Southern Italy’s native farm animals are generally characterized by tolerance to diseases, tolerance to fluctuations in food and water availability, and adaptation to poor management conditions. Unfortunately, many of them have experienced a population decline and several are at risk of extinction. Being aware that the success of conservation depends on understanding the reasons for a breed to be conserved, our aim has always been to concentrate upon the characterization of these gene pools and to assess, if possible, their uniqueness. Understanding the relationship of breeds to various agricultural settings is key to appreciating and fostering performance and survival [
1].
Moreover, concerns regarding animal welfare need to be taken into account when anticipating the probable characteristics of future agricultural systems and potential animal breeding strategies [
2]. It is generally acknowledged that more attention should be paid to the physiological measures of welfare, which include health, tolerance to environmental conditions such as pedoclimatic aspects and enzootic parasites. According to Petazzi
et al. [
3], the choice of genetically adapted breeds is a prerequisite for a basic condition of well-being in whatever environment.
Regarding domestic animal species, research has progressively turned to focusing on molecular polymorphisms, which seem more suitable for selection and breeding strategies. Annotation of protein mutations with their relevance for phenotypic expression has thus been long neglected; but now, an apparently inverse trend seems to be taking place. Recently, growing attention has been focused on functional genetics, an area that is recurrently included amongst animal breeding research priorities. The functional effect of the Hb phenotype on hematological patterns has been demonstrated in humans as well as in mammalian species. Particularly in sheep, individuals carrying extra alpha-globin genes exhibit an overall blood picture mimicking a thalassemia-like syndrome [
12], while positively charged variants have been found to be somehow related to a decreased mean corpuscular volume and hematocrit value [
13].
Based on the above consideration, this work reports some of the results obtained regarding Hb polymorphism in ruminant Mediterranean native breeds obtained over the course of 20 years. The findings have been reviewed in the light of the functional effects of Hb and the common charged variants emphasized.
1.1 Highlights of Ruminant Hemoglobin Molecular Genetics
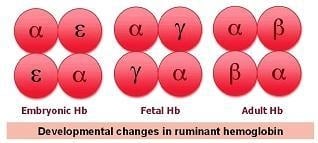
Adult Hb is composed of four subunits, two α-globin subunits and two β-globin subunits, and the interaction between these subunits dictates many oxygen binding characteristics of the protein. Unlike adults, however, embryos and fetuses cannot breathe for themselves, and thus have very different oxygen transport requirements. To accommodate this, the Hb of placental mammals has a high affinity for oxygen before birth, which is helpful for loading oxygen into the blood. After birth and the development of the cardiorespiratory system, Hb oxygen affinity decreases. This change in Hb function occurs because there are different forms of the Hb subunits, which arose from duplications of ancestral genes, and their expression changes throughout development.
According to Goodman
et al. [
20], the amino acid sequences of the α-globins and β-globins are approximately 50% identical, regardless of which vertebrate species is the source, arguing that these two genes are descended from a common ancestor about 450 million years ago, in the ancestral jawed vertebrate. The genes encoding the globins are clustered, with the α-like globin gene cluster on a different chromosome from the β-like globin gene cluster in birds and mammals.
It is generally acknowledged that the α-globin system in mammals is arranged in tandemly duplicated alpha globin genes, which are concurrently expressed, while the genes in the beta globin locus are arranged sequentially from 5’ to 3’: beginning with the gene expressed in embryonic development (called episolon) and ending with the adult beta globin gene. In ruminants as well as in human and in other mammalian species, the α-globin cluster is located on chromosome 25 while the β-globin cluster is positioned on chromosome 15 [
21].
The α-globin gene clusters consist of two a-globin genes (
HBA1 and
HBA2), which are expressed at different levels: the upstream gene being the most efficient. This trend is confirmed in sheep, where extra numeral arrangements, such as α-globin gene triplication and quadruplication, are not unusual, and the α-chain output of the downstream genes progressively decreases [
22,
23].
Also the β-globin system of these small ruminants is worth mentioning. As a result of an unusual evolutionary history, the ovine β-globin gene cluster is differently arranged depending on the A or B haplotypes. In the A sheep as well as in goats, the β-globin locus consists of 12 genes, organized as a triplicated, developmentally expressed four-gene set. Sheep with the B haplotype have a locus arrangement consisting of a duplicated four gene set as the consequence of a recent deletion from a triplicated locus [
24]. Similarly to goats, at birth the A sheep synthesize a juvenile hemoglobin C (HbC), which is produced at birth and exclusively during severe anaemia in adults. The B sheep do not synthesize HbC and continue to produce their adult Hb during anaemia. This is because B sheep lack the beta C gene as well as three other genes present both in A sheep and goats.
1.2 Cattle
Hemoglobin (Hb) was the first cattle protein polymorphism to be found with the alleles
HBBA and
HBBB [
25,
26,
27]. Eight other
HBB alleles have been reported; almost all of them are rare. These alleles are:
HBBC [
28,
29],
HBBD [
30],
HBBDZambia [
31], (
HBBG [
32],
HBBE [
33],
HBBI [
34,
35],
HBBH [
36], and
HBBXBALI [
37]) (
Table 2). At the end of the last century, two new alpha and one beta globin variants were discovered in Italian Podolic cows, a Southern European grey breed that has been autochthonous to Southern Italy since the Early Middle Ages after the migration of Asian populations from the Far East to Italy. Of the new alpha globins, one was allelic to the wild-type gene
HBA1 and the second to the wild type
HBA2. The wild type genes encode for the same alpha globin, which has never been named. Thus, based on structure analyses and in agreement with what had been suggested in the guidelines for gene nomenclature in ruminants [
38], we suggest placing the superscript on the right with the capital letter indicating the point mutation distinguishing the alleles to describe the as-yet unnamed cattle alpha genes. One of the two alpha globin variants presents a single amino acid substitution from asparagine to serine at position 131. In this position, most ruminant species exhibit an asparagine residue (N) [
39], thus the wild-type has been identified with the acronym
HBAN and the related variant as
HBAS; accordingly, the other chain, where histidine is substituted by tyrosine at position 89 [
15] has been named
HBAY. An interesting polymorphism at the
HBB locus was also detected in Italian Podolic cattle [
16]. Structural analyses performed to provide the complete sequence of the beta globin chains and the results pointed out that the taurine
HBBA globin allele identified in Hereford cattle [
27] was co-present with the variant named
HBBAZebu, found in Ongole and Banteng cattle [
37] and characterized by the silent mutation 43Ser→Thr.
