Low Genetic and Parasite Diversity of Invasive Pumpkinseed Lepomis gibbosus (Centrarchidae) Expanding in Türkiye
Abstract
1. Introduction
2. Materials and Methods
2.1. Fish Sampling
2.2. Genetic Analysis of Fish
2.3. Parasite Collection, Processing and Analysis
3. Results
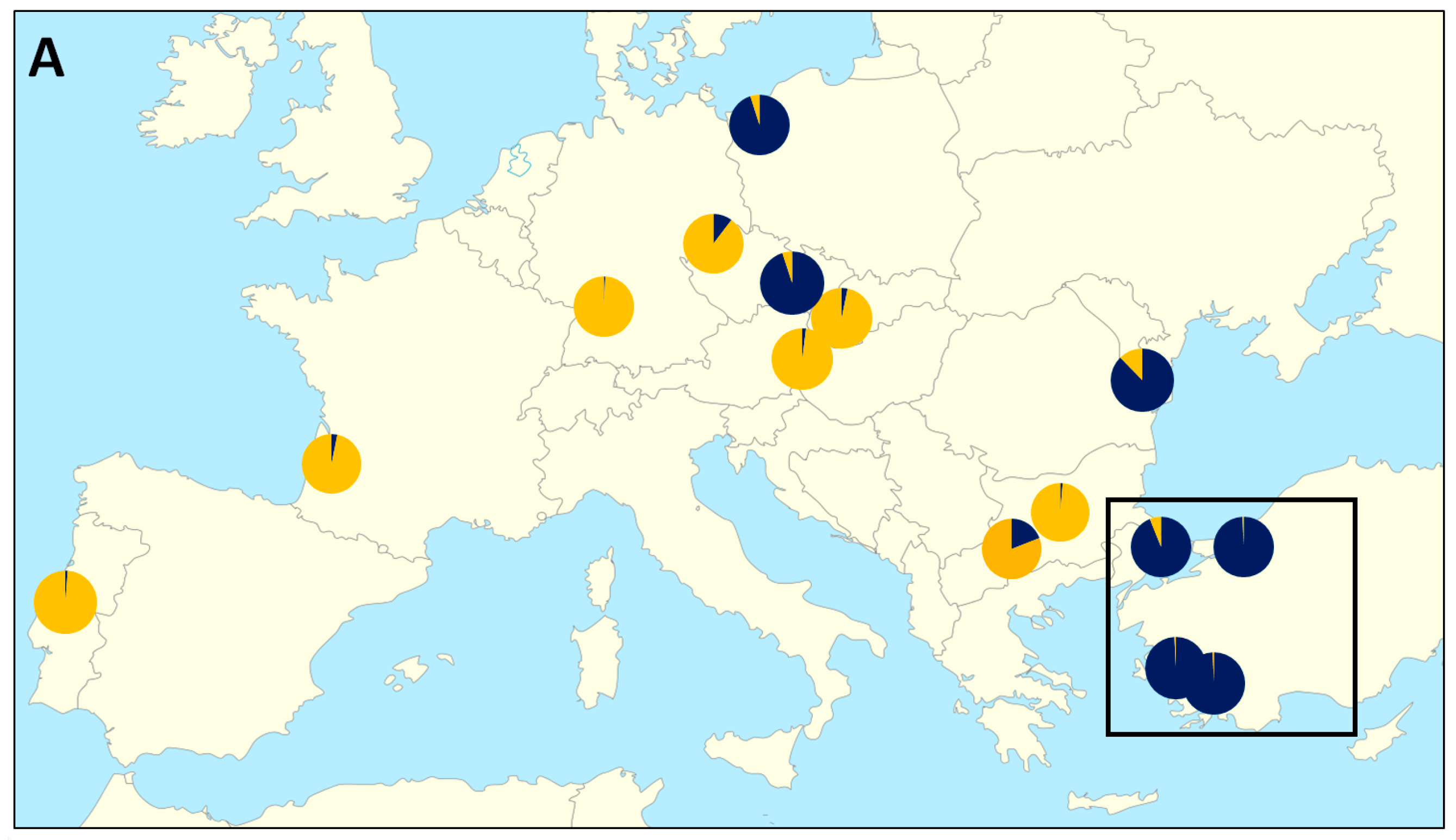
3.1. Genetic Characterisation of Fish Host
3.2. Composition of Parasite Communities
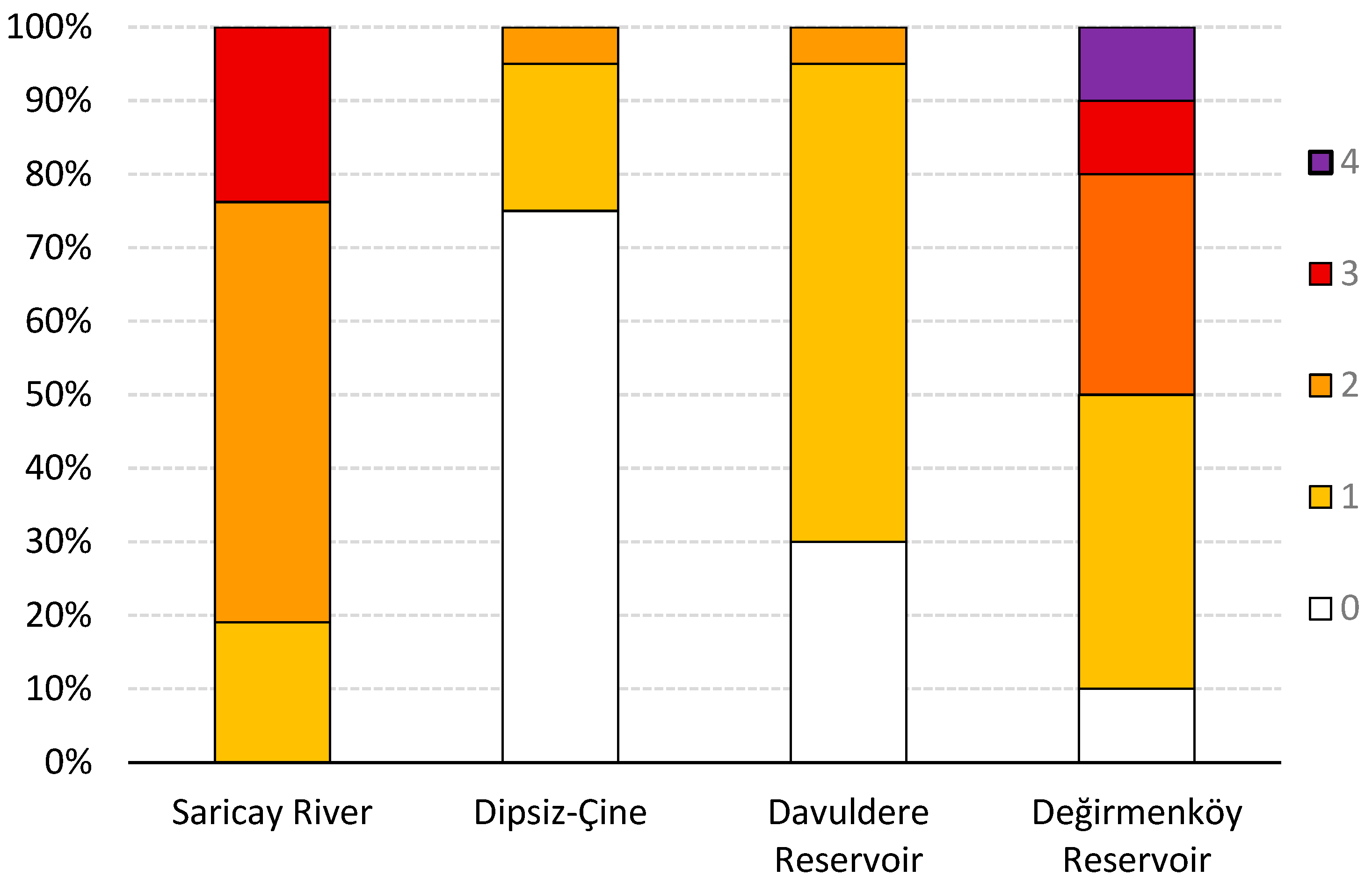
3.3. Parasite Diversity
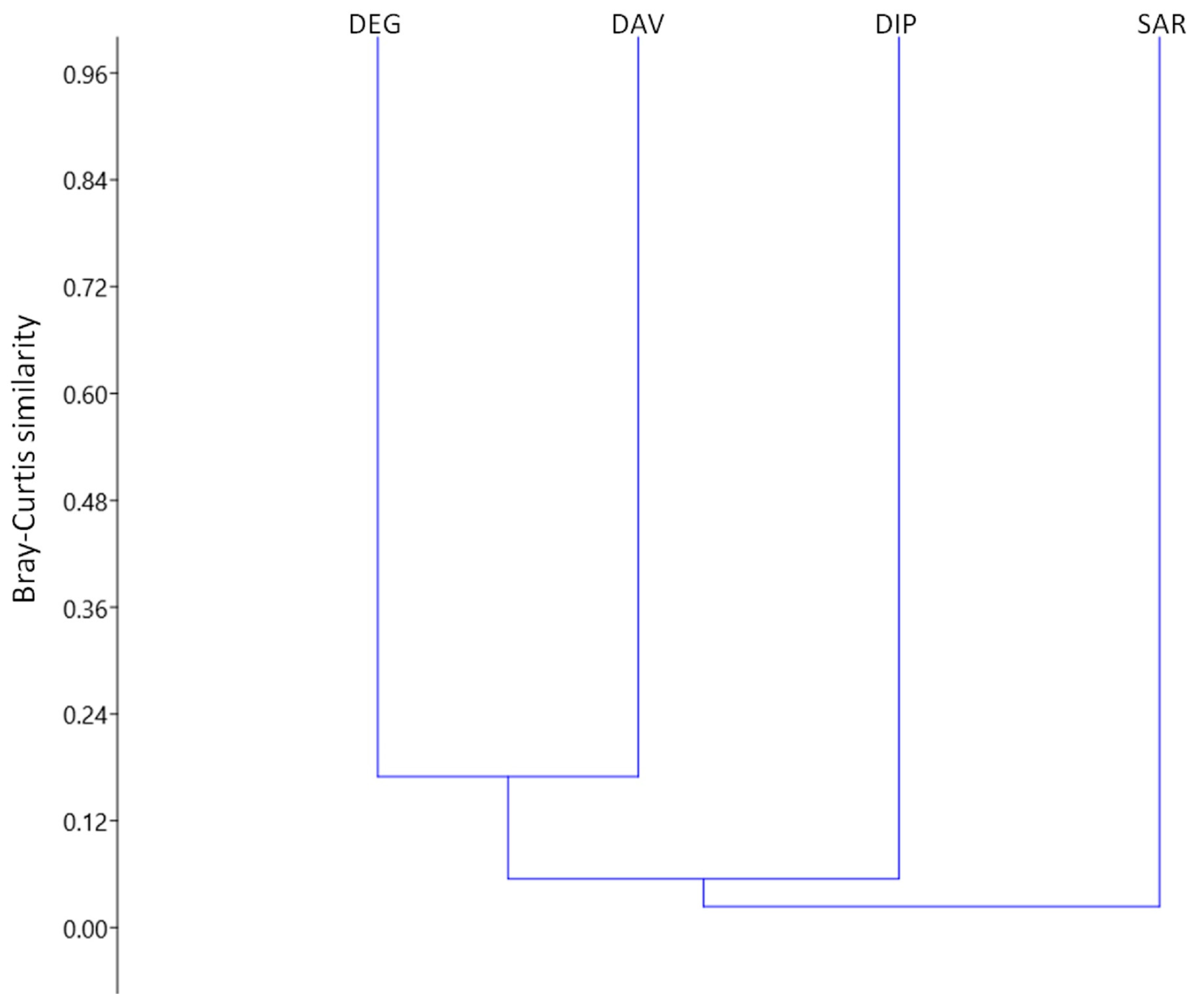
3.4. Similarity in Parasite Communities
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Copp, G.H.; Bianco, P.G.; Bogutskaya, N.G.; Erős, T.; Falka, I.; Ferreira, M.T.; Fox, M.G.; Freyhof, J.; Gozlan, R.E.; Grabowska, J.; et al. To be, or not to be, a non-native freshwater fish? J. Appl. Ichthyol. 2005, 21, 242–262. [Google Scholar] [CrossRef]
- Seebens, H.; Bacher, S.; Blackburn, T.M.; Capinha, C.; Dawson, W.; Dullinger, S.; Genovesi, P.; Hulme, P.E.; van Kleunen, M.; Kühn, I.; et al. Projecting the continental accumulation of alien species through to 2050. Glob. Chang. Biol. 2021, 27, 5970–5982. [Google Scholar] [CrossRef]
- Seebens, H.; Blackburn, T.M.; Dyer, E.E.; Genovesi, P.; Hulme, P.E.; Jeschke, J.M.; Pagad, S.; Pyšek, P.; Winter, M.; Arianoutsou, M.; et al. No saturation in the accumulation of alien species worldwide. Nat. Commun. 2017, 8, 114435. [Google Scholar] [CrossRef] [PubMed]
- Cucherousset, J.; Olden, J.D. Ecological impacts of non-native freshwater fishes. Fisheries 2011, 36, 215–230. [Google Scholar] [CrossRef]
- Gozlan, R.E.; Britton, J.R.; Cowx, I.; Copp, G.H. Current knowledge on non-native freshwater fish introductions. J. Fish Biol. 2010, 76, 751–786. [Google Scholar] [CrossRef]
- Hulme, P.E. Trade, transport and trouble: Managing invasive species pathways in an era of globalisation. J. Appl. Ecol. 2009, 46, 10–18. [Google Scholar] [CrossRef]
- Vitule, J.R.S.; Freire, C.A.; Simberloff, D. Introduction of nonnative freshwater fish can certainly be bad. Fish Fish. 2009, 10, 98–108. [Google Scholar] [CrossRef]
- Leunda, P.M. Impacts of non-native fishes on Iberian freshwater ichthyofauna: Current knowledge and gaps. Aquat. Invasions 2010, 5, 239–262. [Google Scholar] [CrossRef]
- Kuchta, R.; Choudhury, A.; Scholz, T. Asian fish tapeworm: The most successful invasive parasite in freshwaters. Trends Parasitol. 2018, 34, 511–523. [Google Scholar] [CrossRef]
- Shinn, A.P.; Avenant-Oldewage, A.; Bondad-Reantaso, M.G.; Cruz-Laufer, A.J.; García-Vásquez, A.; Hernández-Orts, J.S.; Kuchta, R.; Longshaw, M.; Metselaar, M.; Pariselle, A.; et al. A global review of problematic and pathogenic parasites of farmed tilapia. Rev. Aquacult. 2023, 15, 92–153. [Google Scholar] [CrossRef]
