Optical Temperature Control Unit and Convolutional Neural Network for Colorimetric Detection of Loop-Mediated Isothermal Amplification on a Lab-On-A-Disc Platform
Abstract
1. Introduction
2. Materials and Methods
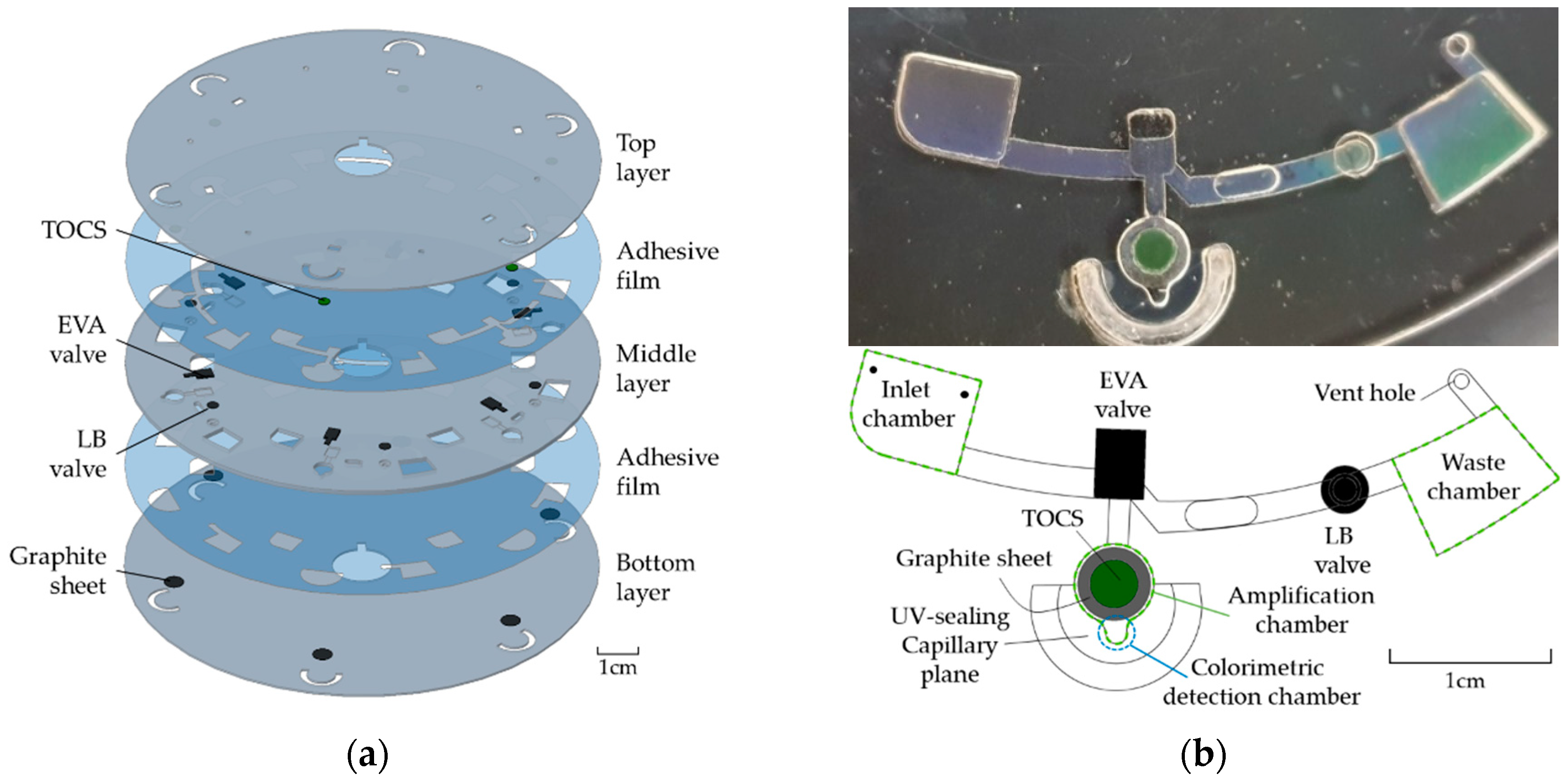
2.1. Disc Fabrication
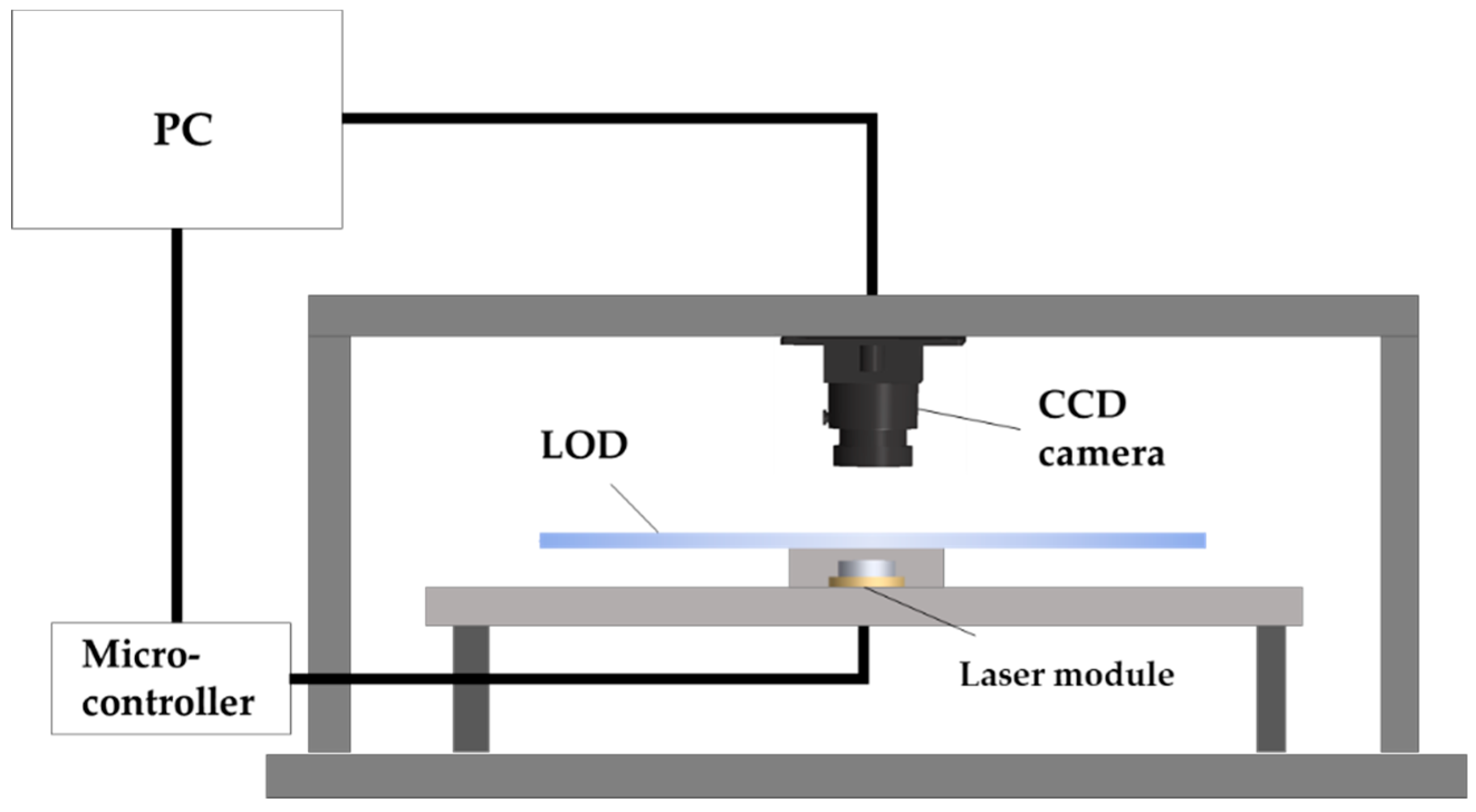