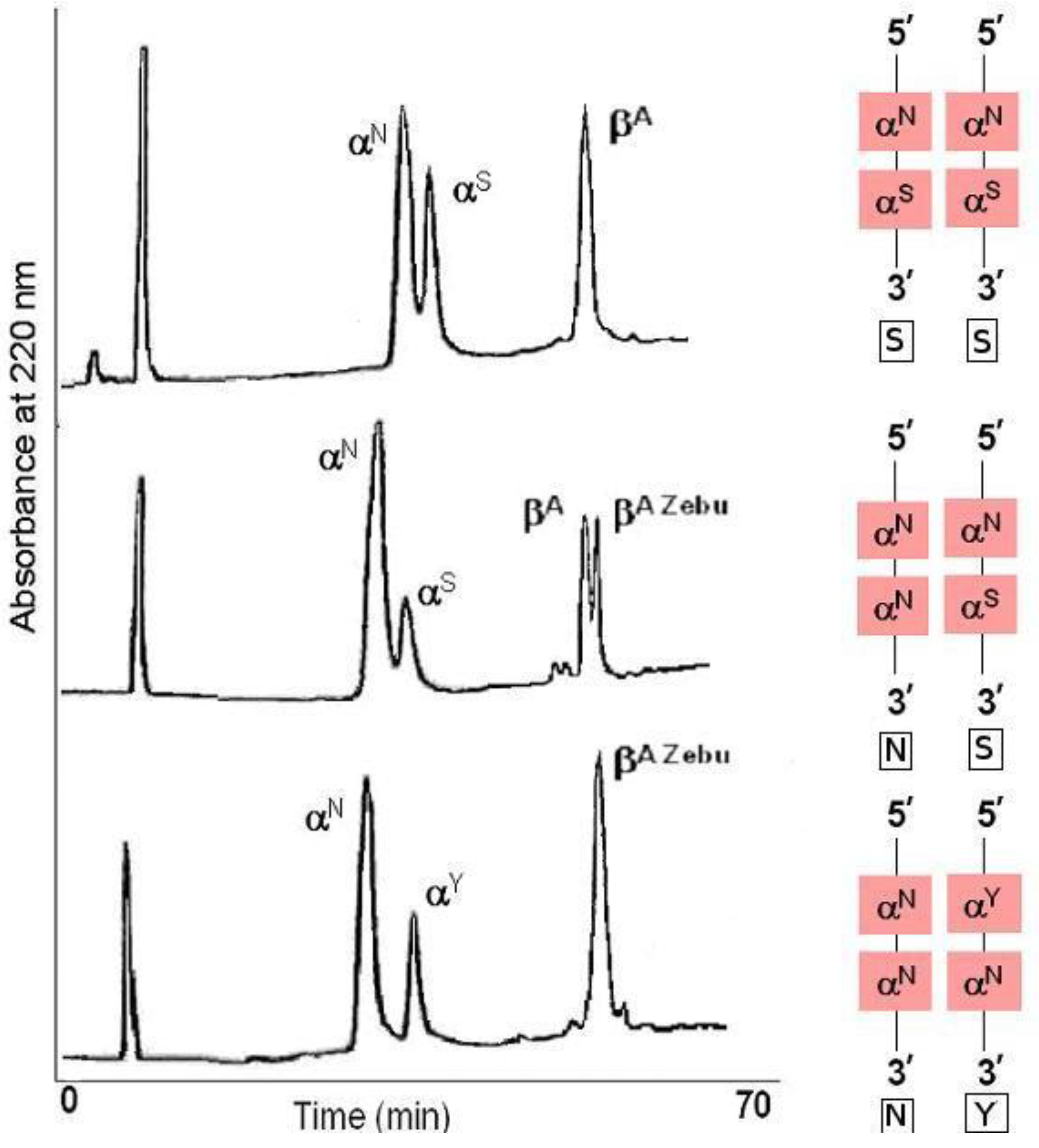
Both the
HBAS and
HBBAZebu encode for silent variants, which do not vary Hb electrophoretical mobility, while the hemoglobins containing the chain encoded by the
HBAY are positively charged and may be electrophoretically detected (
Figure 1).
Table 1 and
Table 2 summarize the alpha and beta globin polymorphism in cattle known so far and the related nomenclature.
Table 1.
Nomenclature of alpha globin genes and products in cattle.
Table 1.
Nomenclature of alpha globin genes and products in cattle.
| | HBA1 Locus | HBA2 Locus |
| Gene Symbol | HBA1N | HBA1Y | HBA2N | HBA2S |
| Globin symbol | IαN | IαY | IIαN | IIαS |
| Hemoglobin name | unnamed | HbY | unnamed | HbS |
Table 2.
Nomenclature of beta globin genes and products in cattle.
Table 2.
