1. Introduction
Healthcare technology continues to evolve toward the objective of constructing a routine health management system. An essential requirement of such a system is continuous physiological signal monitoring [
1]. In the past, conventional methods of physiological measurement often caused discomfort by keeping subjects immovable or attaching various pieces of equipment to their bodies. However, these methods have been improved with the development of electronic technologies. Now, unobtrusive measurement technologies provide the possibility of managing subjects’ health status as they go about their day-to-day lives [
2].
Researchers have been actively developing technologies to detect biological information from individuals without confining them or causing any discomfort. The most commonly studied methods for unobtrusive sensing include capacitive, photoplethysmographic, ballistocardiographic, and seismocardiographic approaches. These could become pervasive in our daily life without causing any discomfort through the use of wearable device (clothes and accessories) or commonly used objects (furniture and tools) [
3,
4]. These novel methods, which unobtrusively monitor the biological activity of subjects during their everyday life, can be used to set up a continuous examination system for long periods. They can also be used effectively for subjects who are difficult to confine physically, such as the elderly, infirm, and infants. In particular, it is expected that unobtrusive biomonitoring for infants could play an important role in managing breathing and cardiovascular problems, which are the main causes of death [
5,
6,
7].
Conventional methods of physiological signal monitoring over long periods have limitations when applied to infants. For example, electrocardiography (ECG) using adhesive electrodes can cause skin allergies or inflammation. In some cases, the skin can be seriously damaged when detaching the electrodes from the body [
6,
8,
9]. In addition, respiratory measurements can cause discomfort and irritation, as the equipment must be fastened firmly to the body [
7]. Unobtrusive methods such as attaching a sensor to the nose [
10] or foot (in the form of a sock) [
11] have been used to measure biological information from infants. However, these methods still require the direct attachment of sensor devices to the body, which can be uncomfortable. Other indirect measuring methods include infrared thermography [
7], piezoelectric force sensors [
9], and Doppler radar [
12]. Further investigations related to the clinical verification of these approaches is ongoing.
Ballistocardiographs (BCGs) represent a possible solution for monitoring infants’ physiological signals. BCGs are non-invasive instruments that measure the reactional motion of the body produced by cardiac contraction. Unlike ECGs, which require electrodes to be attached directly to the patient’s body to detect electric signals, BCGs generally use sensors for force, pressure, vibration, and displacement. These sensors do not require direct attachments, and therefore, they allow the measurement of physiological signals for long periods of time without perturbing the subject [
4,
13]. However, most studies investigating the application of these methods have focused on adults, and few cases have been focused on infants. More studies are necessary to verify the usefulness of BCG-based physiological measurement in infants.
Therefore, this study proposes a non-confining system for monitoring the biological information of infants using BCG technology. In a previous study, we developed a load-cell installed bed to measure the cardiac activity, respiration, and movement of adults [
14,
15]. The basic concepts of the present system, including the load-cell’s arrangement and the circuit design, are based on the former system. However, the detailed hardware specifications differ to account for relatively lightweight infant subjects. Because it is more difficult to obtain complete BCG signals from their immature hearts, which have smaller cardiac output than adults [
16]. In addition, an algorithm that includes an automatic sensor selection with a signal quality check function has been developed to analyze the heart rate (HR) and breathing rate (BR) sagaciously from the four measured load-cell signals. The final objective is to validate and evaluate the feasibility of the infant biological information monitoring system. In experiments on infant subjects, the designed bed was used to acquire biological data, and the biological information from the proposed algorithm was compared with the reference information obtained simultaneously.
3. Results
We analyzed 13 records from four infant participants. The signals collected from the load-cell sensors include the components caused by cardiac activity and respiratory movement. To analyze the biological information, we also developed a MATLAB-based automatic algorithm that has been validated experimentally.
Table 3 and
Table 4 show the results of the developed algorithm with respect to the beat locations for each experiment (see
Section 2.3.3). The algorithm achieved an average detection rate of 76.16% (
Table 3, HR) and 89.35% (
Table 4, BR) and an average positive predictive value of 99.07% (
Table 3, HR) and 96.22% (
Table 4, BR).
