Detection of Gold Nanoparticles Aggregation Growth Induced by Nucleic Acid through Laser Scanning Confocal Microscopy
Abstract
:1. Introduction
2. Preparation of Samples and Methods
2.1. Gold Nanoparticles
2.2. Preparation of the DNA Solution
2.3. LSCM Measurements
2.4. Scanning Electron Microscopy
3. Experimental Results and Discussion
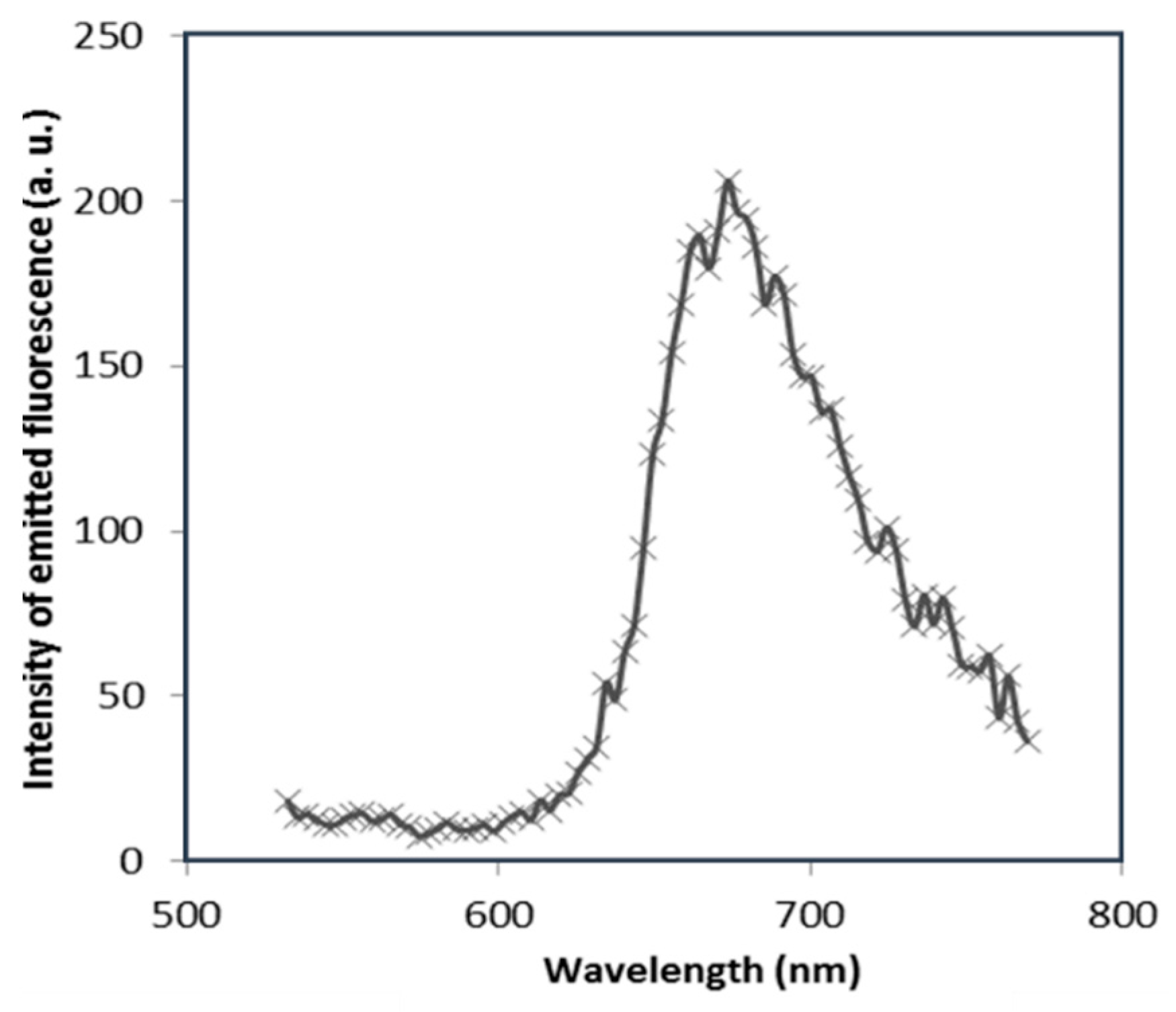
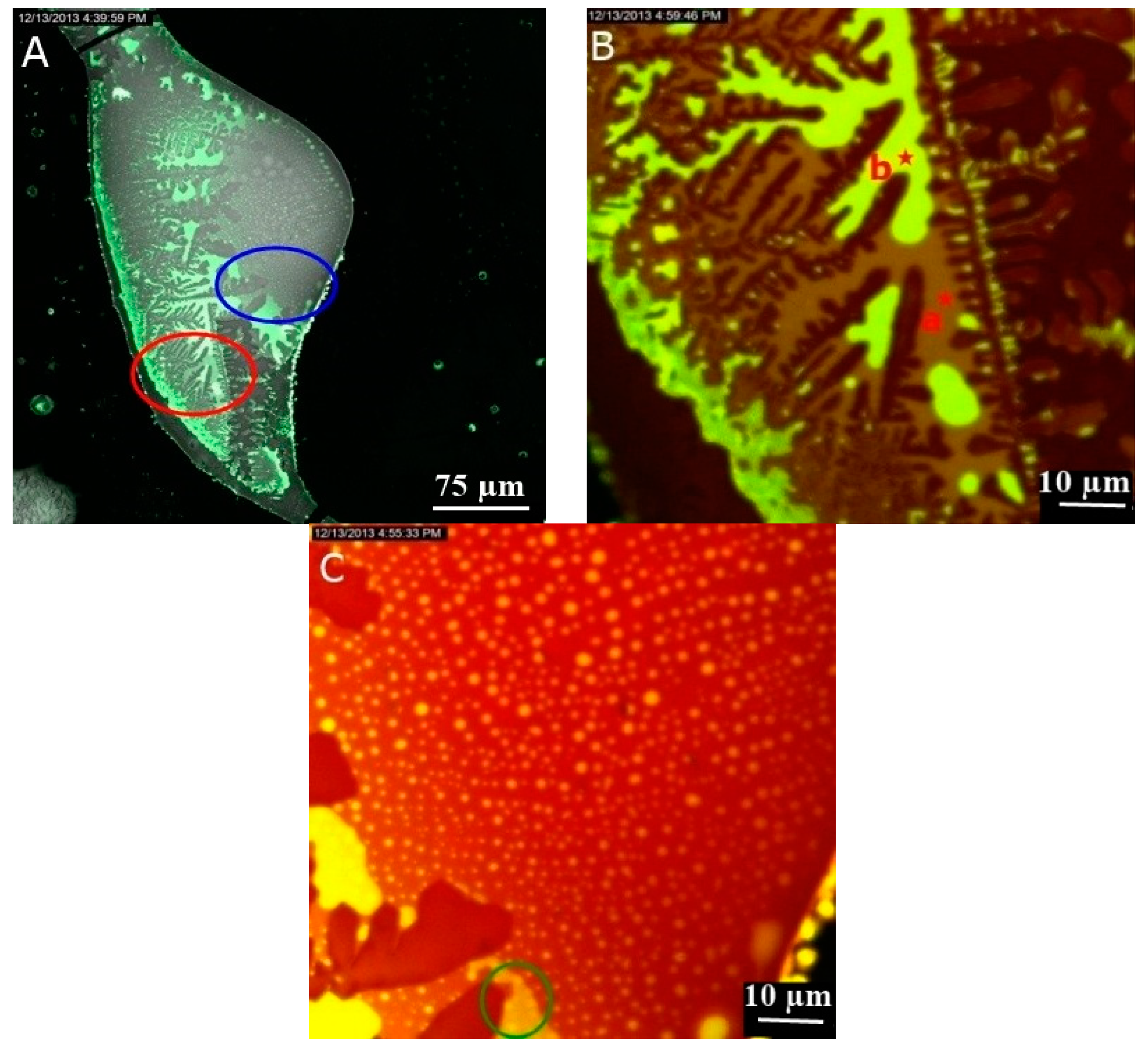
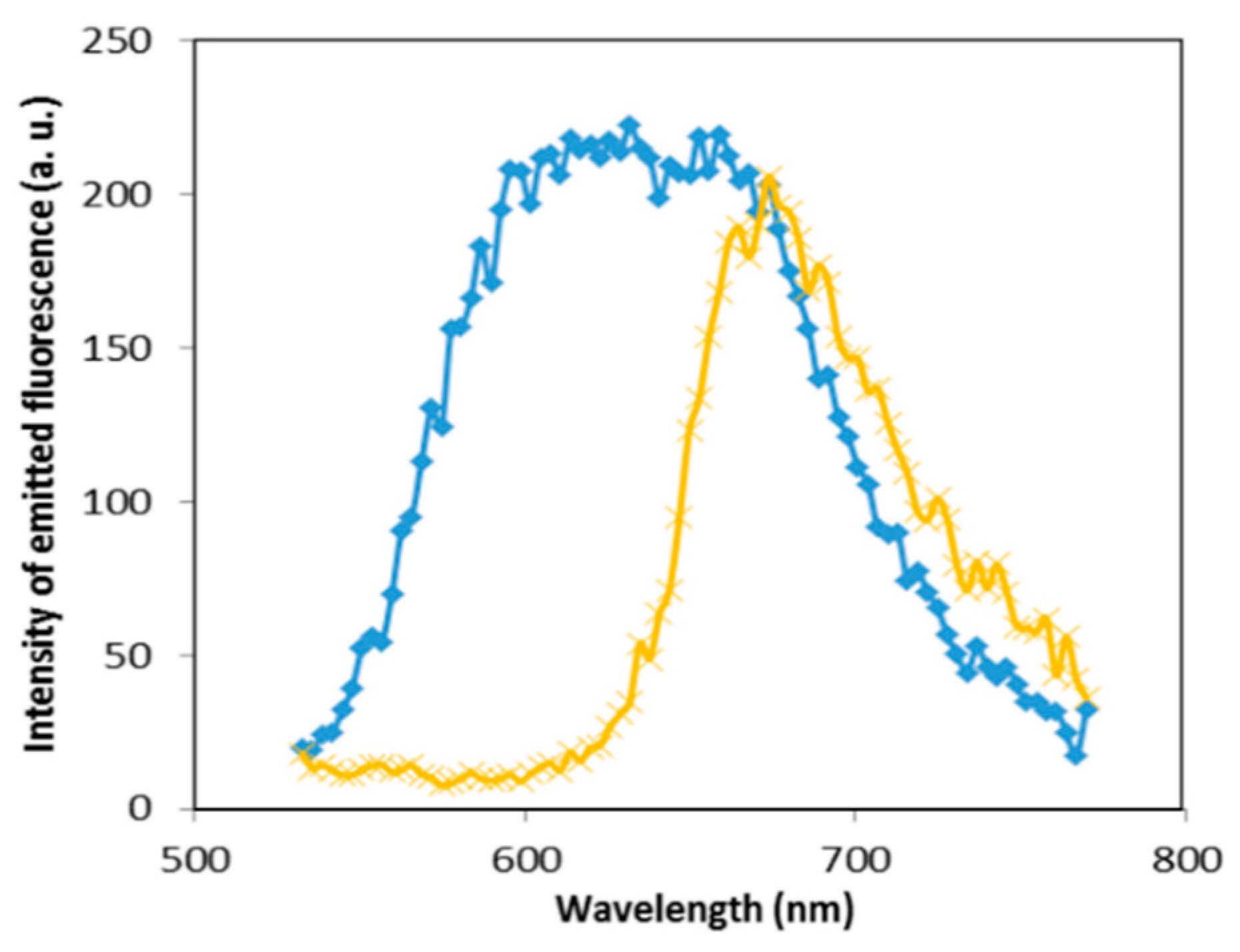