Nomenclature of beta globin genes and products in cattle.
| Gene Symbol | Globin chain responsible for the Hb name | Hb name | Reference |
|---|
| HBBA | βA | HbA | [25,27] |
| HBBAZebu | βAZebu | HbAZebu | [16,37] |
| HBBB | βB | HbB | [26] |
| HBBC | βC | HbC | [28,29] |
| HBBD | βD | HbD | [30] |
| HBBDZambia | βD Zambia | HbDZambia | [31] |
| HBBG | βG | HbG | [32] |
| HBBE | βE | HbE | [33] |
| HBBI | βI | HbI | [34,35] |
| HBBH | βH | HbH | [36] |
| HBBXBALI | βX Bali | HbXBali | [37] |
Figure 1.
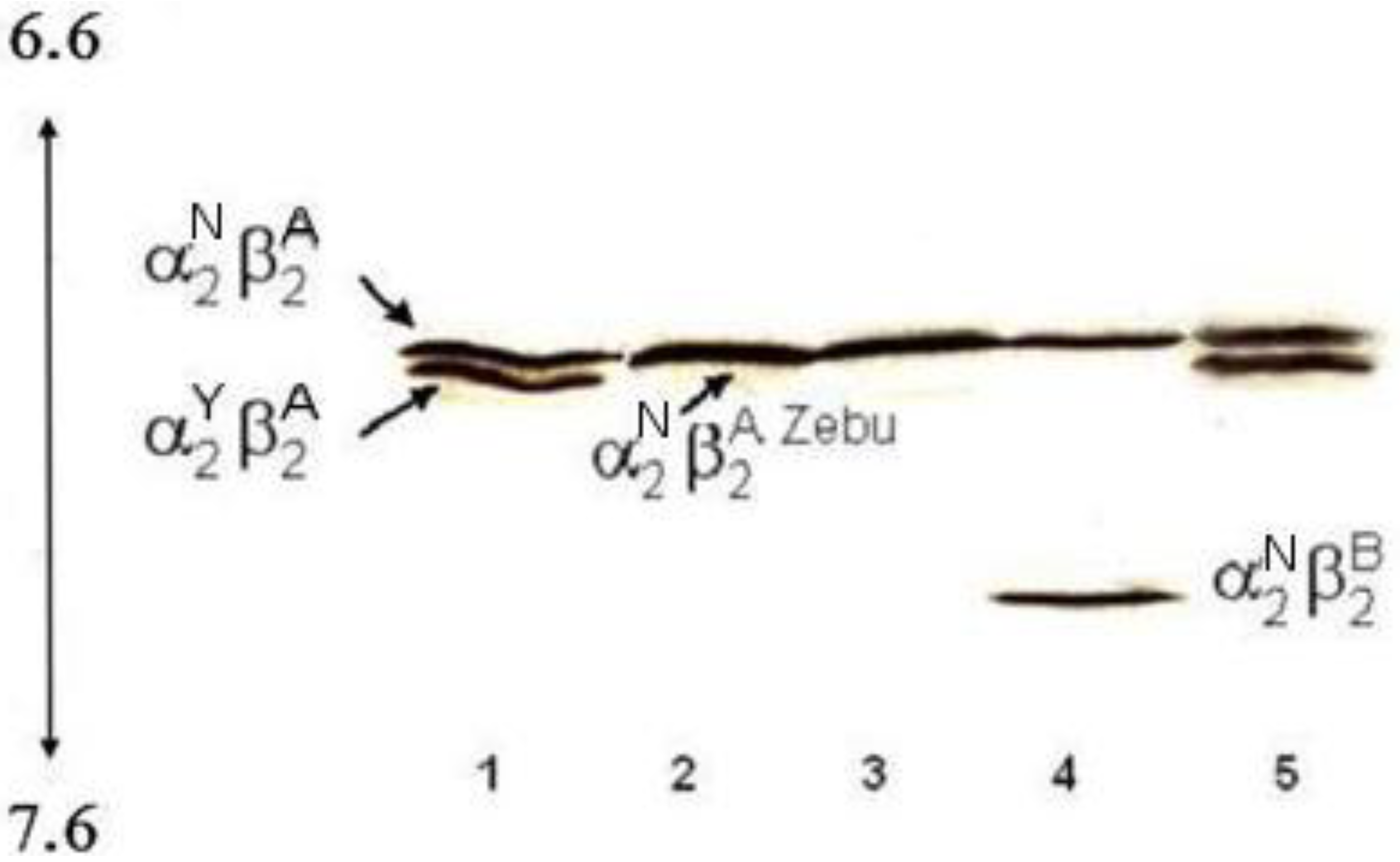
Polyacrylamide gel isoelectric focusing in a narrow pH range, 6.7–7.7, of cattle Hbs carrying different α and β globin chains. Lane 1: two non allelic alpha globins (the wild type IαN and the new variant IIαY) and one beta globin (βA) result in two Hb bands, αN2βA2 and αY2βA2; Lane 2: the same alpha globin encoded by non allelic genes (IαN and IIαN) and two beta globins characterized by the same electrophoretic mobility (βA and βAZebu) result in a single Hb band; Lane 3: the same alpha globin (αN) as in lane 2 and one beta globin (βA) result in single Hb band; Lane 4: the same alpha globin encoded by non allelic genes (IαN and IIαN) and two beta globins (βA and βB) result in two Hb bands, αN2βA2 and αN2βB2; Lane 5: the same as lane 1.
Figure 1.
