Aptamer-Based Biosensors to Detect Aquatic Phycotoxins and Cyanotoxins
Abstract
1. Introduction
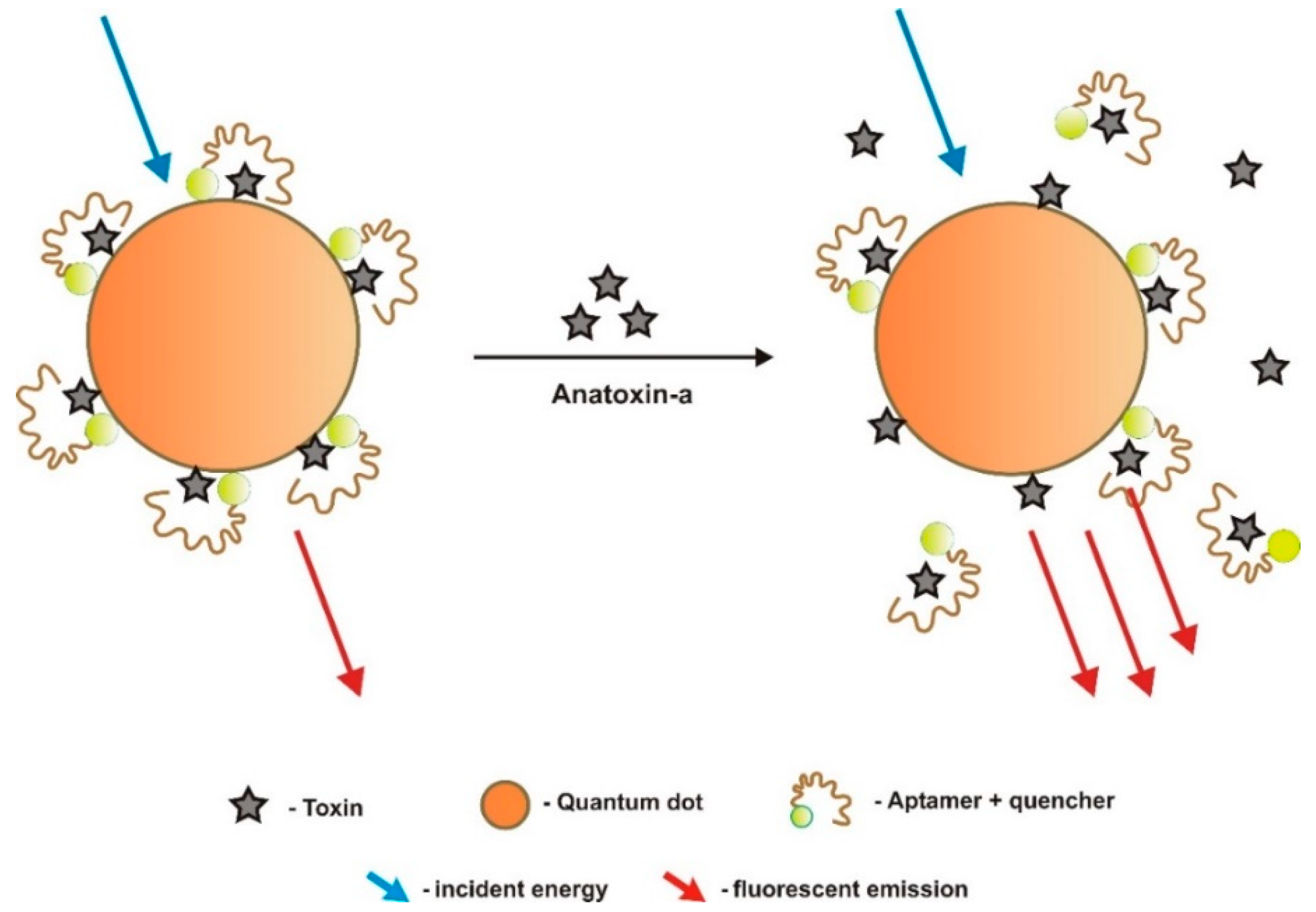
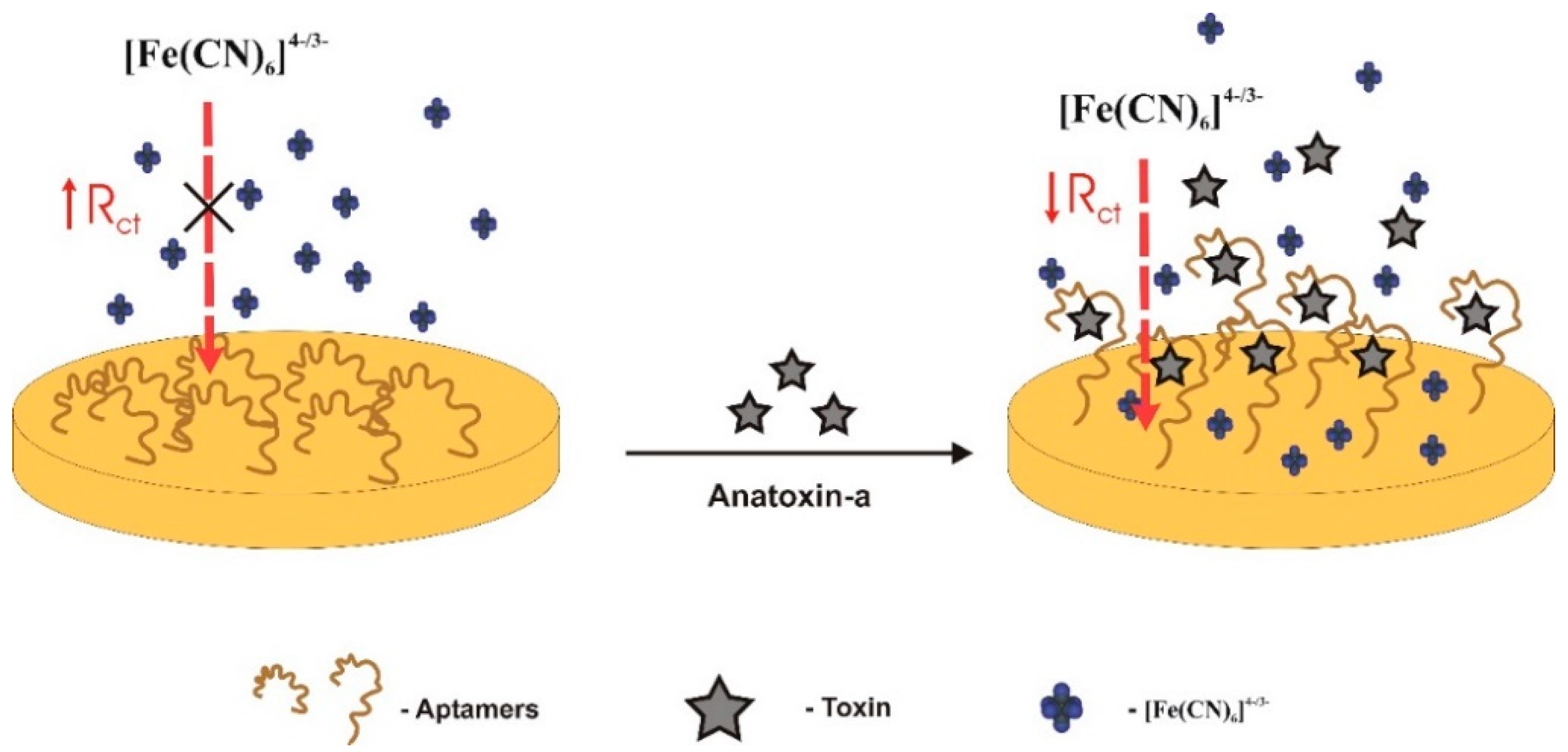
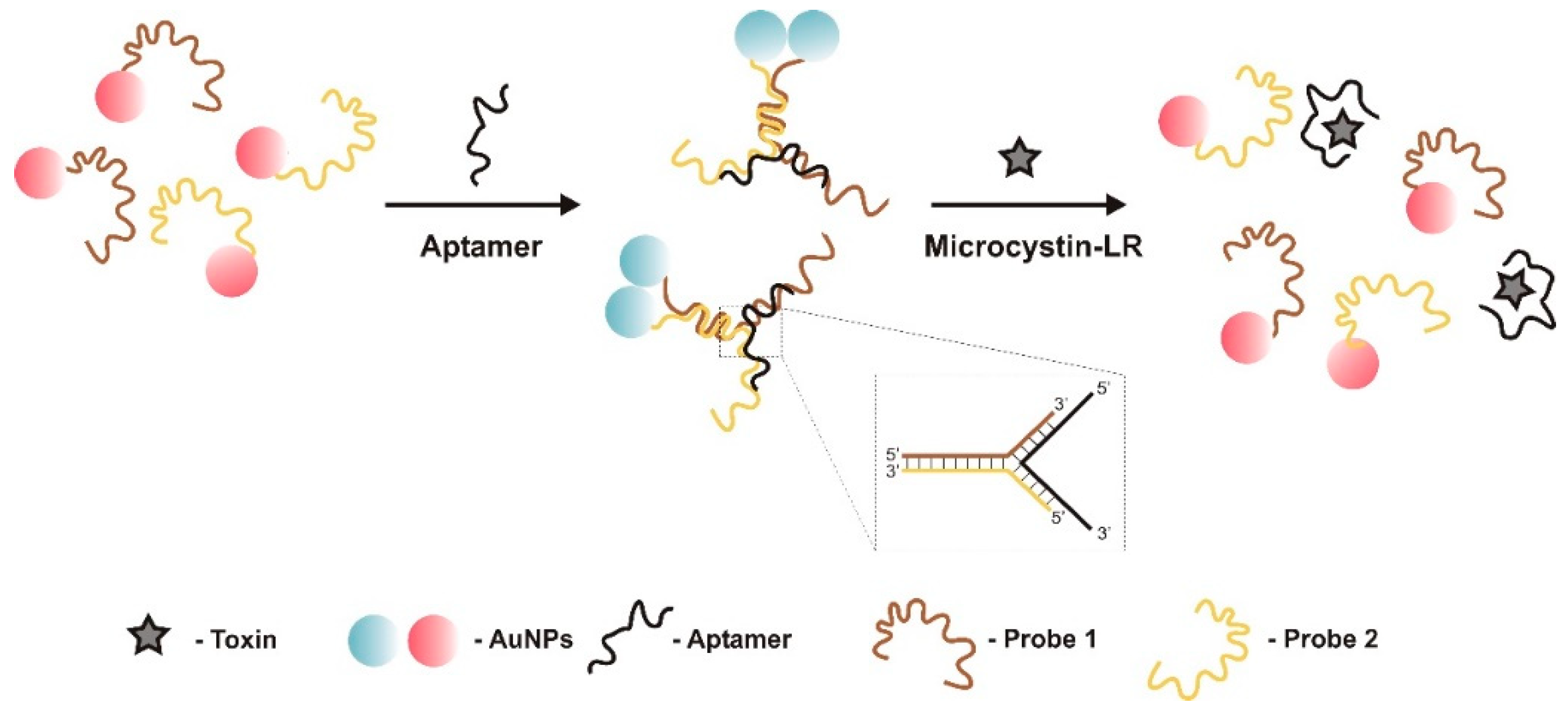
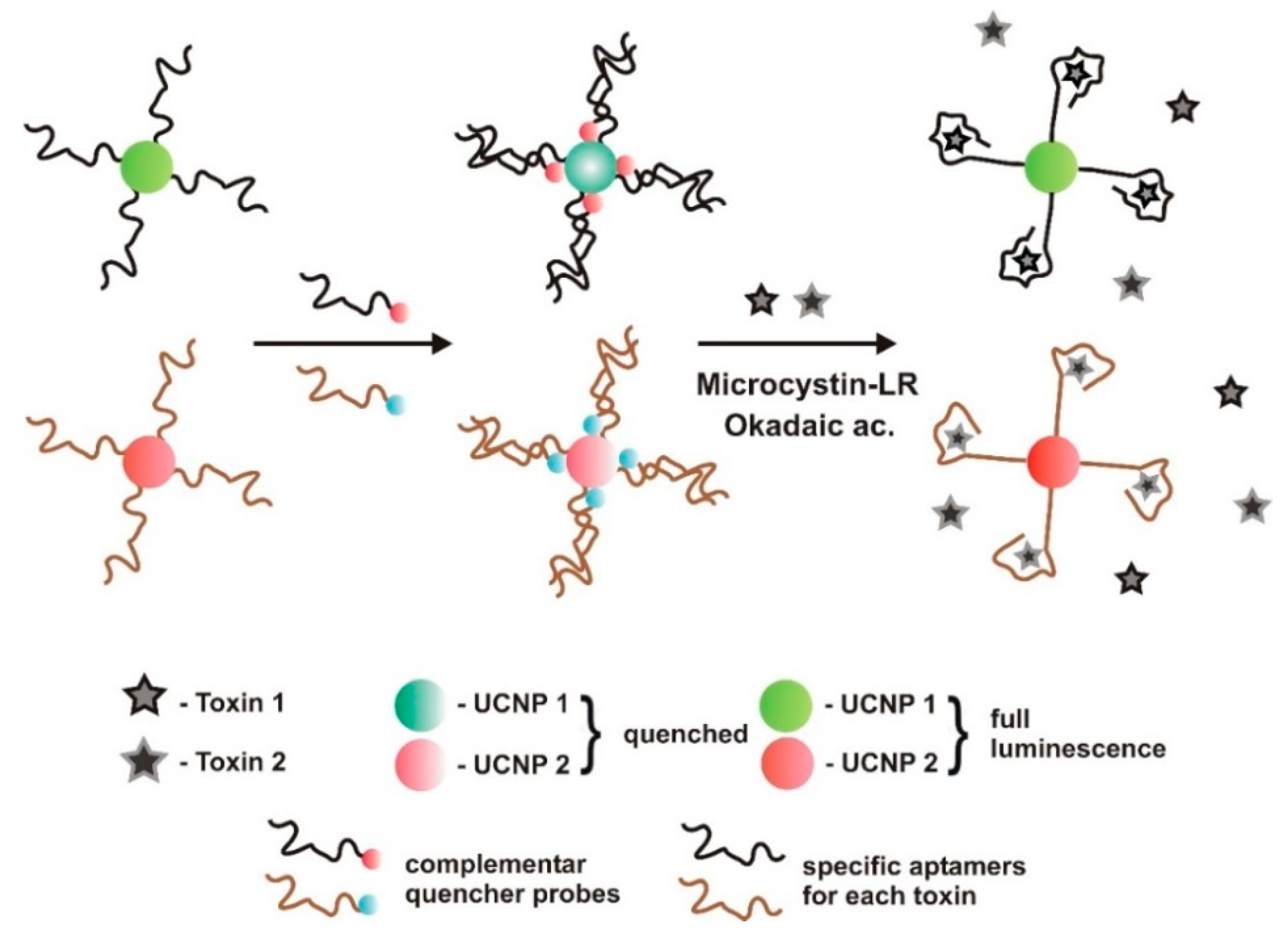
- In target-induced structure switching mode (TISS), the target binds directly to their aptamers, inducing a conformational switch of aptamers, leading to changes on a detectable signal such as weight, fluorescence, color, redox state, conductivity, among others. Most of the sensors analyzed in the present review, and so far developed to detect aquatic toxins, are of this mode [23,28,29,30,31,32,33,34,35,36,38,48,49,50,51,52,53,54,55,56,57].
