1. Introduction
Precision agriculture requires accurate plant or seed distribution across a field. This distribution is to be optimized according to the size and shape of the area in which nutrients and light are provided to plant to obtain the maximum possible yield. These factors are controlled by the spacing between crop rows and the spacing of plants/seeds in a row [
1]. For many crops, row spacing is determined as much by the physical characteristics of agricultural machinery used to work in the field as by the specific biological spacing requirements of the crop [
2]. According to the crop and machinery used, the accuracy of planting by the precision transplanter/seeder to the desired square grid pattern must be adequate for the operation of agricultural machinery in both longitudinal and transverse crop directions.
The current designs of vegetable crop transplanters and seeders utilize several uncoordinated planting modules mounted to a common transport frame. These systems use sub-optimal open-loop methods that neglect the dynamic and kinematic effects of the mobile transport frame and of plant motion relative to the frame and the soil. The current designs also neglect to employ complete mechanical control of the transplant during the entire planting process, producing an error in the final planting position, due to the increased uncertainty of plant location as a result of natural variations in plant size, plant mass, soil traction and soil compaction [
3].
Accurately locating the crop plant, in addition to allowing automatic control of weeds, allows individualized treatment of each plant (e.g., spraying, nutrients). Seeking to ensure minimum physical interaction with plants (i.e., non-contact), different remote sensing techniques have been used for the precise localization of plants in fields. For these localization methods, some authors have decided to address automatic weed control by localizing crop plants with centimetre accuracy during seed drilling [
4] or transplanting [
5,
6] using a global positioning system in real time (RTK-GNSS). These studies, conducted at UC Davis, have shown differences between RTK-GNSS-based expected seed location versus actual plant position. The position uncertainly ranged from 3.0 to 3.8 cm for seeds, and tomato transplants, the mean system RMS was 2.67 cm in the along-track direction. Nakarmi and Tang used an image acquisition platform after planting to estimate the inter-plant distance along the crop rows [
7]. This system could measure inter-plant distance with a minimum error of ±30 cm and a maximum error of ±60 cm.
Today, one of the biggest challenges to agricultural row crop production in industrialized countries is non-chemical control of intra-row (within the crop row) weed plants. Systems such as those developed by Pérez-Ruiz et al. [
8] or the commercial platforms based on computer-controlled hoes developed by Dedousis et al. [
9] are relevant examples of innovative mechanical weeding systems. However, the current effectiveness of mechanical weed removal is constrained by plant spacing, the proximity of the weeds to the plant, the plant height and the operation timing. Other methods for non-chemical weed control, such as the robotic platform developed by Blasco et al. [
10] (capable of killing weeds using a 15-kV electrical discharge), the laser weeding system developed by Shah et al. [
11] or the cross-flaming weed control machine designed for the RHEA project by Frasconi et al. [
12], demonstrate that research to create a robust and efficient system is ongoing. A common feature of all these technological developments is the need for accurate measurement of the distance between plants.
Spatial distribution and plant spacing are considered key parameters for characterizing a crop. The current trend is towards the use of optical sensors or image-based devices for measurements, despite the possible limitations of such systems under uncontrolled conditions such as those in agricultural fields. These image-based tools aim to determine and accurately correlate several quantitative aspects of crops to enable plant phenotypes to be estimated [
13,
14].
Dworak et al. [
15] categorized research studying inter-plant location measurements into two types: airborne and ground-based. Research on plant location and weed detection using airborne sensors has increased due to the increasing potential of unmanned aerial systems in agriculture, which have been used in multiple applications in recent years [
16]. For ground-based research, one of the most widely accepted techniques for plant location and classification is the use of Light Detection and Ranging (LiDAR) sensors [
17]. These sensors provide distance measurements along a line scan at a very fast scanning rate and have been widely used for various applications in agriculture, including 3D tree representation for precise chemical applications [
18,
19] or in-field plant location [
20]. This research continues the approach developed in [
21], in which a combination of LiDAR + IR sensors mounted on a mobile platform was used for the detection and classification of tree stems in nurseries.
Based on the premise that accurate localization of the plant is key for precision chemical or physical removal of weeds, we propose in this paper a new methodology to precisely estimate tomato plant spacing. In this work, non-invasive methods using optical sensors such as LiDAR, infrared (IR) light-beam sensors and RGB-D cameras have been employed. For this purpose, a platform was developed on which different sensor configurations have been tested in two scenarios: North America (UC Davis, CA, USA) and Europe (University of Seville, Andalucia, Spain). The specific objectives, given this approach, were the following:
- -
To design and evaluate the performance of multi-sensor platforms attached to a tractor (a UC Davis platform mounted on the rear of the tractor and a University of Seville platform mounted on the front of the tractor).
- -
To refine the data-processing algorithm to select the most reliable sensor for the detection and localization of each tomato plant.
