Integration of Distributed Services and Hybrid Models Based on Process Choreography to Predict and Detect Type 2 Diabetes
Abstract
1. Introduction
2. Materials and Methods
2.1. Business Context Definition
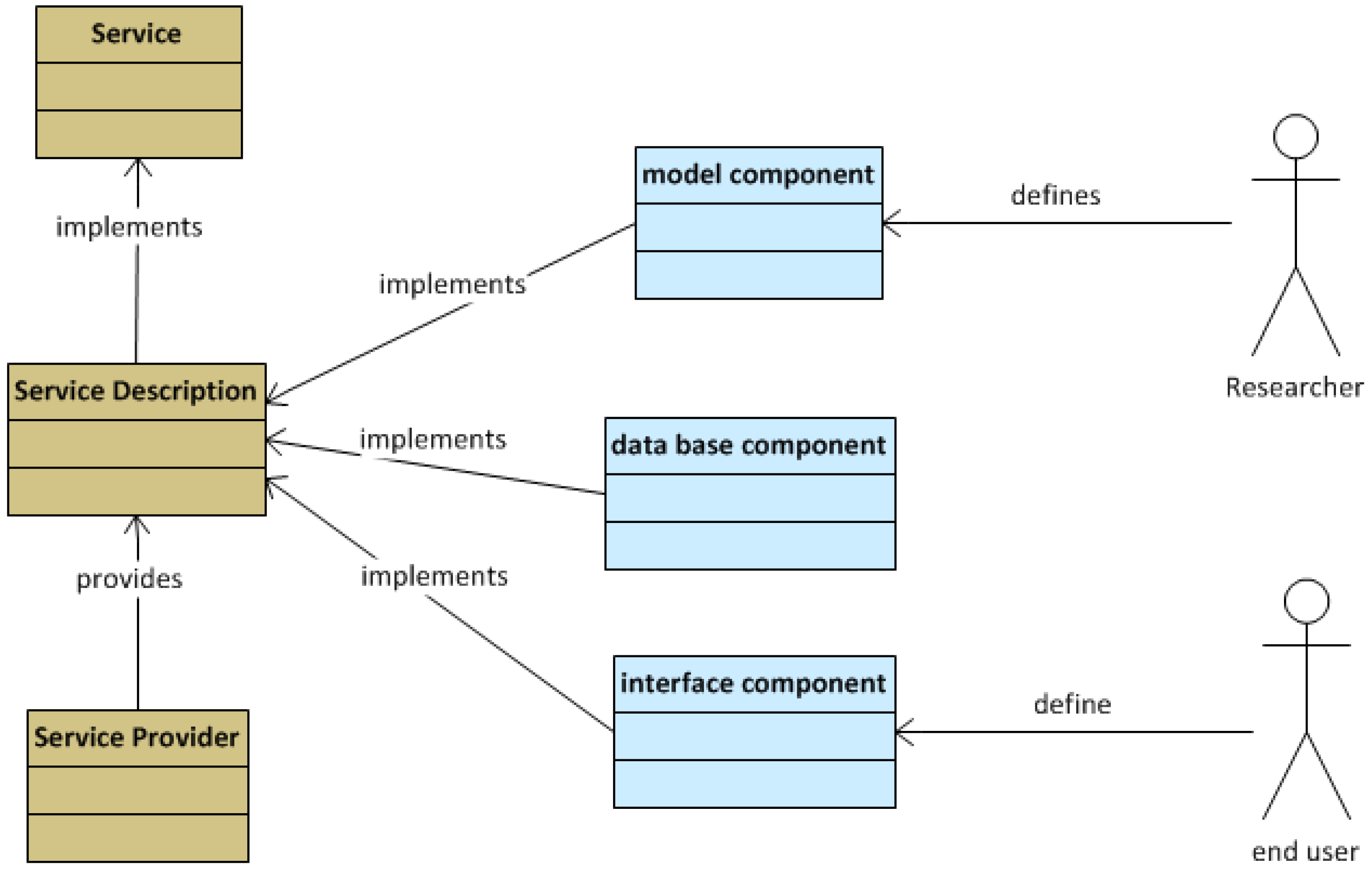
- End users: non-technical end users such as health care professionals, health care managers, patients and citizens; health professionals, including managers and policy makers and medical researchers who are concerned with public health affairs. Good development environments and friendly interfaces will lead to better quality software and will attract professionals to use the tools. Efficient communication among service providers would result in services that better meet end user requirements.
- Service providers are concerned about the commercial exploitation of the system. They need to maintain an effective communication with their end users and a fluent interaction with the runtime environment to explore potential integrations.
- Researchers are mainly concerned with good development environments, a knowledgeable community of developers and access to resources for implementing software and algorithms. The system should support researchers as a major stakeholder and allow them to participate in the system improvement (together with service providers and end users). Two main domains of research are found within this viewpoint: data mining research and software research. The first type is focused on the development of new algorithms and models to perform stratification and variable association analysis. The second type aims to improve the software quality of the services, interfaces and database management.