- Scott, W.B.; Crossman, E.J. Freshwater Fishes of Canada; Fisheries Research Board of Canada: Ottawa, ON, Canada, 1973; pp. 1–184. [Google Scholar]
- Stansch, K. Die Exotischen Zierfische in Wort und Bild; G. Wenzel & Sohn: Braunschweig, Germany, 1914; pp. 1–349. [Google Scholar]
- Kunstler, J. Amiurus nebulosus et Eupomotis gibbosus. Bull. Soc. Acclimat. 1908, 55, 238–244. [Google Scholar]
- Vivier, P. Poissons et crustacés d’eau douce acclimatés en France en eaux libres depuis le début du siècle. Terre Et Vie 1951, 98, 57–82. [Google Scholar]
- Diripasko, O.A.; Demchenko, N.A.; Kulik, P.V.; Zabroda, T.A. An expansion of the pumpkinseed, Lepomis gibbosus (Centrarchidae, Perciformes), area of distribution into the east of Ukraine. Vestnik Zoologii 2008, 42, 269–273, (In Russian with English Summary). [Google Scholar]
- Ribeiro, F.; Leunda, P.M. Non-native fish impacts on Mediterranean freshwater ecosystems: Current knowledge and research needs. Fish. Manag. Ecol. 2012, 19, 142–156. [Google Scholar] [CrossRef]
- Erk’akan, F. The fishes of the Thrace region. Hacettepe Bull. Nat. Sci. Eng. 1983, 12, 39–48. [Google Scholar]
- Barlas, M.; Dirican, S. The fish fauna of the Dipsiz-Çine (Muğla-Aydın) Stream. Gazi Univ. J. Sci. 2004, 17, 35–48. [Google Scholar]
- Barlas, M.; Yilmaz, F.; Dirican, S. New exotic species in the Sarıçay (Milas) and Dipsiz-Çine stream: Lepomis gibbosus (Perciformes: Centrarchidae). In IV. National Ecological and Environmental Congress Book; 5-8 Ekim, Bodrum: Bodrum, Turkey, 2001; pp. 307–312. [Google Scholar]
- Dirican, S.; Barlas, M. Physico-chemical characteristics and fish of Dipsiz and Çine (Mugla-Aydın) Stream. Ekoloji 2005, 14, 25–30. [Google Scholar]
- Ağdamar, S.; Tarkan, A.S.; Keskin, E.; Top Karakuş, N.; Doğaç, E.; Baysal, Ö.; Emiroğlu, Ö. The role of environmental factors and genetic diversity on colonisation success of a non-native fish, Lepomis gibbosus from western part of Türkiye. Biochem. Syst. Ecol. 2015, 58, 195–203. [Google Scholar] [CrossRef]
- Baran, I.; Ongan, T. Gala Gölü’nün limnolojik özellikleri Balıkçılık Sorunları ve Öneriler. In Gala Gölü ve Sorunları Sempozyumu; Doğal Hayatı Koruma Derneği Bilimsel Yayınlar Serisi: İstanbul, Turkey, 1988; pp. 46–54. [Google Scholar]
- Bay, H. Studies on Exotic Pumpkinseed (Lepomis gibbosus L., 1758) Population Living in KEMER Dam and Akçay Stream (Büyük Menderes River, Türkiye). Master’s Thesis, Süleyman Demirel University, Science Institute, Isparta, Turkey, 2010. [Google Scholar]