2. Materials and Methods
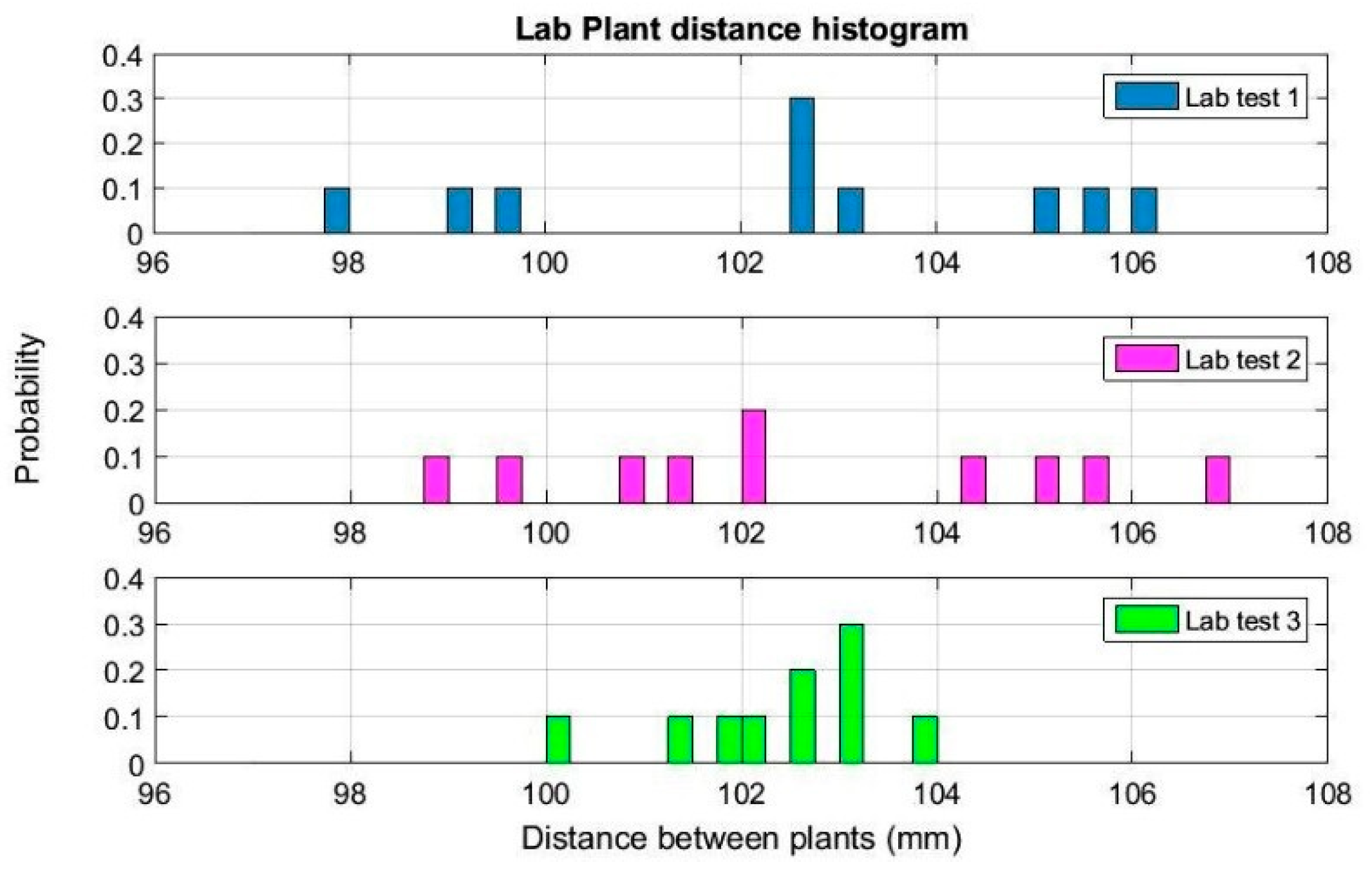
To develop a new sensor platform to measure the space between plants in the same crop row accurately, laboratory and field tests were conducted in Andalucia (Spain) and in California (USA). This allowed researchers to obtain more data under different field conditions and to implement the system improvements required, considering the plant spacing objective. These tests are described below, characterizing the sensors used and the parameters measured.
2.1. Plant Location Sensors
2.1.1. Light-Beam Sensor Specifications
IR light-beam sensors (Banner SM31 EL/RL, Banner Engineering Co., Minneapolis, MN, USA) were used in two configurations: first as a light curtain (with three pairs of sensors set vertically,
Figure 1 central and Figure 4) and later a simpler setup, using only one pair of sensors (
Figure 2), which simplifies the system while still allowing the objective (plant spacing measurement) to be attained. In the light curtain, light-beam sensors were placed transversely in the middle of the platform to detect and discriminate the plant stem in a cross configuration to prevent crossing signals between adjacent sensors. Due to the short range and focus required in laboratory tests, it was necessary to reduce the field of view and the strength of the light signal by masking the emitter and receiver lens with a 3D-printed conical element. In laboratory tests, the height of the first emitter and receiver pair above the platform was 4 cm, and the height of 3D plants (artificial plants were used in laboratory tests; see
Section 2.2) was 13 cm. In the field tests, the sensor was placed 12 cm from the soil (the average height measured manually for real plants in outdoor tests was 19.5 cm) to avoid obstacles in the field (e.g., dirt clods, slight surface undulations). In both cases, the receiver was set to obtain a TTL output pulse each time the IR light-beam was blocked by any part of the plant. The signals generated by the sensors were collected and time-stamped by a microcontroller in real time and stored for off-line analysis. Technical features of the IR light-beam sensors are presented in
Table 1.
