Using Bi-Seasonal WorldView-2 Multi-Spectral Data and Supervised Random Forest Classification to Map Coastal Plant Communities in Everglades National Park
Abstract
1. Introduction
2. Materials and Methods
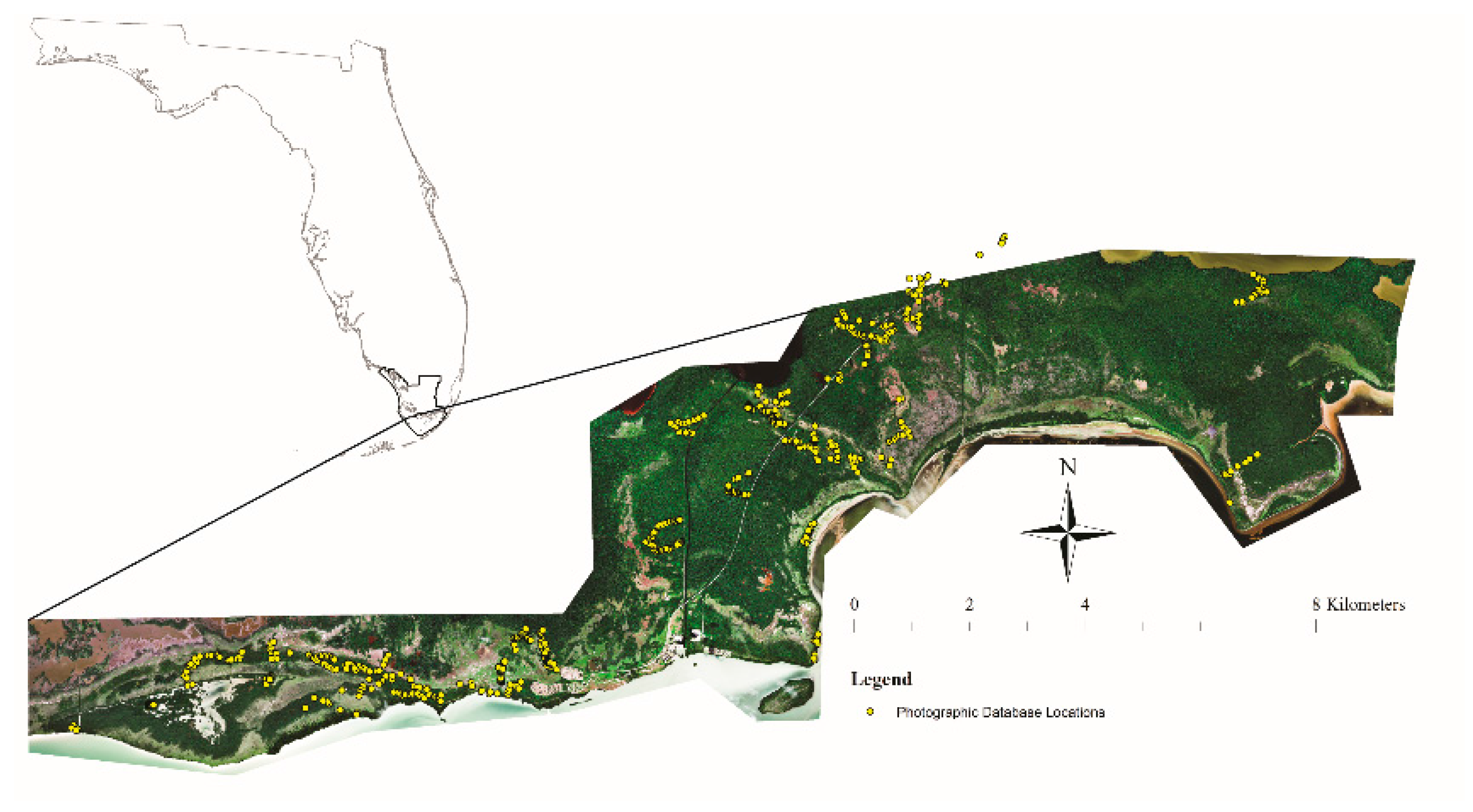
2.1. Study Area
2.2. Community Type Descriptions
- (1)
- Black Mangrove Forest: This forest is dominated by black mangrove (Avicennia germinans (L.)) with few associated woody species [35], except for occasional white mangrove (Laguncularia racemosa (L.) C.F.Gaertn) or red mangrove (Rhizophora mangle L.) found in either the canopy or understory. At times, areas of young black mangrove forest have halophyte species in the understory. Black mangrove forests are considered the most salt-tolerant of the three mangroves found in south Florida [36].
- (2)
- Red Mangrove Forest: This forest is dominated by Rhizophora mangle in the canopy and has little to no understory vegetation [35]. Occasional A. germinans are found scattered throughout; L. racemosa is found even less commonly. Red mangrove forests are considered the most inundation-tolerant of the three mangrove types and are less salt-tolerant than black mangroves [36].