- İlhan, A.; Sarı, H.M.; Kurtul, I.; Akçalı, M. Actual situation of Meriç River’s fish fauna and assessment of possible impacts of alien species on native species. LimnoFish 2020, 6, 75–87. [Google Scholar] [CrossRef]
- Koca, Y.B.; Koca, S.; Yıldız, Ş.; Gürcü, B.; Osanç, E.; Tunçbaş, O.; Aksoy, G. Investigation of histopathological and cytogenetic effects on Lepomis gibbosus (Pisces: Perciformes) in the Çine stream (Aydın/Türkiye) with determination of water pollution. Environ. Toxicol. 2005, 20, 560–571. [Google Scholar] [CrossRef]
- Özcan, G. Distribution of the non-native fish species, pumpkinseed Lepomis gibbosus (Linnaeus, 1758), in Türkiye. Aquat. Invasions 2007, 2, 146–148. [Google Scholar] [CrossRef]
- Özuluğ, M.; Gaygusuz, O.; Gaygusuz, C.G.; Sac, G. New distribution areas of four invasive freshwater fish species from Turkish Thrace. Turk. J. Fish. Aquat. Sci. 2019, 19, 837–845. [Google Scholar] [CrossRef]
- Türker, D.; Ünal, A.; Öktener, A. New locality for the pumpkinseed, Lepomis gibbosus (Linnaeus, 1758) in the Marmara Region, Türkiye. Acta Biol. Turc. 2022, 35, 29–35. [Google Scholar]
- Esmaeili, H.R.; Sayyadzadeh, G.; Eagderi, S.; Abbasi, K. Checklist of freshwater fishes of Iran. FishTaxa 2018, 3, 1–95. [Google Scholar]
- Jang, M.-H.; Kim, J.-G.; Park, S.-B.; Jeong, K.-S.; Cho, G.-I.; Joo, G.-J. The Current Status of the Distribution of Introduced Fish in Large River Systems of South Korea. Int. Rev. Hydrobiol. 2002, 87, 319–328. [Google Scholar] [CrossRef]
- Kizuka, T.; Akasaka, M.; Kadoya, T.; Takamura, N. Visibility from roads predict the distribution of invasive fishes in agricultural ponds. PLoS ONE 2014, 9, e0099709. [Google Scholar] [CrossRef]
- Wang, H.P.; Gao, Z.X.; Rapp, D.; O’Bryant, P.; Yao, H.; Cao, X.J. Effects of temperature and genotype on sex determination and sexual size dimorphism of bluegill sunfish Lepomis macrochirus. Aquaculture 2014, 64–71, 420–421. [Google Scholar] [CrossRef]
- Bock, D.; Caseys, C.; Cousens, R.; Hahn, M.A.; Heredia, S.M.; Hübner, S.; Turner, K.G.; Whitney, K.D.; Rieseberg, L.H. What we still don’t know about invasion genetics. Molec. Ecol. 2015, 24, 2277–2298. [Google Scholar] [CrossRef] [PubMed]
- Cristescu, M.E. Genetic reconstructions of invasion history. Mol. Ecol. 2015, 24, 2212–2225. [Google Scholar] [CrossRef]
- Lambea-Camblor, A.; Morcillo, F.; Muñoz, J.; Perdices, A. Genetic and ecological approaches to introduced populations of pumpkinseed sunfish (Lepomis gibbosus) in Southwestern Europe. Diversity 2023, 15, 1059. [Google Scholar] [CrossRef]