2.2. Experimental Apparatus and Driving Protocol
2.3. Sample Preparation and DNA Extraction
2.4. LAMP Reaction
2.5. Executing Program for Temperature Control and Colorimetric Analysis of the LAMP Reaction
3. Results
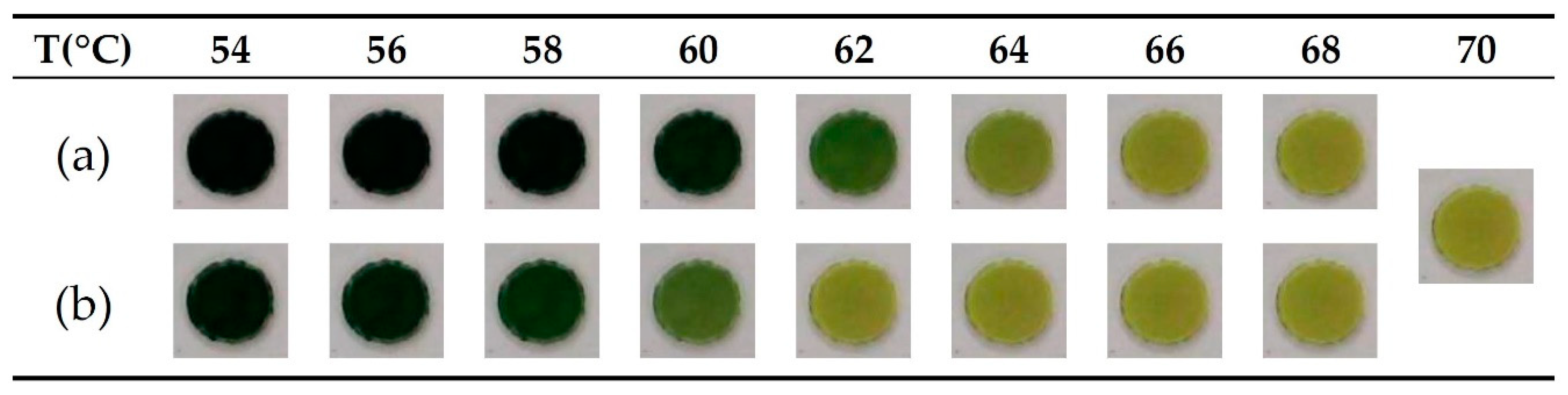
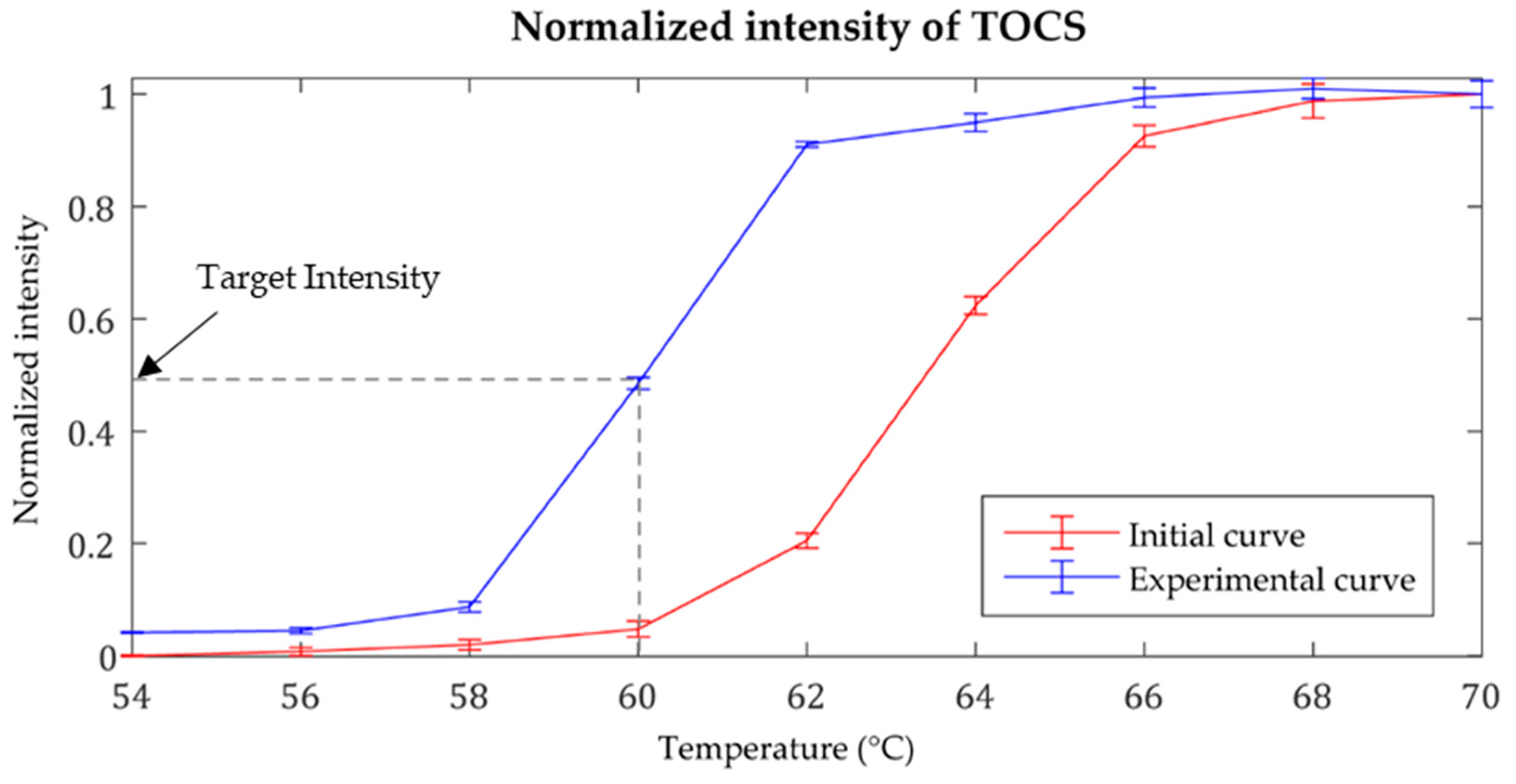