- (3)
- White Mangrove Forest: This forest is dominated by Laguncularia racemosa in the canopy; often halophytes such as Batis maritima L., Sarcocornia perennis (Mill.) A.J. Scott, and Suaeda linearis (Elliott) Moq. are found in the understory. This community is most often found in irregularly flooded areas [35] and is the least salt- and inundation-tolerant of the three mangrove species found in south Florida [36].
- (4)
- Buttonwood/Glycophyte Forest: Buttonwood (Conocarpus erectus L.) is the dominant canopy species of buttonwood forests, but other woody species and a diverse herbaceous understory are also found in this community [37]. Temperature, salinity, tidal fluctuation, substrate, and wave energy influence the size and extent of buttonwood forests [38], which often grade into salt marsh, coastal berm, rockland hammock, coastal hardwood hammock, and coastal rock barren [38]. This community sustains freshwater flooding during the wet season and is dry during the dry season [38]. Buttonwood forests (mean elevation 29 ± 3 cm) maintain an average groundwater table of −33 ± 1 cm and (26–29.5) ± 0.4‰ groundwater salinity [7,37].
- (5)
- Buttonwood/Halophyte Forest: C. erectus is the only canopy tree species in buttonwood/halophyte forests. The understory is comprised of halophytic species such as Batis maritima, Borrichia frutescens (L.) DC., Distichlis spicata (L.) Greene, Sarcocornia perennis, and Suaeda linearis [37]. Buttonwood/halophyte forest (also called buttonwood prairies [37]) (mean elevation 18 ± 3 cm) show a mean groundwater table at −32 ± 2 cm and average groundwater table salinity of 38.8 ± 0.6‰ [37].
- (6)
- Halophyte Prairie: These prairies are comprised of Batis maritima, Borrechia frutescens, D. spicata, Sarcocornia perennis, and Sueda linearis, as well as other less common species; there is no canopy species [37]. Halophyte prairies have marl soils and slightly higher elevation than adjacent black and white mangrove forests [33]. In halophyte prairies, standing water that is brackish to freshwater is present for months during the wet season. Evaporation and lack of drainage cause these communities to become hypersaline during the dry season [32].
- (7)