- Ondračková, M.; Tkachenko, M.; Bartáková, V.; Bryjová, A.; Janáč, M.; Zięba, G.; Pyrzanowski, K.; Kvach, Y. Population genetic structure, parasite infection and somatic condition of pumpkinseed Lepomis gibbosus (Actinopterygii: Centrarchidae) in the Oder river basin. J. Fish Biol. 2023, 102, 426–442. [Google Scholar] [CrossRef] [PubMed]
- Yavno, S.; Gobin, J.; Wilson, C.C.; Vila-Gispert, A.; Copp, G.H.; Fox, M. New and Old World phylogeography of pumpkinseed Lepomis gibbosus (Linnaeus, 1758): The North American origin of introduced populations in Europe. Hydrobiologia 2020, 847, 345–364. [Google Scholar] [CrossRef]
- Ondračková, M.; Bartáková, V.; Kvach, Y.; Bryjová, A.; Trichkova, T.; Ribeiro, F.; Carassou, L.; Martens, A.; Masson, G.; Zechmeister, T.; et al. Parasite infection reflects host genetic diversity in the pumpkinseeds non-native range. Hydrobiologia 2021, 848, 2169–2187. [Google Scholar] [CrossRef]
- Keskin, E.; Ağdamar, S.; Tarkan, A.S. DNA barcoding common non-native freshwater fish species in Türkiye: Low genetic diversity but high population structuring. Mitochondrial DNA 2013, 24, 276–287. [Google Scholar] [CrossRef] [PubMed]
- Goedknegt, M.A.; Feis, M.E.; Wegner, K.M.; Luttikhuizen, P.C.; Buschbaum, C.; Camphuysen, K.C.; van der Meer, J.; Thieltges, D.W. Parasites and marine invasions: Ecological and evolutionary perspectives. J. Sea Res. 2016, 113, 11–27. [Google Scholar] [CrossRef]
- Taraschewski, H. Hosts and parasites as aliens. J. Helminthol. 2006, 80, 99–128. [Google Scholar] [CrossRef]
- Torchin, M.E.; Lafferty, K.D.; Dobson, A.P.; McKenzie, V.J.; Kuris, A.M. Introduced species and their missing parasites. Nature 2003, 421, 628–630. [Google Scholar] [CrossRef]
- Dudliv, I.; Kvach, Y.; Tkachenko, M.Y.; Nazaruk, K.; Ondračková, M. Comparative analysis of parasite load on recently established invasive pumpkinseed Lepomis gibbosus (Actinopterygii: Centrarchidae) in Europe. Acta Parasitol. 2024, 69, 819–830. [Google Scholar] [CrossRef]
- Hockley, F.A.; Williams, C.F.; Reading, A.J.; Taylor, N.G.H.; Cable, J. Parasite fauna of introduced pumpkinseed fish Lepomis gibbosus: First British record of Onchocleidus dispar (Monogenea). Dis. Aquat. Org. 2011, 97, 65–73. [Google Scholar] [CrossRef][Green Version]
- Goswami, U.; Molnár, K.; Cech, G.; Eiras, J.C.; Bandyopadhyay, P.K.; Ghosh, S.; Czeglédi, I.; Székely, C. Evidence of the American Myxobolus dechtiari was introduced along with its host in Europe: Molecular and histological data. Int. J. Parasit. Parasites Wildlife 2021, 15, 51–57. [Google Scholar] [CrossRef]
- Havlátová, L.; Ondračková, M.; Přikrylová, I. Monogenean parasites of Lepomis gibbosus Linnaeus introduced into the River Durance, France. Helminthologia 2015, 52, 323–330. [Google Scholar] [CrossRef]
- Kvach, Y.; Ondračková, M.; Kutsokon, Y.; Dzyziuk, N. New record of monogenean parasites on non-indigenous fishes in the Ukrainian Danube delta. BioInvasions Rec. 2018, 7, 65–72. [Google Scholar] [CrossRef]
- Ondračková, M.; Dávidová, M.; Přikrylová, I.; Pečínková, M. Monogenean parasites of introduced pumpkinseed Lepomis gibbosus (Centrarchidae) in the Danube River basin. J. Helminthol. 2011, 85, 435–441. [Google Scholar] [CrossRef]
- Kvach, Y.; Seifertová, M.; Carassou, L.; Ondračková, M. First record of the American cestode Proteocephalus ambloplitis (Leidy, 1887) (Proteocephalidae) in Europe. J. Helminthol. 2020, 94, e144. [Google Scholar] [CrossRef]
- Kvach, Y.; Jurajda, P.; Bryjová, A.; Trichkova, T.; Ribeiro, F.; Přikrylová, I.; Ondračková, M. European distribution for metacercariae of the North American digenean Posthodiplostomum cf. minimum centrarchi (Strigeiformes: Diplostomidae). Parasitol. Int. 2017, 66, 635–642. [Google Scholar] [CrossRef]
- Stoyanov, B.; Georgieva, S.; Pankov, P.; Kudlai, O.; Kostadinova, A.; Georgiev, B.B. Morphology and molecules reveal the alien Posthodiplostomum centrarchi Hoffman, 1958 as the third species of Posthodiplostomum Dubois, 1936 (Digenea: Diplostomidae) in Europe. Syst. Parasitol. 2017, 94, 1–20. [Google Scholar] [CrossRef]
- MacLeod, C.J.; Paterson, A.M.; Tompkins, D.M.; Duncan, R.P. Parasites lost—Do invaders miss the boat or drown on arrival? Ecol. Lett. 2010, 13, 516–527. [Google Scholar] [CrossRef] [PubMed]
- Kvach, Y.; Ondračková, M.; Janáč, M.; Jurajda, P. Methodological issues affecting the study of fish parasites. II. Sampling method affects ectoparasite studies. Dis. Aquat. Org. 2016, 121, 59–66. [Google Scholar] [CrossRef]
- Kvach, Y.; Ondračková, M.; Janáč, M.; Jurajda, P. Methodological issues affecting the study of fish parasites. I. Duration of live fish storage prior to dissection. Dis. Aquat. Org. 2016, 119, 107–115. [Google Scholar] [CrossRef]
- Ward, R.D.; Zemlak, T.S.; Innes, B.H.; Last, P.; Hebert, P.D.N. DNA barcoding Australia’s fish species. Philos. Trans. R. Soc. B 2005, 360, 1847–1857. [Google Scholar] [CrossRef]
- Weir, B.S.; Cockerham, C.C. Estimating F-statistics for the analysis of population structure. Evolution 1984, 38, 1358–1370. [Google Scholar]
- Belkhir, K.; Borsa, P.; Chikhi, L.; Raufaste, N.; Bonhomme, F. GENETIX 4.05, logiciel sous Windows TM Pour la Génétique des Populations. Laboratoire Génome, Populations, Interactions; CNRS UMR 5000; Université de Montpellier II: Montpellier, France, 1996–2004; Available online: http://www.genetix.univ-montp2.fr/genetix/genetix.htm (accessed on 28 April 2024).
- Raymond, M.; Rousset, F. An exact test for population differentiation. Evolution 1995, 49, 1280–1283. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P. Genalex 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 2006, 6, 288–295. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef]
- Hubisz, M.J.; Falush, D.; Stephens, M.; Pritchard, J.K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour. 2009, 9, 1322–1332. [Google Scholar] [CrossRef] [PubMed]
- Kopelman, N.M.; Mayzel, J.; Jakobsson, M.; Rosenberg, N.A.; Mayrose, I. Clumpak: A program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 2015, 15, 1179–1191. [Google Scholar] [CrossRef] [PubMed]
- Earl, D.A.; vonHoldt, B.M. Structure Harvester: A website and program for visualizing structure output and implementing the Evanno method. Conservation Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Malmberg, G. The excretory systems and the marginal hooks as a basis for the systematics of Gyrodactylus (Trematoda, Monogenea). Arkiv för Zoologi 1970, 23, 1–235. [Google Scholar]
- Cribb, T.H.; Bray, R.A. Gut wash, body soak, blender and heat-fixation: Approaches to the effective collection, fixation and preservation of trematodes of fishes. Syst. Parasitol. 2010, 76, 1–7. [Google Scholar] [CrossRef]
- Georgiev, B.; Biserkov, V.; Genov, T. In toto staining method for cestodes with iron acetocarmine. Helminthologia 1986, 23, 279–281. [Google Scholar]
- Bowles, J.; McManus, D.P. Rapid discrimination of Echinococcus species and strains using a PCR-based RFLP method. Mol. Biochem. Parasitol. 1993, 57, 231–240. [Google Scholar] [CrossRef] [PubMed]
- Carta, L.K.; Li, S. Improved 18S small subunit rDNA primers for problematic nematode amplification. J. Nematol. 2018, 50, 533–542. [Google Scholar] [CrossRef] [PubMed]
- Nunn, G.B. Nematode molecular evolution. Ph.D. Thesis, University of Nottingham, Nottingham, UK, 1992. [Google Scholar]
- Bush, A.O.; Lafferty, K.D.; Lotz, J.M.; Shostak, A.W. Parasitology meets ecology on its own terms: Margolis et al. revisited. J. Parasitol. 1997, 83, 575–583. [Google Scholar] [CrossRef] [PubMed]
- Magurran, A.E. Measuring Biological Diversity; Blackwell Publishing Ltd.: Oxford, UK, 2004; pp. 1–256. [Google Scholar]
- Zander, C.D. Four-year monitoring of parasite communities in gobiid fishes of the south-western Baltic. II. Infracommunity. Parasitol. Res. 2004, 93, 17–29. [Google Scholar] [CrossRef]
- Zander, C.D. Parasite diversity of sticklebacks from the Baltic Sea. Parasitol. Res. 2007, 100, 287–297. [Google Scholar] [CrossRef]
- Aydin, H.; Dilek, M.K.; Aydin, K. Trends in fish and fishery products consumption in Turkey. Turk. J. Fish. Aquat. Sci. 2011, 11, 499–506. [Google Scholar] [CrossRef]
- Al-Hassan, L.A.; Muhsin, K. The presence of Barbus luteus and Heteropneustes fossilis in the Khor al Zubair, in the North-West of the Arabian Gulf. Zool. Middle East 1986, 1, 116–118. [Google Scholar] [CrossRef]
- Ünlü, E.; Çiçek, T.; Değer, D.; Coad, B.W. Range extension of the exotic Indian stinging catfish, Heteropneustes fossilis (Bloch, 1794) (Heteropneustidae) into the Turkish part of the Tigris River watershed. J. Appl. Ichthyol. 2011, 27, 141–143. [Google Scholar] [CrossRef]
- Frankham, R. Inbreeding and extinction: A threshold effect. Conserv. Biol. 1995, 9, 792–799. [Google Scholar] [CrossRef]
- Luikart, G.; Zundel, S.; Rioux, D.; Miquel, C.; Keating, K.A.; Hogg, J.T.; Steele, B.; Foresman, K.; Taberlet, P. Low genotyping error rates and noninvasive sampling in bighorn sheep. J. Wildl. Manag. 2008, 72, 299–304. [Google Scholar] [CrossRef]
- Tarkan, A.S.; Karakus, U.; Top, N.; Keskin, E.; Ünal, E.M.; Britton, J.R. Invasion of pumpkinseed Lepomis gibbosus is facilitated by phenotypic plasticity across its invasion gradient. Biol. Invasions 2021, 23, 3201–3214. [Google Scholar] [CrossRef]
- Çolak, H.S. Metazoan parasites of fish species from Lake Sığırcı (Edirne, Türkiye). Turk. J. Vet. Anim. Sci. 2013, 37, 200–205. [Google Scholar]
- Soylu, E. Metazoan parasites of fish species from Lake Gala (Edirne, Türkiye). Ege J. Fish Aqua. Sci. 2014, 31, 187–193. [Google Scholar]
- Ercan, D.; Andreou, D.; Sana, S.; Öntaş, C.; Baba, E.; Top, N.; Karakuş, U.; Tarkan, A.S.; Gozlan, R.E. Evidence of threat to European economy and biodiversity following introduction of an alien pathogen on the fungal-animal boundary. Emerg. Microbes. Infect. 2015, 4, e52. [Google Scholar] [CrossRef]