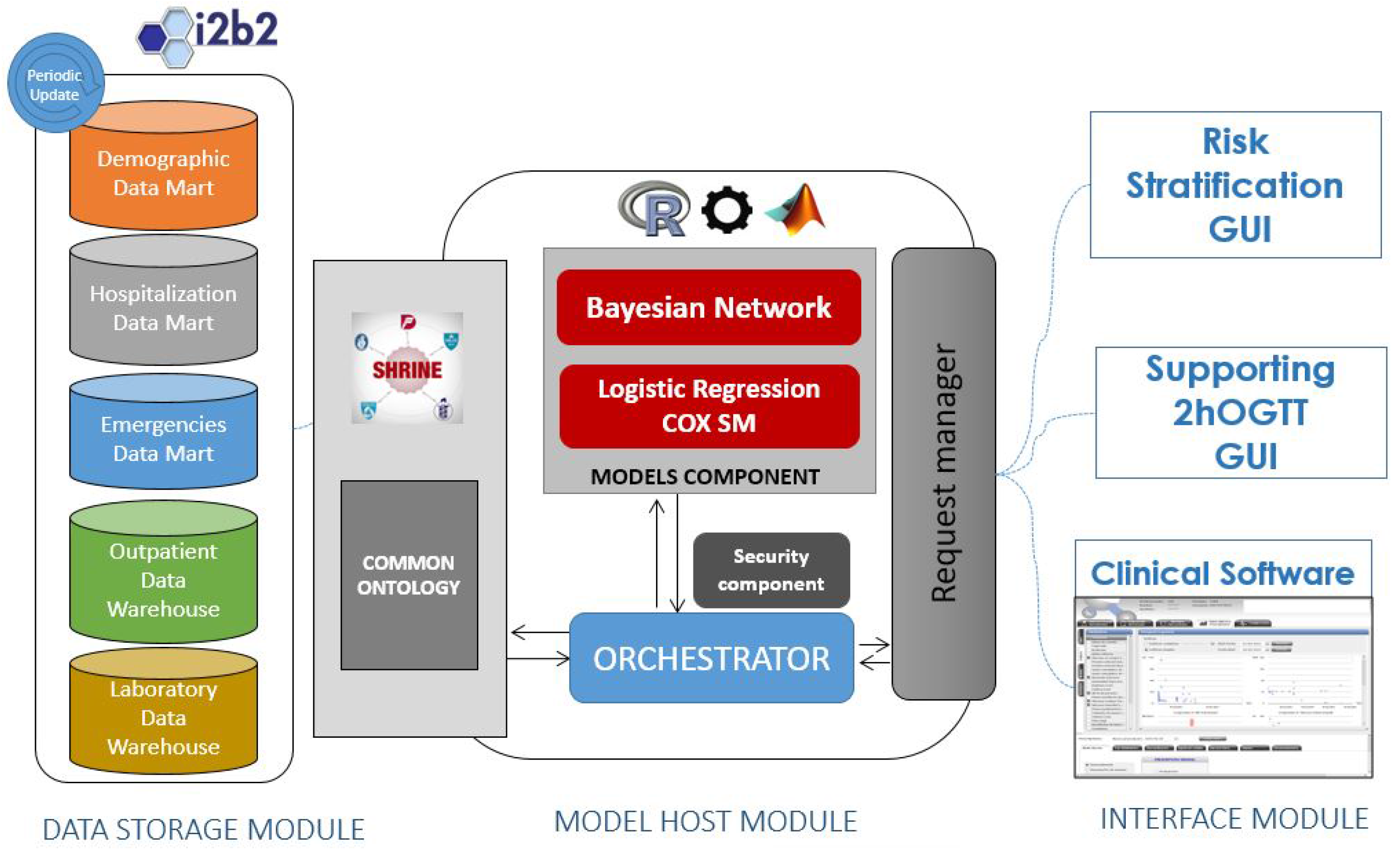
- The Data Storage module is in charge of providing a warehouse for all the data within the system. From a conceptual point of view, the data model is unique for all of the system, containing Electronic Health Records (EHRs) and other kinds of data (logistic and administrative).
- The Model Host module is the core of the system. It is in charge of managing the client requests (user interactions), running the risk scores and querying the data warehouses. It gathers into an application server the tools (models) that will run the algorithms over data from EHRs and provides the services for managing them from the client side. The Model Host module will also contain components to provide horizontal services including security features, tracking and system management.
- The Plug-in module is the part of the system that hosts the user interfaces. These user interfaces are web pages formatted for the intended use for each type of user and scenario. The integration with existing disease management systems is articulated wrapping the interfaces within plug-ins, tailored for each integration case.
2.2. Quality Metrics
- Category 1: a system for running algorithms on demand with a specific running environment regardless of patient health records or additional data than a set of defined parameters.
- Category 2: a system for running algorithms on demand with a specific running environment, which needs patient health records and additional data form a huge amount of variable parameters.
- Category 3: a system for running algorithms on demand at the client side with a specific running environment, which needs raw and pre-processed data.