- Coastal Tropical Hardwood Hammocks: Coastal tropical hardwood hammocks are biodiverse. This community includes a variety of tropical tree and shrub species [7,35] and a different suite of herbaceous species than are found in the mangrove communities [7]. Ground cover is often limited in closed canopy areas and abundant in areas where canopy disturbance has occurred or where this community intergrades with buttonwood forest [7]. Coastal tropical hardwood hammocks are the least salt-tolerant of all the coastal community types and reside at the highest elevation (mean elevation 29 ± 3 cm) [37].
2.3. Classifier Evaluation and Community Prediction for the Study Area
3. Results
3.1. Bi-Seasonal Versus Single Season Signature Assessment
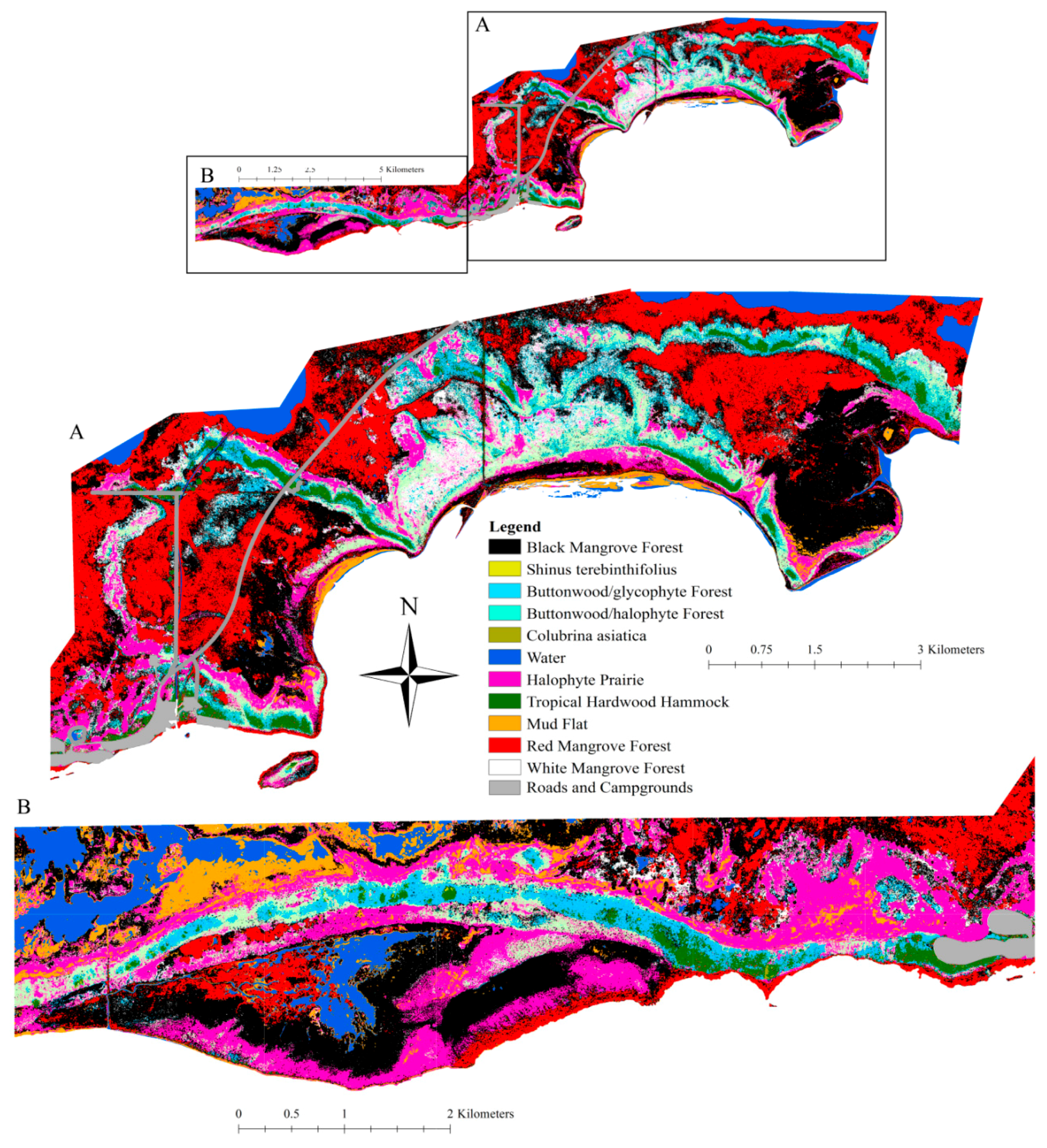
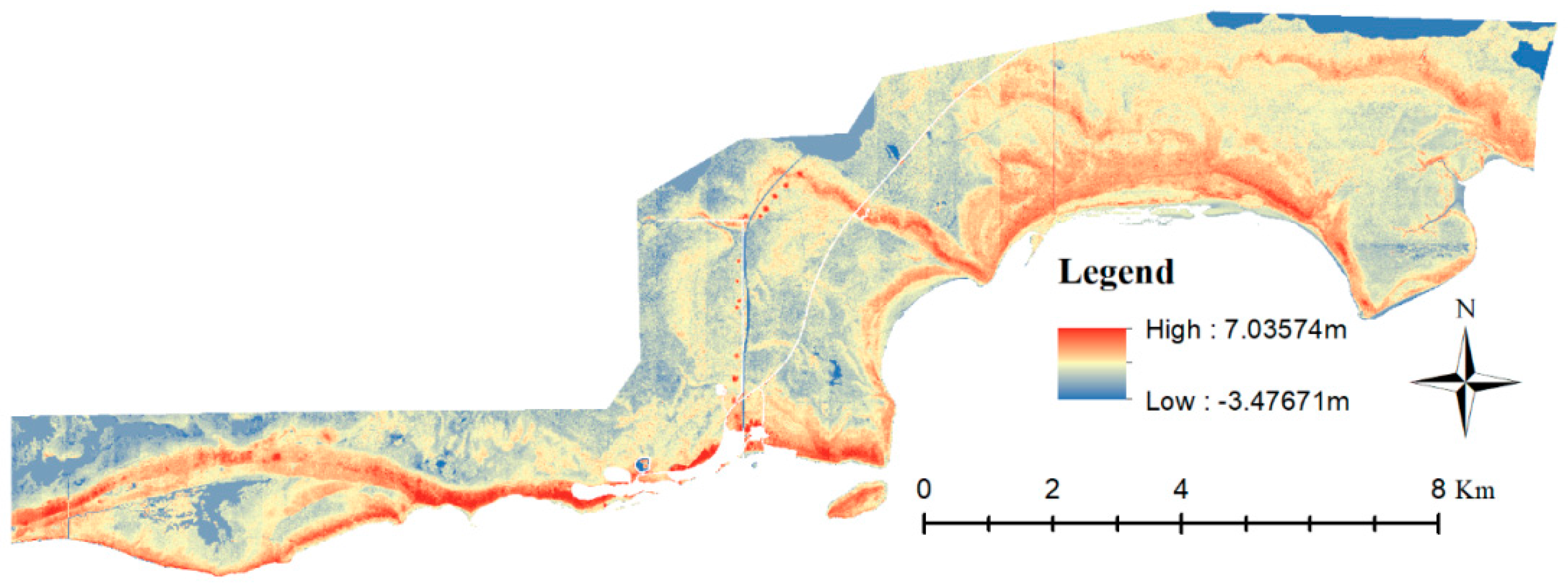
3.2. Vegetation Map
3.3. Community Area and Percent Cover
4. Discussion and Conclusions
Acknowledgments
Author Contributions
Conflicts of Interests
References
- Nicholls, R.J.; Cazenave, A. Sea-level rise and its impact on coastal zones. Science 2010, 328, 1517–1520. [Google Scholar] [CrossRef] [PubMed]
- Terry, J.P.; Chui, T.F.M. Evaluating the fate of freshwater lenses on atoll islands after eustatic sea-level rise and cyclone-driven inundation: A modelling approach. Glob. Planet. Chang. 2012, 88–89, 76–84. [Google Scholar] [CrossRef]
- Kirwan, M.L.; Megonigal, J.P. Tidal wetland stability in the face of human impacts and sea-level rise. Nature 2013, 504, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Li, J.; Sheng, C.; Xu, J.; Wu, L. A review of wetland remote sensing. Sensors 2017, 17, 777. [Google Scholar] [CrossRef] [PubMed]
- Giri, C.; Long, J. Is the geographic range of mangrove forests in the conterminous United States really expanding? Sensors 2016, 16, 2010. [Google Scholar] [CrossRef] [PubMed]
- Kuenzer, C.; Bluemel, A.; Gebhardt, S.; Quoc, T.V.; Dech, S. Remote Sensing of Mangrove Ecosystems: A Review. Remote Sens. 2011, 3, 878–928. [Google Scholar] [CrossRef]
- Saha, A.K.; Saha, S.; Sadle, J.; Jiang, J.; Ross, M.S.; Price, R.M.; Sternberg, L.S.L.O.; Wendelberger, K.S. Sea level rise and South Florida coastal forests. Clim. Chang. 2011, 107, 81–108. [Google Scholar] [CrossRef]
- Zhang, K. Analysis of non-linear inundation from sea-level rise using LIDAR data: A case study for South Florida. Clim. Chang. 2011, 106, 537–565. [Google Scholar] [CrossRef]
- Wang, L.; Silván-Cárdenas, J.L.; Sousa, W.P. Neural network classification of mangrove species from multi-seasonal Ikonos imagery. Photogramm. Eng. Remote Sens. 2008, 74, 921–927. [Google Scholar] [CrossRef]