- Sandwich or sandwich-like sensors mode (SS) takes advantage of the fact that some targets, have two binding sites, which allows them to bind to two recognition elements and form sandwich-like complexes. In the case of aptasensors, this mode can have two formats: aptamer-protein-aptamer and aptamer-protein-antibody. No sensor of this mode was found to have been used to detect aquatic toxins.
- Competitive replacement mode (CR) relies on the replacement of signal modified analytes bound to surface-bound aptamers by the analyte in solution. Signal modified analytes need to be designed and synthesized. After incubation with the non-signal modified target analyte (sample), the signal modified molecules are released from the aptamer into solution, giving rise to a signal related to the target molecule concentration. Another form of this competitive mode consists in replacing the surface bound analyte by the analyte in solution, to modified aptamers that emit or quench a signal [37,60,61,62].
2. Aptasensors for Aquatic Phycotoxins and Cyanotoxins Detection
2.1. Freshwater Toxins
2.1.1. Anatoxin-A (ATX)
2.1.2. Cylindrospermopsin (CYN)
2.1.3. Microcystins
2.2. Marine Toxins
2.2.1. Brevetoxins
2.2.2. Okadaic Acid (OA)
2.2.3. Palytoxin (PTX)
2.2.4. Saxitoxin (STX)
2.2.5. Gonyautoxin (GTX1/4)
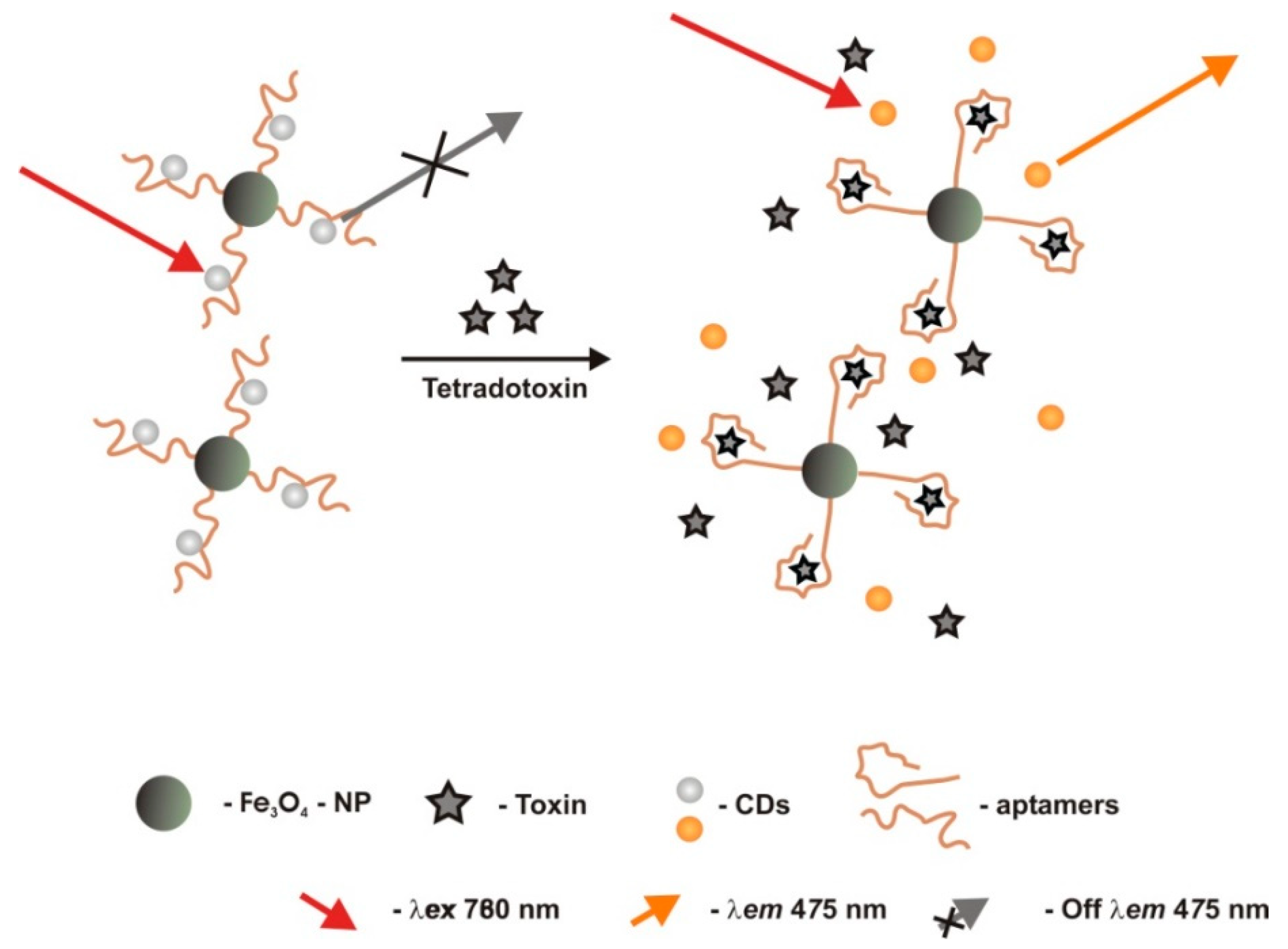
2.2.6. Tetrodotoxin (TTX)
3. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Anderson, D.; Glibert, P.; Burkholder, J. Harmful algal blooms and eutrophication: Nutrient sources, compositions, and consequences. Estuaries 2002, 25, 704–726. [Google Scholar] [CrossRef]
- Mccarthy, M.; Bane, V.; García-Altares, M.; Van Pelt, F.N.A.M.; Furey, A.; O’Halloran, J. Assessment of emerging biotoxins (pinnatoxin G and spirolides) at Europe’s first marine reserve: Lough hyne. Toxicon 2015, 108, 202–209. [Google Scholar] [CrossRef] [PubMed]
- Masó, M.; Garcés, E. Harmful microalgae blooms (HAB): Problematic and conditions that induce them. Mar. Pollut. Bull. 2006, 53, 620–630. [Google Scholar] [CrossRef] [PubMed]
- Smayda, T.J. Reflections on the ballast water dispersal—Harmful algal bloom paradigm. Harmful Algae 2007, 6, 601–622. [Google Scholar] [CrossRef]
- Moreira, C.; Vasconcelos, V.; Antunes, A. Phylogeny and biogeography of cyanobacteria and their produced toxins. Mar. Drugs 2013, 11, 4350–4369. [Google Scholar] [CrossRef] [PubMed]
- Moreira, C.; Ramos, V.; Azevedo, J.; Vasconcelos, V. Methods to detect cyanobacteria and their toxins in the environment. Appl. Microbiol. Biotechnol. 2014, 98, 8073–8082. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.; Pratheepa, V.K.; Botana, L.M.; Vasconcelos, V. Emergent toxins in North Atlantic temperate waters: A challenge for monitoring programs and legislation. Toxins 2015, 7, 859–865. [Google Scholar] [CrossRef] [PubMed]
- Vasconcelos, V. Global changes and the new challenges in the research on cyanotoxin risk evaluation. Limnetica 2015, 34, 149–158. [Google Scholar] [CrossRef]
- Thévenot, D.R.; Toth, K.; Durst, R.A.; Wilson, G.S. Electrochemical biosensors: Recommended definitions and classification. Pure Appl. Chem. 1999, 71, 2333–2348. [Google Scholar] [CrossRef]
- Justino, C.; Duarte, A.; Rocha-Santos, T. Recent progress in biosensors for environmental monitoring: A review. Sensors 2017, 17, 2918. [Google Scholar] [CrossRef] [PubMed]
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Stoltenburg, R.; Reinemann, C.; Strehlitz, B. SELEX—A (r)evolutionary method to generate high-affinity nucleic acid ligands. Biomol. Eng. 2007, 24, 381–403. [Google Scholar] [CrossRef] [PubMed]
- Irvine, D.; Tuerk, C.; Gold, L. Selexion: Systematic evolution of ligands by exponential enrichment with integrated optimization by non-linear analysis. J. Mol. Biol. 1991, 222, 739–761. [Google Scholar] [CrossRef]
- Sampson, T. Aptamers and SELEX: The technology. World Pat. Inf. 2003, 25, 123–129. [Google Scholar] [CrossRef]
- Wang, R.E.; Wu, H.; Niu, Y.; Cai, J. Improving the stability of aptamers by chemical modification. Curr. Med. Chem. 2011, 18, 4126–4138. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Ren, X.J.; Schluesener, H.; Zhang, Z. Aptamers: Selection, modification and application to nervous system diseases. Curr. Med. Chem. 2011, 18, 4159–4168. [Google Scholar] [CrossRef] [PubMed]
- Nonaka, Y.; Yoshida, W.; Abe, K.; Ferri, S.; Schulze, H.; Bachmann, T.T.; Ikebukuro, K. Affinity improvement of a VEGF aptamer by in silico maturation for a sensitive VEGF-detection system. Anal. Chem. 2013, 85, 1132–1137. [Google Scholar] [CrossRef] [PubMed]
- Bullock, T.L.; Sherlin, L.D.; Perona, J.J. Tertiary core rearrangements in a tight binding transfer RNA aptamer. Nat. Struct. Biol. 2000, 7, 497–504. [Google Scholar] [CrossRef] [PubMed]
- Pasternak, A.; Hernandez, F.J.; Rasmussen, L.M.; Vester, B.; Wengel, J. Improved thrombin binding aptamer by incorporation of a single unlocked nucleic acid monomer. Nucl. Acids Res. 2011, 39, 1155–1164. [Google Scholar] [CrossRef] [PubMed]
- Gorenstein, D.G.; He, W.; Volk, D.E.; Elizondo-Riojas, M.-A.; Durland, R.; Engelhardt, J.; Columbia, W. Methods of X-aptamer Generation and Compositions Thereof. U.S. Patent 9,988,623, 5 June 2018. [Google Scholar]
- Lokesh, G.L.; Wang, H.; Lam, C.H.; Thiviyanathan, V.; Ward, N.; Gorenstein, D.G.; Volk, D.E. X-Aptamer Selection and Validation; Humana Press: New York, NY, USA, 2017; pp. 151–174. [Google Scholar]
- Hu, X.; Mu, L.; Wen, J.; Zhou, Q. Immobilized smart RNA on graphene oxide nanosheets to specifically recognize and adsorb trace peptide toxins in drinking water. J. Hazard. Mater. 2012, 213–214, 387–392. [Google Scholar] [CrossRef] [PubMed]
- Cheng, A.K.H.; Sen, D.; Yu, H.Z. Design and testing of aptamer-based electrochemical biosensors for proteins and small molecules. Bioelectrochemistry 2009, 77, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Perumal, V.; Hashim, U. Advances in biosensors: Principle, architecture and applications. J. Appl. Biomed. 2014, 12, 1–15. [Google Scholar] [CrossRef]
- Korotkaya, E.V. Biosensors: Design, classification, and applications in the food industry. Foods Raw Mater. 2014, 2, 161–171. [Google Scholar] [CrossRef]
- Thévenot, D.R.; Toth, K.; Durst, R.A.; Wilson, G.S. Electrochemical biosensors: Recommended definitions and classification. Biosens. Bioelectron. 2001, 16, 121–131. [Google Scholar] [CrossRef]
- Ng, A.; Chinnappan, R.; Eissa, S.; Liu, H.; Tlili, C.; Zourob, M. Selection, characterization, and biosensing application of high affinity congener-specific microcystin-targeting aptamers. Environ. Sci. Technol. 2012, 46, 10697–10703. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Huang, H.; Xu, Y.; Gao, X.; Qiu, B.; Chen, X.; Chen, G. Determination of microcystin-LR in water by a label-free aptamer based electrochemical impedance biosensor. Talanta 2013, 103, 371–374. [Google Scholar] [CrossRef] [PubMed]
- Elshafey, R.; Siaj, M.; Zourob, M. In vitro selection, characterization, and biosensing application of high-affinity cylindrospermopsin-targeting aptamers. Anal. Chem. 2014, 86, 9196–9203. [Google Scholar] [CrossRef] [PubMed]
- Elshafey, R.; Siaj, M.; Zourob, M. DNA aptamers selection and characterization for development of label-free impedimetric aptasensor for neurotoxin anatoxin-a. Biosens. Bioelectron. 2015, 68, 295–302. [Google Scholar] [CrossRef] [PubMed]
- Du, X.; Jiang, D.; Dai, L.; Zhou, L.; Hao, N.; Qian, J.; Qiu, B.; Wang, K. Fabricating photoelectrochemical aptasensor for selectively monitoring microcystin-LR residues in fish based on visible light-responsive BiOBr nanoflakes/N-doped graphene photoelectrode. Biosens. Bioelectron. 2016, 81, 242–248. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Chen, H.; Ma, L.; Liu, D.; Wang, Z. A label-free electrochemical impedance aptasensor for cylindrospermopsin detection based on thionine–graphene nanocomposites. Analyst 2015, 140, 5570–5577. [Google Scholar] [CrossRef] [PubMed]