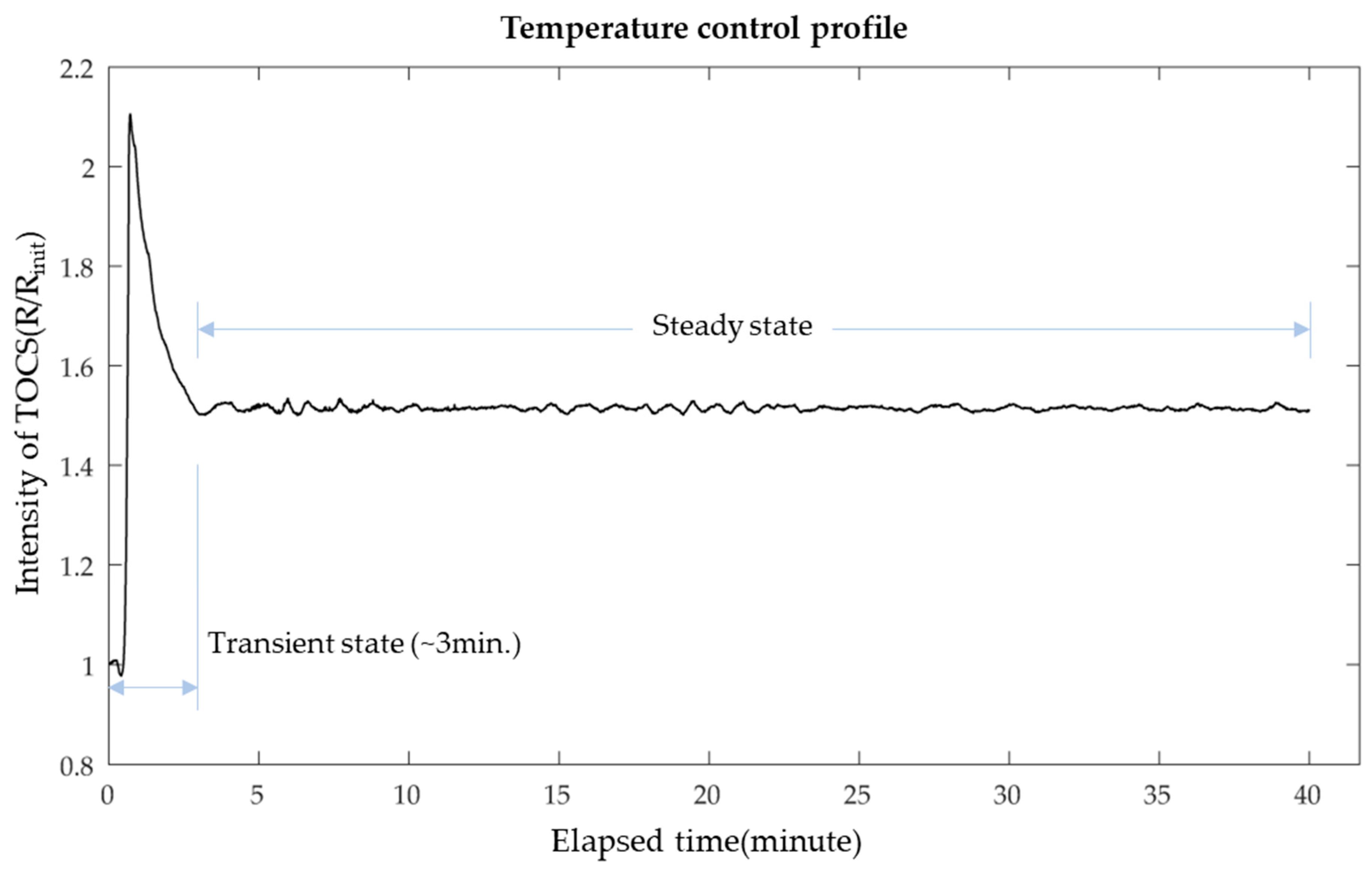
3.1. Temperature Control for the LAMP Reaction
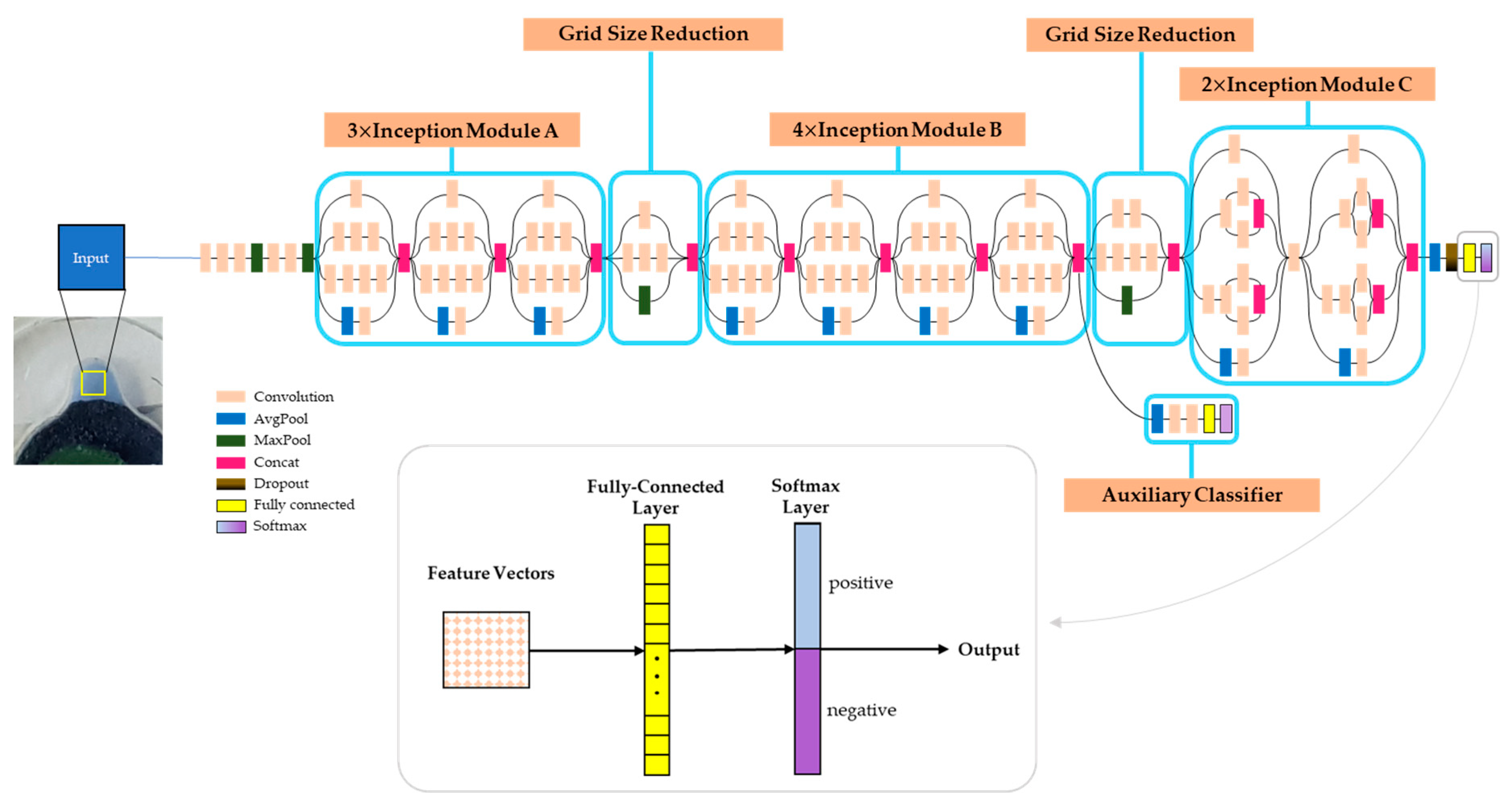
3.2. Analysis of the LAMP Product by CNN (Convolutional Neural Network)
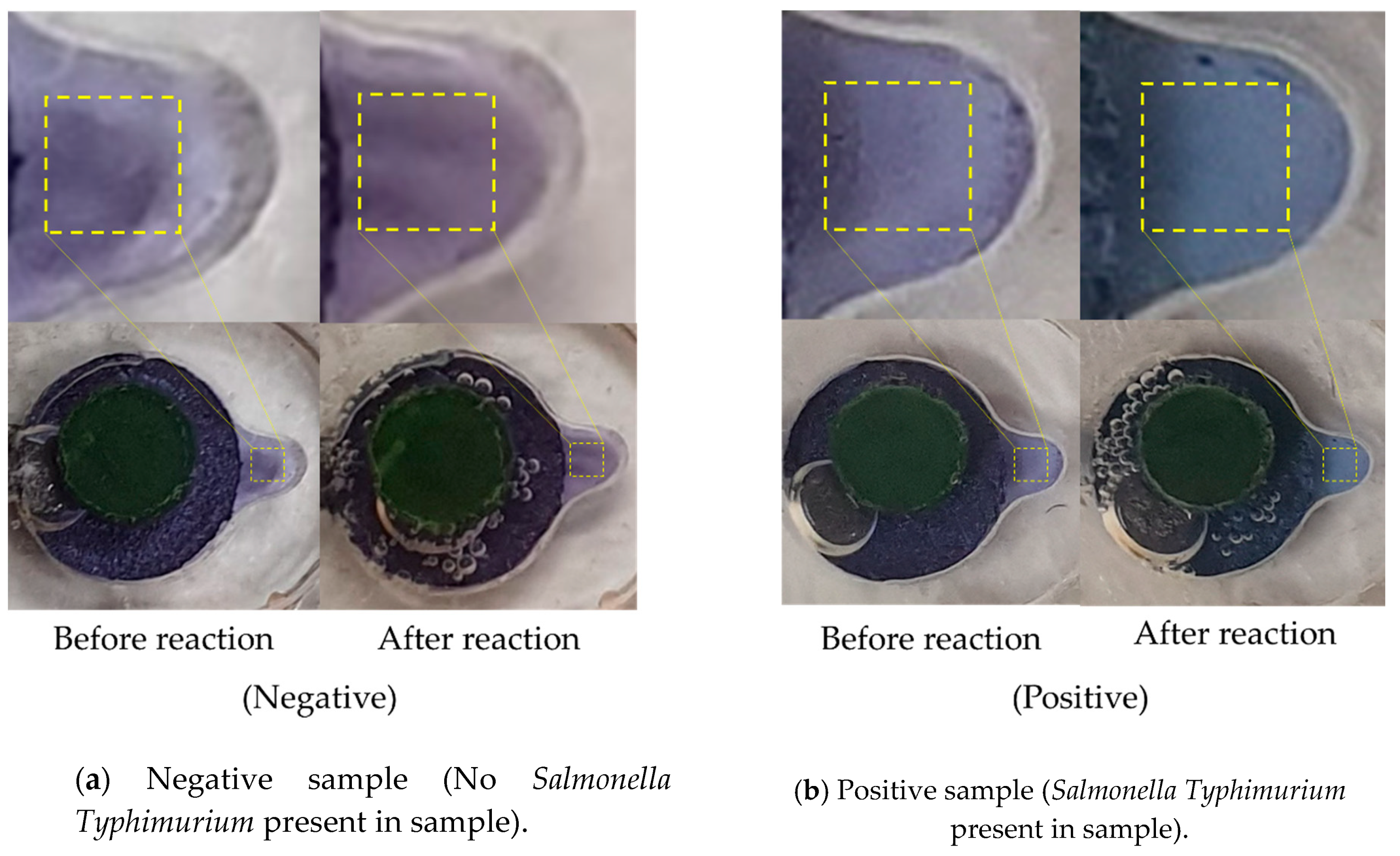
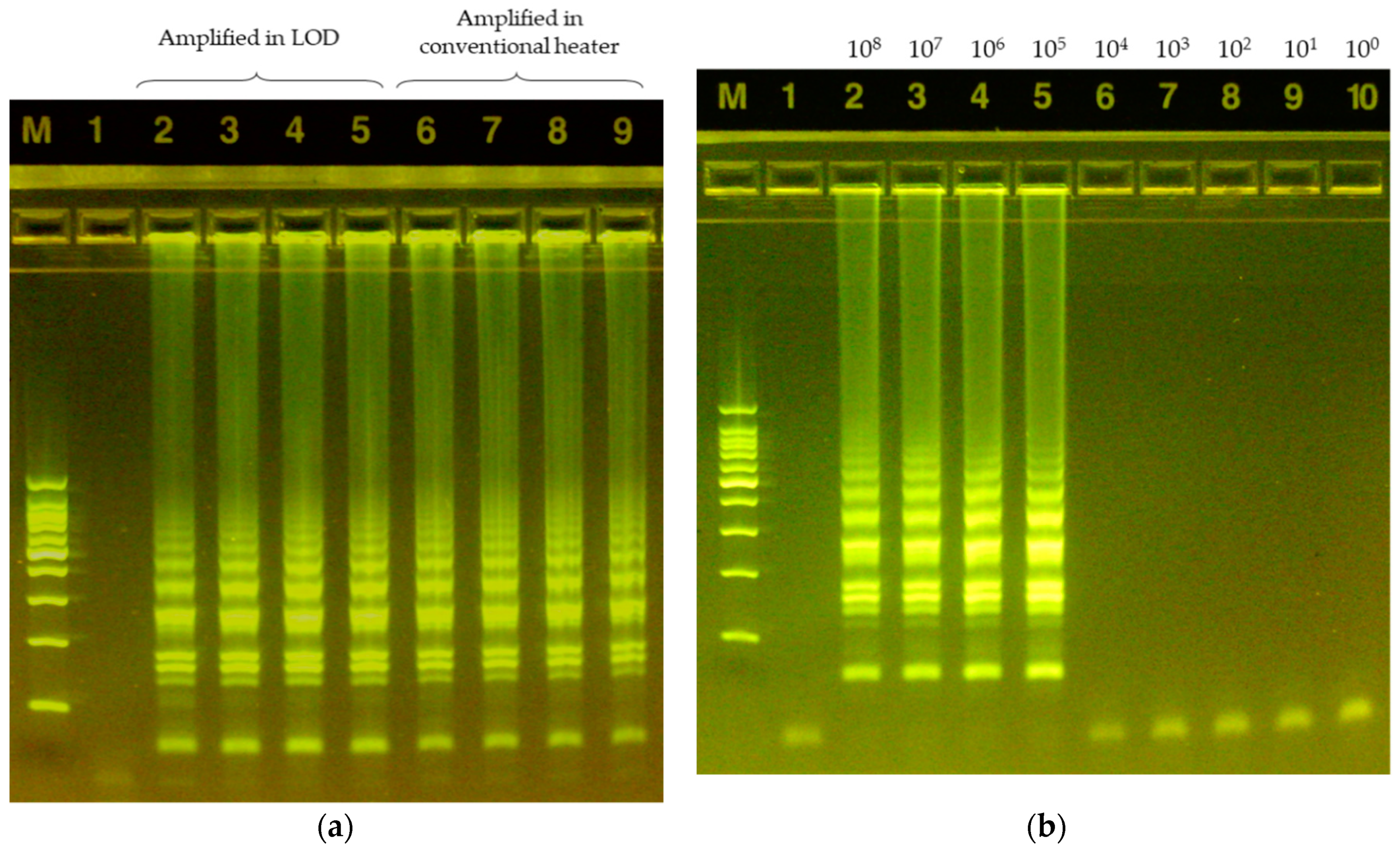
3.3. LAMP Validation
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Fratamico, P.M.; DebRoy, C.; Liu, Y.; Needleman, D.S.; Baranzoni, G.M.; Feng, P. Advances in Molecular Serotyping and Subtyping of Escherichia coli. Front. Microbiol. 2016, 7, 644. [Google Scholar] [CrossRef] [PubMed]
- Dewey-Mattia, D.; Manikonda, K.; Hall, A.J.; Wise, M.E.; Crowe, S.J. Surveillance for foodborne disease outbreaks—United States, 2009–2015. MMWR Surveill. Summ. 2018, 67, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Outbreaks of Foodborne Illness. Available online: https://www.fda.gov/food/recalls-outbreaks-emergencies/outbreaks-foodborne-illness (accessed on 9 May 2019).