2.1.2. Laser Scanner
A LMS 111 LiDAR laser scanner (SICK AG, Waldkirch, Germany), was used in the laboratory and field testing platforms to generate a high-density point cloud on which to perform the localization measurements. Its main characteristics are summarized in
Table 2. The basic operating principle of the LiDAR sensor is the projection of an optical signal onto the surface of an object at a certain angle and range. Processing the corresponding reflected signal allows the sensor to determine the distance to the plant. The LiDAR sensor was interfaced with a computer through an RJ 45 Ethernet port for data recording. Data resolution was greatly affected by the speed of the platform’s movement; thus, maintenance of a constant speed was of key importance for accurate measurements. During data acquisition, two digital filters were activated for optimizing the measured distance values: a fog filter (becoming less sensitive in the near range (up to approximately 4 m)); and an N-pulse-to-1-pulse filter, which filters out the first reflected pulse in case that two pulses are reflected by two objects during a measurement [
22]. Different LiDAR scan orientations were evaluated: scanning vertically with the sensor looking downwards (
Figure 1), scanning with a 45° inclination (push-broom) and a lateral-scanning orientation (side-view).
2.1.3. RGB-D Camera
A Kinect V2 commercial sensor (Microsoft, Redmond, WA, USA), originally designed for indoor video games, was mounted sideways on the research platform during field trials. This sensor captured RGB, NIR and depth images (based on time-of-flight) of tomato plants, although for further analysis, only RGB images were used for the validation of stick/tomato locations obtained from the LiDAR scans, as detailed in
Section 2.4.3. Kinect RGB-captured images have a resolution of 1920 × 1080 pixels and a field of view (FOV) of 84.1 × 53.8°, resulting in an average of approximately 22 × 20 pixels per degree. NIR images and depth camera have a resolution of 512 × 424 pixels, with an FOV of 70 × 60° and a depth-sensing maximum distance of 4.5–5 m. Although systems such as the Kinect sensor were primarily designed for use under controlled light conditions, the second version of this sensor has higher RGB resolution (640 × 480 in v1) and its infrared sensing capabilities were also improved, enabling a more lighting-independent view and supporting its use outdoors under high-illumination conditions. Despite this improvement, we observed that the quality of the RGB images were somewhat affected by luminosity and direct incident light, and therefore, the image must be post-processed to obtain usable results. The images taken by the Kinect sensor were simultaneously acquired and synchronized with the LiDAR scans and the encoder pulses. Because the LabVIEW software (National Instruments, Austin, TX, USA) used for obtaining the scan data was developed to collect three items (the scans themselves, the encoder pulses and the timestamp), a specific Kinect recording software had to be developed to embed the timestamp value in the image data. With the same timestamp for the LiDAR and the image, the data could be matched and the images used to provide information about the forward movement of the platform.
2.2. Lab Platform Design and Tests
To maximize the accuracy of the distance measurements obtained by the sensors, an experimental platform was designed to avoid the seasonal limitations of testing outdoors. Instead of working in a laboratory with real plants, the team designed and created model plants (see
Figure 1) using a 3D printer (Prusa I3, BQ, Madrid, Spain). These plants were mounted on a conveyor chain at a predetermined distance. This conveyor chain system, similar to that of a bicycle, was driven by a small electric motor able to move the belt at a constant speed of 1.35 km·h