2.3. Business Environment Definition
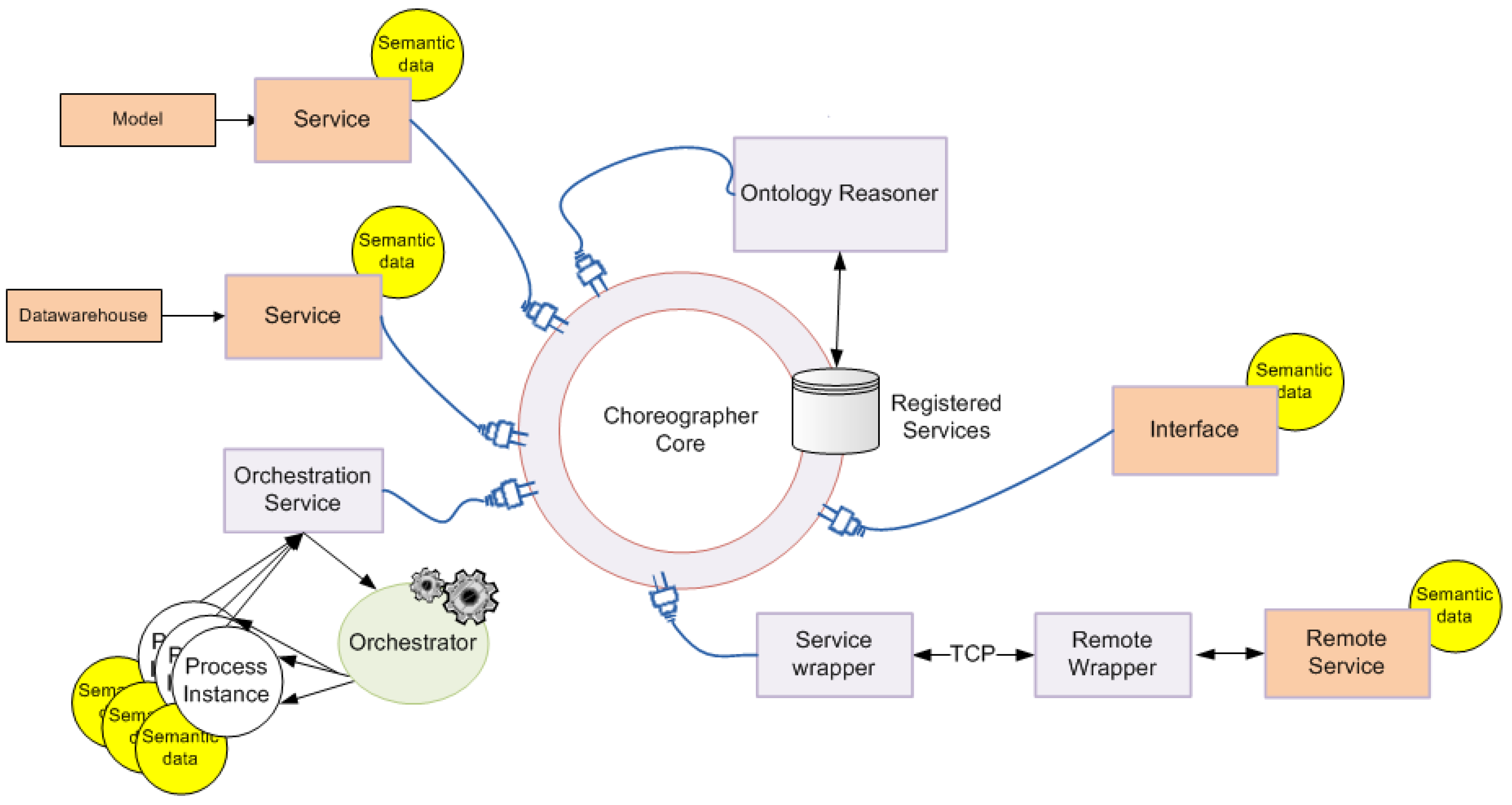
2.4. Service Collaboration Pattern
2.5. Data Warehouse Infrastructure
2.6. T2DM Risk Scores
- is the intercept or prior probability.
- is the intercept baseline hazard rate.
- is the regression coefficient, which denotes the relative weight of the corresponding predictor.
- are the predictors or variables, which can be numerical (continuous) or categorical (0, 1, 2…).
2.7. Design of the Pilot Study
- Training sessions: three group sessions for introducing participants to the tools and learning the actions to visualize data and execute the risk models.
- Evaluation of risk scores and clinical evaluation: evaluation of the tools during sessions of 2 h during eight weeks.
- Data analysis: acquisition of logs, traces and Key Performance Indicators (KPIs) for the technical assessment of the system.
- Computational load (memory footprint on the server).
- Response delay to service request (s).
- Access time to main DB/cache (ms).
- Time usage span (s).
- Maximum response delay.
3. System Architecture Description
3.1. Functional View
3.2. Model Host Component
3.3. Security Component
- Authentication: It must be possible for the service provider to ascertain the identity of the service requester.
- Authorization: The service provider must be able to determine whether the requester has the appropriate rights to invoke the service.
- Message confidentiality: Message contents must only be visible to the intended recipient.
- Message integrity: It must be possible to guarantee that a message has not been altered or tampered with in transport between the service consumer and the service provider.
3.4. Track Component
- Time stamp: dd/mm/yyyy hh:mm:ss.
- Module: the module (view or form) in which the patient is currently.
- Control: the control used: button, label, picture, graph, chart, etc.
- Free text: free text that indicates the interaction or notes for the usability expert.
3.5. Communication Protocol
4. Experimental Results
4.1. Scenarios for T2DM Risk Score Assessment
- Estimate missing variables given available variables measurable with a general practitioner’s visit and laboratory tests in the electronic health record towards risk stratification.
- Estimate the 2h-Oral Glucose Tolerance Test (2h-OGTT) glucose range given all other available variables (supporting a diabetologist to decide whether this test is needed).