Polyacrylamide gel isoelectric focusing in a narrow pH range, 6.7–7.7, of cattle Hbs carrying different α and β globin chains. Lane 1: two non allelic alpha globins (the wild type IαN and the new variant IIαY) and one beta globin (βA) result in two Hb bands, αN2βA2 and αY2βA2; Lane 2: the same alpha globin encoded by non allelic genes (IαN and IIαN) and two beta globins characterized by the same electrophoretic mobility (βA and βAZebu) result in a single Hb band; Lane 3: the same alpha globin (αN) as in lane 2 and one beta globin (βA) result in single Hb band; Lane 4: the same alpha globin encoded by non allelic genes (IαN and IIαN) and two beta globins (βA and βB) result in two Hb bands, αN2βA2 and αN2βB2; Lane 5: the same as lane 1.
1.3 Goats
Goats exhibit a very complex Hb polymorphism due to the presence of a number of allelic and non allelic chains both in the alpha and beta globin systems [
18,
40,
41,
42,
43,
44,
45]. Particularly, the
HBB locus is highly polymorphic, while only four different alpha globin variants have been found so far:
HBA1A and
HBA2T, the most frequent genes (more than 0.98 and almost 0.80, respectively), followed by
HBA2A (about 0.20) and
HBA1B (very rare) [
18]. However, all the possible tetramers resulting from combinations of the heterogeneity of alpha and beta globins resolve in two main electrophoretic zones, A and D, according to their different electrical properties (
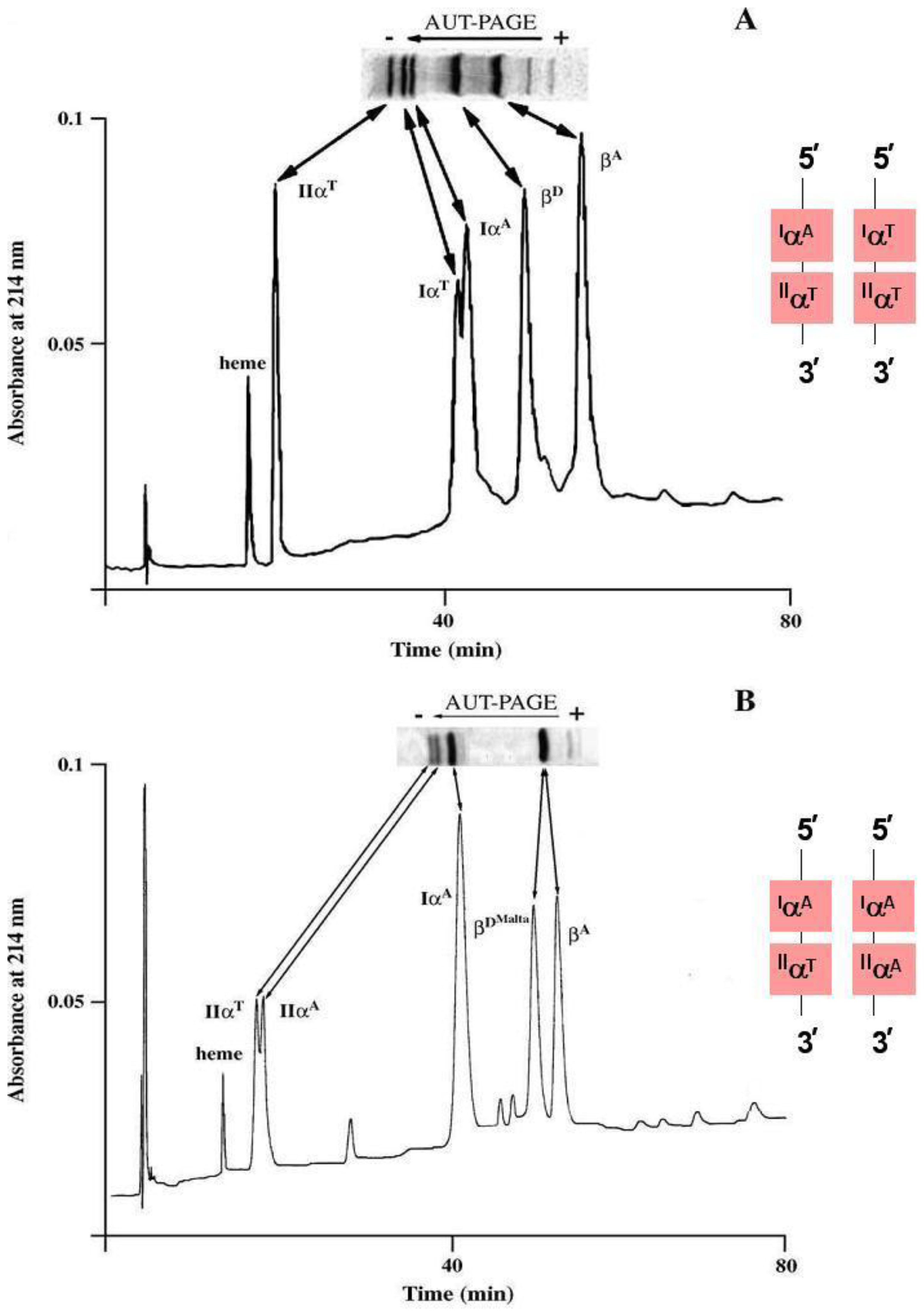
Figure 2). A group of positively charged tetramers such as HbB, HbD and HbD
Malta migrate to the D zone, whose isoelectric point (pI) range is about pH 7.2 (
Table 3). HbB results from an alpha chain variation due to the presence of the rare allele at the
HBA1 locus whereas HbD and HbD
Malta exhibit variation in the beta chain as a consequence of different point mutations at the
HBB locus.
Table 3.
Charged variants at the HBB locus in goats and corresponding hemoglobins with related molecular weight (MW) and isoelectric points (pI).