−1. For the odometry system, the shaft of an incremental optical encoder (63R256, Grayhill Inc., Chicago, IL, USA) was mounted so that it was attached directly to the gear shaft and used to measure the distance travelled, thus serving as a localization reference system. Each channel in this encoder generates 256 pulses per revolution, providing a 3-mm resolution in the direction of travel. The data generated by the light-beam sensors and the cumulative odometer pulse count were collected using a low-cost open-hardware Arduino Leonardo microcontroller (Arduino Project, Ivrea, Italy) programmed in a simple integrated development environment (IDE). This device enabled recording of data that were stored in a text file for further computer analysis. Several repetitions of the tests were made on the platform to optimize the functions of both light-beam and LiDAR sensors. From the three possible LiDAR orientations, lateral scanning was selected for the field trials because it provided the best information on the structure of the plant, as concluded in [
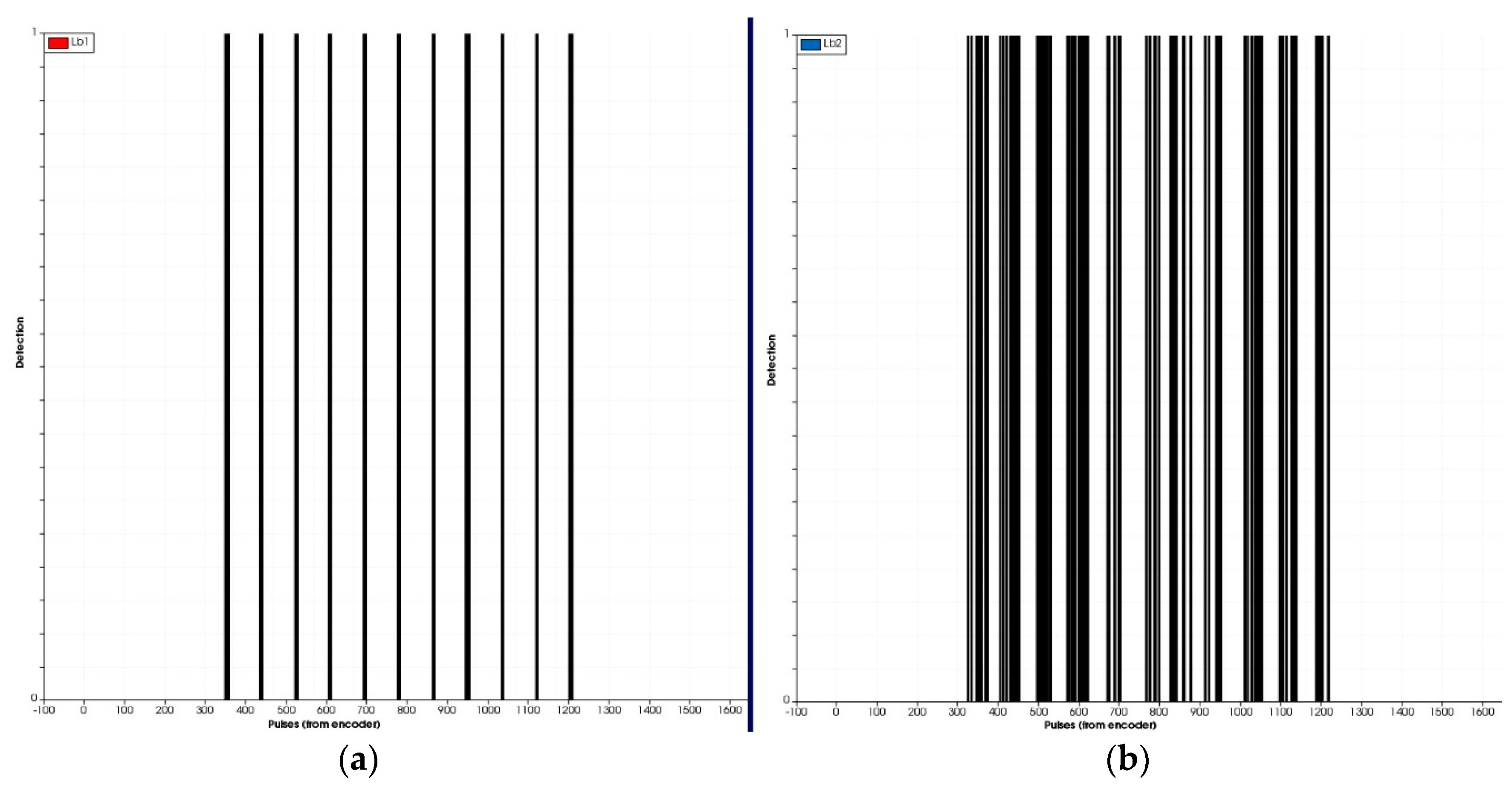
17]. In lab tests, two arrangements of light-beam sensors were assessed: one in a light curtain assembly with three sensor pairs at different heights and another using only one emitter-receiver pair.
2.3. Field Tests
The initial tests, performed in Davis, CA (USA), were used to assess the setup of the light-beam sensor system and detected only the stem of the plants rather than locating it within a local reference system. Once the tomato plants were placed in the field, tests were conducted at the Western Center for Agriculture Equipment (WCAE) at the University of California, Davis campus farm to evaluate the performance of the sensor platform for measuring row crop spacing. For this test, an implement was designed to house the sensors as follows. The same IR light-beam sensor and encoder, both described in
Section 2.1, were used (
Figure 2). The output signals of the sensors were connected to a bidirectional digital module (NI 9403, National Instruments Co., Austin, TX, USA), while the signal encoder was connected to a digital input module (NI 9411, National Instruments Co.). Both modules were integrated into an NI cRIO-9004 (NI 9411, National Instruments Co.), and all data were recorded using LabVIEW (National Instruments Co.). In these early field trials, the team worked on three lines of a small plot of land 20 m in length, where the methodology for detecting the plants within a crop line was tested.
To continue the study of plant localization in a different scenario, additional experiments were designed at the University of Seville, in which a refinement of the LiDAR sensors and data processing were performed. These tests were conducted on several lines of tomato plants manually transplanted from trays, with the plants placed with an approximate, though intentionally non-uniform, spacing of 30 cm. Two of these lines were analysed further, one with 55 tomato plants and the other with 51, and a line of 19 wooden sticks was also placed to provide an initial calibration of the instruments. Due to the initial test conditions, where tomato plants were recently transplanted and had a height of less than 20 cm, the team built an inverted U-shaped platform attached to the front of a small tractor (Boomer 35, New Holland, New Holland, PA, USA,
Figure 3). The choice of the small tractor was motivated by the width of the track, as the wheels of the tractor needed to fit on the sides of the tomato bed, leaving the row of tomatoes clear for scanning and sensors.
As was done in the laboratory platform, the encoder described in
Section 2.1 was used as an odometric system, this time interfaced with an unpowered ground wheel, to determine the instantaneous location of the data along the row.