- Innal, D.; Stavrescu-Bedivan, M.M. A review of current knowledge on parasites of non-indigenous fish species in the inland waters of Türkiye. Transylv. Rev. Syst. Ecol. Res. 2022, 24, 55–74. [Google Scholar]
- Blackburn, T.M.; Ewen, J.G. Parasites as drivers and passengers of human-mediated biological invasions. EcoHealth 2017, 14, 61–73. [Google Scholar] [CrossRef] [PubMed]
- Šimková, A.; Morand, S.; Jobet, E.; Gelnar, M.; Verneau, O. Molecular phylogeny of congeneric monogenean parasites (Dactylogyrus): A case of intrahost speciation. Evolution 2004, 58, 1001–1018. [Google Scholar]
- Beverley-Burton, M. Monogenea and Turbellaria. In Guide to the Parasites of Fishes of Canada, Part I; Margolis, L., Kabata, Z., Eds.; Canadian Special Publication of Fisheries and Aquatic Sciences: Ottawa, ON, Canada, 1984; Volume 74, pp. 5–209. [Google Scholar]
- Roman, E. Parasite fauna of sunfish Lepomis gibbosus (L.), acclimatised in Danube. Doklady Akademii Nauk SSSR 1953, 89, 765–768, (In Russian with English Summary). [Google Scholar]
- Kvach, Y.; Tkachenko, M.; Bartáková, V.; Kutsokon, Y.; Janáč, M.; Demchenko, V.; Ondračková, M. Parasite communities and genetic structure of non-native pumpkinseed, Lepomis gibbosus, in different Black Sea drainages of Ukraine. Knowl. Manag. Aquat. Ecosyst. 2023, 424, 1. [Google Scholar] [CrossRef]
- Rubtsova, N.Y. First Record of Onchocleidus dispar, an alien monogenean from introduced pumpkinseed fish Lepomis gibbosus (Pisces, Centrarchidae) in Ukraine. Sci. Parasitol. 2015, 16, 83–88. [Google Scholar]
- Ondračková, M.; Kvach, Y.; Martens, A.; Jurajda, P. Limited parasite acquisition by non-native Lepomis gibbosus (Actinopterygii: Centrarchidae) at two ponds in the Upper Rhine basin, Germany. J. Helminthol. 2019, 4, 453–460. [Google Scholar] [CrossRef] [PubMed]
- Cech, G.; Sandor, D.; Molnar, K.; Paulus, P.; Papp, M.; Preiszner, B.; Vital, Z.; Varga, A.; Szekely, C. New record of metacercariae of the North American Posthodiplostomum centrarchi (Digenea, Diplostomidae) in pumpkinseed (Lepomis gibbosus) in Hungary. Acta Vet. Hung. 2020, 68, 20–29. [Google Scholar] [CrossRef]
- Locke, S.A.; McLaughlin, D.J.; Marcogliese, D.J. DNA barcodes show cryptic diversity and a potential physiological basis for host specificity among Diplostomoidea (Platyhelminthes: Digenea) parasitizing freshwater fishes in the St. Lawrence River. Canada. Mol. Ecol. 2010, 19, 2813–2827. [Google Scholar] [CrossRef]
- Moravec, F. Parasitic Nematodes of Freshwater Fishes of Europe; Academia: Prague, Czech Republic, 2013; pp. 1–601. [Google Scholar]
- Esposito, A.; Filippi, J.-J.; Gerbaud, C.; Godeaux, Q.; Millot, R.; Agostini, P.-J.; Albertini, C.; Durieux, E.; Foata, J.; Quilichini, Y. Macroparasite Communities with special attention to invasive helminths in European eels Anguilla anguilla from freshwaters and brackish lagoons of a Mediterranean island. Fishes 2023, 8, 375. [Google Scholar] [CrossRef]
- Honcharov, S.L.; Soroka, N.M.; Halat, M.V.; Dubovyi, A.I.; Zhurenko, V.V.; Halushko, I.A. Distribution of the nematodes of the genus Eustrongylides (Nematoda, Dioctophymatidae) in the World. Regul. Mech. Biosyst. 2022, 13, 73–79. [Google Scholar] [CrossRef]

| Locality | SL, mm | TL, mm | W, g | We, g | |
|---|---|---|---|---|---|
| Sarıçay River (n = 21) | M ± sd | 70.4 ± 4.8 | 86.2 ± 5.9 | 11.4 ± 2.6 | 11.0 ± 2.6 |
| Min–max | 61.4–77.4 | 77.3–97.8 | 7.5–16.7 | 7.3–16.3 | |
| Dipsiz-Çine Stream (n = 20) | M ± sd | 48.3 ± 21.8 | 59.0 ± 26.3 | 5.9 ± 11.5 | 5.7 ± 11.2 |
| Min–max | 28.6–110.3 | 35.8–133.9 | 0.7–41.1 | 0.6–39.9 | |
| Davuldere Reservoir (n = 20) | M ± sd | 48.1 ± 6.8 | 59.4 ± 8.1 | 2.9 ± 2.0 | 2.8 ± 2.0 |
| Min–max | 42.7–74.3 | 52.1–90.4 | 1.8–11.3 | 1.6–11.2 | |
| Değirmenköy Reservoir (n = 20) | M ± sd | 57.4 ± 19.4 | 70.4 ± 24.2 | 7.6 ± 9.4 | 7.3 ± 8.9 |
| Min–max | 40.6–98.4 | 50.1–119.7 | 1.6–29.2 | 1.6–28.0 |
| Population | N | NA | I | Ho | He | Uhe | F | HWE | %P |
|---|---|---|---|---|---|---|---|---|---|
| Sarıçay River | 21 | 2.400 | 0.579 | 0.333 | 0.362 | 0.370 | 0.074 | 0.940 | 80 |
| Dipsiz-Çine Stream | 20 | 2.000 | 0.434 | 0.280 | 0.281 | 0.288 | −0.021 | 0.993 | 80 |
| Davuldere Reservoir | 20 | 2.600 | 0.572 | 0.330 | 0.310 | 0.318 | −0.066 | 0.429 | 60 |
