–1979. [Google Scholar] [CrossRef] [PubMed]
- Wolterink, J.; Leiner, T.; Viergever, M.A.; Išgum, I. Automatic Coronary Calcium Scoring in Cardiac CT Angiography Using Convolutional Neural Networks; Springer: Munich, Germany, 2015; Volume 9349. [Google Scholar]
- De Vos, B.D.; Wolterink, J.M.; De Jong, P.A.; Leiner, T.; Viergever, M.A.; Išgum, I. ConvNet-Based Localization of Anatomical Structures in 3-D Medical Images. IEEE Trans. Med. Imaging 2017, 36, 1470–1481. [Google Scholar] [CrossRef]
- Huo, Y.; Xu, Z.; Xiong, Y.; Aboud, K.; Parvathaneni, P.; Bao, S.; Bermudez, C.; Resnick, S.M.; Cutting, L.E.; Landman, B.A. 3D whole brain segmentation using spatially localized atlas network tiles. Neuroimage 2019, 194, 105–119. [Google Scholar] [CrossRef] [PubMed]
- Huang, R.; Xie, W.; Alison Noble, J. VP-Nets: Efficient automatic localization of key brain structures in 3D fetal neurosonography. Med. Image Anal. 2018, 47, 127–139. [Google Scholar] [CrossRef] [PubMed]
- O’Neil, A.Q.; Kascenas, A.; Henry, J.; Wyeth, D.; Shepherd, M.; Beveridge, E.; Clunie, L.; Sansom, C.; Šeduikytė, E.; Muir, K.; et al. Attaining human-level performance with atlas location autocontext for anatomical landmark detection in 3D CT data. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2019; Volume 11131, pp. 470–484. [Google Scholar]
- Mohseni Salehi, S.S.; Khan, S.; Erdogmus, D.; Gholipour, A. Real-Time Deep Pose Estimation With Geodesic Loss for Image-to-Template Rigid Registration. IEEE Trans. Med. Imaging 2019, 38, 470–481. [Google Scholar] [CrossRef]
- Li, X.; Dou, Q.; Chen, H.; Fu, C.W.; Qi, X.; Belavý, D.L.; Armbrecht, G.; Felsenberg, D.; Zheng, G.; Heng, P.A. 3D multi-scale FCN with random modality voxel dropout learning for Intervertebral Disc Localization and Segmentation from Multi-modality MR Images. Med. Image Anal. 2018, 45, 41–54. [Google Scholar] [CrossRef]
- Vesal, S.; Maier, A.; Ravikumar, N. Fully Automated 3D Cardiac MRI Localisation and Segmentation Using Deep Neural Networks. J. Imaging 2020, 6, 65. [Google Scholar] [CrossRef]
- Sokooti, H.; de Vos, B.; Berendsen, F.; Lelieveldt, B.P.F.; Išgum, I.; Staring, M. Nonrigid image registration using multi-scale 3D convolutional neural networks. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2017; Volume 10433, pp. 232–239. [Google Scholar]
- Torng, W.; Altman, R.B. 3D deep convolutional neural networks for amino acid environment similarity analysis. BMC Bioinform. 2017, 18, 302. [Google Scholar] [CrossRef]
- Blendowski, M.; Heinrich, M.P. Combining MRF-based deformable registration and deep binary 3D-CNN descriptors for large lung motion estimation in COPD patients. Int. J. Comput. Assist. Radiol. Surg. 2019, 14, 43–52. [Google Scholar] [CrossRef]
- Lowekamp, B.C.; Chen, D.T.; Ibáñez, L.; Blezek, D. The Design of SimpleITK. Front. Neuroinform. 2013, 7, 45. [Google Scholar] [CrossRef] [PubMed]
- Avants, B.; Tustison, N.; Song, G. Advanced Normalization Tools (ANTS). Insight J. 2009, 2, 1–35. [Google Scholar]
- Chee, E.; Wu, Z. AIRNet: Self-Supervised Affine Registration for 3D Medical Images using Neural Networks. arXiv 2018, arXiv:1810.02583. [Google Scholar]
- Huang, G.; Liu, Z.; van der Maaten, L.; Weinberger, K.Q. Densely Connected Convolutional Networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 2261–2269. [Google Scholar]
- Zhou, S.; Xiong, Z.; Chen, C.; Chen, X.; Liu, D.; Zhang, Y.; Zha, Z.J.; Wu, F. Fast and accurate electron microscopy image registration with 3D convolution. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2019; Volume 11764, pp. 478–486. [Google Scholar]
- Zhao, S.; Lau, T.; Luo, J.; Chang, E.I.C.; Xu, Y. Unsupervised 3D End-to-End Medical Image Registration with Volume Tweening Network. IEEE J. Biomed. Heal. Inform. 2020, 24, 1394–1404. [Google Scholar] [CrossRef]
- Klein, S.; Staring, M.; Murphy, K.; Viergever, M.A.; Pluim, J.P.W. Elastix: A toolbox for intensity-based medical image registration. IEEE Trans. Med. Imaging 2010, 29, 196–205. [Google Scholar] [CrossRef]
- Balakrishnan, G.; Zhao, A.; Sabuncu, M.R.; Dalca, A.V.; Guttag, J. An Unsupervised Learning Model for Deformable Medical Image Registration. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 9252–9260. [Google Scholar]
- Wang, J.; Schaffert, R.; Borsdorf, A.; Heigl, B.; Huang, X.; Hornegger, J.; Maier, A. Dynamic 2-D/3-D rigid registration framework using point-to-plane correspondence model. IEEE Trans. Med. Imaging 2017, 36, 1939–1954. [Google Scholar] [CrossRef]
- Najafabadi, M.M.; Villanustre, F.; Khoshgoftaar, T.M.; Seliya, N.; Wald, R.; Muharemagic, E. Deep learning applications and challenges in big data analytics. J. Big Data 2015, 2, 1. [Google Scholar] [CrossRef]
- Chen, X.W.; Lin, X. Big data deep learning: Challenges and perspectives. IEEE Access 2014, 2, 514–525. [Google Scholar] [CrossRef]
- Vedaldi, A.; Lenc, K. MatConvNet–Convolutional Neural Networks for MATLAB. In Proceedings of the 23rd ACM international Conference on Multimedia, Brisbane, Australia, 12–16 October 2015; pp. 689–692. [Google Scholar]
- Duncan, J.; Ayache, N. Medical image analysis: Progress over two decades and the challenges ahead. IEEE Trans. Pattern Anal. Mach. Intell. 2000, 22, 85–106. [Google Scholar] [CrossRef]
- Iglehart, J.K. Health Insurers and Medical-Imaging Policy—A Work in Progress. N. Engl. J. Med. 2009, 360, 1030–1037. [Google Scholar] [CrossRef]
- Wang, L.; Wang, Y.; Chang, Q. Feature selection methods for big data bioinformatics: A survey from the search perspective. Methods 2016, 111, 21–31. [Google Scholar] [CrossRef] [PubMed]
- Prasoon, A.; Petersen, K.; Igel, C.; Lauze, F.; Dam, E.; Nielsen, M. Deep feature learning for knee cartilage segmentation using a triplanar convolutional neural network. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Quebec City, QC, Canada, 2013; Volume 8150, pp. 246–253. [Google Scholar]
- Hinton, G.E.; Osindero, S.; Teh, Y.-W. A Fast Learning Algorithm for Deep Belief Nets. Neural Comput. 2006, 18, 1527–1554. [Google Scholar] [CrossRef] [PubMed]
- Frackowiak, R.S.J. Functional brain imaging. In Proceedings of the Radiation Protection Dosimetry; Oxford University Press: New York, NY, USA, 1996; pp. 55–61. [Google Scholar]
- Sun, M.J.; Zhang, J.M. Single-pixel imaging and its application in three-dimensional reconstruction: A brief review. Sensors 2019, 19, 732. [Google Scholar] [CrossRef] [PubMed]
- Lyu, M.; Wang, W.; Wang, H.; Wang, H.; Li, G.; Chen, N.; Situ, G. Deep-learning-based ghost imaging. Sci. Rep. 2017, 7, 17865. [Google Scholar] [CrossRef]
- Gupta, S.; Chan, Y.H.; Rajapakse, J.C. Decoding brain functional connectivity implicated in AD and MCI. bioRxiv 2019, 697003. [Google Scholar] [CrossRef]
- Seward, J. Artificial General Intelligence System and Method for Medicine that Determines a Pre-Emergent Disease State of a Patient Based on Mapping a Topological Module. U.S. Patent 9,864,841, 9 January 2018. [Google Scholar]
- Huang, T. Imitating the brain with neurocomputer a “new” way towards artificial general intelligence. Int. J. Autom. Comput. 2017, 14, 520–531. [Google Scholar] [CrossRef]
- Shigeno, S. Brain evolution as an information flow designer: The ground architecture for biological and artificial general intelligence. In Brain Evolution by Design; Springer: Tokyo, Japan, 2017; pp. 415–438. [Google Scholar]
- Mehta, N.; Devarakonda, M.V. Machine Learning, Natural Language Programming, and Electronic Health Records: The next step in the Artificial Intelligence Journey? J. Allergy Clin. Immunol. 2018, 141, 2019–2021. [Google Scholar] [CrossRef]
- Goodfellow, I.J.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial nets. In Proceedings of the Advances in Neural Information Processing Systems, Montreal, QC, Canada, 8–13 December 2014; Volume 3, pp. 2672–2680. [Google Scholar]

| Ref. | Methods | Data | Task | Performance Evaluation |
|---|---|---|---|---|
| Zhou et al. [56] | A 3D variant of FusionNet (One-pass Multi-task Network (OM-Net)) | BRATS 2018 | brain tumor segmentation | 0.916 (WT), 0.827 (TC), 0.807(EC) |
| Chen et al. [57] | Separable 3D U-Net | BRATS 2018 | --do-- | 0.893(WT), 0.830(TC), 0.742(EC) |
| Peng et al. [60] | Multi-Scale 3D U-Nets | BRATS 2015 | --do-- | 0.850(WT), 0.720(TC), 0.610(EC) |
| Kayalıbay et al. [58] | 3D U-Nets | BRATS 2015 | --do-- | 0.850 (WT), 0.872(TC), 0.610(EC) |
| Kamnitsas et al. [54] | 11 layers deep 3D CNN | BRATS 2015 and ISLES 2015 | --do-- | 0.898 (WT), 0.750 (TC), 0.720(EC) |
| Kamnitsas et al. 2016 [53] | 3D CNN in which features extracted by 2D CNNs | BRATS 2017 | --do-- | 0.918 (WT), 0.883(TC), 0.854 (EC) |
| Casamitjana et al. [55] | 3D U-Net followed by fully connected 3D CRF | BRATS 2015 | --do-- | 0.917(WT), 0,836(TC), 0.768(EC) |
| Isensee et al. [59] | 3D U-Nets | BRATS 2017 | --do-- | 0.850(WT), 0.740(TC), 0.640(EC) |
| Ref. | Task | Model | Data | Performance Measures |
|---|---|---|---|---|
| Yang et al. [39] | AD classification | 3D VggNet, 3D Resnet | MRI scans from ADNI dataset (47 AD, 56 NC) | 86.3% AUC using 3D VggNet and 85.4% AUC using 3D ResNet |
| Kruthika et al. [75] | --do-- | 3D capsule network, 3D CNN | MRI scans from ADNI dataset (345 AD, NC, 605, and 991MCI) | Acc. for AD/MCI/NC 89.1% |
| Feng et al. [76] | --do-- | 3D CNN + LSTM | PET + MRI scans from ADNI dataset (93 AD, 100 NC) | Acc. 65.5% (sMCI/NC), 86.4% (pMCI/NC), and 94.8 % (AD/NC) |
| Wegmayr et al. [77] | --do-- | 3D CNN | ADNI and AIBL data sets, 20000 T1 scans | Acc. 72% (MCI/AD), 86 % (AD/NC), and 67 % (MCI/NC) |
| Oh et al. [84] | --do-- | 3D CNN +transfer learning | MRI scans from the ADNI dataset (AD 198, NC 230, pMCI 166, and sMCI 101) at baseline. | 74% (pMCI/sMCI), 86% (AD/NC), 77% (pMCI/NC) |
| Parmar et al. [10] | --do-- | 3D CNN | fMRI scans from ADNI dataset (30 AD, 30 NC) | Classification acc. 94.85 % (AD/NC) |
| Nie et al. [79] | Brain tumor | 3D CNN with learning supervised features | Private, 69 patient (T1 MRI, fMRI, and DTI) | Classification acc. 89.85 % |
| Amidi et al. [85] | Protein shape | 2-layer 3D CNN | 63,558 enzymes from PDB datasets | Classification acc. 78% |
| Zhou et al. [80] | Breast cancer | Weakly supervised 3D CNN | Private, 1537 female patient | Classification acc. 78% 83.7% |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Singh, S.P.; Wang, L.; Gupta, S.; Goli, H.; Padmanabhan, P.; Gulyás, B. 3D Deep Learning on Medical Images: A Review. Sensors 2020, 20, 5097. https://doi.org/10.3390/s20185097
Singh SP, Wang L, Gupta S, Goli H, Padmanabhan P, Gulyás B. 3D Deep Learning on Medical Images: A Review. Sensors. 2020; 20(18):5097. https://doi.org/10.3390/s20185097
Chicago/Turabian StyleSingh, Satya P., Lipo Wang, Sukrit Gupta, Haveesh Goli, Parasuraman Padmanabhan, and Balázs Gulyás. 2020. "3D Deep Learning on Medical Images: A Review" Sensors 20, no. 18: 5097. https://doi.org/10.3390/s20185097
APA StyleSingh, S. P., Wang, L., Gupta, S., Goli, H., Padmanabhan, P., & Gulyás, B. (2020). 3D Deep Learning on Medical Images: A Review. Sensors, 20(18), 5097. https://doi.org/10.3390/s20185097