- Lee, R.Y.; Ou, D.Y.; Shiu, Y.S.; Lei, T.C. Comparisons of using Random Forest and Maximum Likelihood Classifiers with Worldview-2 imagery for classifying Crop Types. In Proceedings of the 36th Asian Conference Remote Sensing Foster, ACRS 2015, Resilient Growth Asia, Quezon City, Philippines, 19–23 October 2015. [Google Scholar]
- Noonan, M.; Chafer, C. A method for mapping the distribution of willow at a catchment scale using bi-seasonal SPOT5 imagery. Weed Res. 2007, 47, 173–181. [Google Scholar] [CrossRef]
- Dymond, C.C.; Mladenoff, D.J.; Radeloff, V.C. Phenological differences in Tasseled Cap indices improve deciduous forest classification. Remote Sens. Environ. 2002, 80, 460–472. [Google Scholar] [CrossRef]
- Gilmore, M.S.; Wilson, E.H.; Barrett, N.; Civco, D.L.; Prisloe, S.; Hurd, J.D.; Chadwick, C. Integrating multi-temporal spectral and structural information to map wetland vegetation in a lower Connecticut River tidal marsh. Remote Sens. Environ. 2008, 112, 4048–4060. [Google Scholar] [CrossRef]
- Baker, C.; Lawrence, R.; Montagne, C.; Patten, D. Mapping wetlands and riparian areas using Landsat ETM+ imagery and decision-tree-based models. Wetlands 2006, 26, 465–474. [Google Scholar] [CrossRef]
- McCarthy, J.M.; Gumbricht, T.; McCarthy, T.; Frost, P.; Wessels, K.; Seidel, F. Flooding Patterns of the Okavango Wetland in Botswana between 1972 and 2000. Ambio 2003, 32, 453–457. [Google Scholar] [CrossRef] [PubMed]
- Rapinel, S.; Hubert-Moya, L.; Clément, B. Combined use of lidar data and multispectral earth observation imagery for wetland habitat mapping. Int. J. Appl. Earth Obs. Geoinf. 2015, 37, 56–64. [Google Scholar] [CrossRef]
- Richards, J.; Gann, D. Vegetation Trends in Indicator Regions of Everglades National Park; Florida International University: Miami, FL, USA, 2015. [Google Scholar]
- Gann, D.; Richards, J.H.; Biswas, H. Consulting Services to Determine the Effectiveness of Vegetation Classification Using World View 2 Satellite Data for the Greater Everglades. GIS Center 2012, 22, 1–62. [Google Scholar]
- Alexander, T.R.; Crook, A.G. Recent vegetational changes in South Florida. In Environments of South Florida: Present and Past; Gleason, P.J., Ed.; Miami Geological Society: Miami, FL, USA, 1974; pp. 61–72. [Google Scholar]
- South Florida Information Access the South Florida Environment: A Region under Stress. Available online: http://sofia.usgs.gov/publications/circular/1134/esns/clim.html (accessed on 21 February 2015).