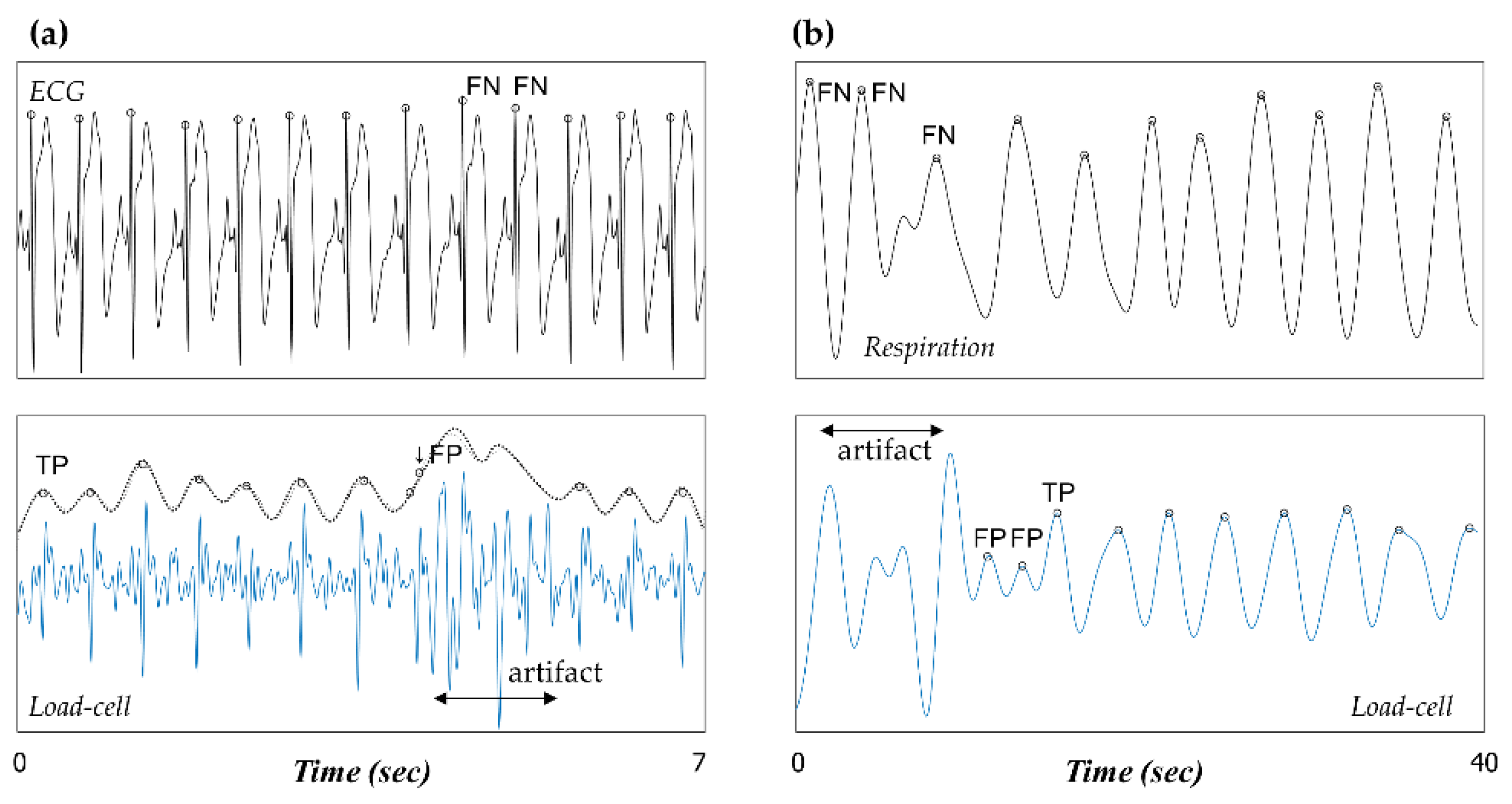
As in the case of many unobtrusive monitoring, the signals acquired by our load-cell system are prone to high-grade noise. The main artifacts are related to movement, which is an inevitable consequence of physiological monitoring. Thus, the most important aspect of signal analysis using unobtrusive sensors is to classify the good-quality signals. In our results, FN is defined as the failure to detect an existing ECG or respiratory peak. It is assumed that most FNs were the result of our algorithm automatically determining an analysis window to be a noise signal and discarding all peaks included in the window. We additionally investigate how FN results inevitably occur in the artifact area and are denoted as FN (inevitable).
However, the number of FPs and FNs gives a limited indication of the accuracy of the estimated HR and BR. We estimate that FN and FP directly affect the lengths of their neighboring beat-to-beat intervals. Therefore, the overall quality of the estimated HR and BR monitoring was assessed by the mean difference and error in computing the intervals extracted from the load-cell sensors and the corresponding intervals given by the reference equipment.
Table 5 and
Table 6 show the adequacy of the proposed algorithm with respect to regular HR or BR monitoring. On average, 73.79% and 84.25% of each signal was identified as being artifact-free and usable for HR or BR estimation, respectively. The results indicate that the mean errors of the estimated HR averaged over 5 s and the estimated BR averaged over 30 s were 2.55% and 2.66%, respectively.
Some other related results cited in the literature are summarized in
Table 7. However, it is impossible to make an objective comparison between our results and those in
Table 7, which were obtained under different experimental circumstances. In particular, most of the results (Records 02–07) were obtained from adult subjects. The differences in subject details, recording times, and sensors used in the study are represented in this table.
4. Discussion and Conclusions
After the basic concept of BCG was introduced in 1877, unobtrusive physiological measurement using BCG technology has become an interesting topic that has attracted much research attention. Recently, a review of numerous studies on BCG has been presented in [
4]. However, it is rare to find studies that have focused on monitoring the biological signals of infants. As mentioned in the introduction section, BCG technology could play a more important role when applied to infants who are difficult to confine physically. We believe that the proposed device and algorithm presented herein are essential for realizing the unobtrusive monitoring of infants.
