Application of SRAP Markers to Identify Gender and Species in Genus Ephedra Tourn. ex L.
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling and Study Area
2.2. Genomic DNA Extraction and Amplification
2.3. Sequencing of SRAP Marker Bands
2.4. SRAP Analysis
2.5. Statistical Analysis
3. Results and Discussion
3.1. Morphometric Features

3.2. SRAP Markers for Molecular Characterization of Ephedra Species/Genders
3.3. SRAP-Based Similarity Indexes (SIs) Between the Inter- and Intraspecific Studied Species
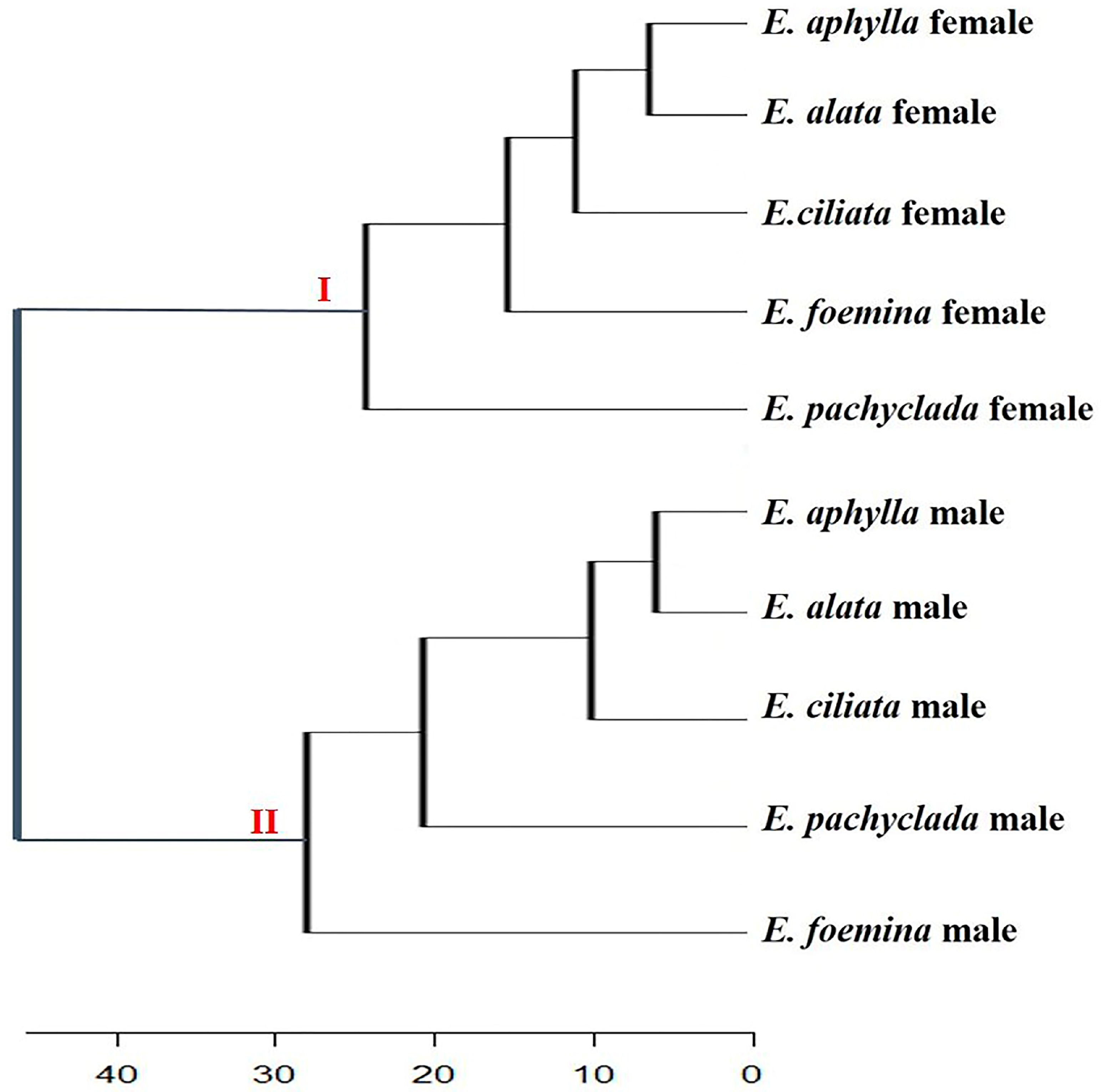
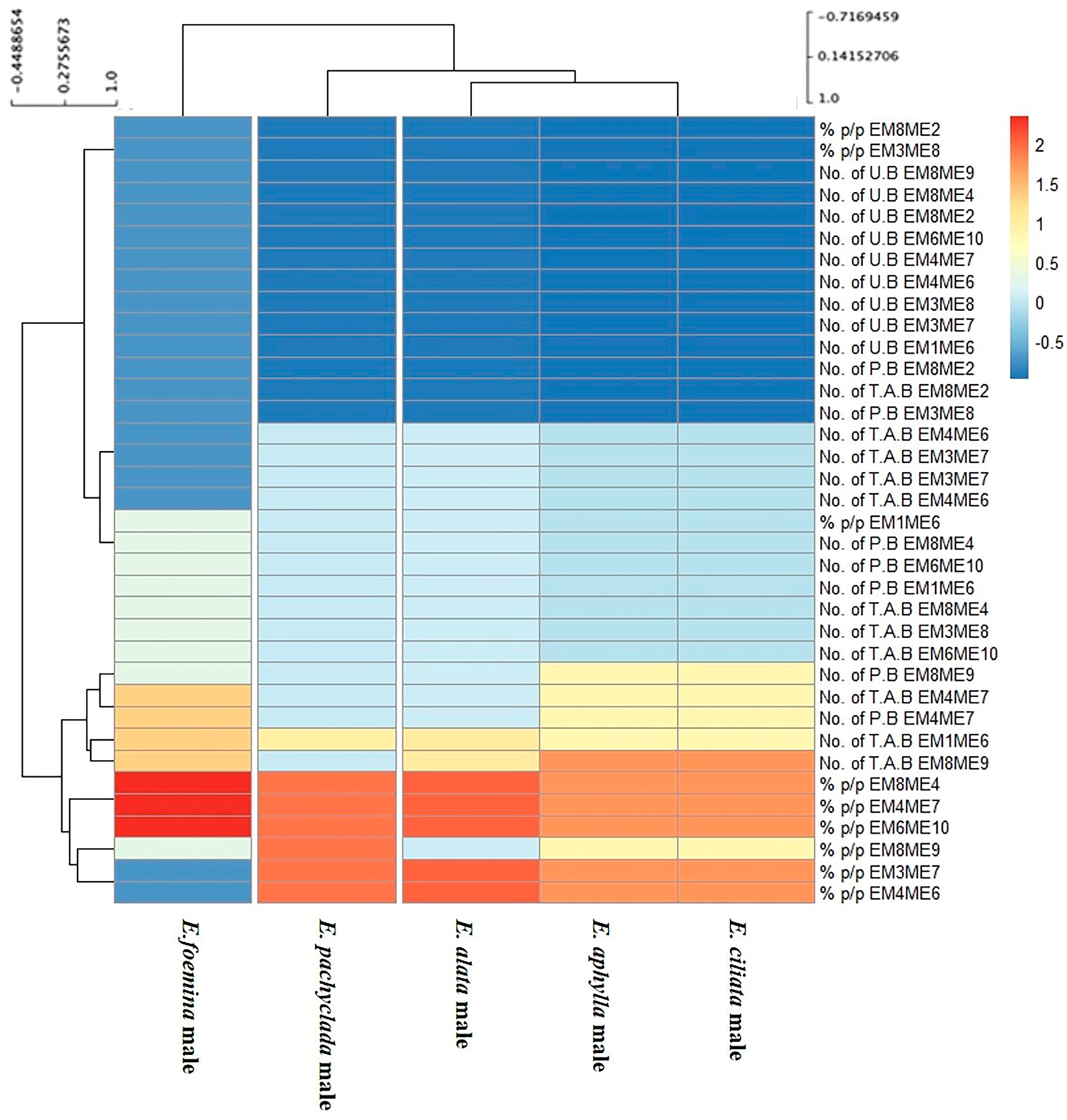
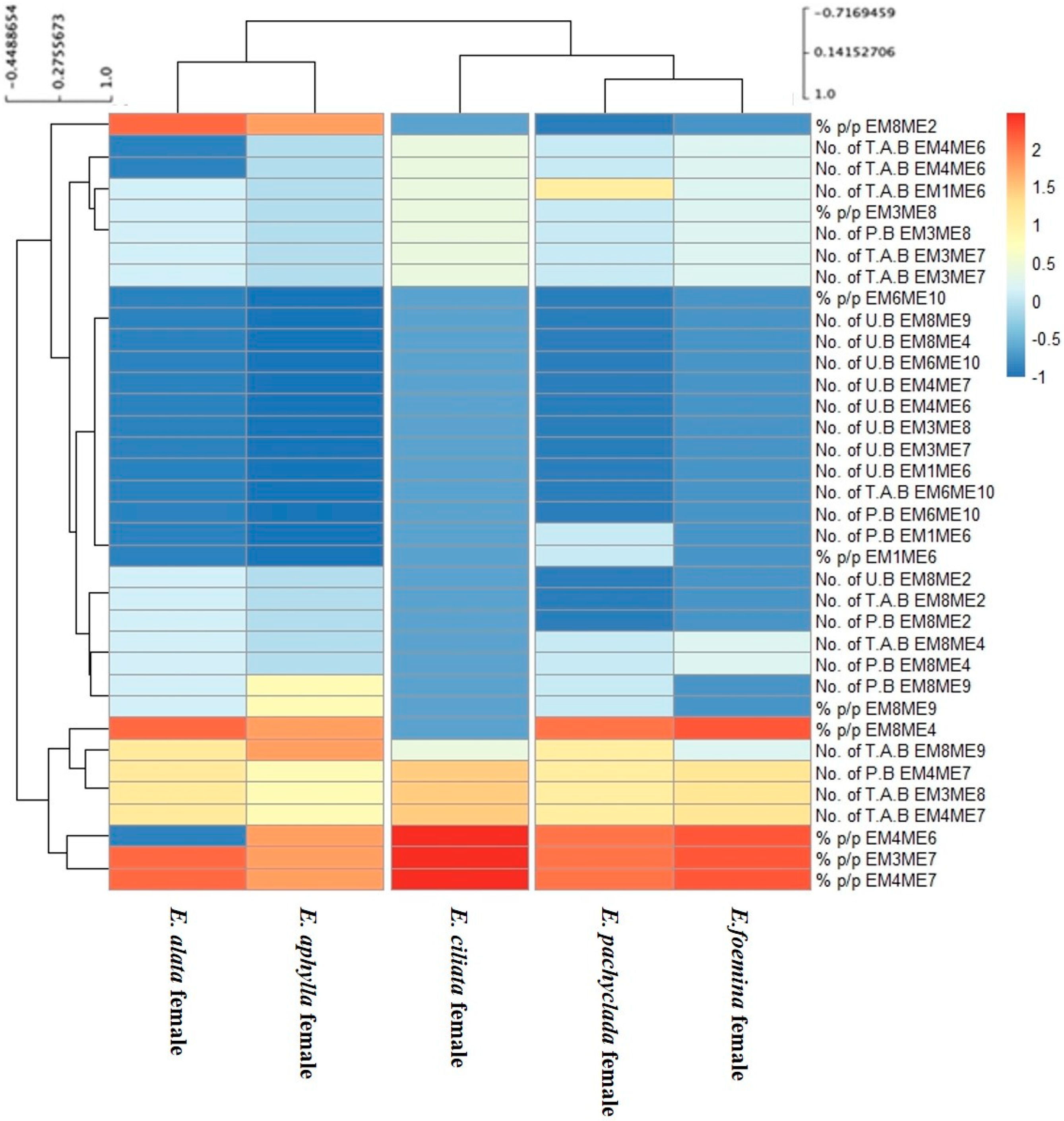
3.4. Heat Map Showing the Genetic Variations of Genders at Inter- and Intra-Specific Levels
3.4.1. Male Genders
3.4.2. Female Genders
3.4.3. Polymorphic Information Content (PIC), Compared with Other Gymnosperms
3.4.4. Marker Index (MI) Compared with Other Gymnosperms
4. Conclusions
5. Recommendation
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| No. of T.A.B. | Number of total amplified bands |
| No of P.B. | Number of polymorphic bands |
| No of U.B. | Number of unique bands |
| % p/p | % of polymorphism/primer |
References
- Mongrand, S.; Badoc, A.; Patouille, B.; Lacomblez, C.; Chavent, M.; Cassagne, C.; Bessoule, J.-J. Taxonomy of gymnospermae: Multivariate analyses of leaf fatty acid composition. Phytochemistry 2001, 58, 101–115. [Google Scholar] [CrossRef] [PubMed]
- Boulos, L. Flora of Egypt Checklist; Al-Hadara Publishing: Cairo, Egypt, 2009; pp. 198–201. [Google Scholar]
- Ickert-Bond, S.M. Systematics of New World Ephedra L. (Ephedraceae): Integrating Morphological and Molecular Data; Arizona State University: Tempe, AZ, USA, 2003; p. 363. [Google Scholar]