- Ross, M.S.; Sah, J.P.; Meeder, J.F.; Ruiz, P.L.; Telesnicki, G. Compositional effects of sea-level rise in a patchy landscape: The dynamics of tree islands in the southeastern coastal everglades. Wetlands 2014, 34, 91–100. [Google Scholar] [CrossRef]
- Munns, R.; Tester, M. Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. [Google Scholar] [CrossRef] [PubMed]
- Munns, R. Comparative physiology of salt and water stress. Plant Cell Environ. 2002, 25, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Sousa, W.P. Distinguishing mangrove species with laboratory measurements of hyperspectral leaf reflectance. Int. J. Remote Sens. 2009, 30, 1267–1281. [Google Scholar] [CrossRef]
- Castañeda-Moya, E.; Twilley, R.R.; Rivera-Monroy, V.H. Allocation of biomass and net primary productivity of mangrove forests along environmental gradients in the Florida Coastal Everglades, USA. For. Ecol. Manag. 2013, 307, 226–241. [Google Scholar] [CrossRef]
- Wendelberger, K.S. Evaluating Plant Community Response to Sea Level Rise and Anthropogenic Drying: Can Life Stage and Competitive Ability be Used as Indicators in Guiding Conservation Actions? Ph.D. Thesis, Biology Department, Florida International University, Miami, FL, USA, 2016. [Google Scholar]
- Holmes, C.W.; Willard, D.; Brewster-Wingard, L.; Wiemer, L.; Marot, M.E. Buttonwood embankment, Northeastern Florida Bay. Available online: http://sofia.usgs.gov/projects/geo_eco_history/btnrdgeabfb1999.html (accessed on 16 November 2015).
- Ross, M.S.; Meeder, J.F.; Sah, J.P.; Ruiz, P.L.; Telesnicki, G.J. The Southeast Saline Everglades revisited: 50 years of coastal vegetation change. J. Veg. Sci. 2000, 11, 101–112. [Google Scholar] [CrossRef]
- Holmes, C.W.; Willard, D.; Brewster-Wingard, L. The Geology of the Buttonwood Ridge and Its Historical Significance. Available online: http://sofia.usgs.gov/projects/geo_eco_history/geoecoab3.html (accessed on 16 November 2015).
- Holmes, C.W.; Marot, M.E. Buttonwood embankment: The historical perspective on its role in Northeastern Florida Bay Hydrology. In Proceedings of the 1999 Florida Bay and Adjacent Marine Systems Science Conference, Key Largo, FL, USA, 1–5 November 1999; pp. 166–168. [Google Scholar]
- Craighead, F.C., Jr. Trees of South Florida Vol 1; University of Miami Press: Coral Gables, FL, USA, 1964. [Google Scholar]
- Olmsted, I.C.; Loope, L.L. Vegetation of the Southern Coastal Region of Everglades National Park between Flamingo and Joe Bay; Report T-620; South Florida Research Center: Homestead, FL, USA, 1981. [Google Scholar]
- Olmsted, I.C.; Loope, L.L. Vegetation along a Microtopographic Gradient in the Estuarine Region of Everglades National Park, Florida, USA; South Florida Research Center: Homestead, FL, USA, 1981. [Google Scholar]
- Florida Climate Center Florida Climate Data. Available online: http://climatecenter.fsu.edu/climate-data-access-tools/climate-data-visualization (accessed on 12 February 2016).
- Rutchey, K.; Schall, T.N.; Doren, R.F.; Atkinson, A.; Ross, M.S.; Jones, D.T.; Madden, M.; Vilchek, L.; Bradley, K.A.; Snyder, J.R.; et al. Vegetation Classification for South Florida Natural Areas; Open-File Report 2006-124; St. Petersburg Coastal and Marine Science Center-USGS: St. Petersburg, FL, USA, 2006.
- Odum, W.E.; McIvor, C.C.; Smith, T.J., III. The Ecology of the Mangroves of South Floirda: A Community Profile; FWS/OBS-81/24; Defense Technical Information Center: Washington, DC, USA, 1982.
- Saha, S.; Sadle, J.; Van Der Heiden, C.; Sternberg, L. Salinity, groundwater, and water uptake depth of plants in coastal uplands of everglades National Park (Florida, USA). Ecohydrology 2015, 8, 128–136. [Google Scholar] [CrossRef]
- Florida Natural Areas Inventory Buttonwood Forest. Available online: http://www.fnai.org/natcom_accounts.cfm (accessed on 12 November 2015).
- Florida Division of Emergency Management. LiDAR Data Inventory. Available online: https://www.floridadisaster.org/dem/ITM/geographic-information-systems/lidar/ (accessed on 15 November 2015).
- Guyon, I.; Elisseeff, A. An Introduction to Variable and Feature Selection. J. Mach. Learn. Res. 2003, 3, 1157–1182. [Google Scholar] [CrossRef]
- Breiman, L.; Friedman, J.; Stone, C.J.; Olshen, R.A. Classification and Regression Trees; CRC Press: Boca Raton, FL, USA, 1984. [Google Scholar]
- Belgiu, M.; Drăguţ, L. Random forest in remote sensing: A review of applications and future directions. ISPRS J. Photogramm. Remote Sens. 2016, 114, 24–31. [Google Scholar] [CrossRef]
- Pal, M. Random forest classifier for remote sensing classification. Int. J. Remote Sens. 2005, 26, 217–222. [Google Scholar] [CrossRef]
- Kuhn, M. Carect: Classification and Regression Training. Available online: https://cran.r-project.org/web/packages/caret/index.html (accessed on 16 November 2015).