4.1.1. Scenario 1: Risk Stratification
4.1.2. Scenario 2: Supporting 2h-OGTT Decision
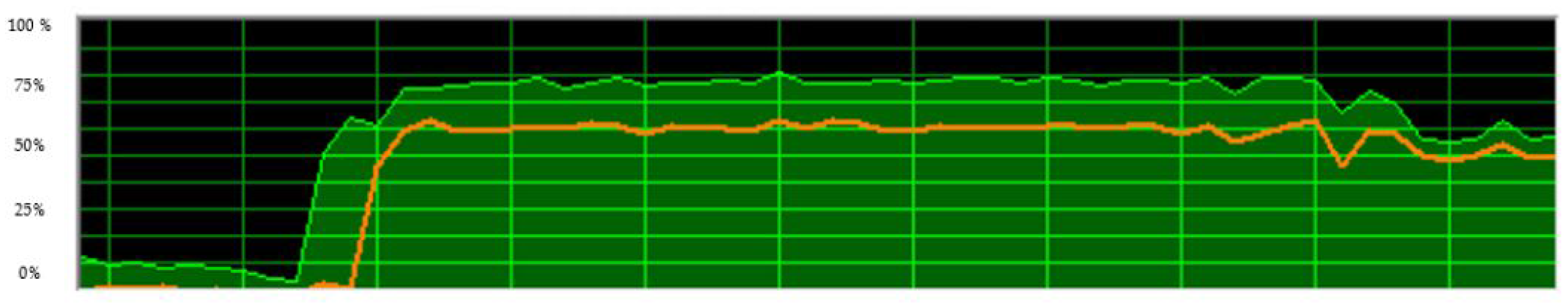
4.2. Technical Assessment
4.3. Map of Evaluations
- Data Warehouses (DW)
- Data Access Layer (Query Engine (QE)): multiple/single subject
- Missing Data Imputation (MDI)
- Risk Score Module (RSM)
- Orchestrator (O)
- Interface Module
- Verification of the model execution:
- -
- Appropriateness of the query
- -
- Units homogenization
- -
- Handling the resultsand storage
- Performance of the model execution:
- -
- Best case vs. worst case
- -
- Latency (time delay of the response)
- -
- Memory Load in the system server
- -
- Central Process Unit (CPU) load
- -
- Network resources
4.4. Verification of the Models’ Execution
Model Integration
5. Discussion
6. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| BN | Bayesian Network |
| EHR | Electronic Health Record |
| DW | Data Warehouse |
| HbA1C | Glycated Hemoglobin |
| HIS | Hospital Information System |
| KPI | Key Performance Indicator |
| T2DM | Type 2 Diabetes Mellitus |
| SOAP | Simple Object Access Protocol |
| RSC | Reference Success Criteria |
| UML | Unified Modeling Language |
| XMSG | X-Message |
| 2h-OGTT | 2 h Oral Glucose Tolerance Test |
References
- Thomas, L.H.P. Update on diabetes classification. Med. Clin. N. Am. 2015, 99, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Kahn, S.E.; Hull, R.L.; Utzschneider, K.M. Mechanisms linking obesity to insulin resistance and type 2 diabetes. Nature 2006, 444, 840–846. [Google Scholar] [CrossRef] [PubMed]
- Guariguata, L.; Whiting, D.; Hambleton, I.; Beagley, J.; Linnenkamp, U.; Shaw, J. Global estimates of diabetes prevalence for 2013 and projections for 2035. Diabetes Res. Clin. Pract. 2014, 103, 137–149. [Google Scholar] [CrossRef] [PubMed]
- Beagley, J.; Guariguata, L.; Weil, C.; Motala, A.A. Global estimates of undiagnosed diabetes in adults. Diabetes Res. Clin. Pract. 2014, 103, 150–160. [Google Scholar] [CrossRef] [PubMed]
- American Diabetes Association. Classification and diagnosis of diabetes. Diabetes Care 2016, 40, S11–S24. [Google Scholar]
- Hippisley-Cox, J.; Coupland, C.; Robson, J.; Sheikh, A.; Brindle, P. Predicting risk of type 2 diabetes in England and Wales: Prospective derivation and validation of QDScore. BMJ 2009, 338, b880. [Google Scholar] [CrossRef] [PubMed]
- Meigs, J.B.; Shrader, P.; Sullivan, L.M.; McAteer, J.B.; Fox, C.S.; Dupuis, J.; Manning, A.K.; Florez, J.C.; Wilson, P.W.; Agostino, R.B.; et al. Genotype score in addition to common risk factors for prediction of type 2 diabetes. N. Engl. J. Med. 2008, 359, 2208–2219. [Google Scholar] [CrossRef] [PubMed]
- Gillies, C.L.; Abrams, K.R.; Lambert, P.C.; Cooper, N.J.; Sutton, A.J.; Hsu, R.T.; Khunti, K. Pharmacological and lifestyle interventions to prevent or delay type 2 diabetes in people with impaired glucose tolerance: Systematic review and meta-analysis. BMJ 2007, 334, 299. [Google Scholar] [CrossRef] [PubMed]
- Noble, D.; Mathur, R.; Dent, T.; Meads, C.; Greenhalgh, T. Risk models and scores for type 2 diabetes: Systematic review. BMJ 2011, 343, d7163. [Google Scholar] [CrossRef] [PubMed]
- Collins, G.S.; Reitsma, J.B.; Altman, D.G.; Moons, K.G. Transparent reporting of a multivariable prediction model for Individual Prognosis or Diagnosis (TRIPOD): The TRIPOD Statement. Ann. Intern. Med. 2015, 162, 55. [Google Scholar]