3.1. Images of LSCM and Fluorescence Spectroscopy
3.1.1. Effect of Gold Nanoparticles on the Fluorescence of NB Bound to Cationic Surfactants
3.1.2. Effect of DNA on GNPs Aggregation
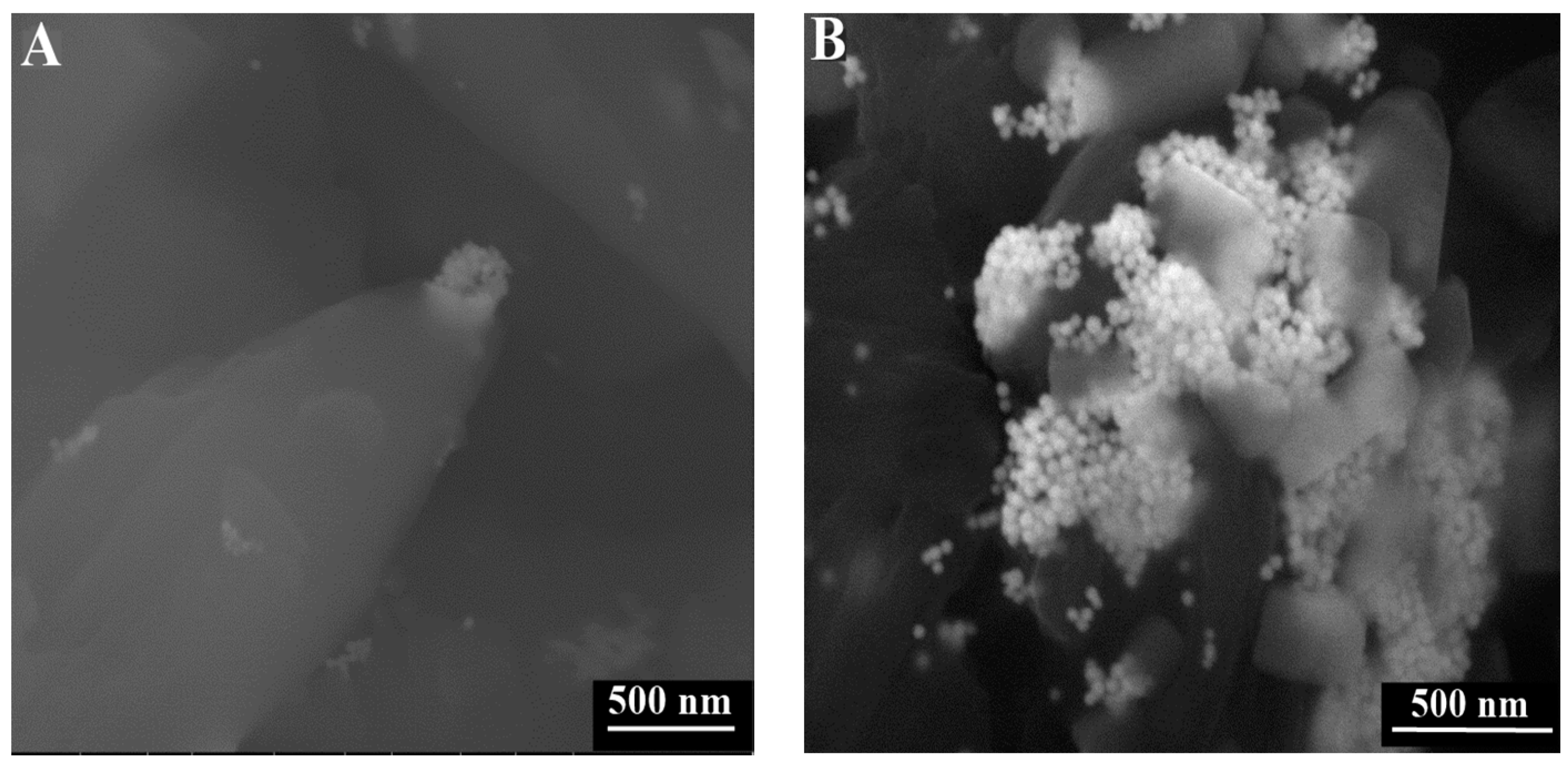
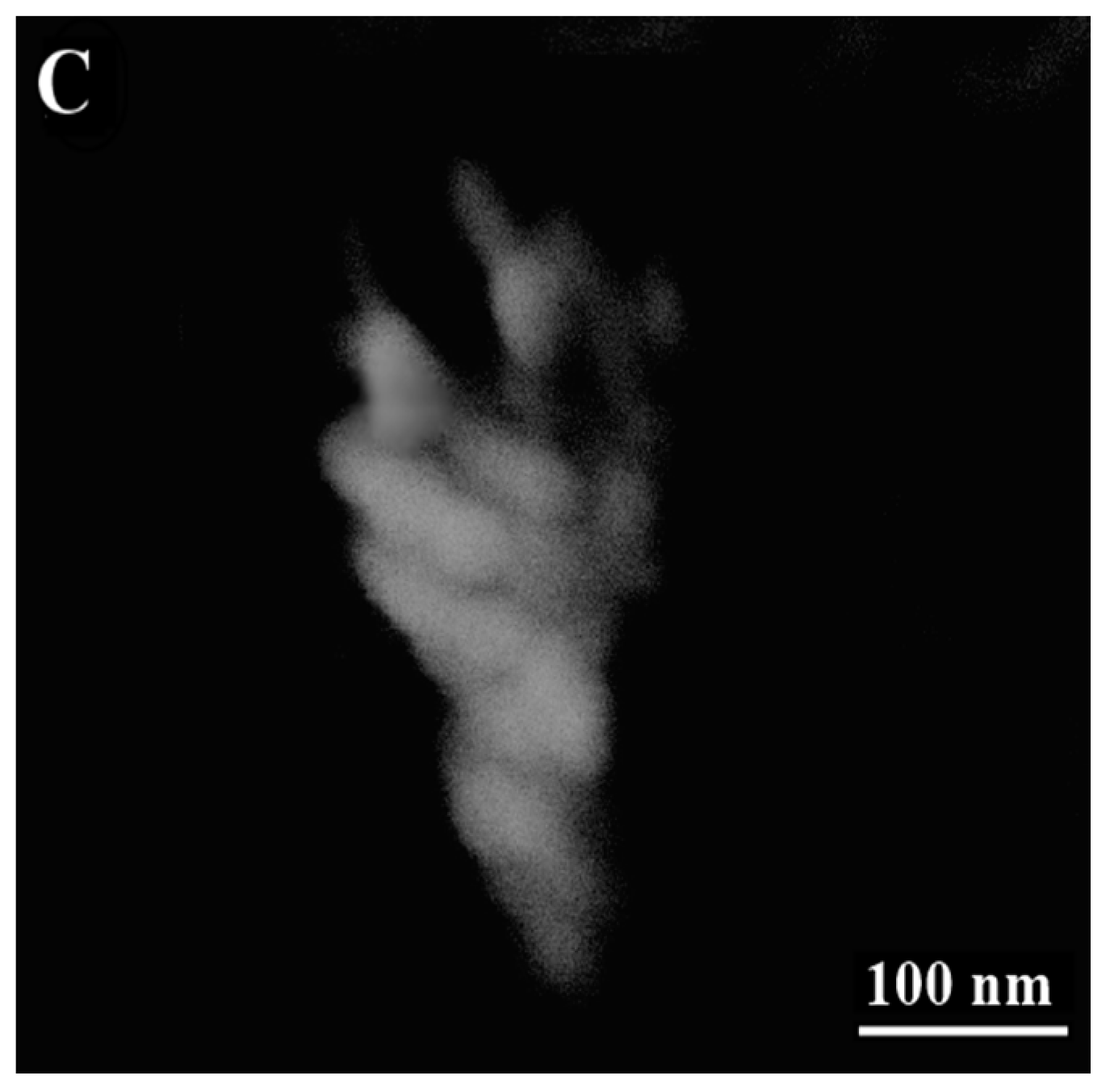
3.2. Images of ESEM
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Appendix

References
- Mirkin, C.A.; Letsinger, R.L.; Mucic, R.C.; Stohoff, J.J. A DNA-based method for rationally