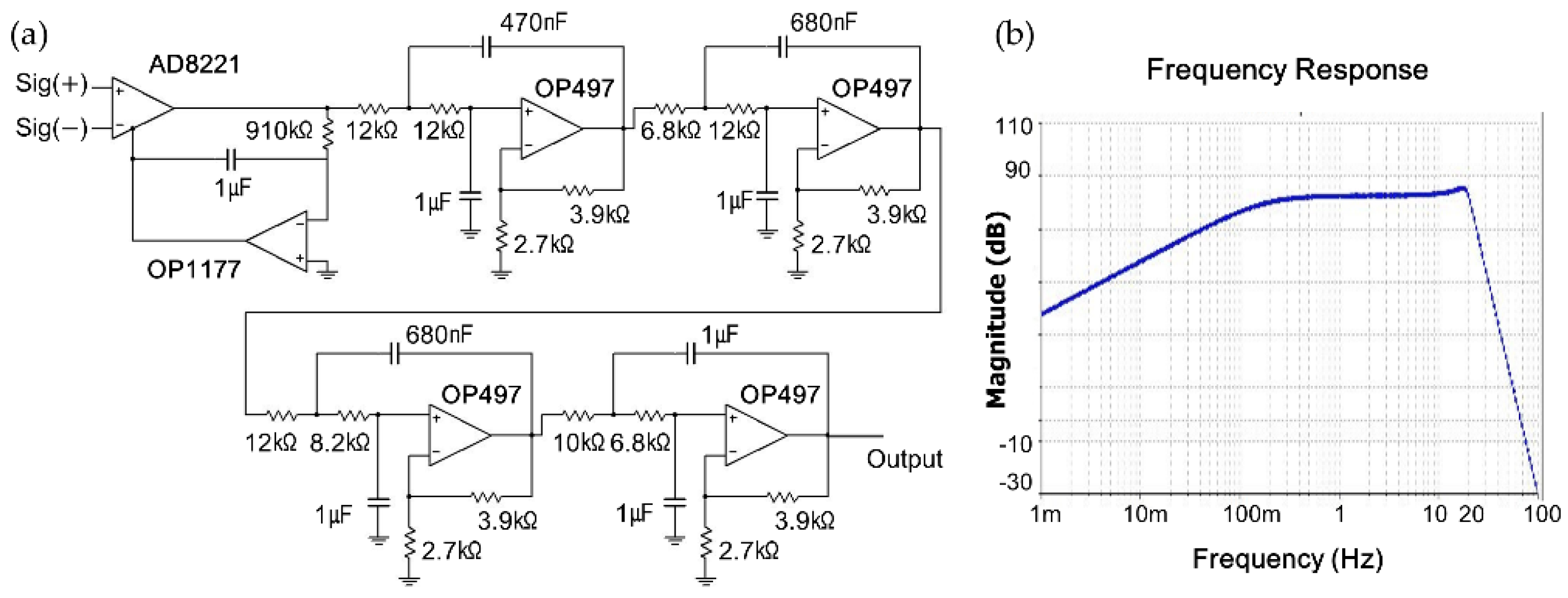
In this study, we developed a load-cell installed bed to measure the cardiac activity, respiration, and movement of infants, as illustrated in
Figure 1. The basic concepts of the present system, including the load-cell’s arrangement and the circuit design, are based on our former system [
14,
15]. However, the detailed hardware specifications differ to account for the relatively low weight of the infant subjects. At the beginning of our study, we attempt to revamp an existing monitoring system. This system was built with four load-cell sensors (MNC-100L, CAS, Yangju-si, Gyeonggi-do, Korea) installed under the legs of the bed. The capacity of the load-cell was 100 kg, and its rated output is 2 mV/V. Although it was well suited for detecting the subtle body vibrations of adults, we could not discern any useful signal for applying the same system to infants. We suppose that the high capacity of the load-cell for covering the entire weight, including that of the bed, leads to lower sensitivity, which is not appropriate for measuring the subtle body vibrations of infants. Considering the few characteristics that affect the load-cell sensitivity (see
Section 2.1), it is relatively easy to reduce the capacity of the load-cell. We designed a simple load-cell system (capacity of 6 kg) while retaining the essential functions, thus we expect a theoretical system capacity of only 24 kg for realizing an improvement in the sensitivity. In the commercialization stage, we plan to incorporate our developed device into the body of a commercial infant’s bed.
We could finally measure cardiac and respiratory signals by using load-cell sensors.
