Electrochemical Affinity Biosensors Based on Disposable Screen-Printed Electrodes for Detection of Food Allergens
Abstract
:1. Introduction
2. Advantages of Electrochemical Biosensors Based on Screen-Printed Electrodes
3. Design of Sensitive Biosensors for Allergen Based on Screen-Printed Electrodes
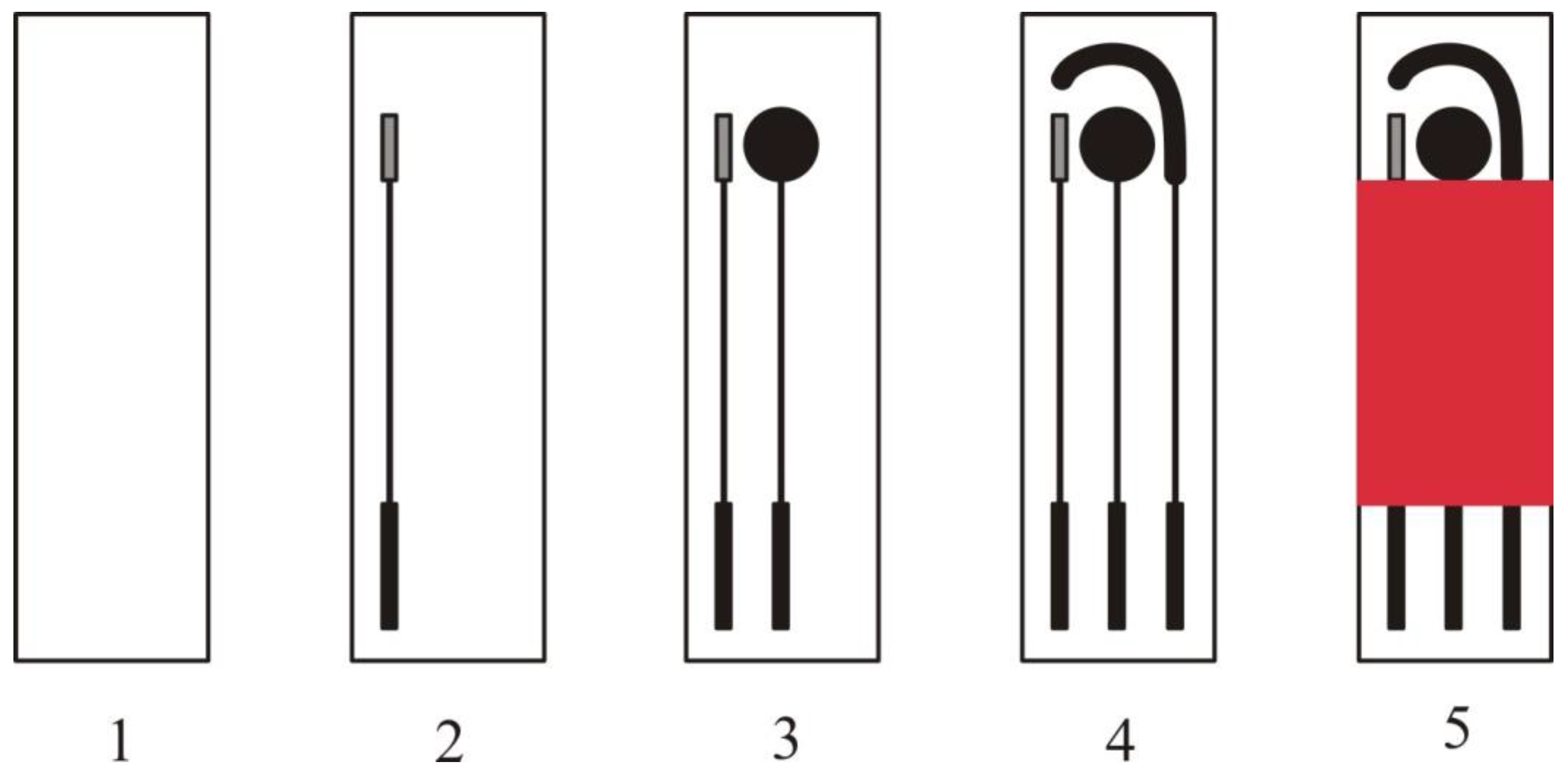
3.1. Bioreceptor Immobilisation and Detection Format
3.2. Aptasensors
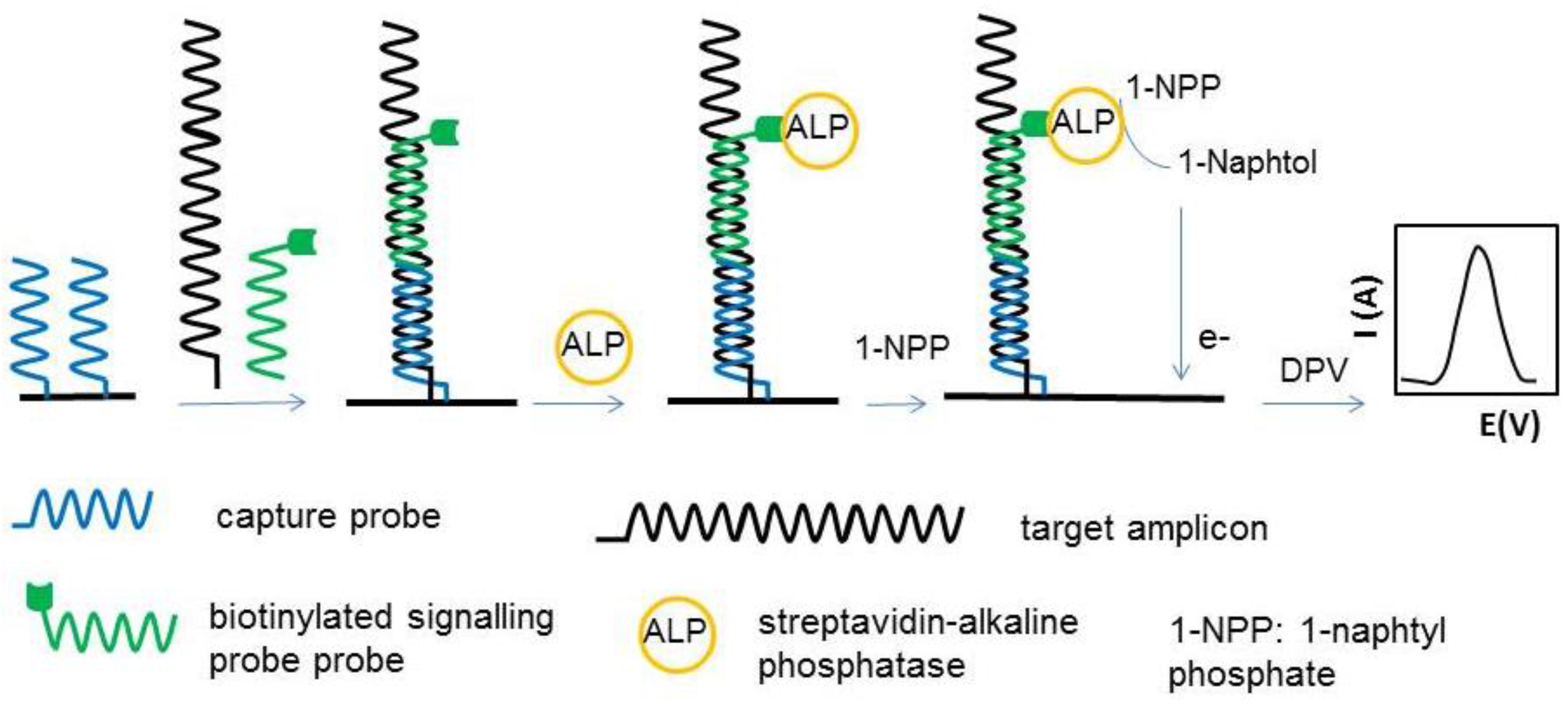
3.3. Genosensors
3.4. Sensors Based on MIPS
3.5. Assay Formats in Affinity Based Biosensors
3.5.1. Sandwich Format
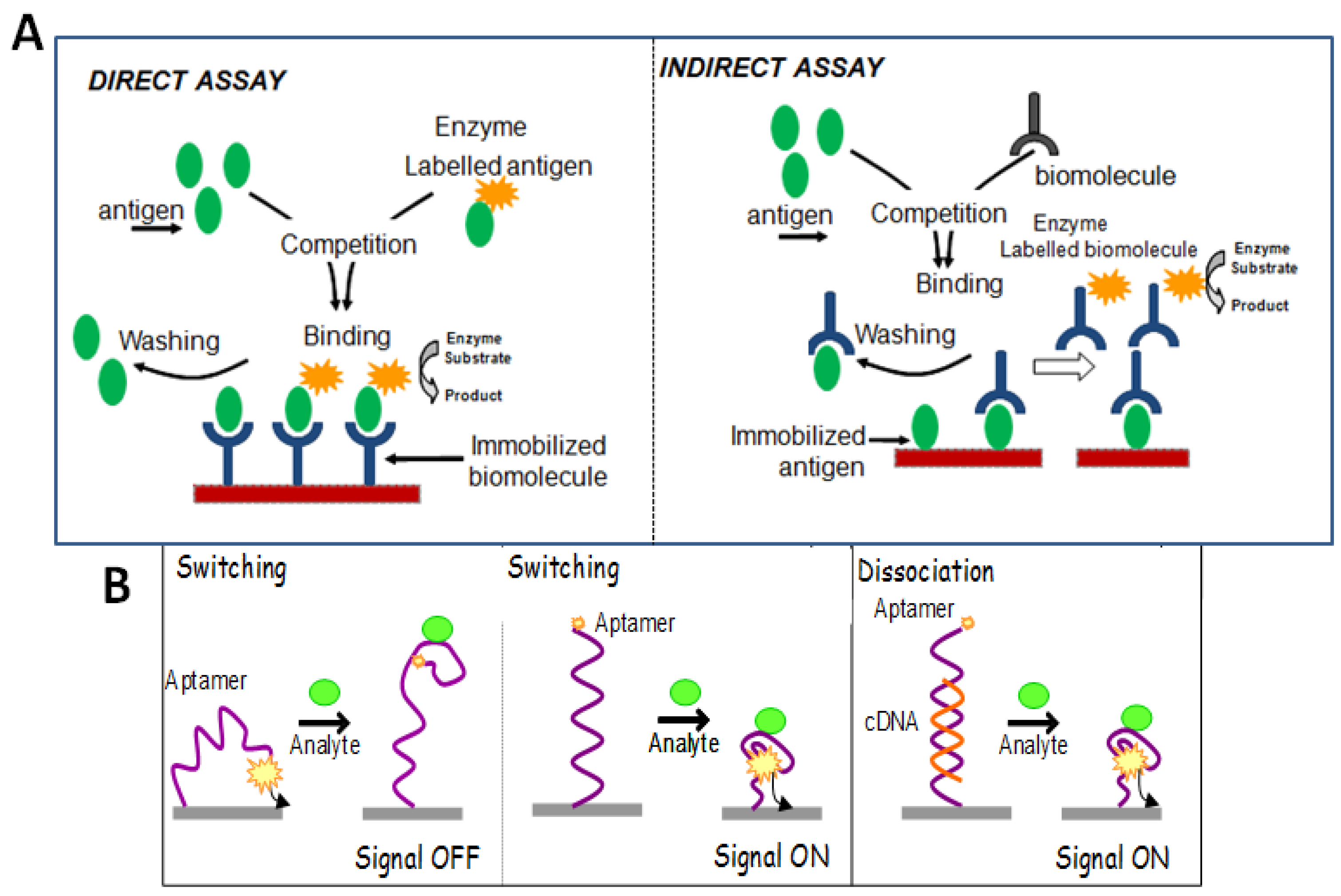
3.5.2. Competitive Binding Format
3.5.3. Non-Competitive Format
3.5.4. Additional Formats in Aptasensing Designs
Target-Induced Structure Switching Format (TISS)
Target-Induced Dissociation/Displacement Format (TID)
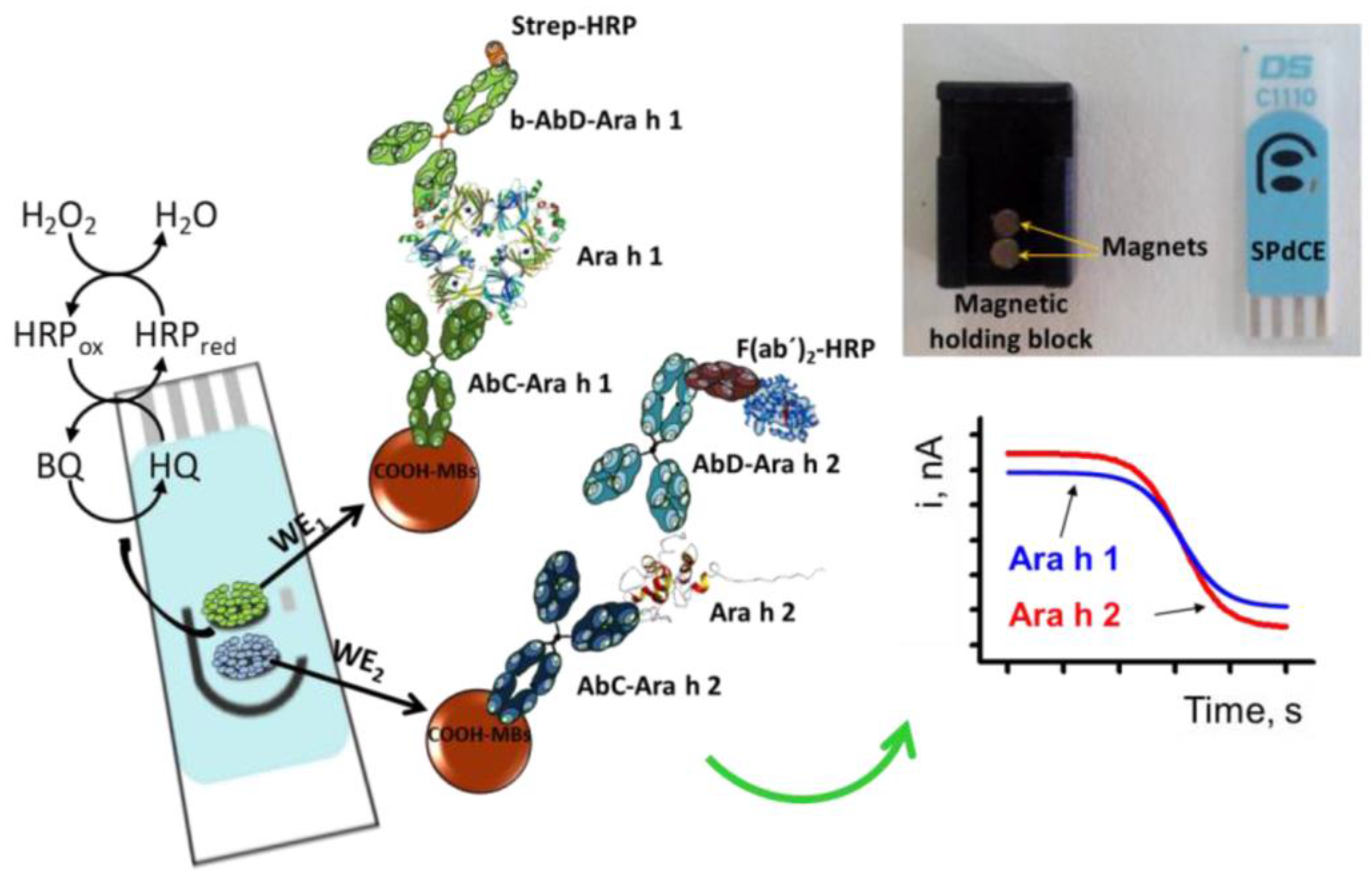
3.6. Assays Coupling Ligand-Modified Magnetic Beads with Electrochemical Detection with SPEs
3.7. Other Tests Coupling Affinity Recognition of Allergens with Electrochemical Detection on SPEs
3.8. Electrochemical Detection Method
4. Applications of Biosensors Based on Screen-Printed Electrodes for Allergen Detection
4.1. Examples of SPE-Based Biosensors for Allergen Detection
4.1.1. Biosensors for Milk Allergens
4.1.2. Biosensors for Egg Allergens
4.1.3. Biosensors for Gluten
4.1.4. Biosensors for Peanut Protein Allergens
4.1.5. Biosensors for Hazelnut Allergens
4.2. Figures of Merit of SPE-Based Biosensors for Allergens
4.3. Challenges Associated with Real Samples Analysis
4.4. Current Trends in Allergen Analysis with SPE-Based Biosensors
- (1)
- Use of nanomaterials such as Au NPs, graphene or carbon nanotubes to increase the effective area, improve conductivity and facilitate the quantitative immobilization of bioreceptors [98].