| Değirmenköy Reservoir | 20 | 2.400 | 0.658 | 0.450 | 0.427 | 0.438 | −0.080 | 0.520 | 80 |
| Locality | N | Prevalence (in %) | Mean Abundance | Species Richness | Mean IC (MI ± sd) | Shannon | Equitability |
|---|---|---|---|---|---|---|---|
| Sarıçay River | 21 | 100 | 124.5 | 3 | 2.05 ± 0.65 | 0.25 | 0.23 |
| Dipsiz-Çine Stream | 20 | 25 | 0.5 | 5 | 0.30 ± 0.56 | 1.47 | 0.91 |
| Davuldere Reservoir | 20 | 70 | 2.5 | 2 | 0.75 ± 0.54 | 0.10 | 0.14 |
| Değirmenköy Reservoir | 20 | 90 | 8.8 | 7 | 1.70 ± 1.10 | 1.24 | 0.64 |
| Parasite Species | Site | Indices | Sarıçay River | Dipsiz-Çine Stream | Davuldere Reservoir | Değirmenköy Reservoir |
|---|---|---|---|---|---|---|
| MYXOZOA | ||||||
| Myxobolus dechtiari | Gills | P, % | 15.0 | |||
| MI ± sd | 15.3 ± 8.1 | |||||
| (min–max) | (6–20) | |||||
| A | 2.30 | |||||
| MONOGENEA | ||||||
| Onchocleidus dispar | Gills | P, % | 70.0 | 55.0 | ||
| MI ± sd | 3.4 ± 2.3 | 1.6 ± 0.8 | ||||
| (min–max) | (1–8) | (1–3) | ||||
| A | 2.40 | 0.90 | ||||
| DIGENEA | ||||||
| Posthodiplostomum centrarchi | Mesentery, liver, spleen, heart, gonads, muscles, head, eyes, coelom | P, % | 100.0 | 5.0 | 60.0 | |
| MI ± sd | 117.5 ± 48.0 | 3.0 | 7.7 ± 11.4 | |||
| (min–max) | (48–223) | (3) | (1–41) | |||
| A | 117.5 | 0.15 | 4.60 | |||
| NEMATODA | ||||||
| Eustrongylides sp. | Mesentery | P, % | 57.1 | 5.0 | 5.0 | 5.0 |
| MI ± sd | 1.6 ± 0.7 | 1.0 | 1.0 | 1.0 | ||
| (min–max) | (1–3) | (1) | (1) | (1) | ||
| A | 0.9 | 0.05 | 0.05 | 0.05 | ||
| Contracaecum sp. | Mesentery | P, % | 10.0 | 10.0 | ||
| MI ± sd | 2.0 ± 1.4 | 1.0 | ||||
| (min–max) | (1–3) | (1) | ||||
| A | 0.2 | 0.1 | ||||
| Paraquimperia tenerrima | Mesentery | P, % | 47.6 | |||
| MI ± sd | 13.9 ± 21.8 | |||||
| (min–max) | (1–70) | |||||
| A | 6.6 | |||||
| Aquariidae gen. sp. | Intestine wall, mesentery | P, % | 20.0 | |||
| MI ± sd | 3.8 ± 2.4 | |||||
| (min–max) | (1–7) | |||||
| A | 0.75 | |||||
| Nematoda gen. sp. | Liver | P, % | 5.0 | |||
| MI ± sd | 1.0 | |||||
| (min–max) | (1) | |||||
| A | 0.05 | |||||
| COPEPODA Lernaea cyprinacea | Fins, mesentery | P, % | 5.0 | 5.0 | ||
| MI ± sd | 1.0 | 1.0 | ||||
| (min–max) | (1) | (1) | ||||
| A | 0.05 | 0.05 | ||||
| Parasite Species | Sarıçay River | Dipsiz-Çine Stream | Davuldere Reservoir | Değirmenköy Reservoir |
|---|---|---|---|---|
| Myxobolus dechtiari | 0.09 | |||
| Onchocleidus dispar | 0.93 | 0.32 | ||
| Posthodiplostomum centrarchi | 0.49 | 0.17 | 0.35 | |
| Eustrongylides sp. | 0.28 | 0.17 | 0.07 | 0.03 |
| Contracaecum sp. | 0.33 | 0.09 | ||
| Paraquimperia tenerrima | 0.23 | |||
| Aquariidae gen. sp. | 0.31 | |||
| Nematoda gen. sp. | 0.17 | |||
| Lernaea cyprinacea | 0.17 | 0.03 |
| Sarıçay River | Dipsiz-Çine Stream | Davuldere Reservoir | Değirmenköy Reservoir | |
|---|---|---|---|---|
| Sarıçay River | 0.003 | 0.001 | 0.067 | |
| Dipsiz-Çine | 0.500 | 0.034 | 0.076 | |
| Davuldere Reservoir | 0.400 | 0.286 | 0.170 | |
| Değirmenköy Reservoir | 0.400 | 0.667 | 0.444 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kvach, Y.; Tkachenko, M.Y.; Giannetto, D.; Míč, R.; Bartáková, V.; Ağdamar, S.; Saç, G.; Özuluğ, M.; Tarkan, A.S.; Ondračková, M. Low Genetic and Parasite Diversity of Invasive Pumpkinseed Lepomis gibbosus (Centrarchidae) Expanding in Türkiye. Diversity 2024, 16, 272. https://doi.org/10.3390/d16050272
Kvach Y, Tkachenko MY, Giannetto D, Míč R, Bartáková V, Ağdamar S, Saç G, Özuluğ M, Tarkan AS, Ondračková M. Low Genetic and Parasite Diversity of Invasive Pumpkinseed Lepomis gibbosus (Centrarchidae) Expanding in Türkiye. Diversity. 2024; 16(5):272. https://doi.org/10.3390/d16050272
Chicago/Turabian StyleKvach, Yuriy, Maria Yu. Tkachenko, Daniela Giannetto, Robert Míč, Veronika Bartáková, Sevan Ağdamar, Gülşah Saç, Müfit Özuluğ, Ali Serhan Tarkan, and Markéta Ondračková. 2024. "Low Genetic and Parasite Diversity of Invasive Pumpkinseed Lepomis gibbosus (Centrarchidae) Expanding in Türkiye" Diversity 16, no. 5: 272. https://doi.org/10.3390/d16050272
APA StyleKvach, Y., Tkachenko, M. Y., Giannetto, D., Míč, R., Bartáková, V., Ağdamar, S., Saç, G., Özuluğ, M., Tarkan, A. S., & Ondračková, M. (2024). Low Genetic and Parasite Diversity of Invasive Pumpkinseed Lepomis gibbosus (Centrarchidae) Expanding in Türkiye. Diversity, 16(5), 272. https://doi.org/10.3390/d16050272