- Liaw, A.; Wiener, M. Classification and Regression by random Forest. R. News 2002, 2, 18–22. [Google Scholar] [CrossRef]
- Tortora, R. A note on sample size estimation for multinomial populations. Am. Stat. 1978, 32, 100–102. [Google Scholar]
- Congalton, R.; Green, K. Sample Design Considerations. Assess. Accuracy Remote Sens. Data 2009, 63–83. [Google Scholar] [CrossRef]
- Olofsson, P.; Foody, G.M.; Stehman, S.V.; Woodcock, C.E. Making better use of accuracy data in land change studies: Estimating accuracy and area and quantifying uncertainty using stratified estimation. Remote Sens. Environ. 2013, 129, 122–131. [Google Scholar] [CrossRef]
- Stehman, S.V. Estimating area from an accuracy assessment error matrix. Remote Sens. Environ. 2013, 132, 202–211. [Google Scholar] [CrossRef]
- R Core Team R. A Language and Environment for Statistical Computing. Available online: http://www.R-project.org/ (accessed on 16 November 2015).
- Hijmans, R.J. Raster: Geographic Data Analysis and Modeling. Available online: https://cran.r-project.org/web/packages/raster/index.html (accessed on 16 November 2015).
- Bivand, R.; Keitt, T.; Rowlingson, B. Rgdal: Bindings for the Geospatial Data Abstraction Library. Available online: https://r-forge.r-project.org/projects/rgdal/ (accessed on 16 November 2015).
- Ross, M.S.; O’Brien, J.J.; da Silveira Lobo Sternberg, L. Sea-level rise and the reduction in pine forests in the Florida Keys. Ecol. Appl. 1994, 4, 144–156. [Google Scholar] [CrossRef]
- Gaiser, E.E.; Zafiris, A.; Ruiz, P.L.; Tobias, F.A.C.; Ross, M.S. Tracking rates of ecotone migration due to salt-water encroachment using fossil mollusks in coastal South Florida. Hydrobiologia 2006, 569, 237–257. [Google Scholar] [CrossRef]
- Desantis, L.R.G.; Bhotika, S.; Williams, K.; Putz, F.E. Sea-level rise and drought interactions accelerate forest decline on the Gulf Coast of Florida, USA. Glob. Chang. Biol. 2007, 13, 2349–2360. [Google Scholar] [CrossRef]
- Williams, K.; Pinzon, Z.S.; Stumpf, R.P.; Raabe, E.A. Sea-Level Rise and Coastal Forests on the Gulf of Mexico. US Geol. Surv. 1999, 1500, 20910. [Google Scholar]
- Krauss, K.W.; From, A.S.; Doyle, T.W.; Doyle, T.J.; Barry, M.J. Sea-level rise and landscape change influence mangrove encroachment onto marsh in the Ten Thousand Islands region of Florida, USA. J. Coast. Conserv. 2011, 15, 629–638. [Google Scholar] [CrossRef]
- NOAA Mean Sea Level Trend Key West, Florida. Available online: https://tidesandcurrents.noaa.gov/sltrends/sltrends_station.shtml?stnid=8724580 (accessed on 20 April 2016).
- Pachauri, R.K.; Allen, M.R.; Barros, V.R.; Broome, J.; Cramer, W.; Christ, R.; Dubash, N.K. Climate Change 2014 Synthesis Report Fifth Assessment Report; IPCC: Geneva, Switzerland, 2014; ISBN 9789291691432. [Google Scholar]
- Davis, S.M.; Childers, D.L.; Lorenz, J.J.; Wanless, H.R.; Hopkins, T.E. A conceptual model of ecological interactions in the mangrove estuaries of the Florida Everglades. Wetlands 2005, 25, 832–842. [Google Scholar] [CrossRef]
- Alongi, D.M. Mangrove forests: Resilience, protection from tsunamis, and responses to global climate change. Estuar. Coast. Shelf Sci. 2008, 76, 1–13. [Google Scholar] [CrossRef]
- Langley, J.A.; McKee, K.L.; Cahoon, D.R.; Cherry, J.A.; Megonigal, J.P. Elevated CO2 stimulates marsh elevation gain, counterbalancing sea-level rise. Proc. Natl. Acad. Sci. USA 2009, 106, 6182–6186. [Google Scholar] [CrossRef] [PubMed]