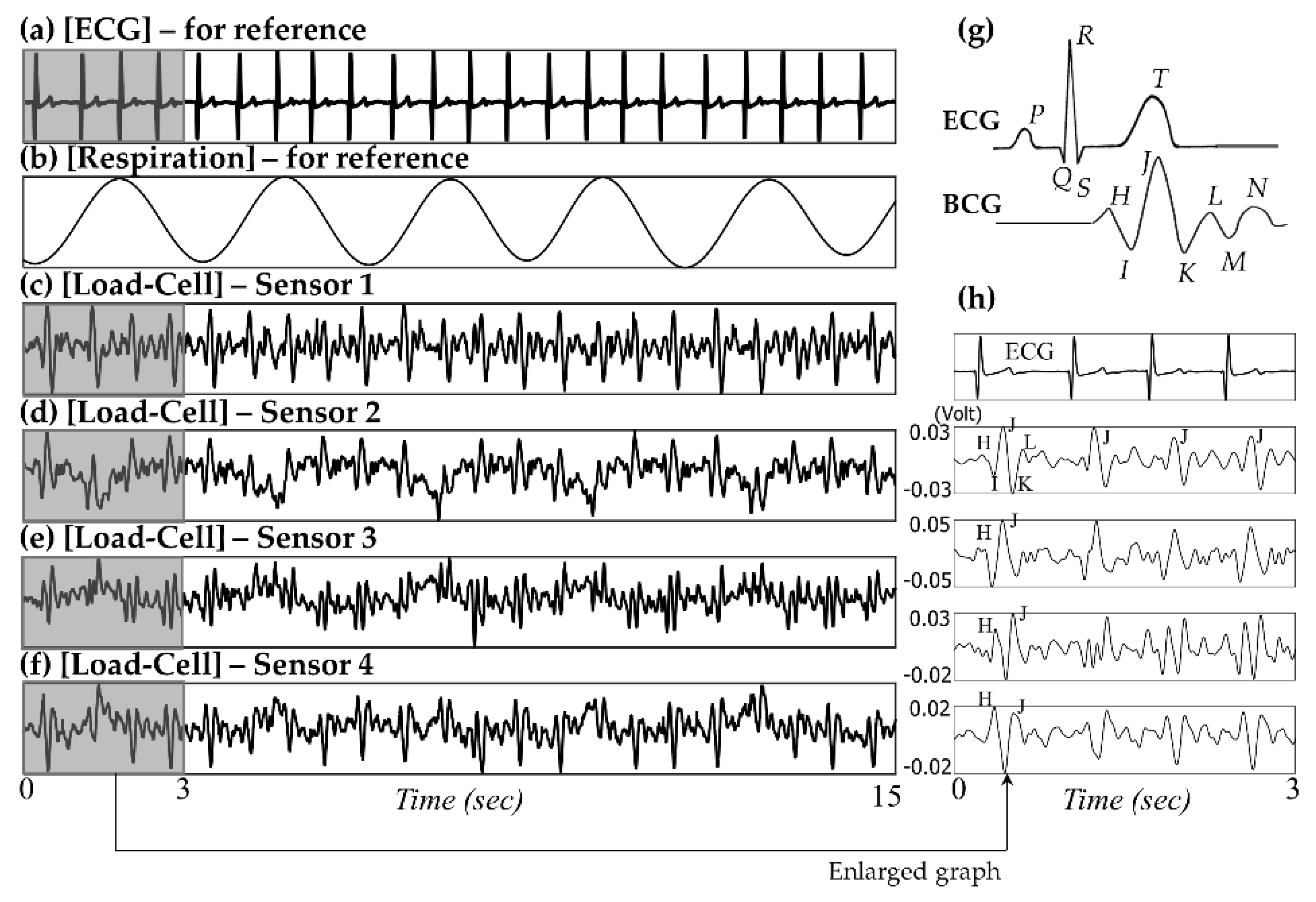
Figure 3 shows a partial example of the monitored signals measured from a 19-month-old male subject. According to the observational record, the subject lay down in bed in a somewhat diagonal direction. His head was located near sensor 1, the region of the right shoulder and heart was located near sensor 2, the region of the hip and right leg was located near sensor 3, and the left leg was located near the middle of sensors 3 and 4 (please refer to the sensor locations in
Figure 1a). Because the chest and stomach are biased to the region of sensors 2 and 3, the respiratory signals are observed as relatively distinct shapes in the corresponding channels. Furthermore, there are phase differences in the recorded respiratory signals between sensors 2 and 3. As already discussed in several literatures, the respiration signals could have phase differences caused by the different movement directions depending on the longitudinal arrangement of sensors [
26]. As mentioned above, sensors 2 and 3 are located near the chest and stomach, respectively. Thus, different timings of expansion and contraction of the stomach and thorax are reflected in the signal [
27].
The amplitudes of the BCG signals vary depending on their location, and the BCG waveform recorded from sensor 2 shows maximum IJ amplitude (
Figure 3h). Because sensor 2 is the nearest to the heart, it can be assumed that the distance between the vibration source and the sensor affects the signal attenuation. Clean BCG waveforms are observed in the signal from sensor 1, and these are particularly less interfered by respiratory movements. In the experiment, the subject’s head was located near sensor 1; we assume that this posture feature is related to the phenomena. We assume that the location of the head affects the interference of the respiratory movement. The relative dominance of cardiac fluctuations in the area near the head is often found from our past experiences, even research approaches for embedding force transducers in pillows to monitor cardiac cycles have been reported [
28].