- Steyerberg, E.W.; Moons, K.G.M.; van der Windt, D.A.; Hayden, J.A.; Perel, P.; Schroter, S.; Riley, R.D.; Hemingway, H.; Altman, D.G. Prognosis Research Strategy (PROGRESS) 3: Prognostic model research. PLoS Med. 2013, 10, e1001381. [Google Scholar] [CrossRef] [PubMed]
- Collins, G.S.; Moons, K.G.M. Comparing risk prediction models. BMJ 2012, 344, e3186. [Google Scholar] [CrossRef] [PubMed]
- Riley, R.D.; Ensor, J.; Snell, K.I.E.; Debray, T.P.A.; Altman, D.G.; Moons, K.G.M.; Collins, G.S. External validation of clinical prediction models using big datasets from e-health records or IPD meta-analysis: Opportunities and challenges. BMJ 2016, 353, i3140. [Google Scholar] [CrossRef] [PubMed]
- Reilly, B.M.; Evans, A.T. Translating clinical research into clinical practice: Impact of using prediction rules to make decisions prediction rules to make decisions. Ann. Intern. Med. 2006, 144, 201–209. [Google Scholar] [CrossRef] [PubMed]
- Altman, D.G.; Vergouwe, Y.; Royston, P.; Moons, K.G. Prognosis and prognostic research: Validating a prognostic model. BMJ 2009, 338, b605. [Google Scholar] [CrossRef] [PubMed]
- Moons, K.G.; Royston, P.; Vergouwe, Y.; Grobbee, D.E.; Altman, D.G. Prognosis and prognostic research: What, why, and how? BMJ 2009, 338, b375. [Google Scholar] [CrossRef] [PubMed]
- Steyerberg, E.W.; Vickers, A.J.; Cook, N.R.; Gerds, T.; Gonen, M.; Obuchowski, N.; Pencina, M.J.; Kattan, M.W. Assessing the performance of prediction models: A framework for some traditional and novel measures. Epidemiology 2010, 21, 128–138. [Google Scholar] [CrossRef] [PubMed]
- Raina, R.; Shen, Y.; Mccallum, A.; Ng, A.Y. Classification with hybrid generative/discriminative models. Proc. Neuro Inf. Process. Syst. 2004, 545–552. [Google Scholar]
- Kayacan, E.; Ulutas, B.; Kaynak, O. Grey system theory-based models in time series prediction. Expert Syst. Appl. 2010, 37, 1784–1789. [Google Scholar] [CrossRef]
- Schmidt, M.I.; Duncan, B.B.; Bang, H.; Pankow, J.S.; Ballantyne, C.M.; Golden, S.H.; Folsom, A.R.; Chambless, L.E. Identifying individuals at high risk for diabetes: The atherosclerosis risk in communities study. Diabetes Care 2005, 28, 2013–2018. [Google Scholar] [CrossRef] [PubMed]
- Talmud, P.J.; Hingorani, A.D.; Cooper, J.A.; Marmot, M.G.; Brunner, E.J.; Kumari, M.; Kivimaki, M.; Humphries, S.E. Utility of genetic and non-genetic risk factors in prediction of type 2 diabetes: Whitehall II prospective cohort study. BMJ 2010, 340, b4838. [Google Scholar] [CrossRef] [PubMed]
- Sackett, D.L. Evidence-based medicine. Semin. Perinatol. 1997, 21, 3–5. [Google Scholar] [CrossRef]
- Hilliard, R. All About IEEE Std 1471. In IEEE Recommended Practice for Architectural Description of Software Intensive Systems (IEEE Std 1471-2000); IEEE Computer Society: New York, NY, USA, 2007. [Google Scholar]
- Kalra, D. ISO/DIS 18308 Requirements for an Electronic Health Record Reference Architecture; International Organization for Standardization: Geneva, Switzerland, 2010. [Google Scholar]
- Segagni, D.; Ferrazzi, F.; Larizza, C.; Tibollo, V.; Napolitano, C.; Priori, S.G.; Bellazzi, R. R engine cell: Integrating R into the i2b2 software infrastructure. J. Am. Med. Inform. Assoc. 2011, 18, 314–317. [Google Scholar] [CrossRef] [PubMed]
- Newcomer, E.; Lomow, G. Understanding SOA With Web Services; Addison-Wesley Professional: Boston, MA, USA, 2004. [Google Scholar]
- Herman, I. Semantic Web. W3C, 2011. Available online: http://www.w3.org/2001/sw/ (accessed on 28 December 2017).