- Ickert-Bond, S.M.; Rydin, C.; Renner, S.S. A fossil-calibrated relaxed clock for Ephedra indicates an Oligocene age for the divergence of Asian and New World clades and Miocene dispersal into South America. J. Syst. Evol. 2009, 47, 444–456. [Google Scholar] [CrossRef]
- González-Juárez, D.E.; Escobedo-Moratilla, A.; Flores, J.; Hidalgo-Figueroa, S.; Martínez-Tagüeña, N.; Morales-Jiménez, J.; Muñiz-Ramírez, A.; Pastor-Palacios, G.; Pérez-Miranda, S.; Ramírez-Hernández, A. A review of the Ephedra genus: Distribution, ecology, ethnobotany, phytochemistry, and pharmacological properties. Molecules 2020, 25, 3283. [Google Scholar] [CrossRef]
- Faried, A.; El-Banhawy, A.; Elqahtani, M. Taxonomic, DNA barcoding and phylogenetic reassessment of the Egyptian Ephedra L. (Ephedraceae). Catrina Int. J. Environ. Sci. 2018, 17, 1–13. [Google Scholar] [CrossRef]
- Khalaf, M.H.; Amer, W.M.; Shaye, N.A.A.; Hassan, M.O.; Gomaa, N.H. Taxonomic Revision of Genus Ephedra Tourn. ex L. in Egypt. with Intra-Gender Diversity in Morphometric Traits and Fatty Acid. Compos. Plants 2024, 13, 2442. [Google Scholar] [CrossRef]
- Yong, Y. Ontogeny and metamorphic patterns of female reproductive organs of Ephedra sinica Stapf (Ephedraceae). Acta Bot. Sin. 2001, 43, 1011–1017. [Google Scholar]
- Rydin, C.; Pedersen, K.R.; Crane, P.R.; Friis, E.M. Former diversity of Ephedra (Gnetales): Evidence from early Cretaceous seeds from Portugal and North America. Ann. Bot. 2006, 98, 123–140. [Google Scholar] [CrossRef]
- Huang, J.; Giannasi, D.E.; Price, R.A. Phylogenetic relationships in Ephedra (Ephedraceae) inferred from chloroplast and nuclear DNA sequences. Mol. Phylogenetics Evol. 2005, 35, 48–59. [Google Scholar] [CrossRef]
- Price, R.A. Systematics of the Gnetales: A review of morphological and molecular evidence. Int. J. Plant Sci. 1996, 157, S40–S49. [Google Scholar] [CrossRef]