- Barr, J.G.; Engel, V.; Fuentes, J.D.; Zieman, J.C.; O’Halloran, T.L.; Smith, T.J.; Anderson, G.H. Controls on mangrove forest-atmosphere carbon dioxide exchanges in western Everglades National Park. J. Geophys. Res. 2010, 115, 1–14. [Google Scholar] [CrossRef]
- McLeod, E.; Chmura, G.L.; Bouillon, S.; Salm, R.; Bjork, M.; Duarte, C.M.; Lovelock, C.E.; Schlesinger, W.H.; Silliman, B.R. A blueprint for blue carbon: Toward an improved understanding of the role of vegetated coastal habitats in sequestering CO2. Front. Ecol. Environ. 2011, 9, 552–560. [Google Scholar] [CrossRef]
- Hernández, C.M.A.; Zaragoza, C.G.; Iriarte-Vivar, S.; Flores-Verdugo, F.J.; Casasola, P.M. Forest structure, productivity and species phenology of mangroves in the La Mancha lagoon in the Atlantic coast of Mexico. Wetl. Ecol. Manag. 2011, 19, 273–293. [Google Scholar] [CrossRef]
- Utrera-López, M.E.; Moreno-Casasola, P. Mangrove litter dynamics in la Mancha Lagoon, Veracruz, Mexico. Wetl. Ecol. Manag. 2008, 16, 11–22. [Google Scholar] [CrossRef]
- Aké-Castillo, J.A.; Vázquez, G.; López-Portillo, J. Litterfall and decomposition of Rhizophora mangle L. in a coastal lagoon in the southern Gulf of Mexico. Hydrobiologia 2006, 559, 101–111. [Google Scholar] [CrossRef]
- Pastor-Guzman, J.; Dash, J.; Atkinson, P.M. Remote sensing of mangrove forest phenology and its environmental drivers. Remote Sens. Environ. 2018, 205, 71–84. [Google Scholar] [CrossRef]
- Colwell, R.K.; Brehm, G.; Cardelús, C.L.; Gilman, A.C.; Longino, J.T. Global warming, elevational range shifts, and lowland biotic attrition in the wet tropics. Science 2008, 322, 258–261. [Google Scholar] [CrossRef] [PubMed]
| December 2011 | April 2013 | Bi-Seasonal Total | |
|---|---|---|---|
| Reflectance Values | 8 (1/band) | 8 (1/band) | 16 |
| Mean | 8 (1/band) | 8 (1/band) | 16 |
| Range | 8 (1/band) | 8 (1/band) | 16 |
| Std Dev | 8 (1/band) | 8 (1/band) | 16 |
| NDVI | 1 | 1 | 2 |
| DTM | 1 | 1 | 1 |
| Total | 34 | 34 | 67 |
| Community | Wet Season | Dry Season | Bi-Seasonal | Wet-Dry | Bi-Wet | Bi-Dry | |
|---|---|---|---|---|---|---|---|
| Class Specific Error | Black mangrove | 16.5 | 9.3 | 8.4 | 7.2 | −8.1 | −0.9 |
| Buttonwood/glyco | 63.6 | 38.7 | 35.7 | 24.9 | −27.9 | −3.0 | |
| Buttonwood/halo | 55.5 | 37.9 | 34.1 | 17.6 | −21.4 | −3.8 | |
| Halophyte prairie | 6.8 | 6.7 | 4.6 | 0.1 | −2.2 | −2.1 | |
| Hardwood hamm | 10.8 | 7.3 | 6.1 | 3.5 | −4.7 | −1.2 | |
| Red mangrove | 18.8 | 16.2 | 13.8 | 2.6 | −5.0 | −2.4 | |
| White mangrove | 37.1 | 27.9 | 25.5 | 9.2 | −11.6 | −2.4 | |
| Mud flat | 12.4 | 22.6 | 10.7 | −10.2 | −1.7 | −11.9 | |
| Water | 0.0 | 0.1 | 0.1 | −0.1 | 0.1 | 0.0 | |
| Overall accuracy | 80.1 ± 1.1 | 84.2 ± 0.9 | 87.2 ± 1.0 | −4.1% | 7.1% | 3.0% |
| Community | Area (ha) | Proportional Area (ha) | Adj. Area (ha) | % Cover of Adj. Area | Adj. Area Std. Error | Adj. Area Lower 95% CI | Adj. Area Upper 95% CI | Proportional Area Bias | Proportion Adj. User’s Accuracy | Adj. User’s Accuracy Std. Error | Adj. User’s Accuracy Lower 95% CI | Adj. User’s Accuracy Upper 95% CI | Proportion Adj. Producer’s Accuracy | Adj. Producer’s Accuracy Std. Error | Adj. Producer’s Accuracy Lower 95% CI | Adj. Producer’s Accuracy Upper 95% CI |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Black mangrove | 2158.5 | 0.31 | 2014.5 | 28.5 | 63.3 | 1890.4 | 2138.6 | 0.85 | 0.85 | 0.05 | 0.75 | 0.95 | 0.91 | 0.03 | 0.85 | 0.97 |
| Invasive species | 16.4 | 0.00 | 227.4 | 3.2 | 41.6 | 145.8 | 309.0 | 0.89 | 0.91 | 0.04 | 0.84 | 0.99 | 0.07 | 0.04 | −0.02 | 0.15 |