- Du, X.; Jiang, D.; Hao, N.; Qian, J.; Dai, L.; Zhou, L.; Hu, J.; Wang, K. Building a three-dimensional nano-bio interface for aptasensing: An analytical methodology based on steric hindrance initiated signal amplification effect. Anal. Chem. 2016, 88, 9622–9629. [Google Scholar] [CrossRef] [PubMed]
- Taghdisi, S.M.; Danesh, N.M.; Ramezani, M.; Ghows, N.; Mousavi Shaegh, S.A.; Abnous, K. A novel fluorescent aptasensor for ultrasensitive detection of microcystin-LR based on single-walled carbon nanotubes and dapoxyl. Talanta 2017, 166, 187–192. [Google Scholar] [CrossRef] [PubMed]
- Eissa, S.; Ng, A.; Siaj, M.; Zourob, M. Label-free voltammetric aptasensor for the sensitive detection of microcystin-LR using graphene-modified electrodes. Anal. Chem. 2014, 86, 7551–7557. [Google Scholar] [CrossRef] [PubMed]
- Eissa, S.; Siaj, M.; Zourob, M. Aptamer-based competitive electrochemical biosensor for brevetoxin-2. Biosens. Bioelectron. 2015, 69, 148–154. [Google Scholar] [CrossRef] [PubMed]
- Bilibana, M.P.; Williams, A.R.; Rassie, C.; Sunday, C.E.; Makelane, H.; Wilson, L.; Ntshongontshi, N.; Jijana, A.N.; Masikini, M.; Baker, P.G.L.; et al. Electrochemical aptatoxisensor responses on nanocomposites containing electro-deposited silver nanoparticles on poly(Propyleneimine) dendrimer for the detection of microcystin-LR in freshwater. Sensors 2016, 16, 1901. [Google Scholar] [CrossRef] [PubMed]
- Borisov, S.M.; Wolfbeis, O.S. Optical biosensors. Chem. Rev. 2008, 108, 423–461. [Google Scholar] [CrossRef] [PubMed]
- Damborsky, P.; Vitel, J.; Katrlik, J. Optical biosensors. Essays Biochem. 2016, 60, 91–100. [Google Scholar] [CrossRef] [PubMed]
- Long, F.; Zhu, A.; Shi, H. Recent advances in optical biosensors for environmental monitoring and early warning. Sensors 2013, 13, 13928–13948. [Google Scholar] [CrossRef] [PubMed]
- Adarsh, N.; Shanmugasundaram, M.; Ramaiah, D. Efficient reaction based colorimetric probe for sensitive detection, quantification, and on-site analysis of nitrite ions in natural water resources. Anal. Chem. 2013, 85, 10008–10012. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Liu, Q. Biosensors and bioelectronics on smartphone for portable biochemical detection. Biosens. Bioelectron. 2016, 75, 273–284. [Google Scholar] [CrossRef] [PubMed]
- Roda, A.; Michelini, E.; Zangheri, M.; Di Fusco, M.; Calabria, D.; Simoni, P. Smartphone-based biosensors: A critical review and perspectives. TrAC Trends Anal. Chem. 2016, 79, 317–325. [Google Scholar] [CrossRef]
- Geng, Z.; Zhang, X.; Fan, Z.; Lv, X.; Su, Y.; Chen, H. Recent progress in optical biosensors based on smartphone platforms. Sensors 2017, 17, 2449. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.J.; Chang, Y.C.; Sun, R.; Li, L. A multichannel smartphone optical biosensor for high-throughput point-of-care diagnostics. Biosens. Bioelectron. 2017, 87, 686–692. [Google Scholar] [CrossRef] [PubMed]
- Han, K.; Liang, Z.; Zhou, N. Design strategies for aptamer-based biosensors. Sensors 2010, 10, 4541–4557. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, C.; Kobayashi, T.; Miyake, M.; Shirai, M.; Miyakea, J. Usage of a DNA aptamer as a ligand targeting microcystin. Mol. Cryst. Liq. Cryst. Sci. Technol. Sect. A Mol. Cryst. Liq. Cryst. 2001, 371, 369–374. [Google Scholar] [CrossRef]
- Li, X.; Cheng, R.; Shi, H.; Tang, B.; Xiao, H.; Zhao, G. A simple highly sensitive and selective aptamer-based colorimetric sensor for environmental toxins microcystin-LR in water samples. J. Hazard. Mater. 2016, 304, 474–480. [Google Scholar] [CrossRef] [PubMed]
- Eissa, S.; Ng, A.; Siaj, M.; Tavares, A.C.; Zourob, M. Selection and identification of DNA aptamers against okadaic acid for biosensing application. Anal. Chem. 2013, 85, 11794–11801. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Wan, Z.; Zhong, L.; Li, X.; Wu, Q.; Wang, J.; Wang, P. Label-free okadaic acid detection using growth of gold nanoparticles in sensor gaps as a conductive tag. Biomed. Microdevices 2017, 19, 2–8. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Yu, J.; Ding, X.; Zhao, G. Photoelectrochemical aptasensor for the sensitive detection of microcystin-LR based on graphene functionalized vertically-aligned TiO2 nanotubes. Electroanalysis 2016, 28, 161–168. [Google Scholar] [CrossRef]
- Lv, J.; Zhao, S.; Wu, S.; Wang, Z. Upconversion nanoparticles grafted molybdenum disulfide nanosheets platform for microcystin-LR sensing. Biosens. Bioelectron. 2017, 90, 203–209. [Google Scholar] [CrossRef] [PubMed]
- Alfaro, K.; Bustos, P.; O Sullivan, C.; Conejeros, P. Facile and cost-effective detection of saxitoxin exploiting aptamer structural switching. Food Technol. Biotechnol. 2015, 53, 337–341. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Hu, B.; Zheng, X.; Cao, Y.; Liu, D.; Sun, M.; Jiao, B.; Wang, L. Gonyautoxin 1/4 aptamers with high-affinity and high-specificity: From efficient selection to aptasensor application. Biosens. Bioelectron. 2016, 79, 938–944. [Google Scholar] [CrossRef] [PubMed]
- Fomo, G.; Waryo, T.T.; Sunday, C.E.; Baleg, A.A.; Baker, P.G.; Iwuoha, E.I. Aptameric recognition-modulated electroactivity of poly(4-styrenesolfonic acid)-doped polyaniline films for single-shot detection of tetrodotoxin. Sensors 2015, 15, 22547–22560. [Google Scholar] [CrossRef] [PubMed]
- Jin, H.; Gui, R.; Sun, J.; Wang, Y. Facilely self-assembled magnetic nanoparticles/aptamer/carbon dots nanocomposites for highly sensitive up-conversion fluorescence turn-on detection of tetrodotoxin. Talanta 2018, 176, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Liu, S.; Lin, M.; Chen, X.; Lin, S.; Du, X.; Li, H.; Ye, H.; Qiu, B.; Lin, Z.; et al. Colorimetric detection of microcystin-LR based on disassembly of orient-aggregated gold nanoparticle dimers. Biosens. Bioelectron. 2015, 68, 475–480. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Duan, N.; Zhang, H.; Wang, Z. Simultaneous detection of microcysin-LR and okadaic acid using a dual fluorescence resonance energy transfer aptasensor. Anal. Bioanal. Chem. 2015, 407, 1303–1312. [Google Scholar] [CrossRef] [PubMed]
- Gu, H.; Duan, N.; Wu, S.; Hao, L.; Xia, Y.; Ma, X.; Wang, Z. Graphene oxide-assisted non-immobilized SELEX of okdaic acid aptamer and the analytical application of aptasensor. Sci. Rep. 2016, 6, 21665. [Google Scholar] [CrossRef] [PubMed]
- Jackson, G.W.; Strych, U.; Frank, E.; Willson, R.C.; Ballerstadt, R.; McNichols, R.J. Portable FRET sensing of proteins, hormones, and toxins using DNA aptamers and quantum dots. In Proceedings of the Nanotechnology 2009: Life Sciences, Medicine, Diagnostics, Bio Materials and Composites, Houston, TX, USA, 3–7 May 2009; pp. 205–208. [Google Scholar]
- Gao, S.; Zheng, X.; Hu, B.; Sun, M.; Wu, J.; Jiao, B.; Wang, L. Enzyme-linked, aptamer-based, competitive biolayer interferometry biosensor for palytoxin. Biosens. Bioelectron. 2017, 89, 952–958. [Google Scholar] [CrossRef] [PubMed]
- Labuda, J.; Brett, A.M.O.; Evtugyn, G.; Fojta, M.; Mascini, M.; Ozsoz, M.; Palchetti, I.; Paleček, E.; Wang, J. Electrochemical nucleic acid-based biosensors: Concepts, terms, and methodology (IUPAC Technical Report). Pure Appl. Chem. 2010, 82, 1161–1187. [Google Scholar] [CrossRef]
- United States Environmental Protection Agency. Toxicological Review of Cyanobacterial Toxins: Anatoxin-A (External Review Draft); United States Environmental Protection Agency: Washington, DC, USA, 2006.
- Gu, K.; Famulok, M. In vitro selection of specific aptamers against microcystin-LR. Zhonghua Yu Fang Yi Xue Za Zhi 2004, 38, 369–373. [Google Scholar] [PubMed]
- Tian, R.-Y.; Lin, C.; Yu, S.-Y.; Gong, S.; Hu, P.; Li, Y.-S.; Wu, Z.-C.; Gao, Y.; Zhou, Y.; Liu, Z.-S.; et al. Preparation of a specific ssDNA aptamer for brevetoxin-2 using SELEX. J. Anal. Methods Chem. 2016, 2016, 9241860. [Google Scholar] [CrossRef] [PubMed]
- Handy, S.M.; Yakes, B.J.; DeGrasse, J.A.; Campbell, K.; Elliott, C.T.; Kanyuck, K.M.; DeGrasse, S.L. First report of the use of a saxitoxin–protein conjugate to develop a DNA aptamer to a small molecule toxin. Toxicon 2013, 61, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Hu, B.; Gao, S.X.; Liu, D.J.; Sun, M.J.; Jiao, B.H.; Wang, L.H. A saxitoxin-binding aptamer with higher affinity and inhibitory activity optimized by rational site-directed mutagenesis and truncation. Toxicon 2015, 101, 41–47. [Google Scholar] [CrossRef] [PubMed]
- Shao, B.; Gao, X.; Yang, F.; Chen, W.; Miao, T.; Peng, J. Screening and structure analysis of the aptamer against tetrodotoxin. J. Chin. Inst. Food Sci. Technol. 2012, 2, 347–351. [Google Scholar]
- Bruchez, M.; Moronne, M.; Gin, P.; Weiss, S.; Alivisatos, A.P. Semiconductor nanocrystals as fluorescent biological labels. Science 1998, 281, 2013–2016. [Google Scholar] [CrossRef] [PubMed]
- Chan, W.C.W.; Nie, S. Quantum dot bioconjugates for ultrasensitive nonisotopic detection. Science 1998, 281, 2016–2018. [Google Scholar] [CrossRef] [PubMed]
- Fawell, J.K.; Mitchell, R.E.; Hill, R.E.; Everett, D.J. The toxicity of cyanobacterial toxins in the mouse: II anatoxin-a. Hum. Exp. Toxicol. 1999, 18, 168–173. [Google Scholar] [CrossRef] [PubMed]
- Humpage, A.R.; Falconer, I.R. Oral toxicity of the cyanobacterial toxin cylindrospermopsin in male Swiss albino mice: Determination of no observed adverse effect level for deriving a drinking water guideline value. Environ. Toxicol. 2003, 18, 94–103. [Google Scholar] [CrossRef] [PubMed]
- Westrick, J.A.; Szlag, D.C.; Southwell, B.J.; Sinclair, J. A review of cyanobacteria and cyanotoxins removal/inactivation in drinking water treatment. Anal. Bioanal. Chem. 2010, 397, 1705–1714. [Google Scholar] [CrossRef] [PubMed]
- Catherine, A.; Bernard, C.; Spoof, L.; Bruno, M. Microcystins and nodularins. In Handbook of Cyanobacterial Monitoring and Cyanotoxin Analysis; John Wiley & Sons, Ltd.: Chichester, UK, 2017; pp. 107–126. ISBN 9517652593. [Google Scholar]
- Botes, D.P.; Wessels, P.L.; Kruger, H.; Runnegar, M.T.C.; Santikarn, S.; Smith, R.J.; Barna, J.C.J.; Williams, D.H. Structural studies on cyanoginosins-LR, -YR, -YA, and -YM, peptide toxins from Microcystis aeruginosa. J. Chem. Soc. Perkin Trans. 1985, 1, 2747. [Google Scholar] [CrossRef]
- Furey, A.; Allis, O.; Ortea, P.; Lehane, M.; James, K. Hepatotoxins: Context and chemical determination. In Freshwater Toxins, Pharmacology, Physiology, and Detection; Botana, L.M., Ed.; CRC Press: Boca Raton, FL, USA, 2008; pp. 845–886. [Google Scholar]
- World Health Organization. Guidelines for Drinking-Water Quality, 2nd ed.; World Health Organization: Geneva, Switzerland, 1998. [Google Scholar]
- World Health Organization. Cyanobacterial Toxins: Microcystin-LR in Drinking Water; Background Document for Development of WHO Guidelines for Drinking-Water Quality; World Health Organization: Geneva, Switzerland, 2003. [Google Scholar]
- Lawton, L.A.; Chambers, H.; Edwards, C.; Nwaopara, A.A.; Healy, M. Rapid detection of microcystins in cells and water. Toxicon 2010, 55, 973–978. [Google Scholar] [CrossRef] [PubMed]
- Gu, K.; Famulok, M. In vitro selection of specific aptamers against microcystin-LR. Zhonghua Yu Fang Yi Xue Za Zhi 2004, 38, 369–373. [Google Scholar] [PubMed]
- Auzel, F. Upconversion and anti-stokes processes with f and d ions in Solids. Chem. Rev. 2004, 104, 139–173. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Duan, N.; Ma, X.; Xia, Y.; Wang, H.; Wang, Z.; Zhang, Q. Multiplexed fluorescence resonance energy transfer aptasensor between upconversion nanoparticles and graphene oxide for the simultaneous determination of mycotoxins. Anal. Chem. 2012, 84, 6263–6270. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Nan, F.; Zhao, J.; Yang, T.; Ge, T.; Jiao, K. A label-free ultrasensitive electrochemical DNA sensor based on thin-layer MoS2 nanosheets with high electrochemical activity. Biosens. Bioelectron. 2015, 64, 386–391. [Google Scholar] [CrossRef] [PubMed]
- Yanfen, F.; Yingping, H.; Jing, Y.; Pan, W.; Genwei, C. Unique ability of BiOBr to decarboxylate d -Glu and d -MeAsp in the photocatalytic degradation of microcystin-LR in water. Environ. Sci. Technol. 2011, 45, 1593–1600. [Google Scholar] [CrossRef] [PubMed]
- Ou, X.; Tan, X.; Liu, X.; Lu, Q.; Chen, S.; Wei, S. A signal-on electrochemiluminescence biosensor for detecting Con A using phenoxy dextran-graphite-like carbon nitride as signal probe. Biosens. Bioelectron. 2015, 70, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Yu, Y.; Li, Q.; Zhou, M.; Cui, H. A homogeneous signal-on strategy for the detection of rpoB genes of Mycobacterium tuberculosis based on electrochemiluminescent graphene oxide and ferrocene quenching. Anal. Chem. 2014, 86, 1608–1613. [Google Scholar] [CrossRef] [PubMed]
- Hirao, A.; Yoo, H.-S. Dendrimer-like star-branched polymers: Novel structurally well-defined hyperbranched polymers. Polym. J. 2011, 43, 2–17. [Google Scholar] [CrossRef]
- Chen, X.; Wang, Y.; Zhang, Y.; Chen, Z.; Liu, Y.; Li, Z.; Li, J. Sensitive electrochemical aptamer biosensor for dynamic cell surface N-glycan evaluation featuring multivalent recognition and signal amplification on a dendrimer-graphene electrode interface. Anal. Chem. 2014, 86, 4278–4286. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Liu, G.; Yuan, R.; Chai, Y.; Gan, X.; Bai, L. Dendrimer functionalized reduced graphene oxide as nanocarrier for sensitive pseudobienzyme electrochemical aptasensor. Biosens. Bioelectron. 2013, 42, 474–480. [Google Scholar] [CrossRef] [PubMed]
- Beakley, L.W.; Yost, S.E.; Cheng, R.; Chandler, B.D. Nanocomposite catalysts: Dendrimer encapsulated nanoparticles immobilized in sol-gel silica. Appl. Catal. A Gen. 2005, 292, 124–129. [Google Scholar] [CrossRef]
- Liu, J.; Liu, J.; Yang, L.; Chen, X.; Zhang, M.; Meng, F.; Luo, T.; Li, M. Nanomaterial-assisted signal enhancement of hybridization for DNA biosensors: A review. Sensors 2009, 9, 7343–7364. [Google Scholar] [CrossRef] [PubMed]
- Gan, X.; Liu, T.; Zhong, J.; Liu, X.; Li, G. Effect of silver nanoparticles on the electron transfer reactivity and the catalytic activity of myoglobin. ChemBioChem 2004, 5, 1686–1691. [Google Scholar] [CrossRef] [PubMed]
- Radisavljevic, B.; Radenovic, A.; Brivio, J.; Giacometti, V.; Kis, A. Single-layer MoS2 transistors. Nat. Nanotechnol. 2011, 6, 147–150. [Google Scholar] [CrossRef] [PubMed]
- Yin, Z.; Li, H.; Li, H.; Jiang, L.; Shi, Y.; Sun, Y.; Lu, G.; Zhang, Q.; Chen, X.; Zhang, H. Single-layer MoS2phototransistors. ACS Nano 2012, 6, 74–80. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Yin, Z.; Du, Y.; Huang, X.; Zeng, Z.; Fan, Z.; Liu, H.; Wang, J.; Zhang, H. Synthesis of few-layer MoS2 nanosheet-coated TiO2 nanobelt heterostructures for enhanced photocatalytic activities. Small 2013, 9, 140–147. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Zeng, Z.; Li, H.; Li, F.; Fan, C.; Zhang, H. Single-layer MoS2 -based nanoprobes for homogeneous detection of biomolecules. J. Am. Chem. Soc. 2013, 135, 5998–6001. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority. Marine biotoxins in shellfish—Saxitoxin group—Scientific opinion of the panel on contaminants in the food chain. EFSA J. 2009, 1019, 1–76. [Google Scholar] [CrossRef]
- Twiner, M.J.; Bottein Dechraoui, M.-Y.; Wang, Z.; Mikulski, C.M.; Henry, M.S.; Pierce, R.H.; Doucette, G.J. Extraction and analysis of lipophilic brevetoxins from the red tide dinoflagellate Karenia brevis. Anal. Biochem. 2007, 369, 128–135. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.; Rodriguez, I.; Barreiro, A.; Kaufmann, M.; Neto, A.I.; Hassouani, M.; Sabour, B.; Alfonso, A.; Botana, L.M.; Vasconcelos, V. New invertebrate vectors of okadaic acid from the North Atlantic waters—Portugal (Azores and Madeira) and Morocco. Toxins 2015, 7, 5337–5347. [Google Scholar] [CrossRef] [PubMed]
- Toh, S.Y.; Citartan, M.; Gopinath, S.C.B.; Tang, T.-H. Aptamers as a replacement for antibodies in enzyme-linked immunosorbent assay. Biosens. Bioelectron. 2015, 64, 392–403. [Google Scholar] [CrossRef] [PubMed]
- Concepcion, J.; Witte, K.; Wartchow, C.; Choo, S.; Yao, D.; Persson, H.; Wei, J.; Li, P.; Heidecker, B.; Ma, W.; et al. Label-free detection of biomolecular interactions using BioLayer interferometry for kinetic characterization. Comb. Chem. High Throughput Screen. 2009, 12, 791–800. [Google Scholar] [CrossRef] [PubMed]
- Deeds, J.R.; Landsberg, J.H.; Etheridge, S.M.; Pitcher, G.C.; Longan, S.W. Non-traditional vectors for paralytic shellfish poisoning. Mar. Drugs 2008, 6, 308–348. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.; Barreiro, A.; Rodriguez, P.; Otero, P.; Azevedo, J.; Alfonso, A.; Botana, L.M.; Vasconcelos, V. New invertebrate vectors for PST, spirolides and okadaic acid in the North Atlantic. Mar. Drugs 2013, 11, 1936–1960. [Google Scholar] [CrossRef] [PubMed]
- Ruttkay-Nedecky, B.; Kudr, J.; Nejdl, L.; Maskova, D.; Kizek, R.; Adam, V. G-quadruplexes as sensing probes. Molecules 2013, 18, 14760–14779. [Google Scholar] [CrossRef] [PubMed]
- Hu, P.; Liu, Z.; Tian, R.; Ren, H.; Wang, X.; Lin, C.; Gong, S.; Meng, X.; Wang, G.; Zhou, Y.; et al. Selection and identification of a DNA aptamer that mimics saxitoxin in antibody binding. J. Agric. Food Chem. 2013, 61, 3533–3541. [Google Scholar] [CrossRef] [PubMed]
- Dörr, F.A.; Kovačević, B.; Maksić, Z.B.; Pinto, E.; Volmer, D.A. Intriguing differences in the gas-phase dissociation behavior of protonated and deprotonated gonyautoxin epimers. J. Am. Soc. Mass Spectrom. 2011, 22, 2011–2020. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.; Rey, V.; Botana, A.; Vasconcelos, V.; Botana, L. Determination of gonyautoxin-4 in echinoderms and gastropod matrices by conversion to neosaxitoxin using 2-mercaptoethanol and post-column oxidation liquid chromatography with fluorescence detection. Toxins 2016, 8, 11. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-W.; Tatavarty, R.; Kim, D.W.; Jung, H.-T.; Gu, M.B. Immobilization-free screening of aptamers assisted by graphene oxide. Chem. Commun. 2012, 48, 2071–2073. [Google Scholar] [CrossRef] [PubMed]
- Pratheepa, V.; Alex, A.; Silva, M.; Vasconcelos, V. Bacterial diversity and tetrodotoxin analysis in the viscera of the gastropods from Portuguese coast. Toxicon 2016, 119, 186–193. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.; Azevedo, J.; Rodriguez, P.; Alfonso, A.; Botana, L.M.; Vasconcelos, V. New gastropod vectors and tetrodotoxin potential expansion in temperate waters of the Atlantic Ocean. Mar. Drugs 2012, 10, 712–726. [Google Scholar] [CrossRef] [PubMed]

| ADVANTAGES AND CHARACTERISTICS OF APTAMERS USE IN SENSORS | AMENABLE to DETECT SMALL MOLECULES, for aptamers are independent of ligand’s size or type, being convenient to detect small or large molecules or even whole cells. Animal immunization for low-molecular-weight targets without immunogenicity is not practical and aquatic toxins are often small molecules. |
| AMENABLE to DETECT TOXINS, which may possibly kill live organisms or cells used to produce antibodies. Toxins do not have a toxic effect on aptamers. | |
| NO ETHICAL CONCERNS related to USE OF ANIMALS since aptamers synthesis is in vitro with no animals involved. | |
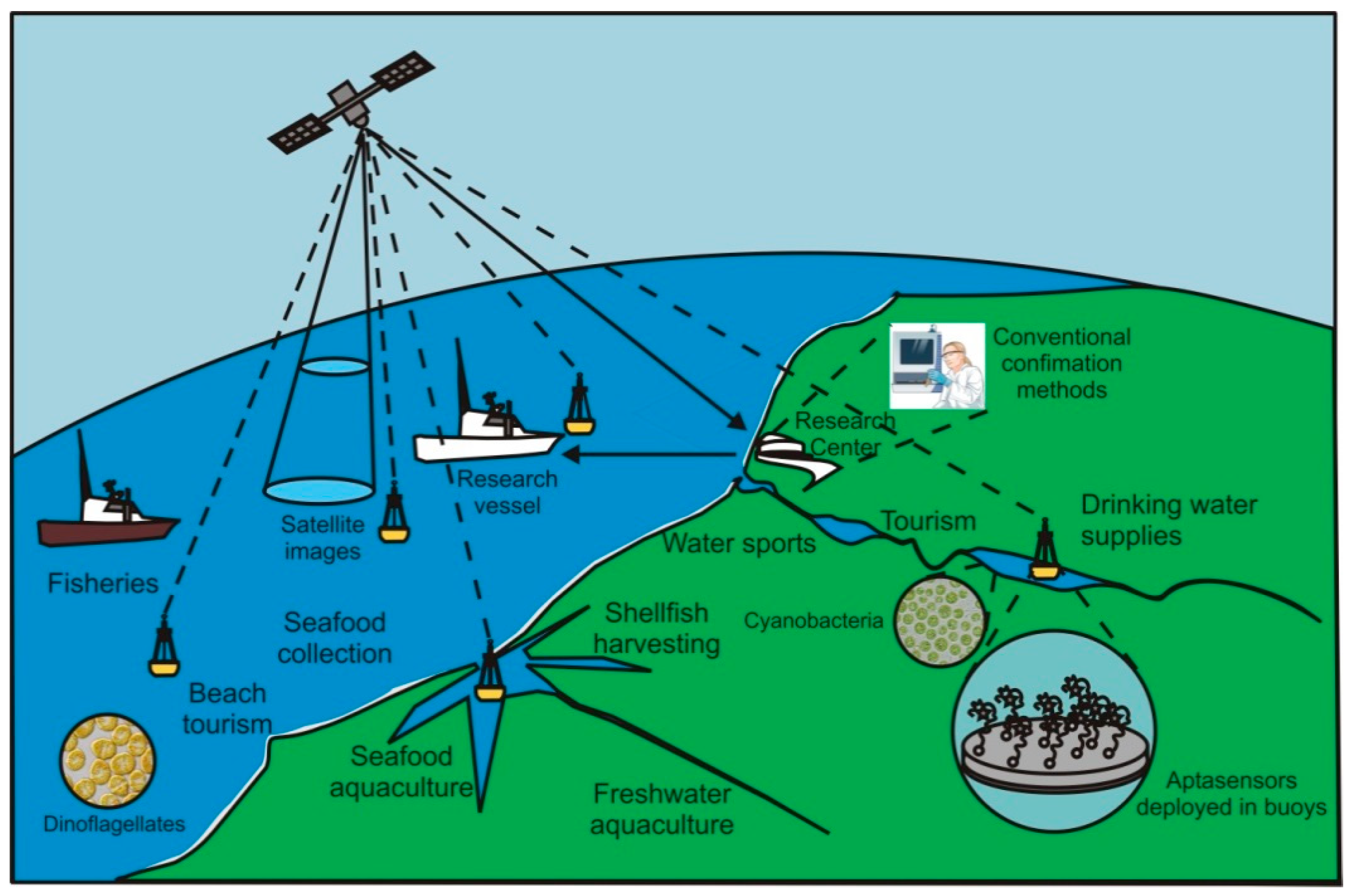
| PORTABILITY and ON-SITE REAL-TIME application are ideal features for risk assessment analysis and risk management of toxic blooms monitoring on water bodies used for drinking, bathing, food collection, and production. | |
| HIGH PERFORMANCE offered by aptamer-based sensors, similar to that offered by antibody-based ones, include selectivity, specificity, and accuracy. Their limits of detection may be well below the limits imposed by the legislation for aquatic toxins. | |
| STABILITY, in which DNA aptamers are very chemically stable; robust at room temperature; and their thermal denaturation is reversible. They are amenable for commercially shipping at ambient temperature and have a long shelf life. Due to its low stability, RNA aptamers are less suitable for environmental monitoring applications, unless stabilization techniques are used to protect them from nucleases. Overall, aptamers are amenable for field studies, deployment in oceanographic buoys or to be stored in the luggage of seafood and freshwater consumers. | |
| VERSATILITY, as they can be synthesized for a wide target diversity, modified by addition of chemical groups (e.g., −SH, −NH2, biotin) and labels (e.g., electrochemical probes, fluorophores, quenchers), adding or cutting nucleotides of the originally selected molecule, and being immobilized on surfaces quite easily, allowing diversity in designing, and detection strategies. | |
| TRAINED PERSONNEL or EXPENSIVE EQUIPMENT in specialized laboratories are not needed, which makes possible the use of sensors by water or food consumers or by low-tech remote laboratories. | |
| REVERSIBILITY of the binding between aptamers and target molecules, allowing multiple uses. | |
| EASY, HANDY and RAPID TO PRODUCE, as aptamers synthesis is very convenient, including the introduction of chemical modifications. They can be developed in weeks while antibodies need months. | |
| MULTIPLEXING ability by using combinations of aptamers marked with different labels in the same aptasensor, allowing for multiple target detection easily. Toxins may occur concomitantly with others. | |
| CONSISTENCY from batch-to-batch production, since aptamers are synthesized in vitro, thereby independent from the intrinsic variability of living organisms. | |
| NO PROPRIETARY in terms of aptamer sequence. Antibodies are often proprietary, while the sequence of aptamers is public to any researcher who wishes to make their synthesis and make use of them. | |
| LOW PRICE, aptamers cost is low when compared to antibodies. Once the nucleic acids sequence is known, chemical or enzymatic syntheses are possible at relatively low cost. | |
| MINIMAL SAMPLE PREPARATION is involved, even for complex matrices, due to their specificity. |
| Target Toxin | Type of Sensor | Design Strategy Mode * | Material Used as Transducer/Electrode/Platform/Redox Markers | Aptamer Immobilization Method | Linear Range of Detection (LRD) | Limit of Detection (LOD) | Reproducibility | Selectivity | Matrix Analyzed and Recover Rate ** | References |
|---|---|---|---|---|---|---|---|---|---|---|
| ATX | Optic (label based)/fluorescence FRET | CR | fluorescent quantum dot nanocrystals and quencher fluorophore dyes | Aptamers were immobilized on streptavidin agarose beads and biotinylated at the 5′ end | Up to 100 μg/mL | - | - | - | - | [61] |
| Electrochemical (label-free)/impedimetric | TISS | gold electrode/[Fe(CN)6]4−/3− (redox marker) | Self-assembled monolayer (SAM) covalently bound through Au-S alkanethiol interaction | 1–100 nM (0.165–16.5 g/L) | 0.5 nM | 6.5% (N = 7) | Good selectivity. CYN, MC-LR | spiked tap water and certified samples 94.8–108.6% (1.4–6.3%) | [31] | |
| CYN | Electrochemical (label-free)/impedimetric | TISS | gold electrode/[Fe(CN)6]4−/3− | SAM covalently bound through Au-S alkanethiol interaction | 0.1–80 nM (0.041–33.15 g/L) | 0.1 nM (0.039 μg/L) | 2.0–9.0% (N = 3) | Good selectivity. CYN, MC-LR, ATX-A | spiked tap water and certified samples 95.8–103.2% (2.0–9.6%) | [30] |
| Electrochemical (label-free)/impedimetric | TISS | thionine–graphene nanocomposite modified glassy carbon electrode (GCE)/[Fe(CN)6]4−/3− | Covalently grafted through glutaraldehyde (cross-linked) | 1–200 nM (0.39–78 g/L) | 0.3 nM (0.117 μg/L) | 1.2% (N = 3) | Good selectivity. OA, MC-LR | spiked lake water 96.3–104.6% (1.3–8.5%) | [33] | |
| MC-LR | Optic (label-free)/Surface plasmon resonance | TISS | sensor chip (BIACORE international AB) | Affinity binding (biotin-streptavidin) | 50.24–1005 M (50–1000 mg/L) | 50.24 nM (50 mg/L) | 10–20% | Poor selectivity. MC-YR, -RR | - | [48] |
| Electrochemical (label-free)/square wave voltammetric | TISS | gold electrode/[Ru-(NH3)6]3+ | SAM covalently bound through AU-S alkanethiol interaction | 0.01–10 nM | 0.0118 nM | - | Good selectivity. MC-YR, -LA | - | [28] | |
| No aptasensor was developed. The adsorption capacity of GO nanosheets was characterized. | TISS | graphene oxide (GO) nanosheets | carboxylic groups of GO are activated by 1-ethyl-3-(3-dimethylamino-propyl) carbodiimide hydrochloride/N-hydroy succinimideDC/NHS) to which NH2-aptamers covalently bind | 0.5–1 ng/L | 0.5 ng/L | - | Good selectivity. MC-RR, -LW, nodularin | spiked tap water 88.3% | [23] | |
| Electrochemical (label-free)/impedimetric | TISS | gold electrode/[Fe(CN)6]4−/3− | SAM covalently bound through AU-S alkanethiol interaction | 0.05–100 nM | 0.018 nM | 3.52% (N = 5). | Good selectivity. against low concentrations of MC-RR | spiked lake, river and tap water 91.2–113.7% (2.6–4.2%) | [29] | |
| Electrochemical (label-free)/square wave voltammetric | TISS | graphene-modified screen-printed carbon electrodes (SPEs)/[Fe(CN)6]4−/3− | physisorption. π-π stacking interactions between the graphene hexagonal cells and the DNA nucleobases. | 0.1–1000 nM | 1.9 pM | - | Good selectivity. OA, MC-LA, -YR | spiked tap water and fish samples 91.7–98.1% (1.67–10.93%) | [36] | |
| Optic (label-based)/colorimetric | TID | polyethylene glycol (PEG) modified AuNPs | Aptamers hybridize to complementary DNA probes immobilized on the surface of AuNPs | 0.1–250 nM | 0.05 nM | 3.6% | Good selectivity. MC-LA, -YR | river, lake, and pond water samples 89–100% and human serum 90–103% | [58] | |
| Optic (label-based)/fluorescence dual FRET | TID | green and red lanthanide upconversion nanoparticles (UCNPs) and quenchers | Affinity binding (biotin-streptavidin) to UCNPs | 0.1–50 μg/L | 0.025 μg/L | 6.47% (N = 7) | Good selectivity. MC-LA, -YR | water, fish, and shrimps 99.1–110.8% | [59] | |
| Electrochemical (label-free)/Amperometric | TISS | graphene co-doped with BiOBr nanoflakes and nitrogen over an indium tin oxide (ITO) electrode | π-π staking interaction between graphene and aptamers | 0.0001–100 nM | 0.03 pM | 3.46% (N = 5) | Good selectivity. MC-LA, -YR | spiked fish collected at local supermarket 97.8–101.6% (2.52–5.14%) | [32] | |
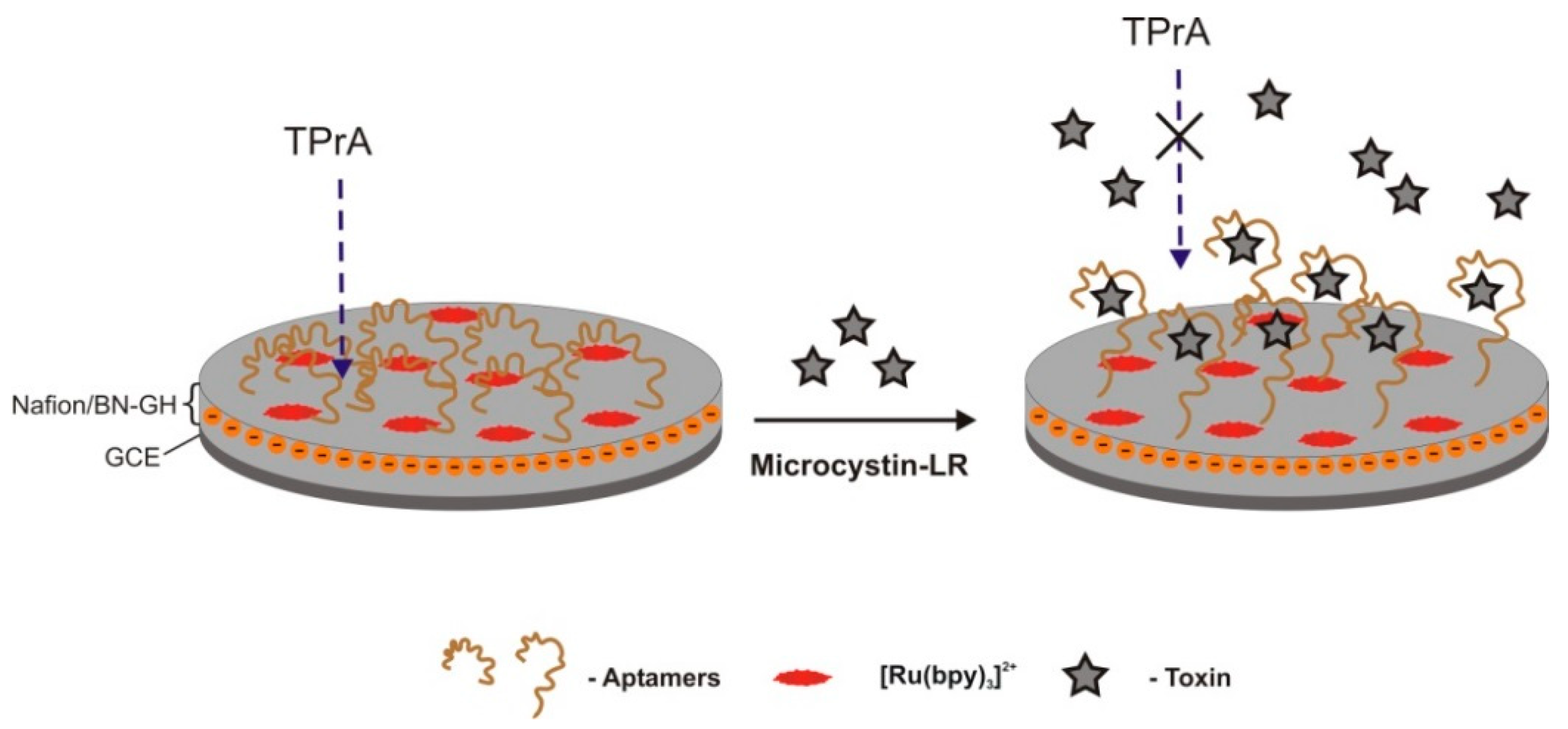
| Optic (label-free)/luminescence | TISS | graphene hydrogel co-doped with boron and nitrogen, self-assembled on a Ru(bpy)32+ immobilization platform/GCE | electrostatic adsorption | 0.0001–1 nM | 0.03 pM | 5.3% (N = 12) | Good selectivity. MC-LA, -YR | human serum, tap water and contaminated water 91.0−104.0% | [34] | |
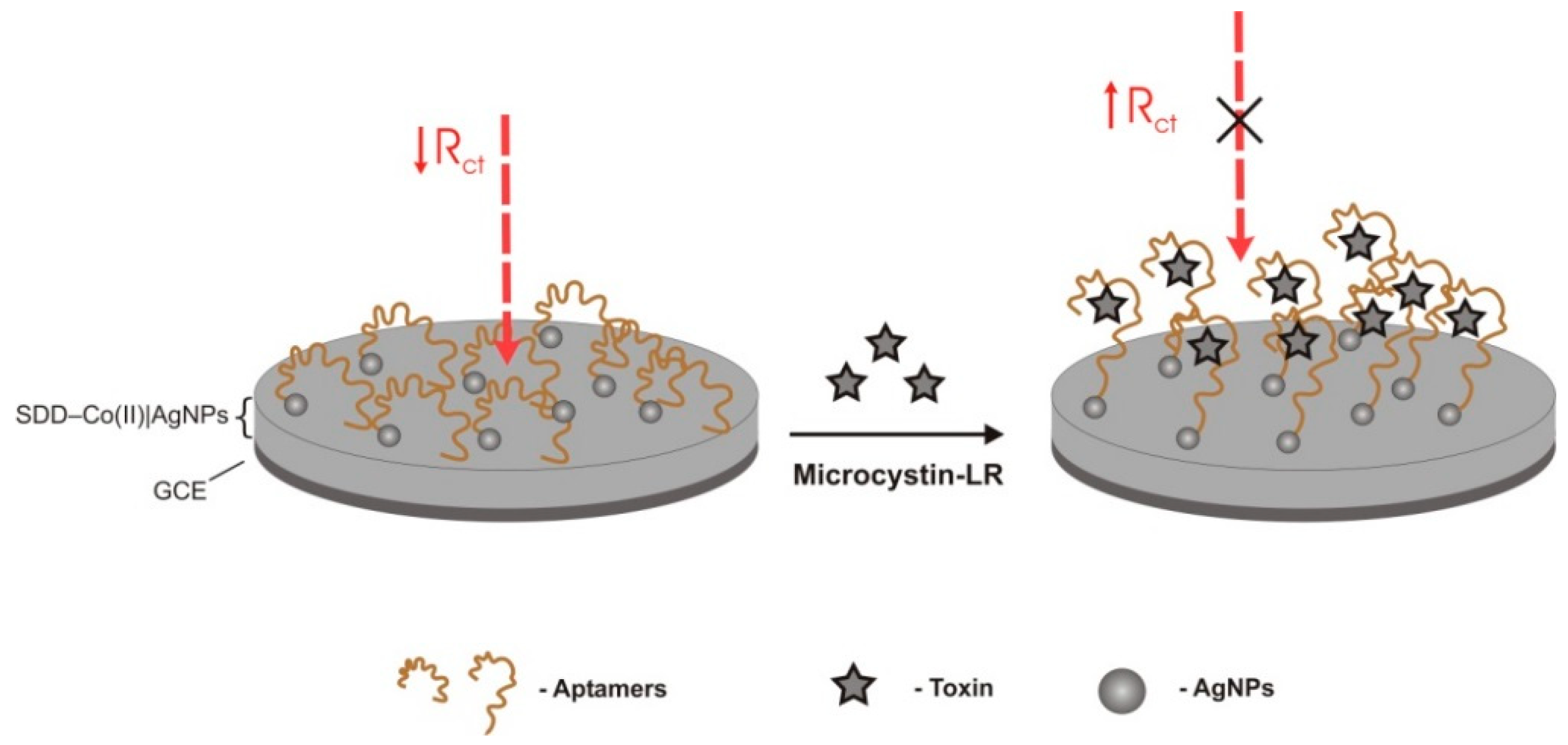
| Electrochemical/impedimetric | TISS | cobalt (II) salicylaldimine metallodendrimer (SDD–Co(II)) doped with electro-synthesized silver NPs over a GCE | SAM covalently bound through Ag-S alkanethiol interaction | 0.1–1.1 μg/L | 0.04 μg/L | - | Good selectivity. Nodularin-R, MC-RR, -YR, 17-estradiol, zearalenone | spiked tap, distilled, and wastewater samples 94–115% (0.96–5.06%) | [38] | |
| Optic (label-based)/colorimetric | TISS | AuNPs | Physisorption at the AuNPs’ surface by coordination between the Au atoms and the N atoms of the exposed bases | 0.5–7500 nM | 0.37 nM | - | Good selectivity. acetamiprid, glyphosate, trichlorfon, clofentezine, atrazine | tap and pond water 95–102% (7.4–10.7%) | [49] | |
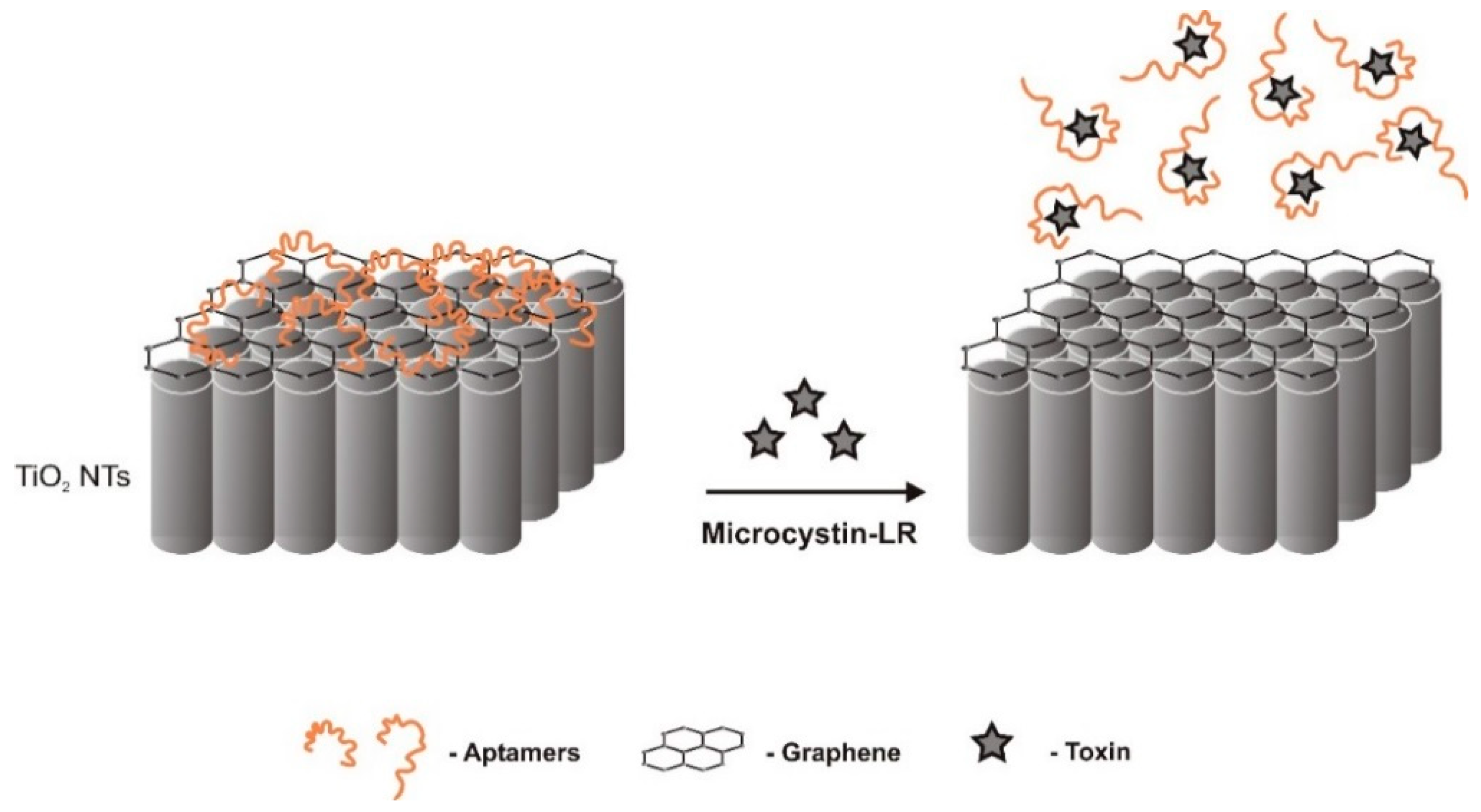
| Electrochemical (label-free)/photoelectrochemical | TISS | titanium dioxide nanotubes photo-anode substrate functionalized with graphene | Physisorption (noncovalent) π-π stacking interactions between the hexagonal cells of graphene and the DNA nucleobases. | 1.0–500 fM | 0.5 fM | 0.3% | Good selectivity. monosultap, atrazine, glyphosate, omethoate, gramoxone, dipterex, and acetamiprid | - | [52] | |
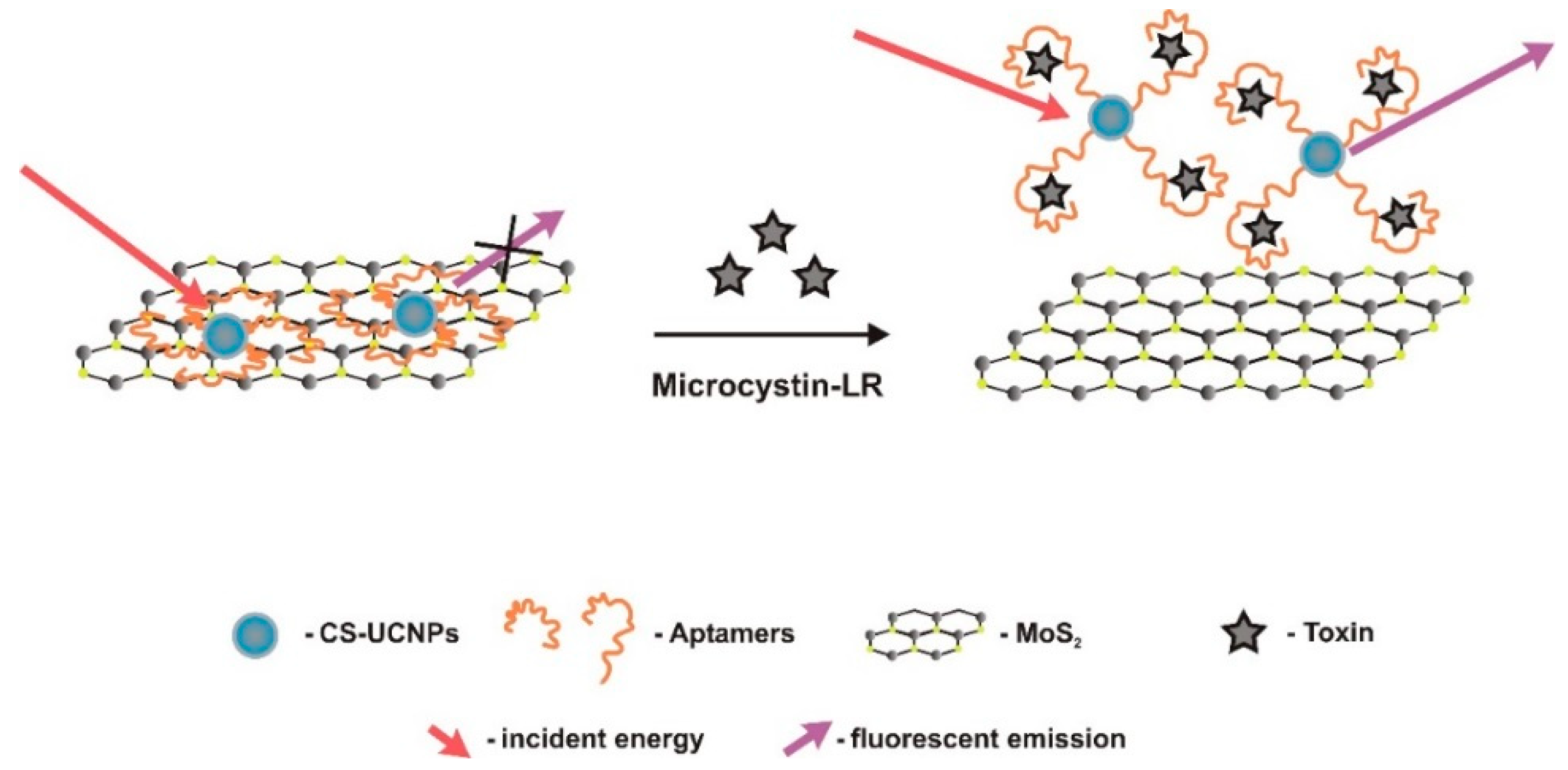
| Optic (label-based)/fluorescent | TISS | lanthanide ions doped core/shell UCNPs (label) grafted on 2D MoS2 nanosheets (platform/quencher) | Affinity binding (biotin-streptavidin) to UCNPs. Van der Waals forces between nucleobases of UCNPs-aptamer and basal plane of MoS2 platform. | 0.01–50 μg/L | 0.002 μg/L | good | Good selectivity. MC-RR, MC-YR, MC-LA, OA | tap water and Tai lake water samples 94–112% (5.3–9.4%) | [53] | |
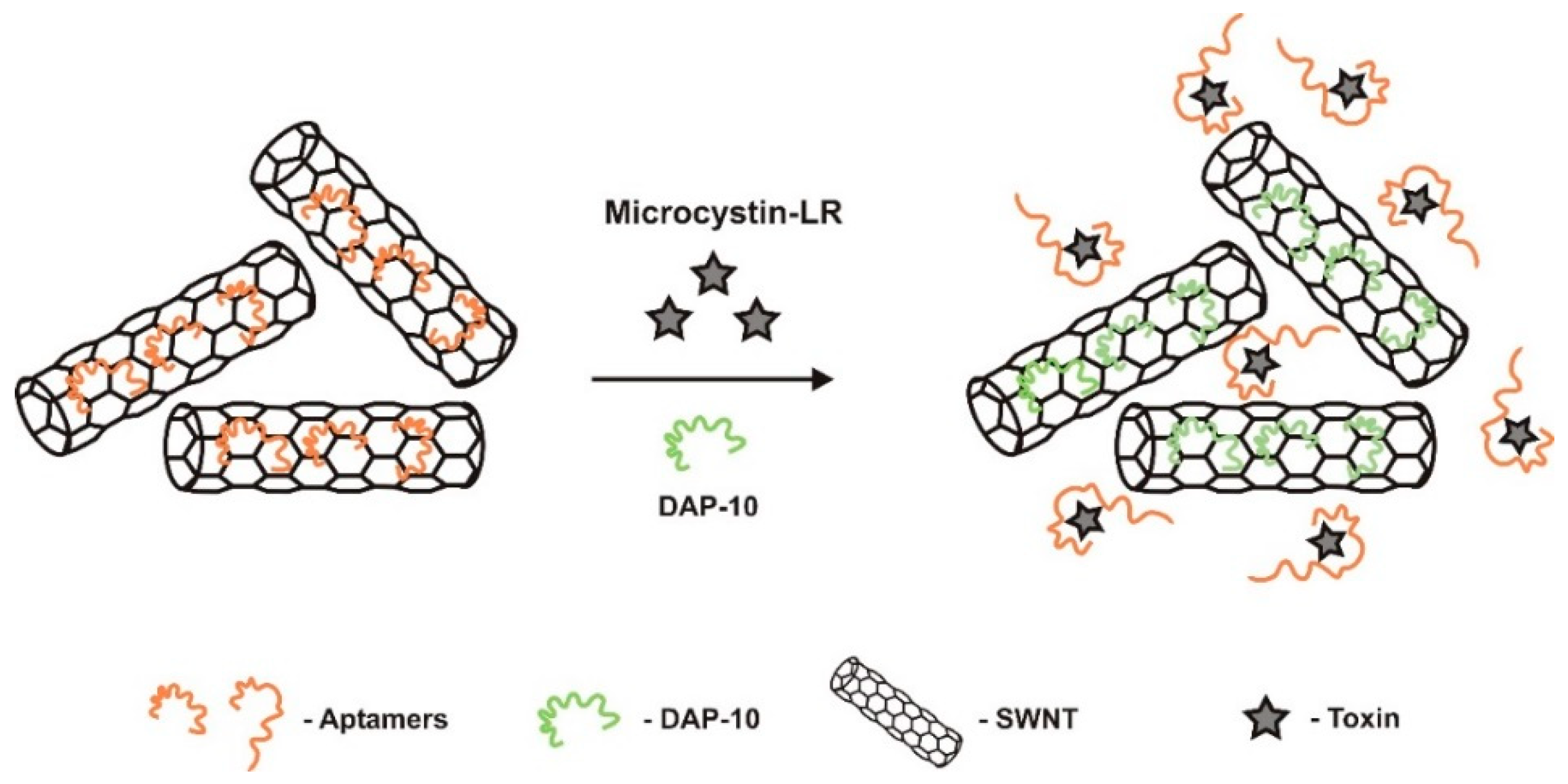
| Optic (label-based)/fluorescent | TISS | single-walled carbon nanotubes (SWNT-immobilizer) dapoxyl (fluorescent dye) | Aptamers interact with SWNTs through π-π stacking interactions | 0.4–1200 nM | 0.138 nM (0.137 μg/L) | - | Good selectivity. MC-LA, ochratoxin A, acetamiprid | water and serum samples 88.46–103.7% (2.6–8.4%) | [35] | |
| MC-LR and MC-LA | Electrochemical/voltammetry | TISS | gold electrode/[Ru-(NH3)6]3+ | Covalent self-assembly via thiol chemistry | 0.01–10 nM | 0.0075 (MC-LR) and 0.0106 nM (-LA) | - | Good selectivity (MC-YR) | - | [28] |
| MC-LR, MC-LA, MC-YR | Electrochemical/voltammetry | TISS | gold electrode/[Ru-(NH3)6]3+ | Covalent self-assembly via thiol chemistry | 0.01–10 nM | 0.0077 (MC-LR); 0.0128 (-LA) and 0.0089 nM (-YR) | - | - | - | [28] |
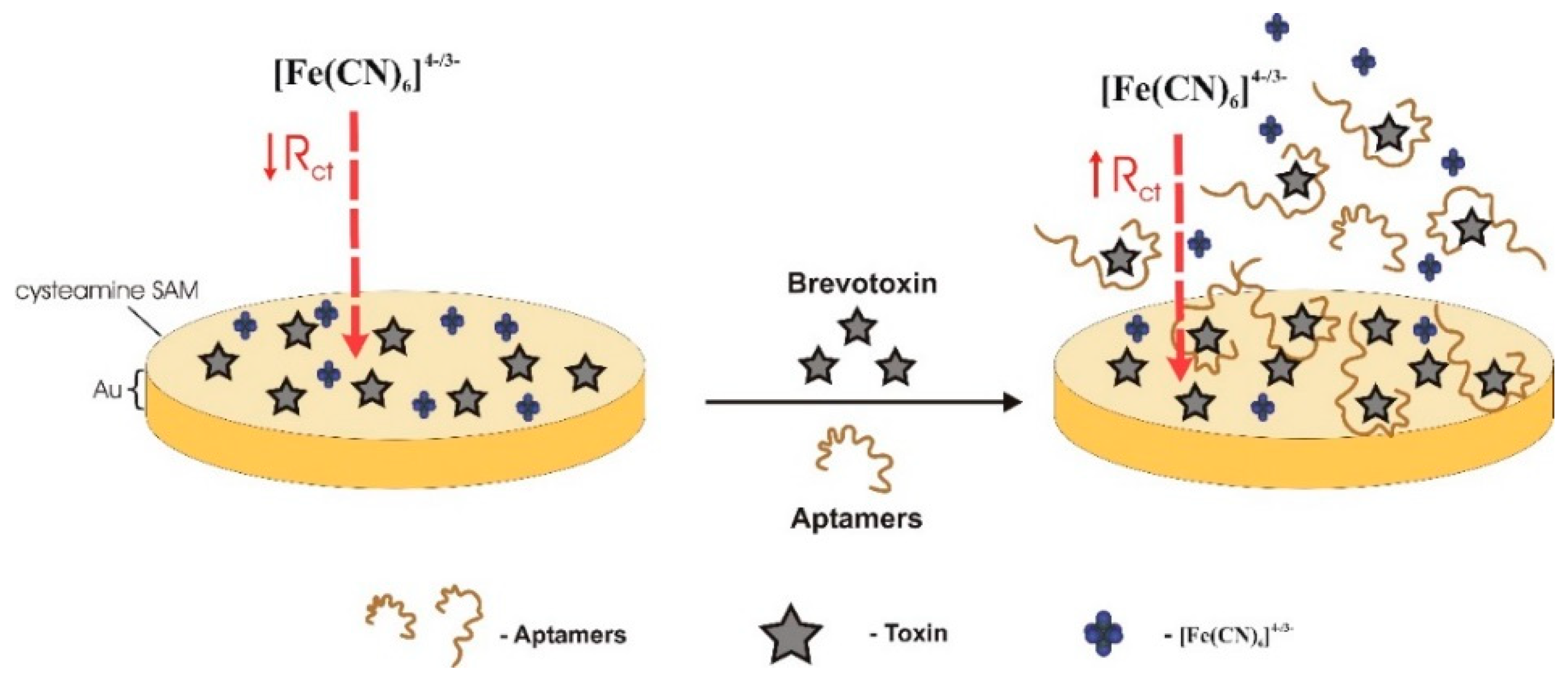
| BTX-2 | Electrochemical (label free)/impedimetric | CR | gold electrode functionalized with a cysteamine self-assembled monolayer (SAM); BTX-2 covalently attached through PDIC/[Fe(CN)6]4−/3− | - | 0.01–2000 μg/L | 0.106 μg/L | - | cross-selectivity to BTX-3 good selectivity to MC-LR and OA | spiked shellfish extracts 102–110% (3.4–6.0%) | [37] |
| OA | Electrochemical (label-free)/impedimetric | TISS | polycrystalline gold electrode/[Fe(CN)6]4−/3− | SAM covalently bound through AU-S alkyldisulfide interaction | 0.1 to 60 μg/L | <0.070 μg/L | 1.0–7.0% (N = 3) | Good selectivity. MC-LR, DTX-1, DTX-2 | uncontaminated shellfish extracts 92% | [50] |
| Optic (label-based)/fluorescence dual FRET | TID | green and red lanthanide UCNPs | Affinity binding (biotin-streptavidin) to UCNPs | 0.1–50 μg/L | 0.05 μg/L | 6.24% (N = 7) | Good selectivity. DTX-1, DTX-2 | water, fish, and shrimps 97.68–120.1% | [59] | |
| Optic (label-free)/colorimetric | TID/CR | biotinylated complimentary sequence to OA aptamer conjugated to avidin-catalase/gold trichloric acid and H2O2 | Affinity binding (biotin-streptavidin) to well-plates | 0.025–10 μg/L | 0.01 μg/L | 2.3–4.5% | Good selectivity. DTX-1, DTX-2, DA and SXT | spiked clam samples 92.86–103.34% (2.28–4.54%) | [60] | |
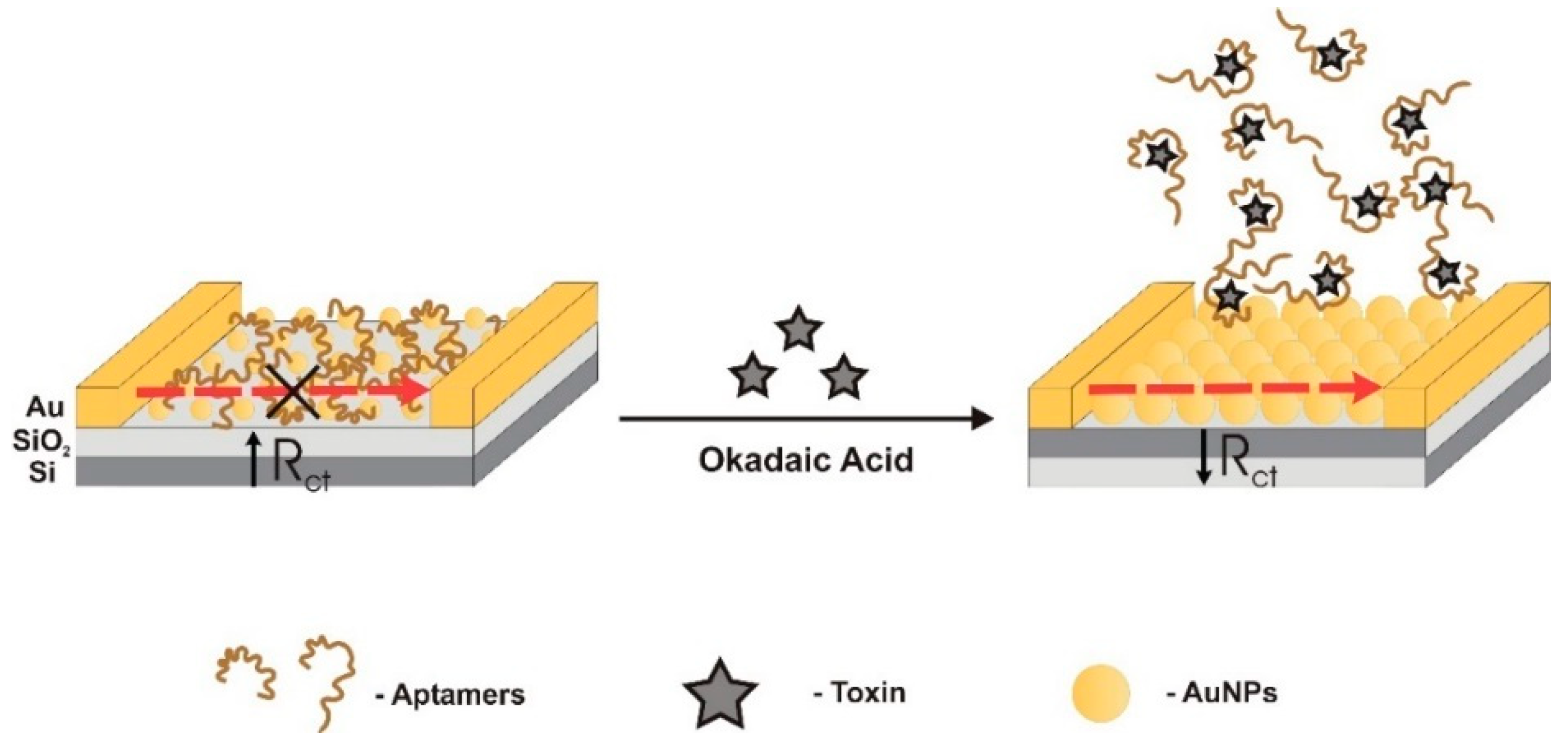
| Electrochemical (label-free)/impedimetric | TISS | AuNPs as conductive bridges | Electrostatic interaction | 5–80 μg/L | 1 μg/L | - | - | - | [51] | |
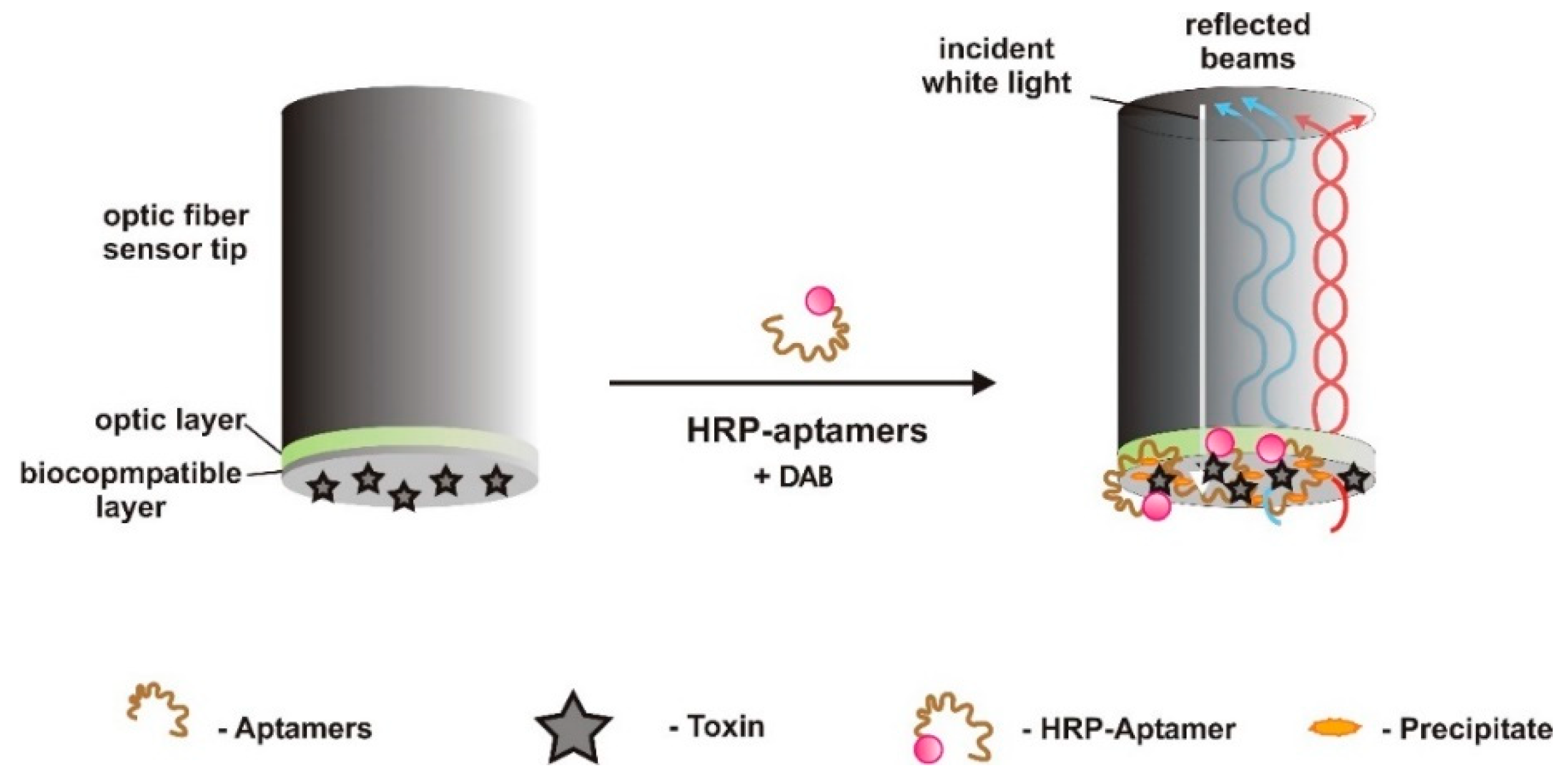
| PTX | Optic (label-free) /biolayer interferometry | CR | PTX immobilized on optic fiber surface (transducer) | - | 0.2–0.7 μg/L | 40 pg/L | - | Good selectivity. OA, STX, MC-LR, BTX-A/B | water, clams, scallops, mussels 100.27–108.24% (2.27–6.76%) | [62] |
| STX | Optic /fluorimetric | TISS | Evagreen (Biotium, Inc) dye for double-stranded DNA/real-time thermocycler | - | 15 μg/L–3 mg/L | 7.5 μg/L | - | Good selectivity. GTX 2/3 | rough shellfish extract | [54] |
| GTX1/4 | Optic (label-free) /biolayer interferometry | TISS | GTX immobilized on optic fiber surface (transducer) | EDC/NHS to which NH2-aptamers covalently and affinity binding (biotin-streptavidin) | 0.2–90 μg/L | 0.05 μg/L | 1.2% | GTX 2/3, neoSTX, STX | spiked shellfish samples 86.70–101.29% | [55] |
| TTX | Electrochemical/impedimetric | TISS | GCE modified with a poly(4-styrenesolfonic acid)-doped polyaniline film, where a TTX-binding aptamer was attached | grafted through a covalent glutaraldehyde cross-link | 0.23–1.07 μg/L | 0.199 μg/L | - | - | - | [56] |
| Optic/fluorimetric | TISS | Fe3O4 nanoparticles modified with the aptamer self-assembled on carbon dots through π-π stacking | carbodiimide-activated covalently bound to Fe3O4 nanoparticles | 0.1–100 μg/L | 0.06 μg/L | - | Aflatoxin B1-B2, botulism neurotoxins A-B, Staphylococcus aureus enterotoxins A-B | human gastric juice, serum and urine 96.00–104.20% (1.44–4.21%) | [57] |
| Aptamer Sequence | Modification 1 (5′-End) | Modification 2 (3′-End) | Affinity (Kd) | References |
|---|---|---|---|---|
| Anatoxin-a (ATX-A) | ||||
| 5′-undisclosed-3′ 1 | None | QSY21−3′ | - | [61] |
| 5′-TGG CGA CAA GAA GAC GTA CAA ACA CGC ACC AGG CCG GAG TGG AGT ATT CTG AGG TCG G-3′ | 5′−HO−(CH2)6−S−S−(CH2)6−disulfide-modified | None | 27.14 ± 5.38 nM | [31] |
| Cylindrospermopsin (CYN) | ||||
| 5′-GGC ATC AGG CAA CAA CCG ATG GTC CGG CCA CCC TAA CAA CCA GCC CAC CCA CCA CCC CGC CG-3′ | 5′−HO−(CH2)6−S−S−(CH2)6−disulfide-modified | None | 20.25 ± 2.7 nM | [30] |
| 5′-ATC AGG CAA CAA CCG ATG GTC CGG CCA CCC TAA CAA CCA GCC CAC CCA CCA CCC CGC CG-3′ | 5′−H2N-C6−Amine modified | None | - | [33] |
| Microcystin-LR (MC-LR) | ||||
| 5′-undisclosed-3′ | 5′−biotin− | - | 103 M−1 | [48] |
| 5′-GGC GCC AAA CAG GAC CAC CAT GAC AAT TAC CCA TAC CAC CTC ATT ATG CCC CAT CTC CGC-3′ | - | - | 50 ± 12 nM | [32,34,35,36,49,52,58,59] |
| 5′−biotin− | NaYF4: Yb, Er/Mn UCNPs | 50 ± 12 nM | [59] | |
| NaYF4: Yb, Tm@NaYF4: Yb core/shell UCNPs-avidin−5′−biotin− | - | 50 ± 12 nM | [53] | |
| 5′−SH(CH2)6− | - | 50 ± 12 nM | [28,38] | |
| 5′-CCG GGG UAG GGA UGG GAG GUA UGG AGG GGU CCU UGU UUC C-3′ (RNA) | 5′−H2N-C6− | - | - | [23,65] |
| 5′-TTT TTG GGT CCC GGG GTA GGG ATG GGA GGT ATG GAG GGG TCC TTG TTT CCC TCT TG-3′ | 5′−SH(CH2)6− | - | - | [29] |
| MC-LR and -LA | ||||
| 5′-CAC GCA CAG AAG ACA CCT ACA GGG CCA GAT CAC AAT CGG TTA GTG AAC TCG TAC GGC GCG-3′ | 5′−SH(CH2)6− | - | 76 ± 13 nM (-LR) 106 ± 17 nM (-LA) | [28] |
| MC-LR, LA and YR | ||||
| 5′-GGA CAA CAT AGG AAA AAG GCT CTG CTA CCG GAT CCC TGT TGT ATG GGC ATA TCT GTT GAT-3′ | 5′-SH(CH2)6− | - | 705 ± 248 nM (-LR) 808 ± 123 nM (-LA) 193 ± 28 nM (-YR) | [28] |
| Brevetoxin (BTX) | ||||
| 5′-GGC CAC CAA ACC ACA CCG TCG CAA CCG CGA GAA CCG AAG TAG TGA TCA TGT CCC TGC GTG-3′ | - | - | 42 nM | [37] |
| 5′-undisclosed-3′ | - | - | - | [66] |
| Okadaic Acid (OA) | ||||
| 5′-GGT CAC CAA CAA CAG GGA GCG CTA CGC GAA GGG TCA ATG TGA CGT CAT GCG GAT GTG TGG-3′ | 5′-HO−(CH2)6−S−S−(CH2)6− | - | 77 nM | [50,51] |
| 5′−biotin− | NaYF4: Yb, Ho UCNPs | 50 ± 12 nM | [59] | |
| 5′-ATT TGA CCA TGT CGA GGG AGA CGC GCA GTC GCT ACC ACC T-3′ | 5′−biotin− | - | 40 ± 13 nM | [60] |
| Palytoxin (PTX) | ||||
| 5′-GGA GGT GGT GGG GAC TTT GCT TGT ACT GGG CGC CCG GTT GAA-3′ | 5′−biotin− | 5′-HRP 2 | 0.843 nM | [62] |
| Saxitoxin (STX) | ||||
| 5′-CCG TGG AAA CAT GTT CAT TGG GCG CAC TCC GCT TTC TGT A-3′ | - | - | ~3.84 M | [54,67] |
| 5′-TTG AGG GTC GCA TCC CGT GGA AAC AGG TTC ATT G-3′ | - | - | 133 nM | [68] |
| Gonyautoxin (GTX) | ||||
| 5′-AAC CTT TGG TCG GGC AAG GTA GGT T-3′ | 5′-Biotin 5′−H2N-C6− | - | 21.9 nM | [55] |
| Tetrodotoxin (TTX) | ||||
| 5′-AAAAATTTCACACGGGTGCCTCGGCTGTCC-3′ | 5′-NH2- | - | - | [56,57,69] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cunha, I.; Biltes, R.; Sales, M.; Vasconcelos, V. Aptamer-Based Biosensors to Detect Aquatic Phycotoxins and Cyanotoxins. Sensors 2018, 18, 2367. https://doi.org/10.3390/s18072367
Cunha I, Biltes R, Sales M, Vasconcelos V. Aptamer-Based Biosensors to Detect Aquatic Phycotoxins and Cyanotoxins. Sensors. 2018; 18(7):2367. https://doi.org/10.3390/s18072367
Chicago/Turabian StyleCunha, Isabel, Rita Biltes, MGF Sales, and Vitor Vasconcelos. 2018. "Aptamer-Based Biosensors to Detect Aquatic Phycotoxins and Cyanotoxins" Sensors 18, no. 7: 2367. https://doi.org/10.3390/s18072367
APA StyleCunha, I., Biltes, R., Sales, M., & Vasconcelos, V. (2018). Aptamer-Based Biosensors to Detect Aquatic Phycotoxins and Cyanotoxins. Sensors, 18(7), 2367. https://doi.org/10.3390/s18072367