During the tests, the platform presented several key points that were addressed: (i) The encoder proved to be sensitive to vibrations and sudden movements, so it was integrated into the axis of rotation of an additional wheel, welded to the structure and dampened as much as possible from the vibrations generated by the tractor. In addition, the team had to reduce the slippage of the encoder wheel on the ground to avoid losing pulses; (ii) Correct orientation of the sensors was also key because the mounting angles of the LiDAR sensors would condition the subsequent analysis of the data obtained in the scans and the determination of which data contributed more valuable information; (iii) The speed of the tractor should be as low as possible remain uniform (during the test the average speed was 0.36 m/s) and maintain a steady course without steering wheel movements to follow a straight path.
2.4. Data-Processing Methodology
To detect precisely and determine properly the distances between plants in both laboratory and field tests, the data provided by the sensors were merged and analysed.
2.4.1. Plant Characterization Using Light-Beam Sensors
The methodology followed to analyse data obtained from the light-beam curtain (which was formed by three light-beam sensors in line) was similar to that described in [
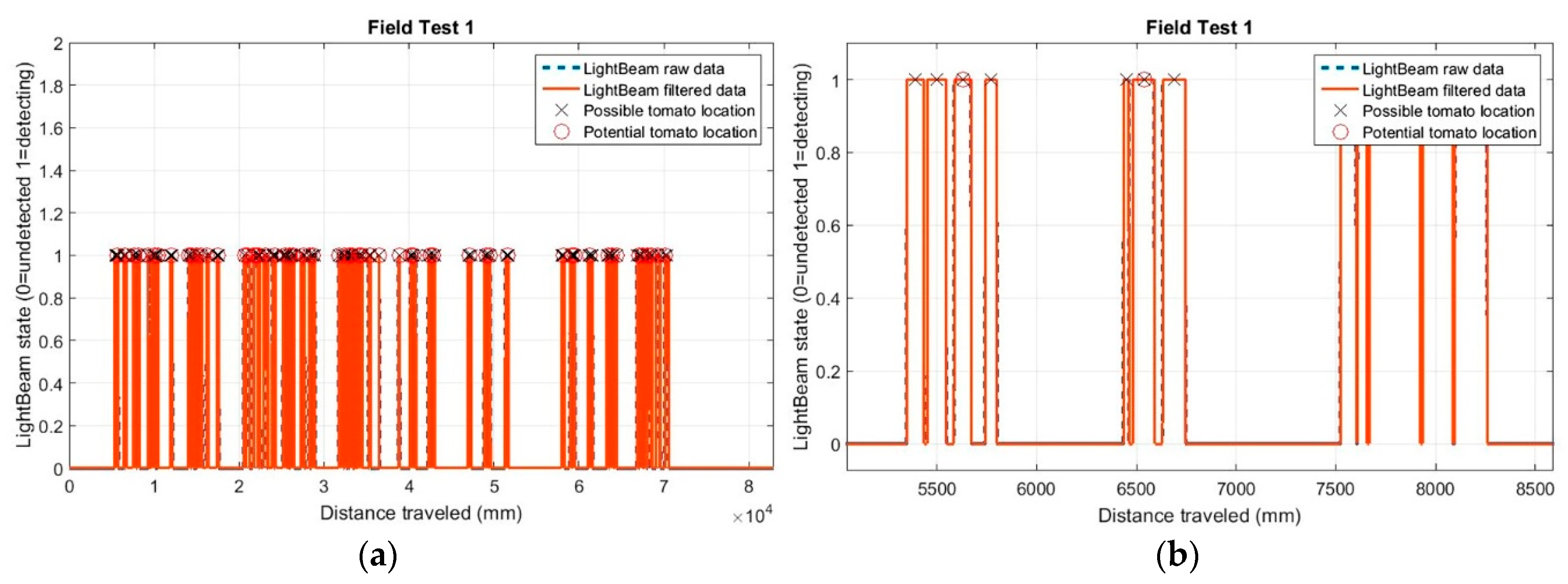
21]. The algorithm outputs the moment that the beam was interrupted and associates the beam with an encoder pulse. Because the 3D plants had a wider shape at the top (leaves) than the bottom (stem), and therefore more interruptions were received, the algorithm had to be adapted to each sensor pair and each height for plant detection. To discriminate correctly between plants for the light curtain case, the developed algorithm implemented a distance range, measured in pulses from the encoder, that allowed the verification of the presence or absence of a plant after the detection of the stem, inferring that interruptions received from the sensors placed at the middle and top heights before and after the stem corresponded to the leaves and the rest of the plant structure, respectively. For the analysis of data obtained from the single pair of IR light-beam sensors, a Matlab routine (MATLAB R2015b, MathWorks, Inc., Natick, MA, USA) was developed. System calibration was performed using 11 artificial plants in the laboratory test and 122 real tomato plants in the UC Davis field test. The methodology used for the detection of tomato plants was based on the following steps:
Selection of Values for the Variables Used by the Programme for Detection:
- a)
Pulse_distance_relation: This variable allowed us to convert the pulses generated by the encoder into the distances travelled by the platforms. In laboratory trials, the encoder was coupled to the shaft that provided motion to the 3D plants, and in the field, it was coupled to a wheel installed inside the structure of the platform. The conversion factors used for the tests were 1.18 and 0.98 mm per pulse for the laboratory and the field, respectively.