- (2)
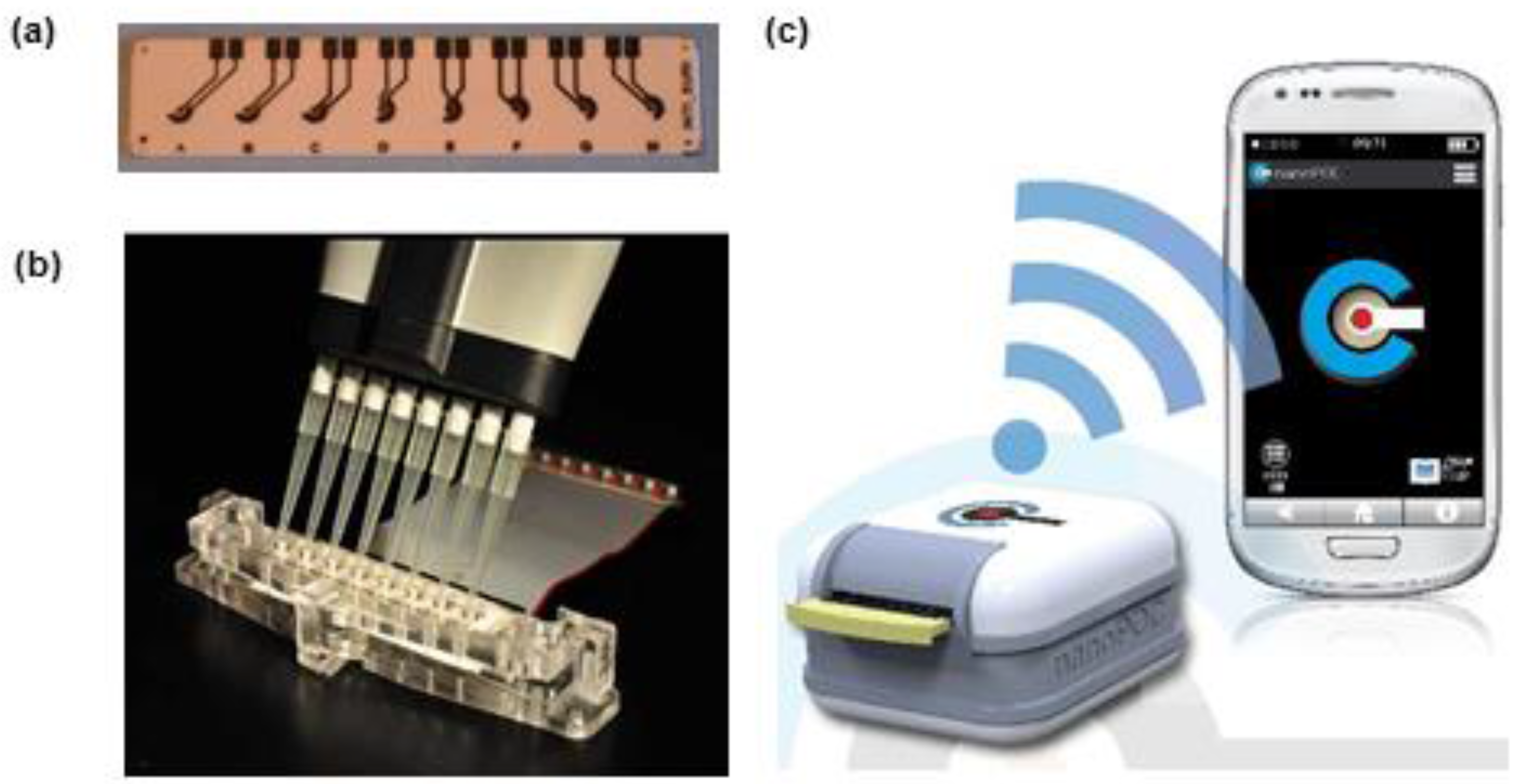
- Development of small portable biosensors controlled by smartphones
- (3)
- Multi-analyte sensors and arrays
5. Conclusions and Perspectives
Author Contributions
Conflicts of Interest
References
- Walker, M.J.; Burns, D.T.; Elliott, C.T.; Gowland, M.H.; Mills, E.N. Is food allergen analysis flawed? Health and supply chain risks and a proposed framework to address urgent analytical needs. Analyst 2016, 141, 24–35. [Google Scholar] [CrossRef]
- Peñas, E.D.L.C.; Uberti, F.; Restani, P. Allergenic Proteins in Enology: A Review on Technological Applications and Safety Aspects. Molecules 2015, 20, 13144–13164. [Google Scholar] [CrossRef]
- Akiyama, H.; Imai, T.; Ebisawa, M. Chapter 4—Japan Food Allergen Labeling Regulation—History and Evaluation. In Advances in Food and Nutrition Research; Steve, L.T., Ed.; Elsevier: Amsterdam, The Netherlands, 2011; Volume 62, pp. 139–171. [Google Scholar]
- European Commission—Health and Food Safety Rapid Alert System for Food and Feed Annual Report 2015. Available online: http://ec.europa.eu/food/safety/docs/rasff_annual_report_2015.pdf (accessed on 16 September 2016).
- Food Allergy Research and Education Allergy Alerts. Available online: https://www.foodallergy.org/alerts/alerts-feed (accessed on 16 September 2016).
- Canadian Food Inspection Agency Food Recall Warnings. Available online: http://www.inspection.gc.ca/about-the-cfia/newsroom/food-recall-warnings/eng/1299076382077/1299076493846 (accessed on 16 September 2016).
- Alves, R.C.; Barroso, M.F.; Gonzalez-Garcia, M.B.; Oliveira, M.B.; Delerue-Matos, M.C. New Trends in Food Allergens Detection: Towards Biosensing Strategies. Crit. Rev. Food Sci. Nutr. 2015, 56, 2304–2319. [Google Scholar] [CrossRef]
- Vo-Dinh, T.; Cullum, B. Biosensors and biochips: Advances in biological and medical diagnostics. Fresenius J. Anal. Chem. 2000, 366, 540–551. [Google Scholar] [CrossRef]
- Clark, L.C.; Lyons, C. Electrode systems for continuous monitoring in cardiovascular surgery. Ann. N. Y. Acad. Sci. 1962, 102, 29–45. [Google Scholar] [CrossRef]
- Campuzano, S.; Montiel, V.R.-V.; Torrente-Rodríguez, R.M.; Reviejo, Á.J.; Pingarrón, J.M. Electrochemical Biosensors for Food Security: Allergens and Adulterants Detection. In Biosensors for Security and Bioterrorism Applications; Nikolelis, P.D., Nikoleli, G.-P., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 287–307. [Google Scholar]
- Vasilescu, A.; Marty, J.-L. Electrochemical aptasensors for the assessment of food quality and safety. TrAC Trend Anal. Chem. 2016, 79, 60–70. [Google Scholar] [CrossRef]
- Trashin, S.; de Jong, M.; Breugelmans, T.; Pilehvar, S.; De Wael, K. Label-Free Impedance Aptasensor for Major Peanut Allergen Ara h 1. Electroanalysis 2015, 27, 32–37. [Google Scholar] [CrossRef]
- Rotariu, L.; Lagarde, F.; Jaffrezic-Renault, N.; Bala, C. Electrochemical biosensors for fast detection of food contaminants – trends and perspective. TrAC Trend Anal. Chem. 2016, 79, 80–87. [Google Scholar] [CrossRef]
- Tudorache, M.; Bala, C. Biosensors based on screen-printing technology, and their applications in environmental and food analysis. Anal. Bioanal. Chem. 2007, 388, 565–578. [Google Scholar] [CrossRef]
- Avramescu, A.; Andreescu, S.; Noguer, T.; Bala, C.; Andreescu, D.; Marty, J.-L. Biosensors designed for environmental and food quality control based on screen-printed graphite electrodes with different configurations. Anal. Bioanal. Chem. 2002, 374, 25–32. [Google Scholar] [CrossRef]
- Murray, R.W.; Ewing, A.G.; Durst, R.A. Chemically modified electrodes Molecular design for electroanalysis. Anal. Chem. 1987, 59, 379A–390A. [Google Scholar]
- Wang, J.; Lu, J.; Hocevar, S.B.; Farias, P.A.M.; Ogorevc, B. Bismuth-Coated Carbon Electrodes for Anodic Stripping Voltammetry. Anal. Chem. 2000, 72, 3218–3222. [Google Scholar] [CrossRef]
- Hayat, A.; Yang, C.; Rhouati, A.; Marty, J.L. Recent advances and achievements in nanomaterial-based, and structure switchable aptasensing platforms for ochratoxin A detection. Sensors 2013, 13, 15187–15208. [Google Scholar] [CrossRef]
- Hayat, A.; Andreescu, S.; Marty, J.-L. Design of PEG-aptamer two piece macromolecules as convenient and integrated sensing platform: Application to the label free detection of small size molecules. Biosens. Bioelectron. 2013, 45, 168–173. [Google Scholar] [CrossRef]
- Hayat, A.; Haider, W.; Rolland, M.; Marty, J.-L. Electrochemical grafting of long spacer arms of hexamethyldiamine on a screen printed carbon electrode surface: Application in target induced ochratoxin A electrochemical aptasensor. Analyst 2013, 138, 2951–2957. [Google Scholar] [CrossRef]
- Metters, J.P.; Kadara, R.O.; Banks, C.E. New directions in screen printed electroanalytical sensors: An overview of recent developments. Analyst 2011, 136, 1067–1076. [Google Scholar] [CrossRef]
- Wilson, R.; Turner, A. Glucose oxidase: An ideal enzyme. Biosens. Bioelectron. 1992, 7, 165–185. [Google Scholar] [CrossRef]
- Newman, J.D.; Turner, A.P. Home blood glucose biosensors: A commercial perspective. Biosens. Bioelectron. 2005, 20, 2435–2453. [Google Scholar] [CrossRef]
- Heller, A.; Feldman, B. Electrochemical glucose sensors and their applications in diabetes management. Chem. Rev. 2008, 108, 2482–2505. [Google Scholar] [CrossRef]
- Li, M.; Li, Y.-T.; Li, D.-W.; Long, Y.-T. Recent developments and applications of screen-printed electrodes in environmental assays—A review. Anal. Chim. Acta 2012, 734, 31–44. [Google Scholar] [CrossRef]
- Fanjul-Bolado, P.; Hernández-Santos, D.; Lamas-Ardisana, P.J.; Martín-Pernía, A.; Costa-García, A. Electrochemical characterization of screen-printed and conventional carbon paste electrodes. Electrochim. Acta 2008, 53, 3635–3642. [Google Scholar] [CrossRef]
- Kadara, R.O.; Jenkinson, N.; Banks, C.E. Screen printed recessed microelectrode arrays. Sens. Actuaturs B Chem. 2009, 142, 342–346. [Google Scholar] [CrossRef]
- Fan, Z.; Ho, J.C.; Takahashi, T.; Yerushalmi, R.; Takei, K.; Ford, A.C.; Chueh, Y.L.; Javey, A. Toward the development of printable nanowire electronics and sensors. Adv. Mater. 2009, 21, 3730–3743. [Google Scholar] [CrossRef]
- Khairy, M.; Kadara, R.O.; Banks, C.E. Electroanalytical sensing of nitrite at shallow recessed screen printed microelectrode arrays. Anal. Meth. 2010, 2, 851–854. [Google Scholar] [CrossRef]
- Šljukić, B.; Malakhova, N.A.; Brainina, K.Z.; Banks, C.E.; Compton, R.G. Screen Printed Electrodes and Screen Printed Modified Electrodes Benefit from Insonation. Electroanalysis 2006, 18, 928–930. [Google Scholar] [CrossRef]
- Choudhry, N.A.; Kampouris, D.K.; Kadara, R.O.; Jenkinson, N.; Banks, C.E. Next generation screen printed electrochemical platforms: Non-enzymatic sensing of carbohydrates using copper (II) oxide screen printed electrodes. Anal. Meth. 2009, 1, 183–187. [Google Scholar] [CrossRef]
- Hayat, A.; Barthelmebs, L.; Sassolas, A.; Marty, J.-L. Development of a novel label-free amperometric immunosensor for the detection of okadaic acid. Anal. Chim. Acta 2012, 724, 92–97. [Google Scholar] [CrossRef]
- Catanante, G.; Mishra, R.K.; Hayat, A.; Marty, J.-L. Sensitive analytical performance of folding based biosensor using methylene blue tagged aptamers. Talanta 2016, 153, 138–144. [Google Scholar] [CrossRef]
- Ocana, C.; Hayat, A.; Mishra, R.; Vasilescu, A.; del Valle, M.; Marty, J.L. A novel electrochemical aptamer-antibody sandwich assay for lysozyme detection. Analyst 2015, 140, 4148–4153. [Google Scholar] [CrossRef]
- Molinari, J.; Moina, C.; Ybarra, G. Electrochemical immunosensor for the determination of β-casein. J. Electrochem. Sci. Eng. 2015, 5, 9–16. [Google Scholar] [CrossRef]
- Saito, M.; Kitsunai, M.; Ahmed, M.U.; Sugiyama, S.; Tamiya, E. Label-free Electrochemical Detection for Food Allergen using Screen Printed Carbon Electrode. Electrochemistry 2008, 76, 606–609. [Google Scholar] [CrossRef]
- Hayat, A.; Barthelmebs, L.; Sassolas, A.; Marty, J.L. An electrochemical immunosensor based on covalent immobilization of okadaic acid onto screen printed carbon electrode via diazotization-coupling reaction. Talanta 2011, 85, 513–518. [Google Scholar] [CrossRef]
- Hayat, A.; Barthelmebs, L.; Marty, J.-L. Electrochemical impedimetric immunosensor for the detection of okadaic acid in mussel sample. Sens. Actuators B Chem. 2012, 171–172, 810–815. [Google Scholar] [CrossRef]
- Ocana, C.; Hayat, A.; Mishra, R.K.; Vasilescu, A.; Del Valle, M.; Marty, J.L. Label free aptasensor for Lysozyme detection: A comparison of the analytical performance of two aptamers. Bioelectrochemistry 2015, 105, 72–77. [Google Scholar] [CrossRef]
- Eissa, S.; Tlili, C.; L'Hocine, L.; Zourob, M. Electrochemical immunosensor for the milk allergen β-lactoglobulin based on electrografting of organic film on graphene modified screen-printed carbon electrodes. Biosens. Bioelectron. 2012, 38, 308–313. [Google Scholar] [CrossRef]
- Eissa, S.; L'Hocine, L.; Siaj, M.; Zourob, M. A graphene-based label-free voltammetric immunosensor for sensitive detection of the egg allergen ovalbumin. Analyst 2013, 138, 4378–4384. [Google Scholar] [CrossRef]
- Rohrbach, F.; Karadeniz, H.; Erdem, A.; Famulok, M.; Mayer, G. Label-free impedimetric aptasensor for lysozyme detection based on carbon nanotube-modified screen-printed electrodes. Anal. Biochem. 2012, 421, 454–459. [Google Scholar] [CrossRef]
- Xie, D.; Li, C.; Shangguan, L.; Qi, H.; Xue, D.; Gao, Q.; Zhang, C. Click chemistry-assisted self-assembly of DNA aptamer on gold nanoparticles-modified screen-printed carbon electrodes for label-free electrochemical aptasensor. Sens. Actuators B Chem. 2014, 192, 558–564. [Google Scholar] [CrossRef]
- Shi, K.; Shiu, K.K. Determination of uric acid at electrochemically activated glassy carbon electrode. Electroanalysis 2001, 13, 1319–1325. [Google Scholar] [CrossRef]
- Banks, C.E.; Crossley, A.; Salter, C.; Wilkins, S.J.; Compton, R.G. Carbon nanotubes contain metal impurities which are responsible for the “electrocatalysis” seen at some nanotube-modified electrodes. Angew. Chem. Int. Edit. 2006, 45, 2533–2537. [Google Scholar] [CrossRef]
- Rick, J.; Chou, T.-C. Using protein templates to direct the formation of thin-film polymer surfaces. Biosens. Bioelectron. 2006, 22, 544–549. [Google Scholar] [CrossRef]
- Poms, R.E.; Klein, C.L.; Anklam, E. Methods for allergen analysis in food: A review. Food Addit. Contam. 2004, 21, 1–31. [Google Scholar] [CrossRef]
- Lu, Y.; Ohshima, T.; Ushio, H. Rapid Detection of Fish Major Allergen Parvalbumin by Surface Plasmon Resonance Biosensor. J. Food Sci. 2004, 69, C652–C658. [Google Scholar] [CrossRef]
- Bremer, M.G.; Smits, N.G.; Haasnoot, W. Biosensor immunoassay for traces of hazelnut protein in olive oil. Anal. Bioanal. Chem. 2009, 395, 119–126. [Google Scholar] [CrossRef]
- Rebe Raz, S.; Liu, H.; Norde, W.; Bremer, M.G. Food allergens profiling with an imaging surface plasmon resonance-based biosensor. Anal. Chem. 2010, 82, 8485–8491. [Google Scholar] [CrossRef]
- Gobi, K.V.; Kim, S.J.; Tanaka, H.; Shoyama, Y.; Miura, N. Novel surface plasmon resonance (SPR) immunosensor based on monomolecular layer of physically-adsorbed ovalbumin conjugate for detection of 2,4-dichlorophenoxyacetic acid and atomic force microscopy study. Sens. Actuators B Chem. 2007, 123, 583–593. [Google Scholar] [CrossRef]
- Hayat, A.; Marty, J.L. Aptamer based electrochemical sensors for emerging environmental pollutants. Front. Chem. 2014, 2, 1–9. [Google Scholar] [CrossRef]
- Cox, J.C.; Ellington, A.D. Automated selection of anti-Protein aptamers. Bioorgan. Med. Chem. 2001, 9, 2525–2531. [Google Scholar] [CrossRef]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef]
- Potty, A.S.R.; Kourentzi, K.; Fang, H.; Schuck, P.; Willson, R.C. Biophysical characterization of DNA and RNA aptamer interactions with hen egg lysozyme. Int. J. Biol. Macromol. 2011, 48, 392–397. [Google Scholar] [CrossRef]
- Tran, D.T.; Janssen, K.P.; Pollet, J.; Lammertyn, E.; Anne, J.; Van Schepdael, A.; Lammertyn, J. Selection and characterization of DNA aptamers for egg white lysozyme. Molecules 2010, 15, 1127–1140. [Google Scholar] [CrossRef]
- Huang, H.; Jie, G.; Cui, R.; Zhu, J.-J. DNA aptamer-based detection of lysozyme by an electrochemiluminescence assay coupled to quantum dots. Electrochem. Comm. 2009, 11, 816–818. [Google Scholar] [CrossRef]
- Nadal, P.; Pinto, A.; Svobodova, M.; Canela, N.; O'Sullivan, C.K. DNA aptamers against the Lup an 1 food allergen. PLoS ONE 2012, 7, e35253. [Google Scholar] [CrossRef]
- Tran, D.T.; Knez, K.; Janssen, K.P.; Pollet, J.; Spasic, D.; Lammertyn, J. Selection of aptamers against Ara h 1 protein for FO-SPR biosensing of peanut allergens in food matrices. Biosens. Bioelectron. 2013, 43, 245–251. [Google Scholar] [CrossRef]
- Pinto, A.; Polo, P.N.; Henry, O.; Redondo, M.C.B.; Svobodova, M.; O’Sullivan, C.K. Label-free detection of gliadin food allergen mediated by real-time apta-PCR. Anal. Bioanal. Chem. 2014, 406, 515–524. [Google Scholar] [CrossRef]
- Amaya-González, S.; de-los-Santos-Álvarez, N.; Miranda-Ordieres, A.J.; Lobo-Castañón, M.J. Sensitive gluten determination in gluten-free foods by an electrochemical aptamer-based assay. Anal. Bioanal. Chem. 2015, 407, 6021–6029. [Google Scholar] [CrossRef]
- Pividori, M.I.; Merkoci, A.; Alegret, S. Electrochemical genosensor design: Immobilisation of oligonucleotides onto transducer surfaces and detection methods. Biosens. Bioelectron. 2000, 15, 291–303. [Google Scholar] [CrossRef]
- Cho, E.J.; Collett, J.R.; Szafranska, A.E.; Ellington, A.D. Optimization of aptamer microarray technology for multiple protein targets. Anal. Chim. Acta 2006, 564, 82–90. [Google Scholar] [CrossRef]
- Bettazzi, F.; Lucarelli, F.; Palchetti, I.; Berti, F.; Marrazza, G.; Mascini, M. Disposable electrochemical DNA-array for PCR amplified detection of hazelnut allergens in foodstuffs. Anal. Chim. Acta 2008, 614, 93–102. [Google Scholar] [CrossRef]
- López, M.S.-P.; Cabanillas, G.F.; Castañón, M.J.L.; López-Ruiz, B. Development of a genosensor for peanut allergen ARA h 2 detection and its optimization by surface response methodology. Biosens. Bioelectron. 2014, 62, 350–356. [Google Scholar] [CrossRef]
- Martín-Fernández, B.; Miranda-Ordieres, A.J.; Lobo-Castañón, M.J.; Frutos-Cabanillas, G.; de-los-Santos-Álvarez, N.; López-Ruiz, B. Strongly structured DNA sequences as targets for genosensing: Sensing phase design and coupling to PCR amplification for a highly specific 33-mer gliadin DNA fragment. Biosens. Bioelectron. 2014, 60, 244–251. [Google Scholar] [CrossRef]
- Whitcombe, M.J.; Chianella, I.; Larcombe, L.; Piletsky, S.A.; Noble, J.; Porter, R.; Horgan, A. The rational development of molecularly imprinted polymer-based sensors for protein detection. Chem. Soc. Rev. 2011, 40, 1547–1571. [Google Scholar] [CrossRef]
- Duan, H.; Wang, X.; Wang, Y.; Sun, Y.; Li, J.; Luo, C. An ultrasensitive lysozyme chemiluminescence biosensor based on surface molecular imprinting using ionic liquid modified magnetic graphene oxide/beta-cyclodextrin as supporting material. Anal. Chim. Acta 2016, 918, 89–96. [Google Scholar] [CrossRef]
- Turner, N.W.; Liu, X.; Piletsky, S.A.; Hlady, V.; Britt, D.W. Recognition of Conformational Changes in β-Lactoglobulin by Molecularly Imprinted Thin Films. Biomacromolecules 2007, 8, 2781–2787. [Google Scholar] [CrossRef]
- Su, W.-X.; Rick, J.; Chou, T.-C. Selective recognition of ovalbumin using a molecularly imprinted polymer. Microchem. J. 2009, 92, 123–128. [Google Scholar] [CrossRef]
- Barnes, A.; Belbruno, J.J. Food Allergen Detection Methods and Systems Using Molecularly Imprinted Polymers. Patent PCT/US2014/048676, 30 April 2015. [Google Scholar]
- Martín-Fernández, B.; Manzanares-Palenzuela, C.L.; López, M.S.-P.; de-los-SantosÁlvarez, N.; López-Ruiz, B. Electrochemical Genosensors in Food Safety Assessment. Crit. Rev. Food Sci. 2015. [Google Scholar] [CrossRef]
- Miranda-Castro, R.; de-los-Santos-Álvarez, N.; Lobo-Castañón, M.J.; Miranda-Ordieres, A.J.; Tuñón-Blanco, P. Structured Nucleic Acid Probes for Electrochemical Devices. Electroanalysis 2009, 21, 2077–2090. [Google Scholar] [CrossRef]
- Del Giallo, M.L.; Lucarelli, F.; Cosulich, E.; Pistarino, E.; Santamaria, B.; Marrazza, G.; Mascini, M. Steric Factors Controlling the Surface Hybridization of PCR Amplified Sequences. Anal. Chem. 2005, 77, 6324–6330. [Google Scholar] [CrossRef]
- Hayat, A.; Barthelmebs, L.; Marty, J.-L. Enzyme-linked immunosensor based on super paramagnetic nanobeads for easy and rapid detection of okadaic acid. Anal. Chem. Acta 2011, 690, 248–252. [Google Scholar] [CrossRef]
- Huang, Y.; Bell, M.C.; Suni, I.I. Impedance biosensor for peanut protein Ara h 1. Anal. Chem. 2008, 80, 9157–9161. [Google Scholar] [CrossRef]
- Xia, Y.; Gan, S.; Xu, Q.; Qiu, X.; Gao, P.; Huang, S. A three-way junction aptasensor for lysozyme detection. Biosens. Bioelectron. 2013, 39, 250–254. [Google Scholar] [CrossRef]
- Xia, J.; Song, D.; Wang, Z.; Zhang, F.; Yang, M.; Gui, R.; Xia, L.; Bi, S.; Xia, Y.; Li, Y.; et al. Single electrode biosensor for simultaneous determination of interferon gamma and lysozyme. Biosens. Bioelectron. 2015, 68, 55–61. [Google Scholar] [CrossRef]
- Peng, Y.; Zhang, D.; Li, Y.; Qi, H.; Gao, Q.; Zhang, C. Label-free and sensitive faradic impedance aptasensor for the determination of lysozyme based on target-induced aptamer displacement. Biosens. Bioelectron. 2009, 25, 94–99. [Google Scholar] [CrossRef]
- Putzbach, W.; Ronkainen, N.J. Immobilization techniques in the fabrication of nanomaterial-based electrochemical biosensors: A review. Sensors 2013, 13, 4811–4840. [Google Scholar] [CrossRef]
- Sassolas, A.; Hayat, A.; Marty, J.-L. Immobilization of enzymes on magnetic beads through affinity interactions. Immobil. Enzymes Cells Third Ed. 2013, 139–148. [Google Scholar]
- Čadková, M.; Metelka, R.; Holubová, L.; Horák, D.; Dvořáková, V.; Bílková, Z.; Korecká, L. Magnetic beads-based electrochemical immunosensor for monitoring allergenic food proteins. Anal. Biochem. 2015, 484, 4–8. [Google Scholar] [CrossRef]
- Rhouati, A.; Hayat, A.; Hernandez, D.B.; Meraihi, Z.; Munoz, R.; Marty, J.-L. Development of an automated flow-based electrochemical aptasensor for on-line detection of ochratoxin A. Sens. Actuators B Chem. 2013, 176, 1160–1166. [Google Scholar] [CrossRef]
- Dominguez, R.B.; Hayat, A.; Sassolas, A.; Alonso, G.A.; Munoz, R.; Marty, J.-L. Automated flow-through amperometric immunosensor for highly sensitive and on-line detection of okadaic acid in mussel sample. Talanta 2012, 99, 232–237. [Google Scholar] [CrossRef]
- Herrasti, Z.; Martínez, F.; Baldrich, E. Electrochemical detection of dopamine using streptavidin-coated magnetic particles and carbon nanotube wiring. Sens. Actuators B Chem. 2014, 203, 891–898. [Google Scholar] [CrossRef]
- Barthelmebs, L.; Hayat, A.; Limiadi, A.W.; Marty, J.-L.; Noguer, T. Electrochemical DNA aptamer-based biosensor for OTA detection, using superparamagnetic nanoparticles. Sens. Actuators B Chem. 2011, 156, 932–937. [Google Scholar] [CrossRef]
- Paleček, E.; Fojta, M. Magnetic beads as versatile tools for electrochemical DNA and protein biosensing. Talanta 2007, 74, 276–290. [Google Scholar] [CrossRef]
- Herrasti, Z.; de la Serna, E.; Ruiz-Vega, G.; Baldrich, E. Developing enhanced magnetoimmunosensors based on low-cost screen-printed electrode devices. Rev. Anal. Chem. 2016, 35, 53–85. [Google Scholar] [CrossRef]
- Ruiz-Valdepeñas Montiel, V.; Campuzano, S.; Conzuelo, F.; Torrente-Rodríguez, R.M.; Gamella, M.; Reviejo, A.J.; Pingarrón, J.M. Electrochemical magnetoimmunosensing platform for determination of the milk allergen β-lactoglobulin. Talanta 2015, 131, 156–162. [Google Scholar] [CrossRef]
- Ruiz-Valdepeñas Montiel, V.; Torrente-Rodríguez, R.M.; Campuzano, S.; Pellicanò, A.; Reviejo, Á.J.; Cosio, M.S.; Pingarrón, J.M. Simultaneous Determination of the Main Peanut Allergens in Foods Using Disposable Amperometric Magnetic Beads-Based Immunosensing Platforms. Chemosensors 2016, 4, 11. [Google Scholar] [CrossRef]
- Kokkinos, C.; Angelopoulou, M.; Economou, A.; Prodromidis, M.; Florou, A.; Haasnoot, W.; Petrou, P.; Kakabakos, S. Lab-on-a-Membrane Foldable Devices for Duplex Drop-Volume Electrochemical Biosensing Using Quantum Dot Tags. Anal. Chem. 2016, 88, 6897–6904. [Google Scholar] [CrossRef]
- Hayat, A.; Marty, J.L. Disposable screen printed electrochemical sensors: Tools for environmental monitoring. Sensors 2014, 14, 10432–10453. [Google Scholar] [CrossRef]
- Liu, H.; Malhotra, R.; Peczuh, M.W.; Rusling, J.F. Electrochemical Immunosensors for Antibodies to Peanut Allergen Ara h2 Using Gold Nanoparticle−Peptide Films. Anal. Chem. 2010, 82, 5865–5871. [Google Scholar] [CrossRef]
- Alves, R.C.; Pimentel, F.B.; Nouws, H.P.A.; Marques, R.C.B.; González-García, M.B.; Oliveira, M.B.P.P.; Delerue-Matos, C. Detection of Ara h 1 (a major peanut allergen) in food using an electrochemical gold nanoparticle-coated screen-printed immunosensor. Biosens. Bioelectron. 2015, 64, 19–24. [Google Scholar] [CrossRef]
- Hansen, J.A.; Wang, J.; Kawde, A.-N.; Xiang, Y.; Gothelf, K.V.; Collins, G. Quantum-Dot/Aptamer-Based Ultrasensitive Multi-Analyte Electrochemical Biosensor. J. Am. Chem. Soc. 2006, 128, 2228–2229. [Google Scholar] [CrossRef]
- Montiel, V.R.-V.; Campuzano, S.; Pellicanò, A.; Torrente-Rodríguez, R.M.; Reviejo, A.J.; Cosio, M.S.; Pingarrón, J.M. Sensitive and selective magnetoimmunosensing platform for determination of the food allergen Ara h 1. Anal. Chim. Acta 2015, 880, 52–59. [Google Scholar] [CrossRef]
- Martín-Fernández, B.; de-los-Santos-Álvarez, N.; Lobo-Castañón, M.J.; López-Ruiz, B. Hairpin-based DNA electrochemical sensor for selective detection of a repetitive and structured target codifying a gliadin fragment. Anal. Bioanal. Chem. 2015, 407, 3481–3488. [Google Scholar] [CrossRef]
- Pilolli, R.; Monaci, L.; Visconti, A. Advances in biosensor development based on integrating nanotechnology and applied to food-allergen management. TrAC Trend Anal. Chem. 2013, 47, 12–26. [Google Scholar] [CrossRef]
- Vasilescu, A.; Wang, Q.; Li, M.; Boukherroub, R.; Szunerits, S. Aptamer-Based Electrochemical Sensing of Lysozyme. Chemosensors 2016, 4, 10. [Google Scholar] [CrossRef]
- Alves, R.C.; Pimentel, F.B.; Nouws, H.P.A.; Correr, W.; González-García, M.B.; Oliveira, M.B.P.P.; Delerue-Matos, C. Detection of the peanut allergen Ara h 6 in foodstuffs using a voltammetric biosensing approach. Anal. Bioanal. Chem. 2015, 407, 7157–7163. [Google Scholar] [CrossRef]
- Restani, P.; Uberti, F.; Danzi, R.; Ballabio, C.; Pavanello, F.; Tarantino, C. Absence of allergenic residues in experimental and commercial wines fined with caseinates. Food Chem. 2012, 134, 1438–1445. [Google Scholar] [CrossRef]
- Abeyrathne, E.D.N.S.; Lee, H.Y.; Ahn, D.U. Egg white proteins and their potential use in food processing or as nutraceutical and pharmaceutical agents—A review. Poultry Sci. 2013, 92, 3292–3299. [Google Scholar] [CrossRef]
- Wang, Y.; Duan, H.; Li, L.; Wang, X.; Li, J.; Bu, Y.; Luo, C. A chemiluminescence sensor for determination of lysozyme using magnetic graphene oxide multi-walled carbon nanotube surface molecularly imprinted polymers. RSC Adv. 2016, 6, 12391–12397. [Google Scholar] [CrossRef]
- Miranda-Castro, R.; de-los-Santos-Álvarez, N.; Miranda-Ordieres, A.; Lobo-Castañón, M. Harnessing Aptamers to Overcome Challenges in Gluten Detection. Biosensors 2016, 6, 16. [Google Scholar] [CrossRef]
- WHO/IUIS Allergen Nomenclature Sub-Committee Allergen Nomenclature. Available online: http://www.allergen.org/search.php?allergenname=&allergensource=Arachis+hypogaea&TaxSource=&TaxOrder=&foodallerg=1&bioname= (accessed on 14 September 2016).
- Zhuang, Y.; Dreskin, S.C. Redefining the major peanut allergens. Immunol. Res. 2013, 55, 125–134. [Google Scholar] [CrossRef]
- Zhou, Y.; Wang, J.-S.; Yang, X.-J.; Lin, D.-H.; Gao, Y.-F.; Su, Y.-J.; Yang, S.; Zhang, Y.-J.; Zheng, J.-J. Peanut Allergy, Allergen Composition, and Methods of Reducing Allergenicity: A Review. Int. J. Food Sci. 2013, 2013, 8. [Google Scholar] [CrossRef]
- International Nut&Dried Fruit, Global Statistical Review 2014–2015. Available online: https://www.nutfruit.org/wp-continguts/uploads/2015/11/global-statistical-review-2014-2015_101779.pdf (accessed on 12 September 2016).
- International Organisation of Wine and Vine-OIV, Resolution OIV-COMEX 502-2012. Available online: http://www.oiv.int/public/medias/2154/oiv-comex-502-2012-en.pdf (accessed on 20 September 2016).
- Scherf, K.A.; Poms, R.E. Recent developments in analytical methods for tracing gluten. J. Cereal Sci. 2016, 67, 112–122. [Google Scholar] [CrossRef]
- Martín-Fernández, B.; de-los-Santos-Álvarez, N.; Martín-Clemente, J.P.; Lobo-Castañón, M.J.; López-Ruiz, B. Challenging genosensors in food samples: The case of gluten determination in highly processed samples. Talanta 2016, 146, 490–495. [Google Scholar] [CrossRef]
- van Eckert, R.; Berghofer, E.; Ciclitira, P.J.; Chirdo, F.; Denery-Papini, S.; Ellis, H.J.; Ferranti, P.; Goodwin, P.; Immer, U.; Mamone, G.; et al. Towards a new gliadin reference material–isolation and characterisation. J. Cereal Sci. 2006, 43, 331–341. [Google Scholar] [CrossRef]
- Amaya-González, S.; de-los-Santos-Álvarez, N.; Miranda-Ordieres, A.J.; Lobo-Castañón, M.J. Aptamer Binding to Celiac Disease-Triggering Hydrophobic Proteins: A Sensitive Gluten Detection Approach. Anal. Chem. 2014, 86, 2733–2739. [Google Scholar] [CrossRef]
- Bazin, I.; Tria, S.A.; Hayat, A.; Marty, J.-L. New biorecognition molecules in biosensors for the detection of toxins. Biosens. Bioelectron. 2017, 87, 285–298. [Google Scholar] [CrossRef]
- Wang, P.; Sun, X.; Su, X.; Wang, T. Advancements of molecularly imprinted polymers in the food safety field. Analyst 2016, 141, 3540–3553. [Google Scholar] [CrossRef]
- Crasson, O.; Rhazi, N.; Jacquin, O.; Freichels, A.; Jérôme, C.; Ruth, N.; Galleni, M.; Filée, P.; Vandevenne, M. Enzymatic functionalization of a nanobody using protein insertion technology. Protein Eng. Des. Sel. 2015, 28, 451–460. [Google Scholar] [CrossRef]
- Cortez-Retamozo, V.; Lauwereys, M.; Hassanzadeh Gh, G.; Gobert, M.; Conrath, K.; Muyldermans, S.; De Baetselier, P.; Revets, H. Efficient tumor targeting by single-domain antibody fragments of camels. Int. J. Cancer 2002, 98, 456–462. [Google Scholar] [CrossRef]

| Allergen/Real Matrix | Biorecognition Details | Detection Method and Electrode Details | Analytical Characteristics | Reference |
|---|---|---|---|---|
| β-casein | Direct detection; Protein A/Ab | SPCE pretreated EC; direct oxidation of casein by DPV | Detection of 1, 10 and 100 µg·mL−1 | [36] |
| β-casein | Competitive; β-casein/Ab/HRP-labeled anti-IgG Ab | SPCE-plasma activated; amperometric reduction of BQ at −0.28 V; | LR: 0–10 ppm | [35] |
| Bovine casein and bovine gamma globulin G (bIgG) in milk | Direct; QD-labeled Ab on screen-printed foldable membrane | Bi-SPCE with bismuth citrate; ASV of Pb and Cd from QD s | DL: 40 ng·mL−1 (casein) | [91] |
| DL 20 ng mL−1 (bIgG) | ||||
| β-lactoglobulin/cake, cheese snacks, sweet biscuits | Direct; Ab/β-lactoglobulin | GR-SPCE; DPV of [Fe(CN)6]3−/4− | LR: 1 pg·mL−1 to 100 ng·mL−1 | [40] |
| DL:0.85 pg·mL−1 | ||||
| β-lactoglobulin/milk | Sandwich; magneto immunoassay; Ab-MBs/β-lactoglobulin/ HRP-Ab | SPCE; amperometry, reduction of BQ at −0.2 V | LR: 2.8–100 ng·mL−1 | [89] |
| DL: 0.8 ng·mL−1 | ||||
| Ovalbumin | Sandwich; magneto immunoassay; Ab-MBs/ovalbumin/HRP-Ab | SPPtE; LSV of H2O2 using thionine as mediator | LR:11–222 nM | [82] |
| DL: 5 nM | ||||
| Ovalbumin/spiked cake extract | Direct; Ab/ovalbumin | GR-SPCE, DPV of Fe(CN)6]3−/4−; | LR: 1 pg·mL−1–0.5 μg·mL−1; | [41] |
| DL: 0.83 pg·mL−1 | ||||
| Genes encoding hazelnut allergens Cor a 1.04 and Cor a 1.03)/hazelnut, chocolate, soymilk, biscuits, lecithin supplements, ketchup, snack, breakfast cereal, peanut butter | Sandwich; tCP/PCR products/bSP/SA-ALP | SPAuE;DPV of α-naphthol | LR: up to 20 nM [Cor a 1.04, Cor a 1.03] | [64] |
| DL: 0.3 nM (Cor a 1.03) 0.1 nM (Cor a 1.04) | ||||
| Gene encoding Ara h 2 peanut protein | Sandwich; tCP/target DNA + bSP/SA-ALP | SPAuE; DPV of 1-naphtophenol | LR: 50 pM–50 nM | [65] |