| Buttonwood/glyco | 429.8 | 0.06 | 615.2 | 8.7 | 32.6 | 551.4 | 679.1 | 0.91 | 0.91 | 0.04 | 0.83 | 0.99 | 0.63 | 0.07 | 0.51 | 0.76 |
| Buttonwood/halo | 653.2 | 0.09 | 667.5 | 9.5 | 4.1 | 659.4 | 675.6 | 1.00 | 1.00 | 0.00 | 1.00 | 1.00 | 0.98 | 0.01 | 0.96 | 1.00 |
| Halophyte prairie | 899.5 | 0.13 | 703.5 | 9.9 | 31.8 | 641.2 | 765.8 | 0.74 | 0.74 | 0.06 | 0.62 | 0.86 | 0.94 | 0.04 | 0.86 | 1.02 |
| Hardwood hamm | 239.2 | 0.03 | 182.9 | 2.6 | 14.6 | 154.2 | 211.6 | 0.62 | 0.62 | 0.07 | 0.49 | 0.76 | 0.82 | 0.11 | 0.61 | 1.02 |
| Red mangrove | 1493.0 | 0.21 | 1470.6 | 20.8 | 46.3 | 1379.8 | 1561.4 | 0.89 | 0.89 | 0.05 | 0.80 | 0.97 | 0.90 | 0.04 | 0.82 | 0.98 |
| White mangrove | 511.6 | 0.07 | 532.6 | 7.5 | 27.4 | 478.9 | 586.3 | 0.91 | 0.91 | 0.04 | 0.83 | 0.99 | 0.87 | 0.08 | 0.71 | 1.03 |
| Mud flat | 253.7 | 0.04 | 240.9 | 3.4 | 7.9 | 225.4 | 256.4 | 0.89 | 0.89 | 0.05 | 0.80 | 0.97 | 0.94 | 0.04 | 0.85 | 1.02 |
| Water | 408.6 | 0.06 | 409.7 | 5.8 | 14.5 | 381.3 | 438.1 | 0.89 | 0.89 | 0.05 | 0.80 | 0.97 | 0.89 | 0.05 | 0.79 | 0.98 |
| Total | 7063.5 | 1.00 | 7064.8 | 100 | Adjusted accuracy | 86.02% | ||||||||||
| Black Mangrove | Buttonwood/gLyco | Buttonwood/hAlo | Water | Halophyte Prairie | Hardwood Hammock | Invasive Species | Mud Flat | Red Mangrove | White Mangrove | |
|---|---|---|---|---|---|---|---|---|---|---|
| Black mangrove | 0.85 | 0.08 | 0.02 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.04 | 0.02 |
| Buttonwood/glyco | 0.06 | 0.89 | 0.06 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 |
| Buttonwood/halo | 0.00 | 0.06 | 0.91 | 0.00 | 0.04 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 |
| Water | 0.00 | 0.00 | 0.00 | 1.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 |
| Halophyte prairie | 0.02 | 0.04 | 0.02 | 0.00 | 0.89 | 0.00 | 0.00 | 0.04 | 0.00 | 0.00 |
| Hardwood hammock | 0.06 | 0.00 | 0.11 | 0.00 | 0.02 | 0.74 | 0.04 | 0.00 | 0.04 | 0.00 |
| Invasive species | 0.02 | 0.00 | 0.15 | 0.00 | 0.02 | 0.06 | 0.62 | 0.00 | 0.13 | 0.00 |
| Mud flat | 0.02 | 0.00 | 0.00 | 0.06 | 0.04 | 0.00 | 0.00 | 0.89 | 0.00 | 0.00 |
| Red mangrove | 0.08 | 0.00 | 0.00 | 0.00 | 0.00 | 0.02 | 0.00 | 0.00 | 0.89 | 0.02 |
| White mangrove | 0.00 | 0.02 | 0.08 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.91 |
| Accuracy: 85.66% | 95% CI: (0.8238, 0.8853) | |||||||||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wendelberger, K.S.; Gann, D.; Richards, J.H. Using Bi-Seasonal WorldView-2 Multi-Spectral Data and Supervised Random Forest Classification to Map Coastal Plant Communities in Everglades National Park. Sensors 2018, 18, 829. https://doi.org/10.3390/s18030829
Wendelberger KS, Gann D, Richards JH. Using Bi-Seasonal WorldView-2 Multi-Spectral Data and Supervised Random Forest Classification to Map Coastal Plant Communities in Everglades National Park. Sensors. 2018; 18(3):829. https://doi.org/10.3390/s18030829
Chicago/Turabian StyleWendelberger, Kristie S., Daniel Gann, and Jennifer H. Richards. 2018. "Using Bi-Seasonal WorldView-2 Multi-Spectral Data and Supervised Random Forest Classification to Map Coastal Plant Communities in Everglades National Park" Sensors 18, no. 3: 829. https://doi.org/10.3390/s18030829
APA StyleWendelberger, K. S., Gann, D., & Richards, J. H. (2018). Using Bi-Seasonal WorldView-2 Multi-Spectral Data and Supervised Random Forest Classification to Map Coastal Plant Communities in Everglades National Park. Sensors, 18(3), 829. https://doi.org/10.3390/s18030829