- Oseev, A.; Lucklum, R.; Zubtsov, M.; Schmidt, M.P.; Mukhin, N.V.; Hirsch, S. SAW-Based Phononic Crystal Microfluidic Sensor—Microscale Realization of Velocimetry Approaches for Integrated Analytical Platform Applications. Sensors 2017, 17, 2187. [Google Scholar] [CrossRef] [PubMed]
- Yager, P.; Edwards, T.; Fu, E.; Helton, K.; Nelson, K.; Tam, M.R.; Weigl, B.H. Microfluidic diagnostic technologies for global public health. Nature 2006, 442, 412–418. [Google Scholar] [CrossRef] [PubMed]
- Packard, M.M.; Wheeler, E.K.; Alocilja, E.C.; Shusteff, M. Performance Evaluation of Fast Microfluidic Thermal Lysis of Bacteria for Diagnostic Sample Preparation. Diagnostics 2013, 3, 105–116. [Google Scholar] [CrossRef]
- Yu, H.; Zhou, G.; Chau, F.S.; Wang, S.; Lee, F. Novel polydimethylsiloxane (PDMS) based microchannel fabrication method for lab-on-a-chip application. Sens. Actuators B Chem. 2009, 137, 754–761. [Google Scholar] [CrossRef]
- Zeng, Y.; Novak, R.; Shuga, J.; Smith, M.T.; Mathies, R.A. High-Performance Single Cell Genetic Analysis Using Microfluidic Emulsion Generator Arrays. Anal. Chem. 2010, 82, 3183–3190. [Google Scholar] [CrossRef]
- Liu, B.; Li, M.; Tian, B.; Yang, X.; Yang, J. A positive pressure-driven PDMS pump for fluid handling in microfluidic chips. Microfluid. Nanofluid. 2018, 22, 94. [Google Scholar] [CrossRef]
- Kim, T.H.; Park, J.; Kim, C.J.; Cho, Y.K. Fully integrated lab-on-a-disc for nucleic acid analysis of food-borne pathogens. Anal. Chem. 2014, 86, 3841–3848. [Google Scholar] [CrossRef]
- Roy, E.; Stewart, G.; Mounier, M.; Malic, L.; Peytavi, R.; Clime, L.; Madou, M.; Bossinot, M.; Bergeron, M.G.; Veres, T. From cellular lysis to microarray detection, an integrated thermoplastic elastomer (TPE) point of care Lab on a Disc. Lab Chip 2015, 15, 406–416. [Google Scholar] [CrossRef]
- Joseph, K.; Ibrahim, F.; Cho, J.; Thio, T.H.G.; Al-Faqheri, W.; Madou, M. Design and Development of Micro-Power Generating Device for Biomedical Applications of Lab-on-a-Disc. PLoS ONE 2015, 10, e0136519. [Google Scholar] [CrossRef]
- Uddin, R.; Burger, R.; Donolato, M.; Fock, J.; Creagh, M.; Hansen, M.F.; Boisen, A. Lab-on-a-disc agglutination assay for protein detection by optomagnetic readout and optical imaging using nano- and micro-sized magnetic beads. Biosens. Bioelectron. 2016, 85, 351–357. [Google Scholar] [CrossRef]
- Nwankire, C.E.; Donohoe, G.G.; Zhang, X.; Siegrist, J.; Somers, M.; Kurzbuch, D.; Monaghan, R.; Kitsara, M.; Burger, R.; Hearty, S.; et al. At-line bioprocess monitoring by immunoassay with rotationally controlled serial siphoning and integrated supercritical angle fluorescence optics. Anal. Chim. Acta 2013, 781, 54–62. [Google Scholar] [CrossRef]
- Hwang, H.; Kim, Y.; Cho, J.; Lee, J.Y.; Choi, M.S.; Cho, Y.K. Lab-on-a-Disc for Simultaneous Determination of Nutrients in Water. Anal. Chem. 2013, 85, 2954–2960. [Google Scholar] [CrossRef]
- Mandal, P.K.; Biswas, A.K.; Choi, K.; Pal, U.K. Methods for Rapid Detection of Foodborne Pathogens: An Overview. Am. J. Food Technol. 2011, 6, 87–102. [Google Scholar] [CrossRef]
- Sayad, A.A.; Ibrahim, F.; Uddin, S.M.; Pei, K.X.; Mohktar, M.S.; Madou, M.; Thong, K.L. A microfluidic lab-on-a-disc integrated loop mediated isothermal amplification for foodborne pathogen detection. Sens. Actuators B Chem. 2016, 227, 600–609. [Google Scholar] [CrossRef]
- Uddin, S.M.; Ibrahim, F.; Sayad, A.A.; Thiha, A.; Pei, K.X.; Mohktar, M.S.; Hashim, U.; Cho, J.; Thong, K.L. A Portable Automatic Endpoint Detection System for Amplicons of Loop Mediated Isothermal Amplification on Microfluidic Compact Disk Platform. Sensors 2015, 15, 5376–5389. [Google Scholar] [CrossRef]
- Norian, H.; Field, R.M.; Kymissis, I.; Shepard, K.L. An integrated CMOS quantitative-polymerase-chain-reaction lab-on-chip for point-of-care diagnostics. Lab Chip 2014, 14, 4076–4084. [Google Scholar] [CrossRef]
- Miao, B.; Peng, N.; Li, L.; Li, Z.; Hu, F.; Zhang, Z.; Wang, C. Centrifugal Microfluidic System for Nucleic Acid Amplification and Detection. Sensors 2015, 15, 27954–27968. [Google Scholar] [CrossRef]
- Steffan, R.J.; Atlas, R.M. Polymerase chain reaction: Applications in environmental microbiology. Annu. Rev. Microbiol. 1991, 45, 137–161. [Google Scholar] [CrossRef]
- Vincent, M.; Xu, Y.; Kong, H.M. Helicase-dependent isothermal DNA amplification. EMBO Rep. 2004, 5, 795–800. [Google Scholar] [CrossRef] [PubMed]
- Mairinger, F.D.; Walter, R.F.H.; Vollbrecht, C.; Hager, T.; Worm, K.; Ting, S.; Wohlschlager, J.; Zarogoulidis, P.; Zarogoulidis, K.; Schmid, K.W. Isothermal multiple displacement amplification: A methodical approach enhancing molecular routine diagnostics of microcarcinomas and small biopsies. Onco Targets Ther. 2014, 7, 1441–1447. [Google Scholar] [CrossRef] [PubMed]