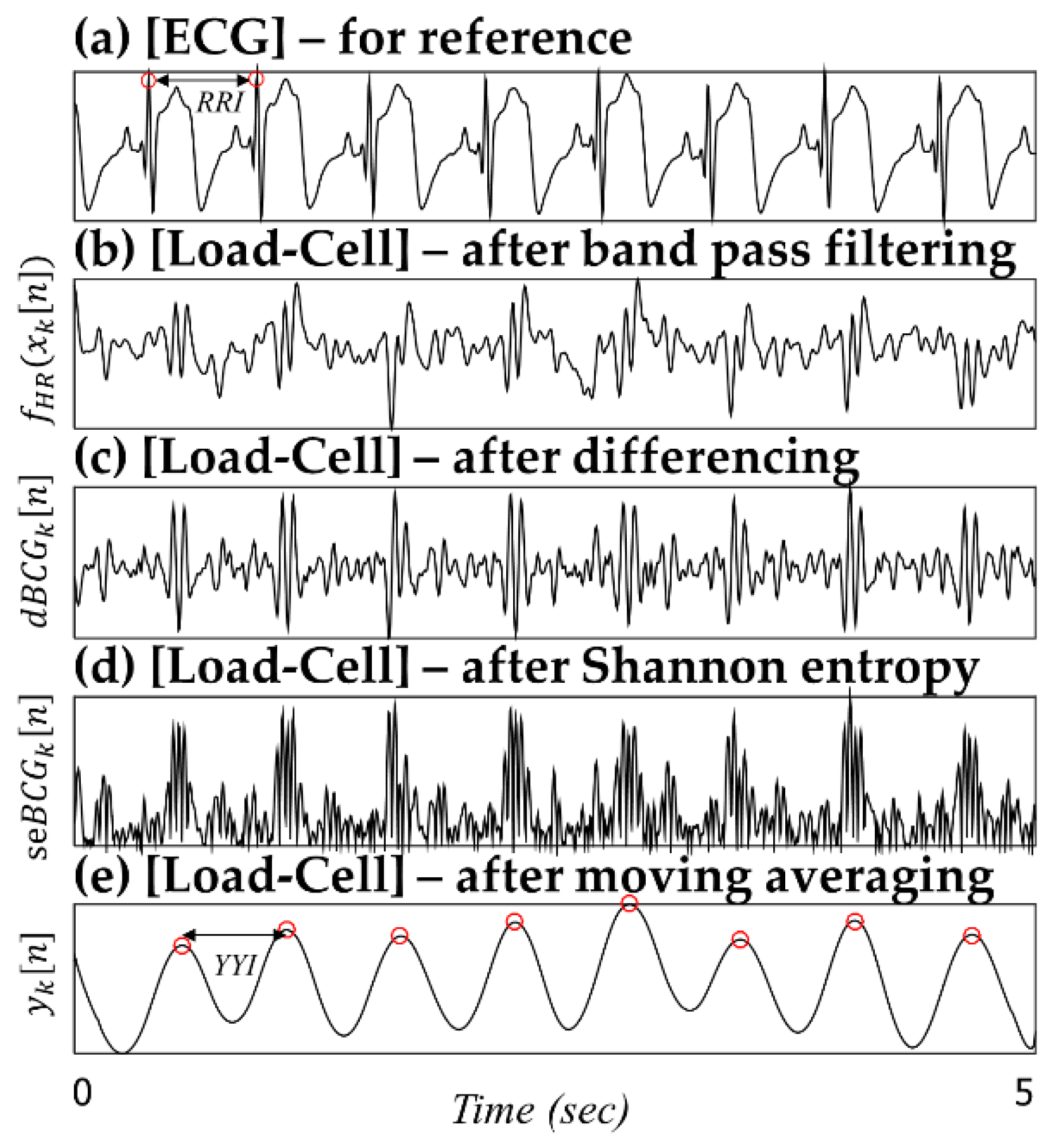
The biggest challenge in monitoring the heart rate using BCG technology is finding features. Because it is generally known that the J peak has the largest amplitude, algorithms for estimating the heart rate by detecting the J peak have been developed. However, unlike an ECG signal having one special fiducial point, that is, QRS complex, the several waveform peaks of the BCG signal (labeled H through N) have similar size and shape. Furthermore, because the BCG signal waveform varies a lot depending on the sensor used and the position at which it was measured, it is not easy to develop a robust algorithm that considers all these characteristics. Brüser
et al. developed and verified a new algorithm that can measure the beat-to-beat interval that corresponds to a heartbeat, unlike the conventional method for finding the location of the J peak [
25]. It is difficult to expect the J peak to be dominant for an infant BCG signal processed in this study because of the nature of a premature heart. We also thought that it was more important to find the corresponding interval to obtain the correct average heartbeat rate instead of finding the exact location of the J peak. Thus, we developed a modified algorithm based on Brüser’s method.