- Lluch-Ariet, M. The MOSAIC System—Contributions to Efficient and Secure Exchange of Networked Clinical Data. Ph.D. Thesis, Polytechnic University of Catalonia, Barcelona, Spain, 2016. [Google Scholar]
- Roset, R.; Lurgi, M.; Croitoru, M.; Lluch-Ariet, B.H.M.; Lewis, P. Visual mapping tool for database interoperability: The healthagents case. In Proceedings of the Third Conceptual Structures Tool Interoperability Workshop, Toulouse, France, 7 July 2008; pp. 44–54. [Google Scholar]
- Murphy, S.N.; Weber, G.; Mendis, M.; Gainer, V.; Chueh, H.C.; Churchill, S.; Kohane, I. Serving the enterprise and beyond with informatics for integrating biology and the bedside (i2b2). J. Am. Med. Inform. Assoc. 2010, 17, 124–130. [Google Scholar] [CrossRef] [PubMed]
- Murphy, S.; Churchill, S.; Bry, L.; Chueh, H.; Weiss, S.; Lazarus, R.; Zeng, Q.; Dubey, A.; Gainer, V.; Mendis, M.; et al. Instrumenting the healthcare enterprise for discovery research in the genomic era. Genome Res. 2009, 19, 1675–1681. [Google Scholar] [CrossRef] [PubMed]
- Lindstrom, J.; Tuomilehto, J. The diabetes risk score: A practical tool to predict type 2 diabetes risk. Diabetes Care 2003, 26, 725–731. [Google Scholar] [CrossRef] [PubMed]
- Alssema, M.; Vistisen, D.; Heymans, M.W.; Nijpels, G.; Glümer, C.; Zimmet, P.Z.; Shaw, J.E.; Eliasson, M.; Stehouwer, C.D.A.; Tabák, A.G.; et al. The evaluation of screening and early detection strategies for type 2 diabetes and impaired glucose tolerance (DETECT-2) update of the Finnish diabetes risk score for prediction of incident type 2 diabetes. Diabetologia 2010, 54, 1004–1012. [Google Scholar] [CrossRef] [PubMed]
- Mann, D.M.; Bertoni, A.G.; Shimbo, D.; Carnethon, M.R.; Chen, H.; Jenny, N.S.; Muntner, P. Comparative validity of 3 diabetes mellitus risk prediction scoring models in a multiethnic US cohort: The Multi-Ethnic Study of Atherosclerosis. Am. J. Epidemiol. 2010, 171, 980–988. [Google Scholar] [CrossRef] [PubMed]
- Stern, M.P. Identification of persons at high risk for type 2 diabetes mellitus: Do we need the oral glucose tolerance test? Ann. Intern. Med. 2002, 136, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Abdul-Ghani, M.A.; Abdul-Ghani, T.; Stern, M.P.; Karavic, J.; Tuomi, T.; Bo, I.; DeFronzo, R.A.; Groop, L. Two-step approach for the prediction of future type 2 diabetes risk. Diabetes Care 2011, 34, 2108–2112. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.; Simmons, R.K.; Harding, A.H.; Wareham, N.J.; Griffin, S.J. A simple risk score identifies individuals at high risk of developing type 2 diabetes: A prospective cohort study. Fam. Pract. 2008, 25, 191–196. [Google Scholar] [CrossRef] [PubMed]
- Guasch-Ferré, M.; Bulló, M.; Costa, B.; Martínez-Gonzalez, M.Á.; Ibarrola-Jurado, N.; Estruch, R.; Barrio, F. A risk score to predict type 2 diabetes mellitus in an elderly Spanish Mediterranean population at high cardiovascular risk. PLoS ONE 2012, 7, e33437. [Google Scholar] [CrossRef] [PubMed]
- Wilson, P.W.F. Prediction of incident diabetes mellitus in middle-aged adults: The Framingham Offspring Study. Arch. Intern. Med. 2007, 167, 1068–1074. [Google Scholar] [CrossRef] [PubMed]
- Franzin, A.; Sambo, F.; Camillo, B.D. bnstruct: An R package for Bayesian Network structure learning in the presence of missing data. Bioinformatics 2017, 53, 1250–1252. [Google Scholar] [CrossRef] [PubMed]
- Sambo, F.; Camillo, B.D.; Franzin, A.; Facchinetti, A.; Hakaste, L.; Kravic, J.; Fico, G.; Tuomilehto, J.; Groop, L.; Gabriel, R.; et al. A Bayesian Network analysis of the probabilistic relations between risk factors in the predisposition to type 2 diabetes. In Proceedings of the 2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Milan, Italy, 25–29 August 2015. [Google Scholar]
- Rood, B.; Lewis, M.J. Grid resource availability prediction-based scheduling and task replication. J. Grid Comput. 2009, 7, 479–500. [Google Scholar] [CrossRef]
- Ramakrishnan, L.; Reed, D.A. Predictable quality of service atop degradable distributed systems. Clust. Comput. 2013, 16, 321–334. [Google Scholar] [CrossRef]
- Kianpisheh, S.; Kargahi, M.; Charkari, N.M. Resource availability prediction in distributed systems: An approach for modeling non-stationary transition probabilities. IEEE Trans. Parallel Distrib. Syst. 2017, 28, 2357–2372. [Google Scholar] [CrossRef]
- Heuer, A.; Kaufmann, T.; Weyer, T. Extending an IEEE 42010-compliant viewpoint-based engineering-framework for embedded systems to support variant management. In Proceedings of the 4th International Embedded Systems Symposium, Paderborn, Germany, 17–19 June 2013; Springer: Berlin/Heidelberg, Germany, 2013; pp. 283–292. [Google Scholar]
- Dagliati, A.; Sacchi, L.; Bucalo, M.; Segagni, D.; Zarkogianni, K.; Millana, A.M.; Cancela, J.; Sambo, F.; Fico, G.; Barreira, M.T.M.; et al. A data gathering framework to collect type 2 diabetes patients data. In Proceedings of the 2014 IEEE-EMBS International Conference on Biomedical and Health Informatics (BHI), Valencia, Spain, 1–4 June 2014; pp. 244–247. [Google Scholar]
- Weber, G.M.; Murphy, S.N.; McMurry, A.J.; MacFadden, D.; Nigrin, D.J.; Churchill, S.; Kohane, I.S. The Shared Health Research Information Network (SHRINE): A prototype federated query tool for clinical data repositories. J. Am. Med. Inform. Assoc. 2009, 16, 624–630. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Millana, A.; Fico, G.; Fernández-Llatas, C.; Traver, V. Performance assessment of a closed-loop system for diabetes management. Med. Biol. Eng. Comput. 2015, 53, 1295–1303. [Google Scholar] [CrossRef] [PubMed]
- Site, F.W. Foundation for Intelligent Physical Agents. Available online: http://www.pa.org/ (accessed on 28 December 2017).