- b)
Detection_filter: To eliminate possible erroneous detections, especially during field trials due to the interaction of leaves, branches and even weeds, the detections were first filtered. We filtered every detection that corresponded to an along-track distance of less than 4 mm while the sensor was active (continuous detection).
- c)
Theoretical_plant_distance: The value for the theoretical distance between plants in a crop line. The value set during testing was 100 mm and 380 mm for the laboratory and the field, respectively.
- d)
Expected_plant_distance: Expected distance between plants in a crop line was defined as the theoretical plant distance plus an error of 20%.
Importing of raw data recorded by the sensors (encoder and existence “1” or absence “0” of detection by the IR sensors). The conversion factor (pulse_distance_relation) provided the distance in mm for each encoder value.
Data were filtered by removing all detections whose length or distance travelled, while the sensors were active, was less than the set value (detection_filter). Thus, potential candidates were selected by registering the following:
- a)
The distance at the start of the detection;
- b)
The distance at the end of the detection;
- c)
The distance travelled during detection and (iv) the mean distance during the detection, which was considered the location of the stem of the plant.
Distance evaluation between the current candidate and the previous potential plant:
- a)
If the evaluated distance was greater than the value set (expected_plant_distance), we considered this candidate as a potential new plant, registering in a new matrix: the number of the plant, the detections that defined it, the midpoint location and the distance from the previous potential plant.
- b)
If the evaluated distance was less than the set value (expected_plant_distance), plant candidate data was added to the previous potential plant, recalculating all components for this potential plant. The new midpoint was considered the detection closest to the theoretical midpoint.
2.4.2. Plant Characterization Using a Side-View LiDAR
For the analysis of the data obtained from the LiDAR, it is important to mention the high complexity of its data, in both volume and format, compared with those data obtained by the light-beam. This is reflected in the following section, which explains the proposed methodology for obtaining both the aerial point clouds of the tomato rows referenced to the encoder sensor and the tomato plant identification. This is a prerequisite for tomato plant localization. For this purpose, it was necessary to pre-process the data, followed by a transformation and translation from the LiDAR sensor to the scanned point.
Pre-Processing of Data
(i) Data pre-processing was performed at the LiDAR sensor.
An off-line Matlab process was used with the actual field data collected during the field experiments. Data were filtered to eliminate false positives or those that did not contribute relevant information, considering only those detections with a distance greater than 0.05 m. Later, the resulting detections were transformed from polar to Cartesian coordinates using a horizontal orientation coordinate system as a reference.
(ii) From the LiDAR sensor to the scanned point: transformations and data delimitation.
To transform the horizontal LiDAR coordinates to the actual LiDAR orientation (lateral in our case), the following steps were followed:
- a)
The starting points were the Cartesian coordinates obtained using the horizontal orientation as a reference .
- b)
To integrate the scan from LiDAR into the platform coordinate system, a different transformation
was applied (see Equation (1)), considering the actual orientation of the LiDAR (see
Table 3). Each LiDAR scanned point in the platform coordinate system
was obtained:
Once transformed, the x translation was applied to coordinates obtained for the actual LiDAR orientation. The encoder values recorded at each scan time were used to update the point cloud x coordinate related to the tractor advance. Additionally, the height values (z coordinate) were readjusted by subtracting the minimum obtained.
Plant Localization
The 3D point cloud processing was performed at each stick or tomato row. Thus, using manual distance delimitation, point clouds were limited to the data above the three seedbeds used during the tests.