| DL: 10 pM; | ||||
| RSD: 7.22% (n = 8) | ||||
| S: 3 µA/nM | ||||
| Ara h 1/food extracts (hazelnuts; peanuts, chocolate bars; chocolate chip cookies; peanut creams; peanut oil) and saliva | Sandwich; cAb-MB/Ara h 1/bAb/SA-HRP | SPCE, amperometry, HQ detection at −0.2 V | LR: 20.8–1000 ng·mL−1 | [96] |
| DL: 6.3 ng·mL−1 | ||||
| Ara h 1/cookies and chocolate | Sandwich; cAb/Ara h 1/b-Ab/SA-ALP | AuNPs-modified SPCE; LSW for stripping enzymatically deposited Ag | LR: 12.6–2000 ng mL−1 | [94] |
| DL: 3.8 ng·mL−1 | ||||
| dual detection of Ara h 1 and Ara h 2/ wheat flour, hazelnuts; peanuts; chocolate bars with roasted peanuts and peanut cream; | Sandwich; Magneto-immunoassay; cAb-MBs/Ara h 1/b-Ab/SA-HRP-polymer (Ara h 1); cAb-MBs/Ara h 2/Ab/F(ab’)2-HRP (Ara h 2) | Dual SPCE; amperometry, HQ detection at −0.2 V | Ara h 1: | [90] |
| LR: 60–1000 ng·mL−1 | ||||
| DL: 18 ng·mL−1 | ||||
| Ara h 2: | ||||
| LR: 0.25–5 ng·mL−1 | ||||
| DL: 0.07 ng·mL−1; | ||||
| Ara h 6/chocolate and cookies | Sandwich; cAb/Ara h 1/b-Ab/SA-ALP | AuNPs-modified SPCE; LSW of enzymatically deposited Ag | LR: 1–100 ng·mL−1 | [100] |
| DL: 0.27 ng·mL−1; | ||||
| RSD ≤ 9.8% (n = 6) | ||||
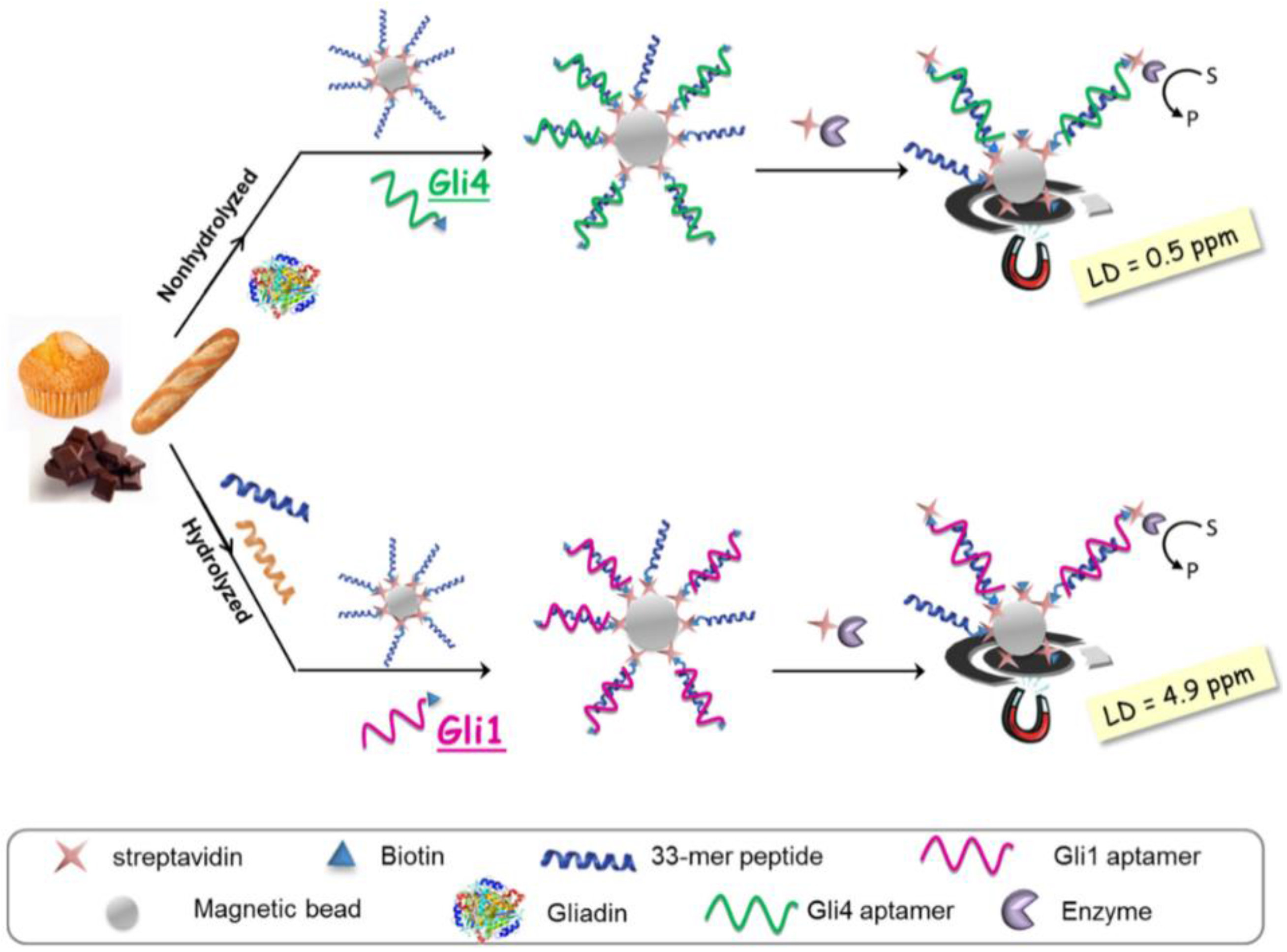
| Gluten/gluten-free foods (including heat-treated and hydrolysed) | Competitive; 33-mer peptide-modified MBs/b-APT+gliadin/SA-HRP. APT: Gli 1 or Gli 4 | SPCE; chrono-amperometry at 0 V of TMBox | DL: 4.9 ng·mL−1 (Gli 1); | [61] |
| DL: 0.5 ng·mL−1 (Gli 4) | ||||
| Sequence encoding the 33-mer peptide in wheat/wheat flour | Sandwich; tCP/target DNA + FITC-SP/anti-FITC-Fab–HRP | SPAuE; chronoamperometry at −0.2 V of TMBox for 60 s | LR: up to 50 nM; | [66] |
| DL: 0.3 nM | ||||
| DNA encoding gliadin fragment | Sandwich; tCP/target DNA + b-SP/SA-HRP | SPAuE; chrono- amperometry at 0 V of TMBox | LR: 5–50 nM | [97] |
| DL: 1 nM | ||||
| RSD (n = 3) = 21% (10 nM) and 4.3% (5 nM). | ||||
| Lysozyme/wine | Direct; APT Cox/lysozyme and APT Tran/lysozyme | SPCE/EIS | Apt Cox: | [39] |
| LR: 0.1–0.8 μM | ||||
| DL: 100 nM | ||||
| Apt Tran: | ||||
| LR: 0.025–0.8 μM | ||||
| DL:25 nM | ||||
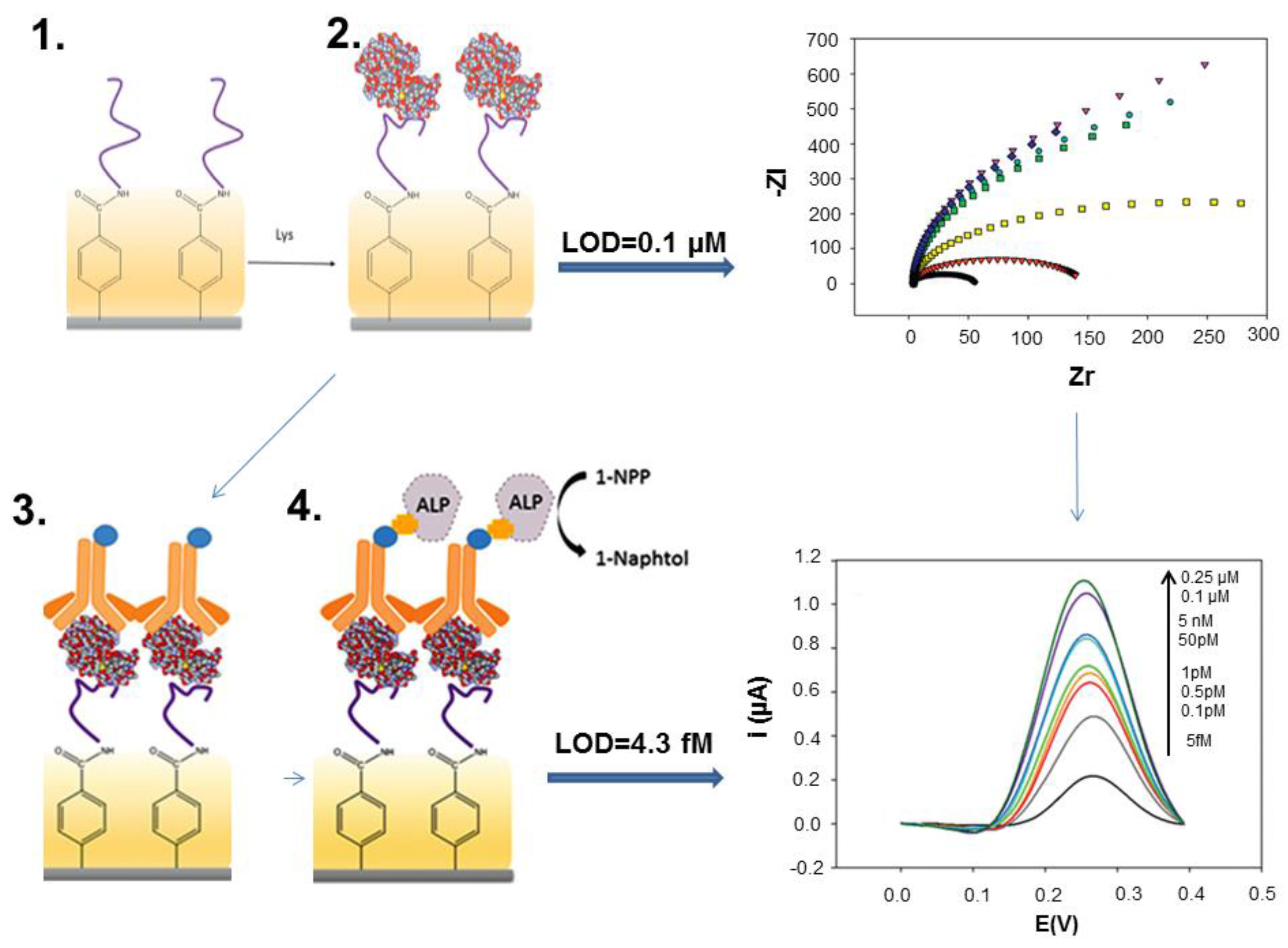
| Lysozyme/wine | Sandwich; APT/lysozyme/ALP-Ab | SPCE; DPV, 1-naphtol | LR: 5 fM– 5 nM | [34] |
| DL: 4.3 fM | ||||
| Lysozyme/egg white | Direct; APT/lysozyme | AuNP-SPCE; SWV, [Ru(NH3)6]3+ | LR:1–50.0 pg·mL−1 | [43] |
| DL: 0.3 pg·mL−1 | ||||
| Lysozyme | Direct; APT/lysozyme | MWCNT–SPCE, EIS [Fe(CN)6]3−/4− | DL: 12.09 ng mL−1 (862 nM) | [42] |
| Lysozyme | Direct; MIP/lysozyme | SPPtE, CV | 1 µg·mL−1 leads to 30.3% reduction in current, compared to 4.5% for control | [46] |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vasilescu, A.; Nunes, G.; Hayat, A.; Latif, U.; Marty, J.-L. Electrochemical Affinity Biosensors Based on Disposable Screen-Printed Electrodes for Detection of Food Allergens. Sensors 2016, 16, 1863. https://doi.org/10.3390/s16111863
Vasilescu A, Nunes G, Hayat A, Latif U, Marty J-L. Electrochemical Affinity Biosensors Based on Disposable Screen-Printed Electrodes for Detection of Food Allergens. Sensors. 2016; 16(11):1863. https://doi.org/10.3390/s16111863
Chicago/Turabian StyleVasilescu, Alina, Gilvanda Nunes, Akhtar Hayat, Usman Latif, and Jean-Louis Marty. 2016. "Electrochemical Affinity Biosensors Based on Disposable Screen-Printed Electrodes for Detection of Food Allergens" Sensors 16, no. 11: 1863. https://doi.org/10.3390/s16111863
APA StyleVasilescu, A., Nunes, G., Hayat, A., Latif, U., & Marty, J.-L. (2016). Electrochemical Affinity Biosensors Based on Disposable Screen-Printed Electrodes for Detection of Food Allergens. Sensors, 16(11), 1863. https://doi.org/10.3390/s16111863