- González-Vélez, H.; Mier, M.; Julià-Sapé, M.; Arvanitis, T.N.; García-Gómez, J.M.; Robles, M.; Lewis, P.H.; Dasmahapatra, S.; Dupplaw, D.; Peet, A.; et al. HealthAgents: Distributed multi-agent brain tumor diagnosis and prognosis. Appl. Intell. 2009, 30, 191–202. [Google Scholar] [CrossRef]
- Bellazzi, R. Big data and biomedical informatics: A challenging opportunity. IMIA Yearb. 2014, 9, 8–13. [Google Scholar] [CrossRef] [PubMed]
- Maximilien, E.M.; Singh, M.P. A framework and ontology for dynamic web services selection. IEEE Internet Comput. 2004, 8, 84–93. [Google Scholar] [CrossRef]
- Pryor, G.; Lucey, B.; Maddipatla, S.; McClanahan, C.; Melonakos, J.; Venugopalakrishnan, V.; Patel, K.; Yalamanchili, P.; Malcolm, J. High-level GPU computing with Jacket for MATLAB and C/C++. In Proceedings of the Modeling and Simulation for Defense Systems and Applications VI; International Society for Optics and Photonics: Bellingham, WA, USA, 2011; Volume 8060, p. 806005. [Google Scholar]





| RSC ID | RSC Description |
|---|---|
| RSC#1 | Supporting rich human computer interaction |
| RSC#2 | Supporting intelligent hardware abstraction |
| RSC#3 | Enabling system-driven interaction |
| RSC#4 | Supporting continuity of care |
| RSC#5 | Supporting end user security and privacy |
| RSC#6 | Supporting the update, set up and management of system components |
| RSC#7 | Supporting remote/local operation |
| RSC#8 | Supporting data granted access to perform CRUD (Create, Replace, Update, Delete) operations |
| RSC#9 | Interface existing information systems |
| RSC#10 | Supporting service providers to offer system services |
| RSC#11 | Allowing users to find system tools |
| RSC#12 | Supporting exploitation of different business models |
| RSC#13 | Capturing and utilizing user feedback |
| RSC#14 | Supporting rapid development of new models |
| RSC#15 | Model-based development of services through integrated model transformation tools |
| RSC#16 | Supporting on-line elicitation of requirements and the collection of runtime feedback from users of risk score services |
| RSC#17 | Supporting advanced search, reuse and sharing of service components and resources |
| RSC#18 | Supporting customization of system services |
| Types of Services | Description |
|---|---|
| Module-to-Module (B2B) | Services that are provided by one system module to other module(s) of a different type (e.g., a web service provider from the models requires data from a remote database service provider). |
| Module-to-Client (B2C) | Services that are provided by a module to client stakeholders (e.g., a web service provider provides remote execution of a model). |
| Open Source | Structured Storage | Unstructured Storage | Scalability | Ontologies | |
|---|---|---|---|---|---|
| MongoDB | YES | NO | YES (Json) | YES | NO |
| Hadoop | YES | YES | YES | YES | NO |
| OracleDB | NO | YES | YES -for Oracle NoSQL | YES | NO |
| MySQL | YES | YES | NO | Compromised | NO |
| SQLServer | NO | YES | NO | Compromised | NO |
| I2B2 | YES | YES | NO | Compromised | YES |
| Cassandra | YES | NO | YES | YES | NO |
| Risk Score Name and Validation Study | Mathematical Model | Performance (C statistic) | Predictors |
|---|---|---|---|
| Findrisc [32,33] | Weighted Logistic Regression | 85% | Age, AHT medication, FG, BMI, Waist |
| ARIC [20,34] | Logistic Regression | 80% | Age, Ethnicity, FG, HDL, Triglyceride, Blood Pressure, FHD, Waist, Height |
| San Antonio [35,36] | Linear Regression | 84% | Age, Gender, Ethnicity, FG, BMI, HDL, Blood Pressure, FHD |
| Cambridge [21,37] | Logistic Regression | 75% | Age, Gender, AHT, Steroids, BMI, FHD, Smoking habit |
| PREDIMED [38] | Multivariate Cox Survival Model | 78% | AHT, FG, Blood Pressure, FHD, Smoker, Alcohol Intake |
| Framingham [34,39] | Logistic Regression | 84% | Age, Gender, AHT, FG, BMI, HDL, Triglyceride, Blood Pressure, FHD, Waist |
| MOSAIC [40] | Bayesian Network | 79% | Age, Gender, FG, Smoker, Alcohol, AHT, LL, Physical Activity, Triglyceride, HDL, BMI, Waist, Stroke, FHD |