Table 3.
Charged variants at the HBB locus in goats and corresponding hemoglobins with related molecular weight (MW) and isoelectric points (pI).
| β-globin | Mutation versus βA | MW | Hb name | Hb pI |
|---|
| βA [40] | | 16,021.4 | HbA | 6.75 |
| βD [41] | 20Asp > His | 16,043.46 | HbD | 7.17 |
| βDMalta [44] | 69Asp > Gly | 15,963.37 | HbDMalta | 7.14 |
| βE [41] | 86Gln > His; 103Lys > Arg; 124Leu > Val | 16,044.40 | HbE | 6.79 |
Figure 2.
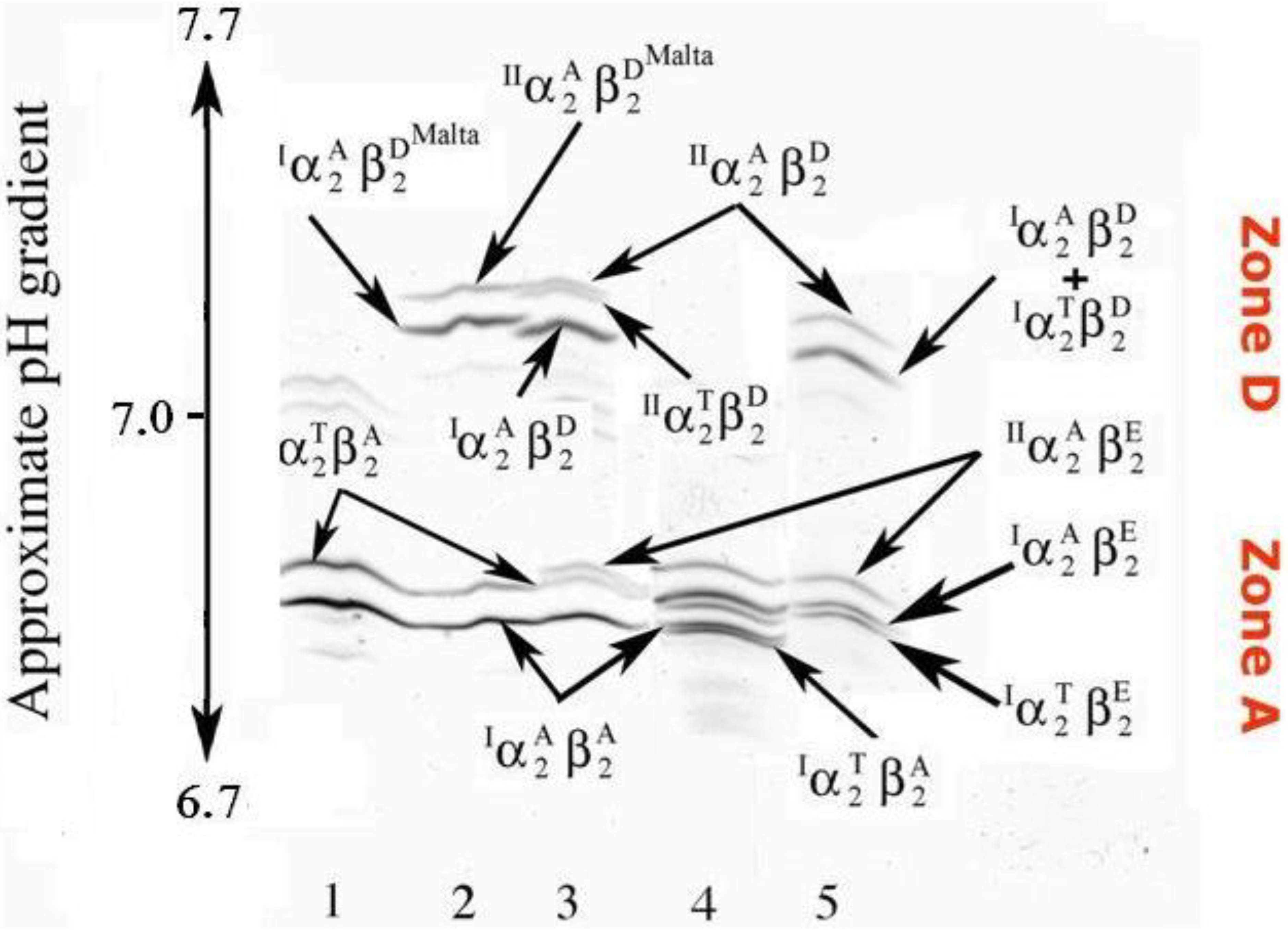
Polyacrylamide gel isoelectric (PAGIF) focusing on narrow pH range, 6.7–7.7, of goat hemoglobins carrying different α and β globin chains. Lane 1: two non allelic alpha globins (IαA and IIαT) and one beta globin (βA) resulting in two Hb bands, IαA2βA2 and IIαT2βA2; Lane 2: two non allelic alpha globins (IαA and IIαT) and two beta globins (βA and βDMalta) resulting in a twin couple of Hb bands, the former couple migrating to zone A (IαA2βA2 and IIαT2βA2) and the latter to zone D (IαA2β DMalta 2 and IIαT2β DMalta2); Lane 3: one non allelic alpha globin ( IαA), two allelic alpha globins ( IIαA and IIαT) and two beta globins (βA and βD) resulting in a twin triplet of Hb bands, with the former triplet migrating to zone A (IαA2βA2, IIαT2βA2 and IIαA2βA2) and the latter to zone D (IαA2βD2, IIαT2βA2 and IIαA2βD2); Lane 4: two allelic alpha globins (the wild type IαA and the new variant IαT), a non allelic alpha (IIαA) and two beta (βA and βE) resulting in six Hbs all migrating to zone A; Lane 5: the same alpha globins as in lane 4 (IαA, IαT and IIαA) but different beta globins (βE and βD) resulting in six Hbs, where the three tetramers containing the βE globin (IαT2βE2, IαA2βE2 and IIαA2βE2) migrate to zone A and the others containing the βD globin (IαT2βD2, IαA2βD2 and IIαA2βD2) migrate to zone D.
Figure 2.
Polyacrylamide gel isoelectric (PAGIF) focusing on narrow pH range, 6.7–7.7, of goat hemoglobins carrying different α and β globin chains. Lane 1: two non allelic alpha globins (IαA and IIαT) and one beta globin (βA) resulting in two Hb bands, IαA2βA2 and IIαT2βA2; Lane 2: two non allelic alpha globins (IαA and IIαT) and two beta globins (βA and βDMalta) resulting in a twin couple of Hb bands, the former couple migrating to zone A (IαA2βA2 and IIαT2βA2) and the latter to zone D (IαA2β DMalta 2 and IIαT2β DMalta2); Lane 3: one non allelic alpha globin ( IαA), two allelic alpha globins ( IIαA and IIαT) and two beta globins (βA and βD) resulting in a twin triplet of Hb bands, with the former triplet migrating to zone A (IαA2βA2, IIαT2βA2 and IIαA2βA2) and the latter to zone D (IαA2βD2, IIαT2βA2 and IIαA2βD2); Lane 4: two allelic alpha globins (the wild type IαA and the new variant IαT), a non allelic alpha (IIαA) and two beta (βA and βE) resulting in six Hbs all migrating to zone A; Lane 5: the same alpha globins as in lane 4 (IαA, IαT and IIαA) but different beta globins (βE and βD) resulting in six Hbs, where the three tetramers containing the βE globin (IαT2βE2, IαA2βE2 and IIαA2βE2) migrate to zone A and the others containing the βD globin (IαT2βD2, IαA2βD2 and IIαA2βD2) migrate to zone D.
![Diversity 02 00679 g002]()
1.4. Sheep
Four different α-globin [
46,
47,
48] and eight β-globin chains [
14,
49,
50,
51,
52,
53,
54,
55] have been described in domestic sheep (
Ovis aries). As for the β-globin chains, β
C and β
F are respectively encoded by the dormant gene
HBBC [
24] and by the gamma gene
HBBF switched on during foetal life [
55]. The remaining six globins are controlled by the
HBB locus. Out of them, two are encoded by the
HBBA and
HBBG alleles located in the switching A haplotype while four are encoded by the
HBBB,
HBBE,
HBBH,
HBBI alleles in the
B non-switching haplotype [
56]. As to the α-globin chains, the qualitative polymorphism is additionally complicated by the presence of a quantitative variation related to the fact that non allelic loci produce unequal amounts of α-globin (
Table 4).
Table 4.
Nomenclature of alpha globin genes and products, and amino acid exchanges involved in the different variants, referred to IαL globin in sheep.
Table 4.
Nomenclature of alpha globin genes and products, and amino acid exchanges involved in the different variants, referred to IαL globin in sheep.
| | | | Amino Acid position |
|---|
| Locus | Allele | Globin | 8 | 15 | 113 |
|---|
| HBA1 | HBA1L | IαL | S (Ser) | G (Gly) | L (Leu) |
| HBA1D | IαD | | D (Asp) | |
| HBA1A | IαA | A (Ala) | | |
| HBA2 | HBA2L | IIαL | | | |
| HBA2H | IIαH | | | H (His) |
| HBA3 | HBA3L | III αL | | | |
| HBA3H | IIIαH | | | H (His) |
| HBA4 | HBA4H | IVαH | | | H (His) |
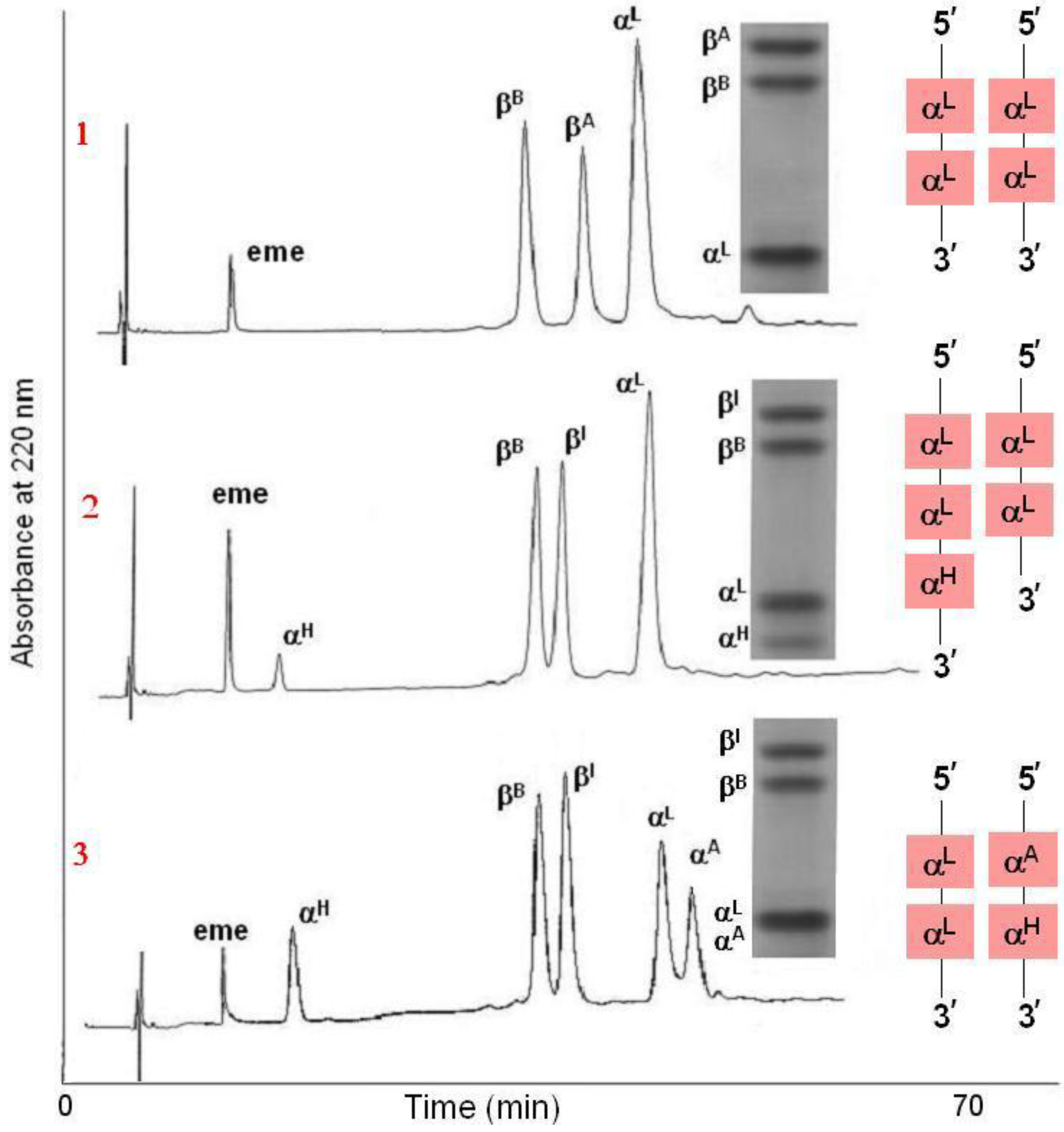
The genetic polymorphisms result in four electrophoretically separable protein zones, C, B, A, and F, according to their different electrical properties (
Figure 3). The less and the most positively charged tetramers are the fetal Hb, HbF (α
2γ
2), and the juvenile Hb, HbC (α
2Cβ
2) (
Figure 3, Lane 9). Most adult tetramers resulting from different variants both in the alpha and beta globin systems are found in the A or B zone, except for α
2Dβ
2A, which migrates to the F zone (
Figure 3, Lane 2 and 4).
Figure 3.
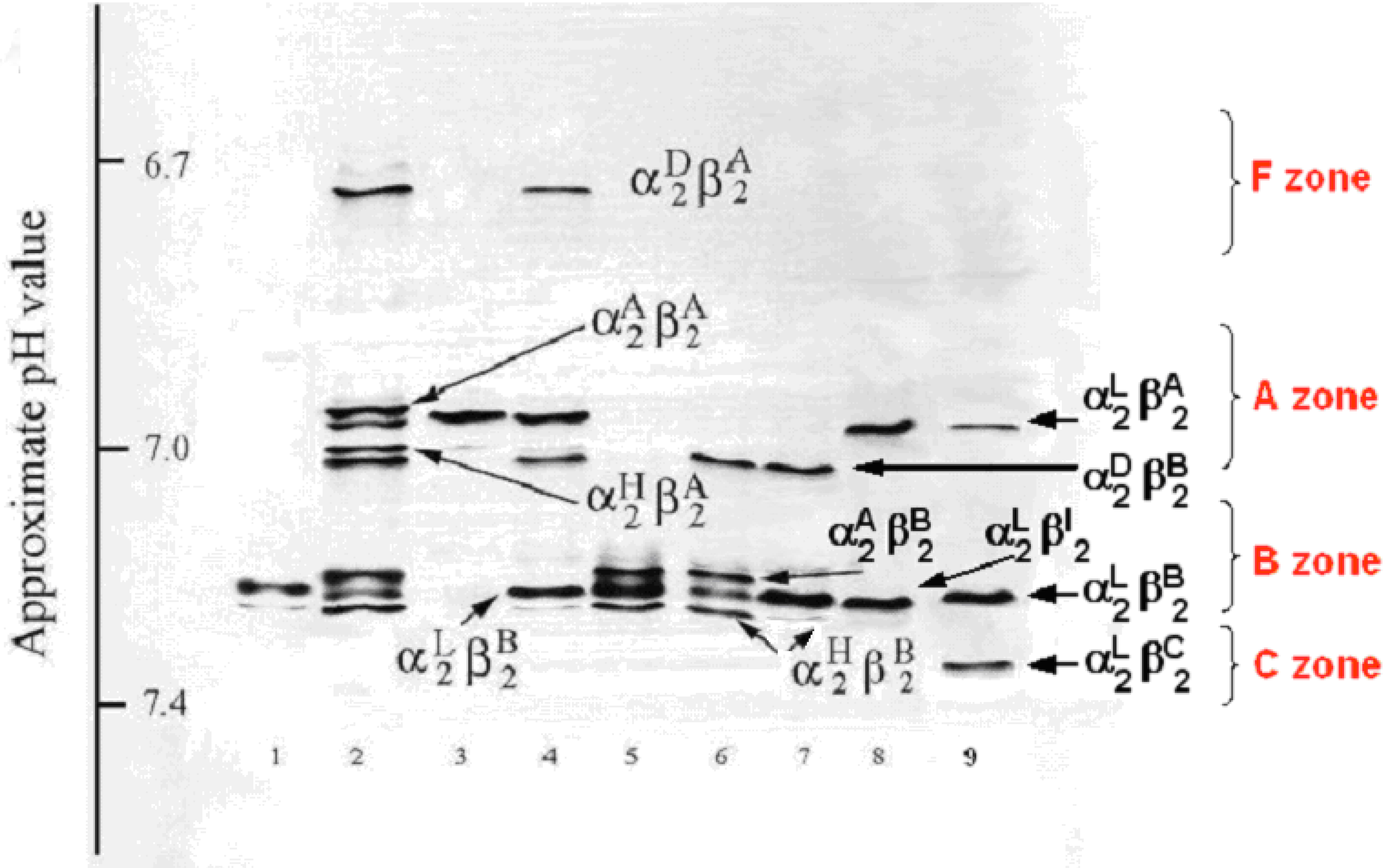
Polyacrylamide gel isoelectric focusing (PAGIF) showing adult Hb band patterns in Apulian sheep carrying different α and β globin chains. Lane 1: HBALL/LH and HBBB/B genotype. Two α (IαL, IIαH) and one β (βB) globins resulting in two bands, IIαH2βB2 and IαL2βB2.; Lane 2: HBADLH/AH and HBBA/B genotype. Four α (IαA, IαD, IIαL, II+IIIαH) and two β (βA, βB) globins resulting in eight bands, II+IIIαH2βB2, IIαL2βB2, IαA2βB2, IαD2βB2, II+IIIαH2βA2, IIαL2βA2, IαA2βA2 and IαD2βA2; Lane 3: HBALL/LLH and HBAA/A genotype. Two α (I+IIαL, IIIαH) and one β (βA) globins resulting in two bands, IIIαH2βA2 and I+IIαL2βA2; Lane 4: HBADLH/LL and HBBA/B genotype. Three α (IαD, I+IIαL, IIIαH) and two β (βA, βB) globins resulting in six bands, IIIαH2βB2, I+IIαL2βB2, IαD2βB2, IIIαH2βA2, I+IIαL2βA2 and IαD2βA2; Lane 5: HBALL/AH and HBBB/B genotype. Three α (IαA, I+IIαL, IIαH) and one β (βB) globins resulting in three bands, IIαH2βB2, I+IIαL2βB2 and IαA2βB2; Lane 6: HBADLLH/AH and HBBB/B genotype. Four α (IαA, IαD, II+IIIαL, II+IVαH) and one β (βB) globins resulting in four bands, II+IVαH2βB2, II+IIIαL2βB2, IαA2βB2 and IαD2βB2; Lane 7: HBADLLH/LL and HBBB/B genotype. Three α (IαD, IαL, IVαH) and one β (βB) globins resulting in three bands, I+II+IIIαL2βB2, IαDβB2 and IVαHβB2, the third band is very faint due to the low output of the HBA4gene; Lane 8: HBALL/LL and HBBA/I genotype. One α (I+IIαL) and two β (βA, βI) globins resulting in two bands α2Lβ2A and α2Lβ2I (α2Lβ2I band overlaps with α2Lβ2B); Lane 9: HBALL/LL and HBBA/B genotype. One α (IαL) and three β (βA, βB, βC) globins resulting in three bands, αL2βA2 , αL2βB2 and αL2βC2.
Figure 3.
Polyacrylamide gel isoelectric focusing (PAGIF) showing adult Hb band patterns in Apulian sheep carrying different α and β globin chains. Lane 1: HBALL/LH and HBBB/B genotype. Two α (IαL, IIαH) and one β (βB) globins resulting in two bands, IIαH2βB2 and IαL2βB2.; Lane 2: HBADLH/AH and HBBA/B genotype. Four α (IαA, IαD, IIαL, II+IIIαH) and two β (βA, βB) globins resulting in eight bands, II+IIIαH2βB2, IIαL2βB2, IαA2βB2, IαD2βB2, II+IIIαH2βA2, IIαL2βA2, IαA2βA2 and IαD2βA2; Lane 3: HBALL/LLH and HBAA/A genotype. Two α (I+IIαL, IIIαH) and one β (βA) globins resulting in two bands, IIIαH2βA2 and I+IIαL2βA2; Lane 4: HBADLH/LL and HBBA/B genotype. Three α (IαD, I+IIαL, IIIαH) and two β (βA, βB) globins resulting in six bands, IIIαH2βB2, I+IIαL2βB2, IαD2βB2, IIIαH2βA2, I+IIαL2βA2 and IαD2βA2; Lane 5: HBALL/AH and HBBB/B genotype. Three α (IαA, I+IIαL, IIαH) and one β (βB) globins resulting in three bands, IIαH2βB2, I+IIαL2βB2 and IαA2βB2; Lane 6: HBADLLH/AH and HBBB/B genotype. Four α (IαA, IαD, II+IIIαL, II+IVαH) and one β (βB) globins resulting in four bands, II+IVαH2βB2, II+IIIαL2βB2, IαA2βB2 and IαD2βB2; Lane 7: HBADLLH/LL and HBBB/B genotype. Three α (IαD, IαL, IVαH) and one β (βB) globins resulting in three bands, I+II+IIIαL2βB2, IαDβB2 and IVαHβB2, the third band is very faint due to the low output of the HBA4gene; Lane 8: HBALL/LL and HBBA/I genotype. One α (I+IIαL) and two β (βA, βI) globins resulting in two bands α2Lβ2A and α2Lβ2I (α2Lβ2I band overlaps with α2Lβ2B); Lane 9: HBALL/LL and HBBA/B genotype. One α (IαL) and three β (βA, βB, βC) globins resulting in three bands, αL2βA2 , αL2βB2 and αL2βC2.
![Diversity 02 00679 g003]()