- Toley, B.J.; Covelli, I.; Belousov, Y.; Ramachandran, S.; Kline, E.; Scarr, N.; Vermeulen, N.; Mahoney, W.; Lutz, B.R.; Yager, P. Isothermal strand displacement amplification (iSDA): A rapid and sensitive method of nucleic acid amplification for point-of-care diagnosis. Analyst 2015, 140, 7540–7549. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Dong, T. Multifunctional Sample Preparation Kit and On-Chip Quantitative Nucleic Acid Sequence-Based Amplification Tests for Microbial Detection. Anal. Chem. 2012, 84, 8541–8548. [Google Scholar] [CrossRef] [PubMed]
- Østerberg, F.W.; Rizzi, G.; Donolato, M.; Bejhed, R.S.; Mezger, A.; Strömberg, M.; Nilsson, M.; Strømme, M.; Svedlindh, P.; Hansen, M.F. On-Chip Detection of Rolling Circle Amplified DNA Molecules from Bacillus Globigii Spores and Vibrio Cholerae. Small 2014, 10, 2877–2882. [Google Scholar] [CrossRef] [PubMed]
- Tsaloglou, M.N.; Watson, R.J.; Rushworth, C.M.; Zhao, Y.; Niu, X.; Sutton, J.M.; Morgan, H. Real-time microfluidic recombinase polymerase amplification for the toxin B gene of Clostridium difficile on a SlipChip platform. Analyst 2015, 140, 258–264. [Google Scholar] [CrossRef] [PubMed]
- Kurn, N.; Chen, P.; Heath, J.D.; Kopf-Sill, A.; Stephens, K.M.; Wang, S. Novel Isothermal, Linear Nucleic Acid Amplification Systems for Highly Multiplexed Applications. J. Clin. Chem. 2005, 51, 1973–1981. [Google Scholar] [CrossRef]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, e63. [Google Scholar] [CrossRef]
- Duan, Y.B.; Ge, C.Y.; Zhang, X.K.; Wang, J.X.; Zhou, M.G. Development and Evaluation of a Novel and Rapid Detection Assay for Botrytis cinerea Based on Loop-Mediated Isothermal Amplification. PLoS ONE 2014, 9, e111094. [Google Scholar] [CrossRef]
- Mori, Y.; Nagamine, K.; Tomita, N.; Notomi, T. Detection of Loop-Mediated Isothermal Amplification Reaction by Turbidity Derived from Magnesium Pyrophosphate Formation. Biochem. Biophys. Res. Commun. 2001, 289, 150–154. [Google Scholar] [CrossRef]
- Goto, M.; Honda, E.; Ogura, A.; Nomoto, A.; Hanaki, K.I. Colorimetric detection of loop-mediated isothermal amplification reaction by using hydroxy naphthol blue. Biotechniques 2009, 46, 167–172. [Google Scholar] [CrossRef] [PubMed]
- Fischbach, J.; Xander, N.C.; Frohme, M.; Glökler, J.F. Shining a light on LAMP assays’ A comparison of LAMP visualization methods including the novel use of berberine. Biotech 2015, 58, 189–194. [Google Scholar] [CrossRef] [PubMed]
- Choi, M.S.; Yoo, J.C. Automated Centrifugal-Microfluidic Platform for DNA Purification Using Laser Burst Valve and Coriolis Effect. Appl. Biochem. Biotechnol. 2015, 175, 3778–3787. [Google Scholar] [CrossRef] [PubMed]
- Ko, J.; Yoo, J.C. Loop-Mediated Isothermal Amplification Using a Lab-on-a-Disc Device with Thin-film Phase Change Material. Sensors 2018, 186, 54–65. [Google Scholar] [CrossRef] [PubMed]
- McLinden, T.; Sargeant, J.M.; Thomas, M.K.; Papadopoulos, A.; Fazil, A. Component costs of foodborne illness: A scoping review. BMC Public Health 2014, 14, 509. [Google Scholar] [CrossRef] [PubMed]
- Bell, R.L.; Jarvis, K.G.; Ottesen, A.R.; McFarland, M.A.; Brown, E.W. Recent and emerging innovations in Salmonella detection: A food and environmental perspective. Microb. Biotechnol. 2016, 9, 279–292. [Google Scholar] [CrossRef] [PubMed]
- Cho, A.R.; Dong, H.J.; Cho, S. Rapid and Sensitive Detection of Salmonella Spp. by Using a Loop-Mediated Isothermal Amplification Assay in Duck Carcass Sample. Korean J. Food Sci. Anim. Resour. 2013, 33, 655–663. [Google Scholar] [CrossRef][Green Version]
- Fu, M.F.F. Advanced High Strength Natural Fibre Composites in Construction, 1st ed.; ELSEVIER: Amsterdam, The Netherlands, 2016. [Google Scholar]
- Rachmadi, R.F.; Purnama, I. Vehicle color recognition using convolutional neural network. arXiv 2015, arXiv:1510.07391. [Google Scholar]
- Wang, P.T.; Chou, J.J.; Tseng, C.W. Colorimetric Characterization of Color Image Sensors Based on Convolutional Neural Network Modeling. Sens. Mater. 2019, 31, 1513–1522. [Google Scholar] [CrossRef]
- Xia, X.; Xia, C.; Bing, X.; Nan, X. Inception-v3 for flower classification. In Proceedings of the 2017 2nd International Conference on Image, Vision and Computing (ICIVC), Chengdu, China, 2–4 June 2017; pp. 783–787. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the inception architecture for computer vision. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 2818–2826. [Google Scholar]
- Review: Inception-v3—1st Runner Up (Image Classification) in ILSVRC 2015. Available online: https://medium.com/@sh.tsang/review-inception-v3-1st-runner-up-image-classification-in-ilsvrc-2015-17915421f77c (accessed on 5 July 2019).