- Rydin, C.; Källersjö, M.; Friis, E.M. Seed plant relationships and the systematic position of Gnetales based on nuclear and chloroplast DNA: Conflicting data, rooting problems, and the monophyly of conifers. Int. J. Plant Sci. 2002, 163, 197–214. [Google Scholar] [CrossRef]
- Jia, H.; Zhao, L.; Wang, Y.; Wu, H.; Zhao, H.; Zhu, Y.; Jiao, Y.; Wang, G.; Zhou, C.; Huang, C. Comparative transcriptome analysis reveals sex-biased expression of hormone-related genes at an early stage of sex differentiation in red bayberry (Morella rubra). Horticulturae 2022, 8, 183. [Google Scholar] [CrossRef]
- Rydin, C.; Khodabandeh, A.; Endress, P.K. The female reproductive unit of Ephedra (Gnetales): Comparative morphology and evolutionary perspectives. Bot. J. Linn. Soc. 2010, 163, 387–430. [Google Scholar] [CrossRef]
- Boecklen, W.J.; Hoffman, M.T. Sex-biased herbivory in Ephedra trifurca: The importance of sex-by-environment interactions. Oecologia 1993, 96, 49–55. [Google Scholar] [CrossRef]
- Wu, H.; Yu, Q.; Ran, J.-H.; Wang, X.-Q. Unbiased subgenome evolution in allotetraploid species of Ephedra and its implications for the evolution of large genomes in gymnosperms. GBE 2021, 13, evaa236. [Google Scholar] [CrossRef] [PubMed]
- Rydin, C.; Korall, P. Evolutionary relationships in Ephedra (Gnetales), with implications for seed plant phylogeny. Int. J. Plant Sci. 2009, 170, 1031–1043. [Google Scholar] [CrossRef]
- Ickert-Bond, S.M.; Wojciechowski, M.F. Phylogenetic relationships in Ephedra (Gnetales): Evidence from nuclear and chloroplast DNA sequence data. Syst. Bot. 2004, 29, 834–849. [Google Scholar] [CrossRef]
- Hong, K.; Radian, Y.; Manda, T.; Xu, H.; Luo, Y. The development of plant genome sequencing technology and its conservation and application in endangered gymnosperms. Plants 2023, 12, 4006. [Google Scholar] [CrossRef]
- Fossdal, C.G.; Krokene, P.; Olsen, J.E.; Strimbeck, R.; Viejo, M.; Yakovlev, I.; Mageroy, M.H. Epigenetic stress memory in gymnosperms. Plant Physiol. 2024, 195, 1117–1133. [Google Scholar] [CrossRef]
- Freitag, H.; Maier-Stolte, M. A new species and a new combination in the genus Ephedra from Arabia. Edinb. J. Bot. 1992, 49, 89–93. [Google Scholar] [CrossRef]
- Maryam, M.J.J.; Awan, F.S.; Ahmad, S.; Khan, I.A. Development of a molecular method for sex identification in date palm (Phoenix dactylifera L.) plantlets using novel sex-linked microsatellite markers. 3 Biotech. 2016, 6, 22. [Google Scholar] [CrossRef]
- González, D.; Vovides, A.P. A modification to the SCAR (Sequence Characterized Amplified Region) method provides phylogenetic insights within Ceratozamia (Zamiaceae). Rev. Mex. Biodiv. 2012, 83, 929–938. [Google Scholar] [CrossRef]
- Hajari, E.; Hannweg, K.; Nonyane, D.; Shezi, Z.; de Jager, K.; du Preez, R. Sex determination in Kei-apple (Dovyalis caffra) using molecular markers and flow cytometry. J. S. Afr. bot. 2019, 127, 278–283. [Google Scholar] [CrossRef]
- Li, G.; Quiros, C.F. Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: Its application to mapping and gene tagging in Brassica. Theor. Appl. Genet. 2001, 103, 455–461. [Google Scholar] [CrossRef]
- Zhou, Y.; Wang, X.; Zhang, X. Development and application of a SRAP marker for the identification of sex in Buchloe dactyloides. Euphytica 2011, 181, 261–266. [Google Scholar] [CrossRef]
- Robarts, D.W.; Wolfe, A.D. Sequence-related amplified polymorphism (SRAP) markers: A potential resource for studies in plant molecular biology. Appl. Plant Sci. 2014, 2, 1400017. [Google Scholar] [CrossRef] [PubMed]
- Vos, P.; Hogers, R.; Bleeker, M.; Reijans, M.; Lee, T.v.d.; Hornes, M.; Friters, A.; Pot, J.; Paleman, J.; Kuiper, M. AFLP: A new technique for DNA fingerprinting. Nucleic Acids Res. Spec. Publ. 1995, 23, 4407–4414. [Google Scholar] [CrossRef]
- Ferriol, M.; Picó, B.; Nuez, F. Genetic diversity of a germplasm collection of Cucurbita pepo using SRAP and AFLP markers. Theor. Appl. Genet. 2003, 107, 271–282. [Google Scholar] [CrossRef]
- Inan, N.; Yildiz, M.; Sensoy, S.; Kafkas, S.; Abak, K. Efficacy of ISSR and SRAP techniques for molecular characterization of some Cucurbita genotypes including naked (hull-less) seed pumpkin. J. Anim. Plant Sci. 2012, 22, 126–136. [Google Scholar]
- Amer, W.M.; Al Shaye, N.A.; Hassan, M.O.; Khalaf, M.H. Heteroblastic Inflorescence of Lamium amplexicaule L. in Egyptian Flora. Plants 2023, 12, 1028. [Google Scholar] [CrossRef]
- Walas, Ł.; Mandryk, W.; Thomas, P.A.; Tyrała-Wierucka, Ż.; Iszkuło, G. Sexual systems in gymnosperms: A review. Basic. Appl. Ecol. 2018, 31, 1–9. [Google Scholar] [CrossRef]
- Khalaf, M.H.; Amer, W.M.; Shaye, N.A.A.; Hassan, M.O.; Gomaa, N.H. Anatomical Tool as an Additional Approach for Identifying Pharmaceutically Important Ephedra Species (Ephedraceae) at the Gender Identity Level in Egypt. Biology 2024, 13, 947. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Ji, F.; Hou, F.; Shi, Q.; Xing, G.; Chen, H.; Weng, Y.; Kang, X. Morphological, palynological and molecular assessment of Hemerocallis core collection. Sci. Hortic. 2021, 285, 110181. [Google Scholar] [CrossRef]
- Wang, S.; Li, Y.; Li, Z.; Chang, L.; Li, L. Identification of an SCAR marker related to female phenotype in Idesia polycarpa Maxim. Genet. Mol. Res. 2015, 14, 2015–2022. [Google Scholar] [CrossRef] [PubMed]
- Zheng, C.; Zhu, X.; Fang, L.; Chen, Y.; Wu, W. Analysis of sex and strains of Ficus awkeotsang Makino by SRAP. Chin. J. Trop. Crops 2009, 12, 1740–1745. [Google Scholar]
- Wu, W.; Zhu, X.; Chen, Y. Identification of a sex-associated SRAP marker in Ficus pumila L. Chin. J. Appl. Environ. Biol. 2008, 5, 021. [Google Scholar]
- Kumar, J.; Agrawal, V. Assessment of genetic diversity, population structure and sex identification in dioecious crop, Trichosanthes dioica employing ISSR, SCoT and SRAP markers. Heliyon 2019, 5, e01346. [Google Scholar] [CrossRef]
- Zapata-Valenzuela, J.; Isik, F.; Maltecca, C.; Wegrzyn, J.; Neale, D.; McKeand, S.; Whetten, R. SNP markers trace familial linkages in a cloned population of Pinus taeda—Prospects for genomic selection. Tree Genet. Genomes 2012, 8, 1307–1318. [Google Scholar] [CrossRef]
- Wu, Y.; Zhou, Q.; Huang, S.; Wang, G.; Xu, L. SNP development and diversity analysis for Ginkgo biloba based on transcriptome sequencing. Trees 2019, 33, 587–597. [Google Scholar] [CrossRef]
- Chybicki, I.; Oleksa, A.; Burczyk, J. Increased inbreeding and strong kinship structure in Taxus baccata estimated from both AFLP and SSR data. Heredity 2011, 107, 589–600. [Google Scholar] [CrossRef]
- Scotti, I.; Magni, F.; Paglia, G.; Morgante, M. Trinucleotide microsatellites in Norway spruce (Picea abies): Their features and the development of molecular markers. Theor. Appl. Genet. 2002, 106, 40–50. [Google Scholar] [CrossRef]
- Yibrah, H.S.; Manders, G.; Clapham, D.H.; Von Arnold, S. Biological factors affecting transient transformation in embryogenic suspension cultures of Picea abies. J. Plant Physiol. 1994, 144, 472–478. [Google Scholar] [CrossRef]
- Kyoda, S.; Setoguchi, H. Phylogeography of Cycas revoluta Thunb. (Cycadaceae) on the Ryukyu Islands: Very low genetic diversity and geographical structure. Plant Syst. Evol. 2010, 288, 177–189. [Google Scholar] [CrossRef]
| Species | Sex | Locality | GPS Coordinates | Date of Collection | |
|---|---|---|---|---|---|
| N | E | ||||
| E. alata | Male | Abo Zeinema, plain | 29°02′31″ | 33°06′30″ | 14 June 2023 |
| Female | Wadi bed of Wadi Feiran | 28°43′06″ | 33°37′06″ | 15 May 2023 | |
| E. aphylla | Female and Male | Wadi bed of wadi Al-Arbaeen | 28°55′58″ | 33°94′58″ | 17 August 2023 |
| E. ciliata | Male | Wadi bed of wadi Lethi, | 28°33′07″ | 33°58′24″ | 15 July 2023 |
| Female | Monastery at St. Catherine | 28°33′25″ | 33°58′23″ | 25 August 2023 | |
| E. foemina | Female and Male | Gabal Mousa, rocky ridge | 28°54′13″ | 33°57′55″ | 16 July 2023 |
| E. pachyclada | Female | Abu Walia, Rocky Ridge | 28°53′55″ | 33°51′39″ | 16 August 2023 |
| Male | Abu Walia, Rocky Ridge | 28°53′60″ | 33°50′40″ | 16 August 2023 | |
| Primer Code | Primer Sequence | Molecular Weight bp |
|---|---|---|
| EM1ME6 | Em-1: GACTGCGTACGAATTAAT Me-6: TGA GTC CAA ACC GGA CA | 75–145 |
| EM3ME7 | Em-3: GACTGCGTACGAATTGAC Me-7: TGA GTC CAA ACC GGA CG | 100 |
| EM3ME8 | Em-3: GACTGCGTACGAATTGAC Me-8: TGA GTC CTT TCC GGT GC | 55–85 |
| EM4ME6 | Em-4: GACTGCGTACGAATTTGA Me-6: TGA GTC CAA ACC GGA CA | 120 |
| EM4ME7 | Em-4: GACTGCGTACGAATTTGA Me-7: TGA GTC CAA ACC GGA CG | 75–90 |
| EM6ME10 | Em-6: GACTGCGTACGAATTGCA Me-10: TGAGTCCAAACCGGTGC | 95 |
| EM8ME2 | Em-8: GACTGCGTACGAATTCAC Me-4: TGAGTCCAAACCGGACC | 70–85 |
| EM8ME4 | Em-8: GACTGCGTACGAATTCAC Me-4: TGAGTCCAAACCGGACC | 95 |
| EM8ME9 | Em-8: GACTGCGTACGAATTCAC Me-9: TGAGTCCAAACCGGTCC | 85–240 |
| Species and Genders | E. alata | E. aphylla | E. ciliata | E. foemina | E. pachyclada | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Female | Male | Female | Male | Female | Male | Female | Male | Female | Male | ||
| E. alata | female | 1.0 | |||||||||
| male | 0.85 | 1.0 | |||||||||
| E. aphylla | female | 0.76 | 0.83 | 1.0 | |||||||
| male | 0.81 | 0.96 | 0.88 | 1.0 | |||||||
| E. ciliata | female | 0.85 | 0.91 | 0.83 | 0.96 | 1.0 | |||||
| male | 0.78 | 0.92 | 0.84 | 0.96 | 0.92 | 1.0 | |||||
| E. foemina | female | 0.85 | 0.91 | 0.75 | 0.88 | 0.91 | 0.92 | 1.0 | |||
| male | 0.84 | 0.81 | 0.72 | 0.78 | 0.81 | 0.83 | 0.81 | 1.0 | |||
| E. pachyclada | female | 0.76 | 0.91 | 0.75 | 0.88 | 0.91 | 0.92 | 0.91 | 0.81 | 1.0 | |
| male | 0.84 | 0.81 | 0.72 | 0.78 | 0.81 | 0.83 | 0.90 | 0.82 | 0.81 | 1.0 | |
| Index | Value | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| EM1ME6 | EM3ME7 | EM3ME8 | EM4ME6 | EM4ME7 | EM6ME10 | EM8ME2 | EM8ME4 | EM8ME9 | |
| H | 0.32 | 0.18 | 0.38 | 0.32 | 0.18 | 0.5 | 0.18 | 0.18 | 0.42 |
| PIC | 0.27 | 0.16 | 0.30 | 0.27 | 0.16 | 0.38 | 0.16 | 0.16 | 0.33 |
| E | 1.6 | 0.9 | 1.5 | 0.8 | 1.8 | 0.5 | 0.2 | 0.9 | 2.1 |
| Hav | 0.016 | 0.018 | 0.019 | 0.032 | 0.009 | 0.05 | 0.009 | 0.018 | 0.014 |
| MI | 0.026 | 0.016 | 0.028 | 0.026 | 0.016 | 0.025 | 0.0018 | 0.016 | 0.03 |
| D | 0.37 | 0.2 | 0.45 | 0.38 | 0.19 | 0.78 | 0.99 | 0.2 | 0.52 |
| R | 0.8 | 0.2 | 1 | 0.4 | 0.4 | 1 | 0.4 | 0.2 | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shaye, N.A.A.; Amer, W.M.; Hassan, M.O.; Gomaa, N.H.; Khalaf, M.H. Application of SRAP Markers to Identify Gender and Species in Genus Ephedra Tourn. ex L. Diversity 2025, 17, 600. https://doi.org/10.3390/d17090600
Shaye NAA, Amer WM, Hassan MO, Gomaa NH, Khalaf MH. Application of SRAP Markers to Identify Gender and Species in Genus Ephedra Tourn. ex L. Diversity. 2025; 17(9):600. https://doi.org/10.3390/d17090600
Chicago/Turabian StyleShaye, Najla A. Al, Wafaa M. Amer, Mahmoud O. Hassan, Nasr H. Gomaa, and Maha H. Khalaf. 2025. "Application of SRAP Markers to Identify Gender and Species in Genus Ephedra Tourn. ex L." Diversity 17, no. 9: 600. https://doi.org/10.3390/d17090600
APA StyleShaye, N. A. A., Amer, W. M., Hassan, M. O., Gomaa, N. H., & Khalaf, M. H. (2025). Application of SRAP Markers to Identify Gender and Species in Genus Ephedra Tourn. ex L. Diversity, 17(9), 600. https://doi.org/10.3390/d17090600