| REQUEST:,20/09/2017 13:03:55.802 {“IdMessage”: “b8df1baf178043539cad47beea3a51e2”, “sender”: “ModelsService”, “receiver”: “ModelService.R”, “Credential”: token, “language”: “XMSG-SOAP”, “inresponse”, “Content”: { “method”: “LaunchRScript”, “parameters”: { “script”: { “type”: “xs:string”, “value”: “setwd”(“C:\CHOREOGRAPHER\WorkingDirectoryR”), v0=c(SubjID = 21, SEX = 2, AGE = 76, ETHNIC = 1, WAIST = 100, PULSE = 80, DBP = 70,,HOMA_B = 0, HOMA_IR = 0, GLUC0 = 5, TRIG = 1.5909090, MS = 2, PHYSICAL_WORK = 1,CURR_SMOKE = 2, MAR_MARR = 1, MAR_DIV = 2, MAR_WID = 2, PROF_NONE = 2), v0df=as.data.frame(t(v0)), v1=c(SubjID = 22, SEX = 1, AGE = 54, ETHNIC = 1, WAIST = 111, PULSE = 80, DBP = 80,,HOMA_B = 0, HOMA_IR = 0, GLUC0 = 7.222222, TRIG = 1.05681, MS = 1, PHYSICAL_WORK = 1,CURR_SMOKE = 2, MAR_MARR = 2, MAR_DIV = 2, MAR_WID = 1, PROF_NONE = 2), v1df=as.data.frame(t(v1))}, “resultObjects”: {“type”: “xs:string”, “value”: “pd1, pd2”}}}} |
| RESPONSE:,20/09/2017 13:03:55.834 {“IdMessage”: “e624f00474074b5fad810c014ff4a62e”, “sender”: “ModelService.R”, “receiver”: “ModelsService”, “Credential”: token, “language”: “XMSG-SOAP”, “inresponse”: “b8df1baf178043539cad47beea3a51e2”, “Content”: {,“method”: “LaunchRScriptResponse”, “parameters”: “LaunchRScriptResult”: {“type”: “xs:string”, “value”: “[0,000212 0,000446 0,000731 0,001271 0,002155 0,004113 0,0054412 0,0064554 0,008481 0,0122504 0,018581; 0,001776 0,0037354 0,00611205 0,010603 0,017916 0,0339384 0,044676 0,052806 0,068869 0,098103 0,1453966}}}} |
| Gender | Male (2)/Female (6) | |
| Age (Years) | 42 ± 13 | |
| Professional Experience (Years) | 14 ± 10 | |
| ICT Literacy (Self-reported) | High = 3; Medium = 3; Low = 2; | |
| Number of Patients Assisted | Overall | 319.33 ± 247.66 |
| T2DM Patients | 127.44 ± 75.22 | |
| High risk of developing T2DM | 48.00 ± 33.79 | |
| Indicator of Use | Mean | SD | Min | Max |
|---|---|---|---|---|
| Number of users per day | 2.5 | 16.43 | 1 | 4 |
| Duration of sessions (min) | 26.16 | 13.72 | 0.25 | 45.93 |
| Number of patients evaluated per doctor | 6.25 | 4.97 | 1 | 15 |
| Number of patients evaluated per day | 10.71 | 12.18 | 0 | 26 |
| Number of sessions per doctor (user) | 1.82 | 1.16 | 1 | 5 |
| Prediction Risk Score | |||||
| n | Latency (s) | CPU(%) | Memory (kB) | Bandwith (kbps) | |
| Best Case | 1 | 0.016 | 20.20 | 374,012 | 9.8 |
| Worst Case | 8080 | 25.876 | 60.50 | 463,853 | 173.35 |
| Data Imputation Model | |||||
| Input Vars | Latency (s) | CPU(%) | Memory (kB) | Bandwith (kbps) | |
| Best Case | 20 | 1.486 | 48.50 | 360,416 | 40.23 |
| Worst Case | 0 | 1.860 | 49.5 | 360,748 | 63.56 |
| Database Module Performance | ||||||
|---|---|---|---|---|---|---|
| Service | Number of Subjects | Time to Setup (min) | Latency per Patient (s) | CPU (%) | Memory (kb) | Bandwidth (kbps) |
| Emergency | 658 | 79 | 7.412 | 43.70 | 137,733 | 720 |
| Outpatient | 1020 | 67 | 1.766 | |||
| Laboratory | 6402 | 248 | 3.462 | |||
| Regular Queries | - | - | 0.254 | 60.20 | 80,457 | 72,459 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Martinez-Millana, A.; Bayo-Monton, J.-L.; Argente-Pla, M.; Fernandez-Llatas, C.; Merino-Torres, J.F.; Traver-Salcedo, V. Integration of Distributed Services and Hybrid Models Based on Process Choreography to Predict and Detect Type 2 Diabetes. Sensors 2018, 18, 79. https://doi.org/10.3390/s18010079
Martinez-Millana A, Bayo-Monton J-L, Argente-Pla M, Fernandez-Llatas C, Merino-Torres JF, Traver-Salcedo V. Integration of Distributed Services and Hybrid Models Based on Process Choreography to Predict and Detect Type 2 Diabetes. Sensors. 2018; 18(1):79. https://doi.org/10.3390/s18010079
Chicago/Turabian StyleMartinez-Millana, Antonio, Jose-Luis Bayo-Monton, María Argente-Pla, Carlos Fernandez-Llatas, Juan Francisco Merino-Torres, and Vicente Traver-Salcedo. 2018. "Integration of Distributed Services and Hybrid Models Based on Process Choreography to Predict and Detect Type 2 Diabetes" Sensors 18, no. 1: 79. https://doi.org/10.3390/s18010079
APA StyleMartinez-Millana, A., Bayo-Monton, J.-L., Argente-Pla, M., Fernandez-Llatas, C., Merino-Torres, J. F., & Traver-Salcedo, V. (2018). Integration of Distributed Services and Hybrid Models Based on Process Choreography to Predict and Detect Type 2 Diabetes. Sensors, 18(1), 79. https://doi.org/10.3390/s18010079