- Zhang, G.; Brown, E.W.; González-Escalona, N. Comparison of real-time PCR, reverse transcriptase real-time PCR, loop-mediated isothermal amplification, and the FDA conventional microbiological method for the detection of Salmonella spp. in produce. J. Appl. Enciron. Microbiol. 2011, 77, 6495–6501. [Google Scholar] [CrossRef]
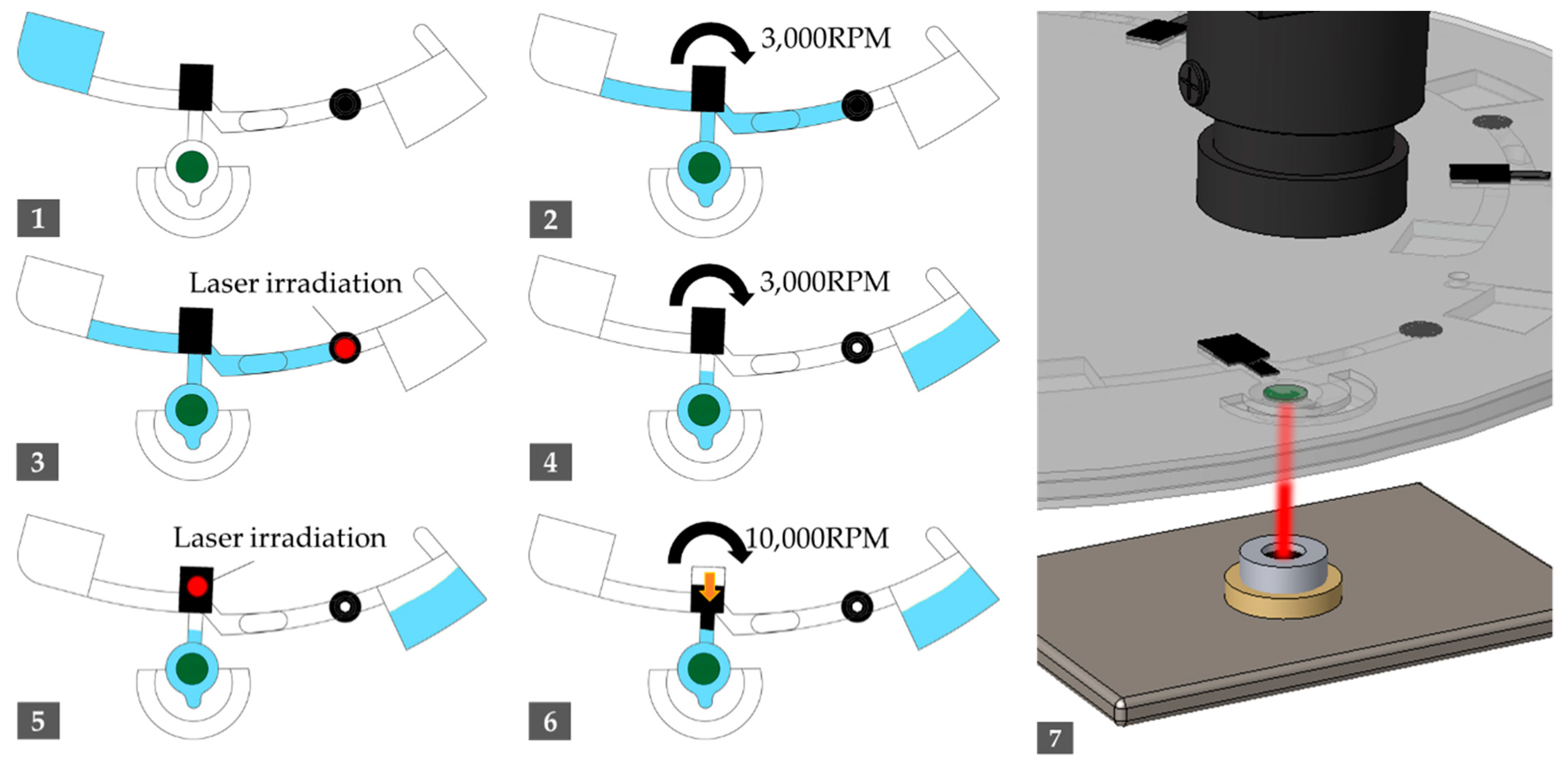


| Step | Operation | Spinning Speed (rpm) | Duration (s) |
|---|---|---|---|
| 1 | Sample injection | - | - |
| 2 | Spin out to move samples in the channel | 3000 | 30 |
| 3 | Irradiate laser to LB valve | - | 10 |
| 4 | Spin out the residual to waste chamber | 3000 | 10 |
| 5 | Irradiate laser to melting EVA valve | - | 60 |
| 6 | Spin out to push the melted EVA valve in radial way | 10,000 | 60 |
| 7 | Irradiate laser to amplification chamber in adjusted duty ratio | - | 2400 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lim, D.Y.S.; Seo, M.-J.; Yoo, J.C. Optical Temperature Control Unit and Convolutional Neural Network for Colorimetric Detection of Loop-Mediated Isothermal Amplification on a Lab-On-A-Disc Platform. Sensors 2019, 19, 3207. https://doi.org/10.3390/s19143207
Lim DYS, Seo M-J, Yoo JC. Optical Temperature Control Unit and Convolutional Neural Network for Colorimetric Detection of Loop-Mediated Isothermal Amplification on a Lab-On-A-Disc Platform. Sensors. 2019; 19(14):3207. https://doi.org/10.3390/s19143207
Chicago/Turabian StyleLim, Da Ye Seul, Moo-Jung Seo, and Jae Chern Yoo. 2019. "Optical Temperature Control Unit and Convolutional Neural Network for Colorimetric Detection of Loop-Mediated Isothermal Amplification on a Lab-On-A-Disc Platform" Sensors 19, no. 14: 3207. https://doi.org/10.3390/s19143207
APA StyleLim, D. Y. S., Seo, M.-J., & Yoo, J. C. (2019). Optical Temperature Control Unit and Convolutional Neural Network for Colorimetric Detection of Loop-Mediated Isothermal Amplification on a Lab-On-A-Disc Platform. Sensors, 19(14), 3207. https://doi.org/10.3390/s19143207