assembling nanoparticles into macroscopic materials. Nature 1996, 382, 607–609. [Google Scholar] [CrossRef] [PubMed]
- Rahban, M.; Divsalar, A.; Saboury, A.A.; Golestani, A. Nanotoxicity and Spectroscopy Studies of Silver Nanoparticle: Calf Thymus DNA and K562 as Targets. J. Phys. Chem. C 2010, 114, 5798–5803. [Google Scholar] [CrossRef]
- Basu, S.; Jana, S.; Pande, S.; Pal, T. Interaction of DNA bases with silver nanoparticles: Assembly quantified through SPRS and SERS. J. Colloid Interface Sci. 2008, 321, 288–293. [Google Scholar] [CrossRef] [PubMed]
- Niemeyer, M.; Mirkin, C.A. NanoBiotechnology: Concepts, Methods and Applications; Wiley-VCH: Weinheim, Germany, 2004. [Google Scholar]
- Storhoff, J.J.; Mirkin, C.A. Programmed materials synthesis with DNA. Chem. Rev. 1999, 99, 1849–1862. [Google Scholar] [CrossRef] [PubMed]
- Niemeyer, C.M. Nanopartikel, Proteine und Nucleinsäuren: Die Biotechnologie Begegnet den Materialwissenschaften. Angew. Chem. 2001, 113, 4254–4287. (In Germany) [Google Scholar] [CrossRef]
- Braun, E.; Eichen, Y.; Sivan, U.; Ben-Yoseph, G. DNA-templated assembly and electrode attachment of a conducting silver wire. Nature 1998, 391, 775–778. [Google Scholar] [CrossRef] [PubMed]
- Single Stranded DNA Induced Assembly of Gold Nanoparticles. Available online: http://dspace.mit.edu/handle/1721.1/3949 (accessed on 31 July 2015).
- Ganech, K.N.; Sastry, M. DNA-amine interactions: From monolayers to nanoparticles. J. Indian Inst. Sci. 2002, 82, 105–112. [Google Scholar]
- Nakao, H.; Shiigi, H.; Yamamoto, Y.; Tokonami, S.; Nagaoka, T.; Sugiyama, S.; Ohtani, T. Highly Ordered Assemblies of Au Nanoparticles Organized on DNA. Nano Lett. 2003, 3, 1391–1394. [Google Scholar] [CrossRef]
- Lee, J.S.; Han, M.S.; Mirkin, C.A. Colorimetric Detection of Mercuric Ion (Hg2+) in Aqueous Media Using DNA-Functionalized Gold Nanoparticles. Angew. Chem. 2007, 119, 4171–4174. [Google Scholar] [CrossRef]
- Kanayama, N.; Takarada, T.; Maeda, M. Rapid naked-eye detection of mercury ions based on non-crosslinking aggregation of double-stranded DNA-carrying gold nanoparticles. Chem. Commun. 2011, 47, 2077–2086. [Google Scholar] [CrossRef] [PubMed]
- De la Rica, R.; Stevens, M. Plasmonic ELISA for the ultrasensitive detection of disease biomarkers with the naked eye. Nat. Nanotechnol. 2012, 7, 821–824. [Google Scholar] [CrossRef] [PubMed]
- Tsai, S.; Chen, Y.; Liaw, J. Compound Cellular Imaging of Laser Scanning Confocal Microscopy by Using Gold Nanoparticles and Dyes. Sensors 2008, 8, 2306–2316. [Google Scholar] [CrossRef]
- Durr, N.J.; Larson, T.; Smith, D.K.; Korgel, B.A.; Sokolov, K.; Ben-Yarkar, A. Two-Photon Luminescence Imaging of Cancer Cells Using Molecularly Targeted Gold Nanorods. Nano Lett. 2007, 7, 941–945. [Google Scholar] [CrossRef] [PubMed]
- Aioub, M.; Kang, B.; Mackey, M.A.; El-Sayed, M.A. Biological Targeting of Plasmonic Nanoparticles Improves Cellular Imaging via the Enhanced Scattering in the Aggregates Formed. J. Phys. Chem. Lett. 2014, 5, 2555–2561. [Google Scholar] [CrossRef] [PubMed]
- Mitra, R.K.; Sinha, S.S.; Maiti, S.; Pal, S.K. Interactions of Nile Blue with Micelles, Reverse Micelles and a Genomic DNA. J. Fluoresc. 2008, 18, 423–432. [Google Scholar] [CrossRef] [PubMed]
- Mitra, R.K.; Sinha, S.S.; Maiti, S.; Pal, S.K. Sequence Dependent Ultrafast Electron Transfer of Nile Blue in Oligonucleotides. J. Fluoresc. 2009, 19, 353–361. [Google Scholar] [CrossRef] [PubMed]
- Jim, J.; Lee, N.; Le, E.; Yoon, S. Surface Modification of Citrate-Capped Gold Nanoparticles Using CTAB Micelles. Bull. Korean Chem. Soc. 2014, 35, 2567–2569. [Google Scholar]
- Jana, N.R.; Gearheart, L.; Murphy, C.J. Seeding Growth for Size Control of 5–40 nm Diameter Gold Nanoparticles. Langmuir 2001, 17, 6782–6786. [Google Scholar] [CrossRef]
- Oregon Medical Laser Center. Nile Blue. Available online: http://omlc.org/spectra/PhotochemCAD/html/068.html (accessed on 23 June 2015).
- Gao, Q.; Zheng, Y.; Song, C.; Lu, L.; Tian, X.; Xu, A. Selective and sensitive colorimetric detection of copper ions based on anti-aggregation of the glutathioneinduced aggregated gold nanoparticles and its application for determining sulfide anions. RSC Adv. 2013, 3, 21424–21430. [Google Scholar] [CrossRef]
- Kang, K.A.; Wang, J.; Jasinski, J.B.; Achilefu, S. Fluorescence Manipulation by Gold Nanoparticles: From Complete Quenching to Extensive Enhancement. J. Nanobiotechnol. 2011, 9, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Schneider, G.; Decher, G.; Nerambourg, N.; Praho, R.; Werts, M.H.V.; Blanchard-Desce, M. Distance-dependent fluorescence quenching on gold nanoparticles ensheathed with layer-by-layer assembled polyelectrolytes. Nano Lett. 2006, 6, 530–536. [Google Scholar] [CrossRef] [PubMed]
- My, N.; Wc, L. Fluorescence enhancements of fiber-optic biosensor with metallic nanoparticles. Opt. Expr. 2009, 17, 5867–5878. [Google Scholar]
- Kang, K.A.; Hong, B. Biocompatible nano-metal particle fluorescence enhancers. Crit. Rev. Eukaryot. Gene Expr. 2006, 16, 45–60. [Google Scholar] [CrossRef] [PubMed]
- Gary, R.; Amelio, D.; Garofalo, F.; Petriashvili, G.; de Santo, M.P.; Kwong Ip, Y.; Barberi, R. Endothelial-like nitric oxide synthase immunolocalization by using gold nanoparticles and dyes. Biomed. Opt. Expr. 2015, 6, 4738–4748. [Google Scholar] [CrossRef] [PubMed]
- Livolant, F.; Leforestier, A. Condensed phases of DNA: Structures and phase transitions. Prog. Polymer. Sci. 1996, 21, 1115–1164. [Google Scholar] [CrossRef]
- Saminathan, M.; Thomas, T.; Shirahata, A.; Pillai, C.K.S.; Thomas, T.J. Polyamine structural effects on the induction and stabilization of liquid crystalline DNA: Potential applications to DNA packaging, gene therapy and polyamine therapeutics. Nucleic Acids Res. 2002, 30, 3722–3731. [Google Scholar] [CrossRef] [PubMed]
- Thanh, N.T.K.; Vernhet, A.; Rosenzweig, Z. Gold Nanoparticles in Bioanalytical Assays and Sensors. Springer Ser. Chem. Sens. Biosens. 2005, 3, 261–277. [Google Scholar]
- Sun, L.; Sun, Y.; Zhang, F.X.; Yang, T.; Guo, C.; Liu, Z.; Li, Z. Atomic force microscopy and surface-enhanced Raman scattering detection of DNA based on DNA–nanoparticle complexes. Nanotechnology 2009, 20, 125502–125514. [Google Scholar] [CrossRef] [PubMed]


© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gary, R.; Carbone, G.; Petriashvili, G.; De Santo, M.P.; Barberi, R. Detection of Gold Nanoparticles Aggregation Growth Induced by Nucleic Acid through Laser Scanning Confocal Microscopy. Sensors 2016, 16, 258. https://doi.org/10.3390/s16020258
Gary R, Carbone G, Petriashvili G, De Santo MP, Barberi R. Detection of Gold Nanoparticles Aggregation Growth Induced by Nucleic Acid through Laser Scanning Confocal Microscopy. Sensors. 2016; 16(2):258. https://doi.org/10.3390/s16020258
Chicago/Turabian StyleGary, Ramla, Giovani Carbone, Gia Petriashvili, Maria Penelope De Santo, and Riccardo Barberi. 2016. "Detection of Gold Nanoparticles Aggregation Growth Induced by Nucleic Acid through Laser Scanning Confocal Microscopy" Sensors 16, no. 2: 258. https://doi.org/10.3390/s16020258
APA StyleGary, R., Carbone, G., Petriashvili, G., De Santo, M. P., & Barberi, R. (2016). Detection of Gold Nanoparticles Aggregation Growth Induced by Nucleic Acid through Laser Scanning Confocal Microscopy. Sensors, 16(2), 258. https://doi.org/10.3390/s16020258