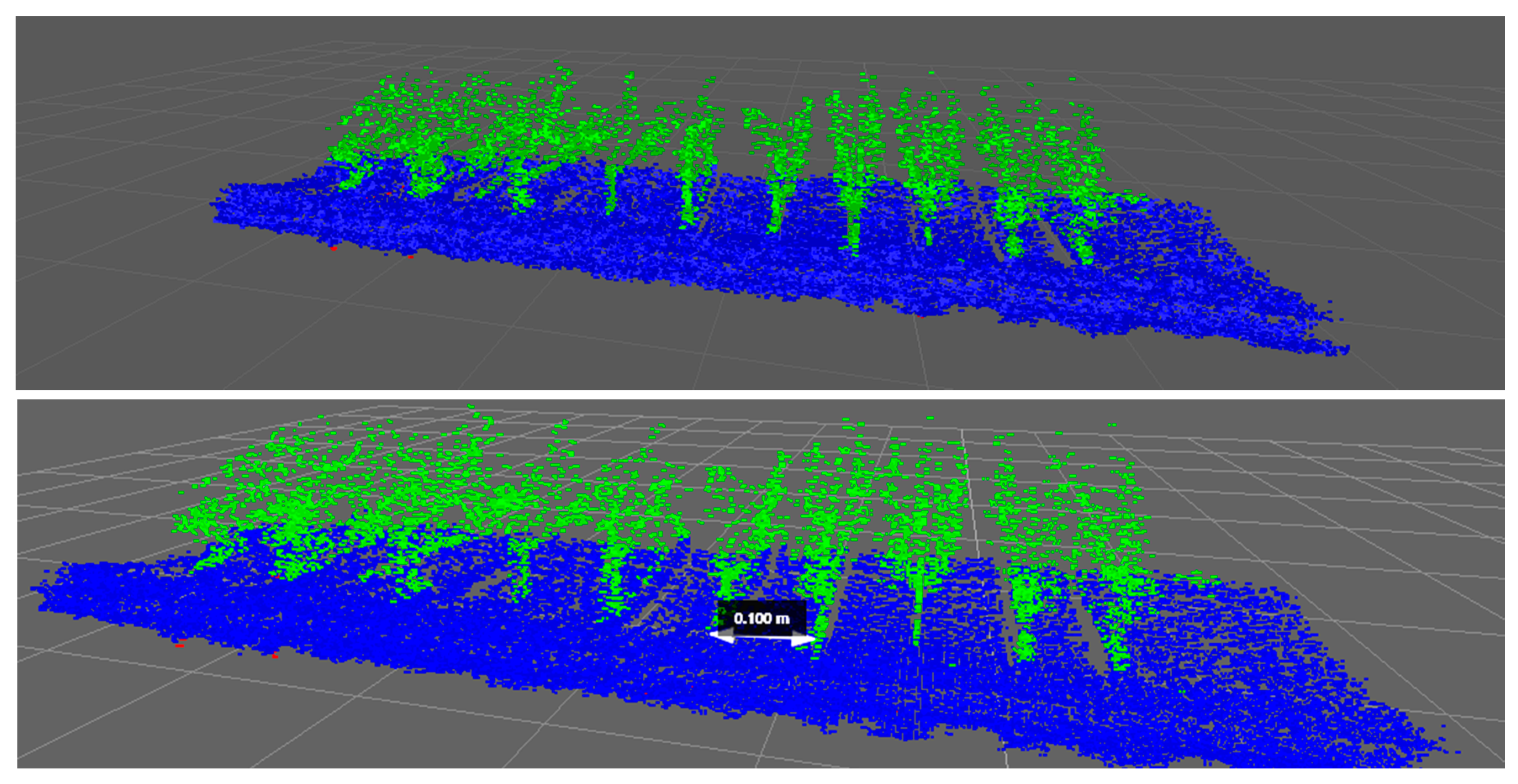
(i) Aerial Point Cloud Extraction
The aerial point data cloud was extracted using a succession of pre-filters. First, all points that did not provide new information were removed using a gridding filter, reducing the size of the point cloud. A fit plane function was then applied to distinguish the aerial points from the ground points. In detail, the applied pre-filters were:
- a)
Gridding filter: Returns a downsampled point cloud using a box grid filter. GridStep specifies the size of a 3D box. Points within the same box are merged to a single point in the output (see
Table 4).
- b)
pcfitplane [
23]: This Matlab function fits a plane to a point cloud using the M-estimator SAmple Consensus (MSAC) algorithm. The MSAC algorithm is a variant of the RANdom SAmple Consensus (RANSAC) algorithm. The function inputs were: the distance threshold value between a data point and a defined plane to determine whether a point is an inlier, the reference orientation constraint and the maximum absolute angular distance. To perform plane detection or soil detection and removal, the evaluations were conducted at every defined evaluation interval (see
Table 4).
Table 4 shows the parameter values chosen during the aerial point cloud extraction. The chosen values were selected by trial and error, selecting those that yielded better results without losing much useful information.
(ii) Plant Identification and Localization
• Plant Clustering
A k-means clustering [
24] was performed on the resulting aerial points to partition the point cloud data into individual plant point cloud data. The parameters used to perform the k-means clustering were as follows:
- ○
An initial number of clusters: Floor ((distance_travelled_mm/distance_between_plants_theoretical) + 1) × 2
- ○
The squared Euclidean distance for the centroid cluster. Each centroid is the mean of points in the cluster.
- ○
The squared Euclidean distance measure and the k-means++ algorithm were used for cluster centre initialization.
- ○
The clustering was repeated five times using the initial cluster centroid positions from the previous iteration.
- ○
Method for choosing initial cluster centroid positions: Select k seeds by implementing the k-means++ algorithm for cluster centre initialization.
A reduction in the number of clusters was determined directly by evaluating the cluster centroids. If pair of centroids were closer than the
min_distance_between_plants, the process was repeated by reducing the number of clusters by one (
Table 5).
A clustering size evaluation was performed, excluding clusters smaller than min_cluster_size.