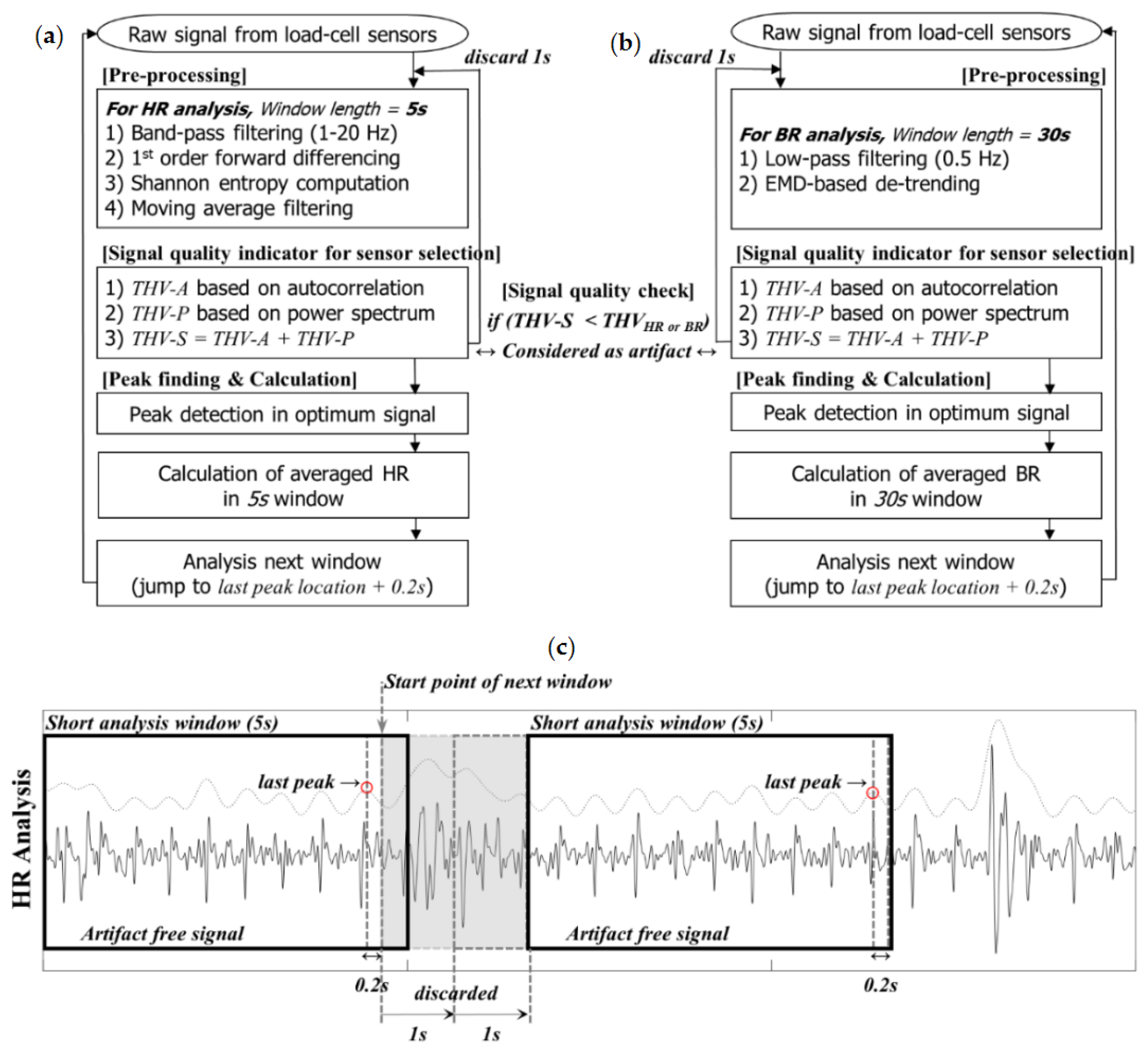
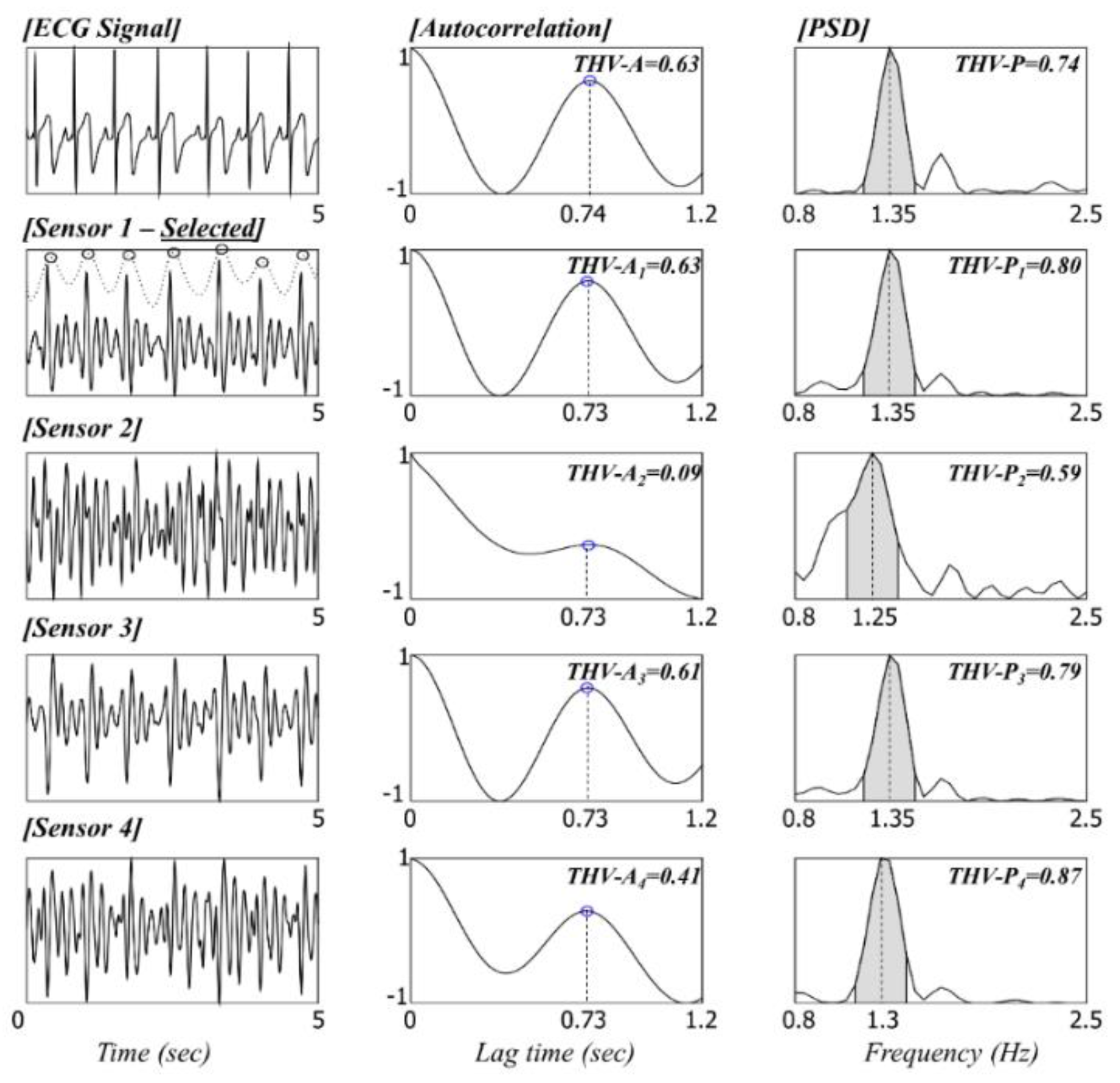
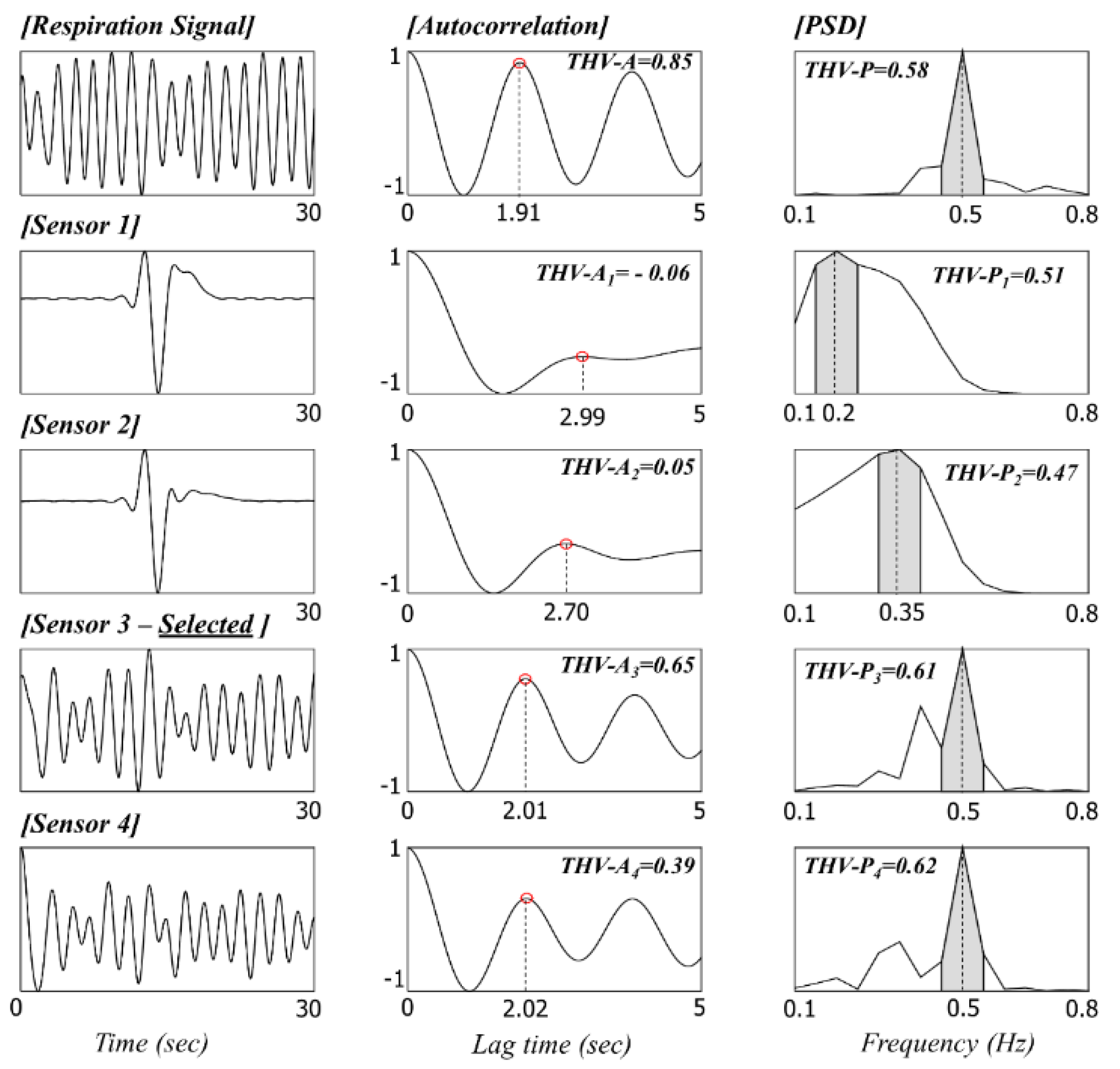
In this study, the sensor selection process that detects valid signals from multiple sensors was considered important, unlike in previous studies. By combining the advantages of existing methods [
5,
29], we developed an automatic sensor selection method using an indicator that can estimate the artifact level using the auto correlation coefficient value and area rate of PSD. We assessed the coverage and mean relative error of experiments by varying the threshold value of the indicator that decides the artifact level. There is a trade-off: if the threshold value is set high, the error as well as the coverage decrease because of the increased number of dropped signals. The threshold value from our empirical experience based on the result of this study is 0.65 for HR analysis and 1.25 for BR analysis.
The performance of our algorithm was evaluated using experimental results, and it is summarized in
Table 3,
Table 4,
Table 5 and
Table 6.
Table 3 and
Table 4 show the result of evaluating how precisely the algorithm can detect the peak location in terms of TP, FP, and FN. The peak was detected at 76.16% and 89.35% in contrast to the reference signal (ECG for heartbeat and chest-belt for respiration, respectively). The increased number of FNs in HR analysis is inevitable because of the nature of the algorithm, which automatically runs the entire process of checking the quality and selecting the optimum sensor. In fact, the inevitable number of FNs that occurred in the area determined to be an artifact area was around 55% (=7469/13,417) of the total number of FNs in the HR analysis. This serves as contrasting evidence that only a clear signal was analyzed using a strict standard that is underpinned by high PPV values. Apart from traditional methods for finding the location of the J peak, the detected beat-to-beat intervals that correspond to the heartbeat and respiratory cycle were evaluated by averaging the error of the preset window. The averaged relative differences and errors are summarized in
Table 5 and
Table 6, respectively.
We also summarized several related studies in
Table 7. The application of BCGs to infants is very rare, and it is difficult to directly compare our method with other BCG-based techniques applied to a similar group of subjects. Erkinjuntti
et al. reported the feasibility of the static charge sensitive bed method for the monitoring of around 40 neonates [
16]. However, they showed just a partial example of the simultaneous recording of signals and did not present algorithms for HR and BR analysis. Wang
et al. investigated a piezoelectric film (PVDF) sensor system; however, their experiments only lasted 10 min on average, and their method showed only mean peak detection errors including FP and FN. Thus, we could estimate the accuracy of the estimated HR and BR signals to have mean errors of 8.24% and 4.41%, respectively [
9]. For adults, BCG-derived HR and BR measurements have achieved lower errors than in our study. Other sensors such as a static charge sensor, piezoelectric sensor (PVDF and EMFi), acceleration sensor, and optical sensor have been used for detecting BCG signals when lying down in bed. A direct comparison is difficult because each method has its own advantages. However, because the load-cell sensor used in this study has the stability in terms of long-term monitoring with high accuracy as well as can simultaneously detect static/dynamic changes, we decided that it was suitable for monitoring the biological signals of infants for a long period of time when they were lying in bed. Studies for comparing the features of various sensors processed with standardized conditions in a controlled situation are needed.
We designed a load-cell-based physiological signal monitoring bed for infants and investigated whether BCGs and respiratory signals could be obtained accurately. By analyzing the data derived from 13 experiments with four infants, we verified that the proposed BCG-based technology could estimate both HR and BR information with average errors of 2.55% and 2.66%, respectively, compared with reference equipment. Although our results demonstrate that the proposed method shows acceptable performance, there are two main limitations in our study—insufficient recording time and too few measurements. Future research projects should increase the number of subjects in the experiment and extend the age distribution. The recording times should be increased to cover various occurrences of all possible types and combinations of movements during sleep and wakefulness. However, despite these limitations, the preliminary results provide a positive feasibility for future studies of BCG-based measurements in infants. We believe that the proposed device and algorithm presented here are essential steps toward substantiating the unobtrusive physiological measurement for infants. The proposed technology could be used for the continuous observation of infants, especially to detect respiratory distress and cardiac abnormalities. Also we expect extensive applications in the field of sleep research for analyzing sleep efficiency and structures of infants.