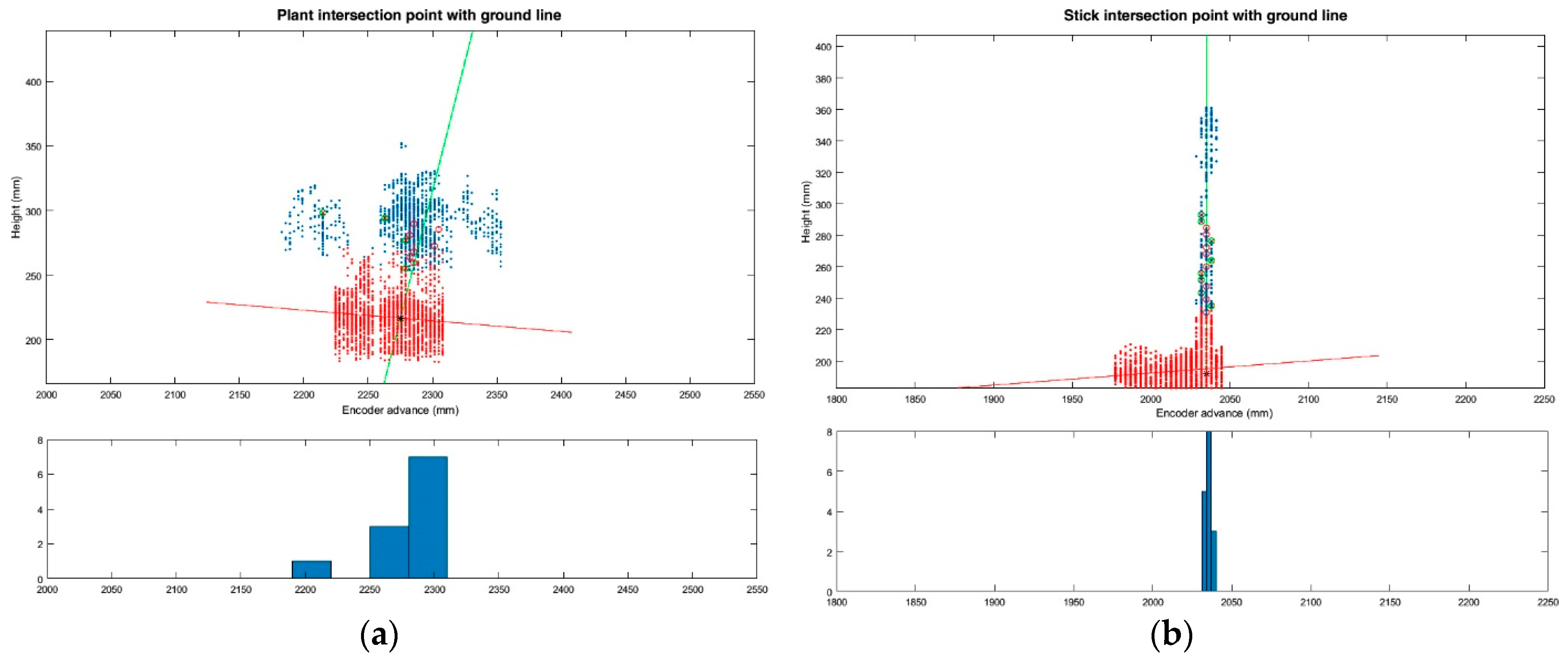
(iii) Plant Location
Three different plant locations were considered:
2.4.3. Validation of Plant Location Using RGB Kinect Images
To obtain the distance between the stems of two consecutive plants using the Kinect camera, it is necessary to characterize the plants correctly and then locate the stems with RGB images from the Kinect camera. This characterization and location of the stem was conducted as follows: a sequence of images of the entire path was obtained (~250 images in each repetition), where the camera’s shooting frequency was established steadily in at 1-s intervals. Obtaining the string with the timestamp of each image was a key aspect of the routine developed in Matlab, as this string would later be used for integration with the LiDAR measurement. The relationship between the timestamp and its corresponding encoder value was used to spatially locate each image (x-axis, corresponding to the tractor advance).
Image processing techniques applied in the characterization of tomato plants from Kinect images generally followed these steps:
- (i)
According to [
25], the first step in most works regarding image analysis is the pre-processing of the image. In this work, an automatic cropping of the original images was performed (
Figure 4a), defining a ROI. Because the test was conducted under unstructured light conditions, the white balance and the saturation of the cropped image (
Figure 4b) were modified;
- (ii)
Next, the image segmentation step of an object-based image analysis (OBIA) was performed to generate boundaries around pixel groups based on their colour. In this analysis, only the green channel was evaluated to retain most of the pixels that define the plant, generating a mask that isolates them (
Figure 4c) and classifying the pixels as plant or soil pixels. In addition, morphological image processing (erosion and rebuild actions) was performed to eliminate green pixels that were not part of the plant;
- (iii)
Each pair of consecutive images was processed by a routine developed in Matlab using the Computer Vision System Toolbox. This routine performs feature detection and extraction based on the Speeded-Up Robust Features “SURF” algorithm [
26] to generate the key points and obtain the pairings between characteristics of the images.
Once the plant was identified in each image, the location of the stem of each plant was defined according to the following subroutine:
- (1)
Calculating the distance in pixels between two consecutive images, as well as the encoder distance between these two images, the “forward speed” in mm/pixel was obtained for each image;
- (2)
For each image, and considering that the value of the encoder corresponds to the centre of the image, the distance in pixels from the stem to the centre of the image was calculated. Considering whether it was to the left or to the right of the centre, this distance was designated positive or negative, respectively;
- (3)
The stem location was calculated for each image using the relation shown in Equation (2) below;
- (4)
Plant location was obtained for each image in which the plant appeared; the average value of these locations was used to calculate the distance between plants: