Molecular Species Delimitation Using COI Barcodes of Mealybugs (Hemiptera: Pseudococcidae) from Coffee Plants in Espírito Santo, Brazil
Abstract
1. Introduction
2. Materials and Methods
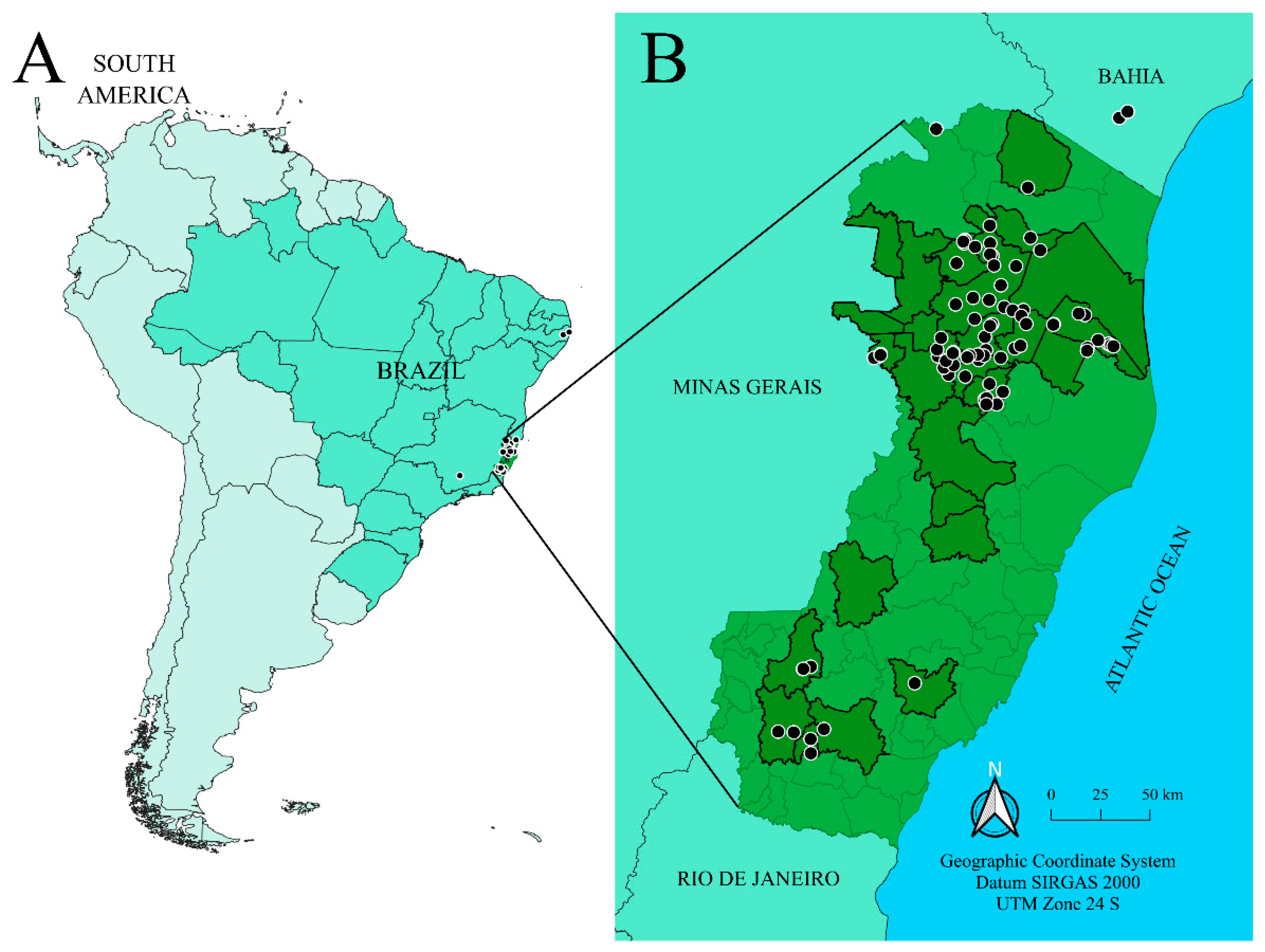
2.1. Sample Collection and Morphological Identification
2.2. DNA Extraction, Amplification, and Sequencing
2.3. Data Assembly and Analysis
2.4. Phylogenetic Reconstruction and Species-Delimitation Methods
3. Results and Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cure, J.R.; Rodríguez, D.; Gutierrez, A.P.; Ponti, L. The Coffee Agroecosystem: Bio-Economic Analysis of Coffee Berry Borer Control (Hypothenemus hampei). Sci. Rep. 2020, 10, 12262. [Google Scholar] [CrossRef] [PubMed]
- Davis, A.P.; Mieulet, D.; Moat, J.; Sarmu, D.; Haggar, J. Arabica-like Flavour in a Heat-Tolerant Wild Coffee Species. Nat. Plants 2021, 7, 413–418. [Google Scholar] [CrossRef] [PubMed]
- Jaramillo, J.; Muchugu, E.; Vega, F.E.; Davis, A.; Borgemeister, C.; Chabi-Olaye, A. Some Like It Hot: The Influence and Implications of Climate Change on Coffee Berry Borer (Hypothenemus hampei) and Coffee Production in East Africa. PLoS ONE 2011, 6, e24528. [Google Scholar] [CrossRef] [PubMed]
- Jayakumar, M.; Rajavel, M.; Surendran, U.; Gopinath, G.; Ramamoorthy, K. Impact of Climate Variability on Coffee Yield in India—With a Micro-Level Case Study Using Long-Term Coffee Yield Data of Humid Tropical Kerala. Clim. Chang. 2017, 145, 335–349. [Google Scholar] [CrossRef]
- Martins, M.Q.; Fortunato, A.S.; Rodrigues, W.P.; Partelli, F.L.; Campostrini, E.; Lidon, F.C.; DaMatta, F.M.; Ramalho, J.C.; Ribeiro-Barros, A.I. Selection and Validation of Reference Genes for Accurate RT-QPCR Data Normalization in Coffea Spp. under a Climate Changes Context of Interacting Elevated [CO2] and Temperature. Front. Plant Sci. 2017, 8, 307. [Google Scholar] [CrossRef]
- Vega, F.E.; Rosenquist, E.; Collins, W. Global Project Needed to Tackle Coffee Crisis. Nature 2003, 425, 343. [Google Scholar] [CrossRef]
- Correa, L.R.; Souza, B.; Santa-Cecília, L.V.C.; Prado, E. Estudos biológicos de cochonilhas do gênero Planococcus (Hemiptera: Pseudococcidae) em diferentes hospedeiros. Arq. Inst. Biol. 2011, 78, 233–240. [Google Scholar] [CrossRef]
- Läderach, P.; Ramirez–Villegas, J.; Navarro-Racines, C.; Zelaya, C.; Martinez–Valle, A.; Jarvis, A. Climate Change Adaptation of Coffee Production in Space and Time. Clim. Chang. 2017, 141, 47–62. [Google Scholar] [CrossRef]
- Pham, Y.; Reardon-Smith, K.; Mushtaq, S.; Cockfield, G. The Impact of Climate Change and Variability on Coffee Production: A Systematic Review. Clim. Chang. 2019, 156, 609–630. [Google Scholar] [CrossRef]
- Chemura, A.; Mudereri, B.T.; Yalew, A.W.; Gornott, C. Climate Change and Specialty Coffee Potential in Ethiopia. Sci. Rep. 2021, 11, 8097. [Google Scholar] [CrossRef]
- Culik, M.P.; Martins, D.D.S.; Gullan, P.J. First Records of Two Mealybug Species in Brazil and New Potential Pests of Papaya and Coffee. J. Insect Sci. 2006, 6, 23. [Google Scholar] [CrossRef] [PubMed]
- Santa-Cecília, L.V.C.; Souza, B.; de Souza, J.C.; Prado, E.; Junior, A.M.; Fornazier, M.J.; Carvalho, G.A. Cochonilhas-Farinhentas em Cafeeiros: Bioecologia, Danos e Métodos de Controle; Epamig: Belo Horizonte, Brazil, 2007. [Google Scholar]
- Costa, M.B.; Souza, B.; Santa-Cecília, L.V.C.; Prado, E. Tabela de Vida de Fertilidade de Planococcus citri (Risso) e Planococcus minor (Maskell) (Hemiptera: Pseudococcidae) Em Cafeeiro. Coffee Sci. 2016, 11, 204–210. [Google Scholar]
- Rondelli, V.M.; Peronti, A.L.B.G.; Dias, J.R.M.; Fogaça, I.; Dos Santos, I.L.V.; Nery, A.G. New Records of Mealybugs (Hemiptera: Pseudococcidae) Infesting Rosettes of Conilon Coffee Plants in the State of Rondônia, South-Western Amazon, Brazil. Fla. Entomol. 2018, 101, 705. [Google Scholar] [CrossRef]
- He, Y.B.; Wan, X.W.; Liu, Y.H.; Sun, G.M.; Zhan, R.L. Mitochondrial COI from Dysmicoccus brevipes (Hemiptera: Pseudococcidae) Suggests Cryptic Lineage and Pinpoints the Source of the Introduction to China. Fla. Entomol. 2012, 95, 183–191. [Google Scholar] [CrossRef]
- Pacheco da Silva, V.C.; Bertin, A.; Blin, A.; Germain, J.-F.; Bernardi, D.; Rignol, G.; Botton, M.; Malausa, T. Molecular and Morphological Identification of Mealybug Species (Hemiptera: Pseudococcidae) in Brazilian Vineyards. PLoS ONE 2014, 9, e103267. [Google Scholar] [CrossRef]
- Daane, K.M.; Middleton, M.C.; Sforza, R.F.H.; Kamps-Hughes, N.; Watson, G.W.; Almeida, R.P.P.; Correa, M.C.G.; Downie, D.A.; Walton, V.M. Determining the Geographic Origin of Invasive Populations of the Mealybug Planococcus ficus Based on Molecular Genetic Analysis. PLoS ONE 2018, 13, e0193852. [Google Scholar] [CrossRef] [PubMed]
- Cid, M.; Fereres, A. Characterization of the Probing and Feeding Behavior of Planococcus citri (Hemiptera: Pseudococcidae) on Grapevine. Ann. Entomol. Soc. Am. 2010, 103, 404–417. [Google Scholar] [CrossRef]
- Correa, M.C.G.; Zaviezo, T.; Le Maguet, J.; Herrbach, E.; Malausa, T. Characterization of Microsatellite DNA Libraries from Three Mealybug Species and Development of Microsatellite Markers for Pseudococcus viburni (Hemiptera: Pseudococcidae). Bull. Entomol. Res. 2014, 104, 213–220. [Google Scholar] [CrossRef]
- Correa, M.C.G.; Lombaert, E.; Malausa, T.; Crochard, D.; Alvear, A.; Zaviezo, T.; Palero, F. Mealybug Species from Chilean Agricultural Landscapes and Main Factors Influencing the Genetic Structure of Pseudococcus viburni. Sci. Rep. 2015, 5, 16483. [Google Scholar] [CrossRef]
- Brahmachari, V.; Kohli, S.; Gulati, P. In Praise of Mealybugs. J. Genet. 2018, 97, 379–389. [Google Scholar] [CrossRef]
- Heya, H.M.; Khamis, F.M.; Onyambu, G.K.; Akutse, K.S.; Mohamed, S.A.; Kimathi, E.K.; Ombura, F.L.O.; Ekesi, S.; Dubois, T.; Subramanian, S.; et al. Characterization and Risk Assessment of the Invasive Papaya Mealybug, Paracoccus marginatus, in Kenya under Changing Climate. J. Appl. Entomol. 2020, 144, 442–458. [Google Scholar] [CrossRef]
- Andrews, K.R.; Gerritsen, A.; Rashed, A.; Crowder, D.W.; Rondon, S.I.; van Herk, W.G.; Vernon, R.; Wanner, K.W.; Wilson, C.M.; New, D.D.; et al. Wireworm (Coleoptera: Elateridae) Genomic Analysis Reveals Putative Cryptic Species, Population Structure, and Adaptation to Pest Control. Commun. Biol. 2020, 3, 489. [Google Scholar] [CrossRef] [PubMed]
- Park, D.-S.; Leem, Y.J.; Hahn, K.-W.; Suh, S.-J.; Hong, K.-J.; Oh, H.-W. Molecular Identification of Mealybugs (Hemiptera: Pseudococcidae) Found on Korean Pears. J. Econ. Entomol. 2010, 103, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Wang, D.; Guo, G.; Hu, Y.; Wei, J.; Liu, J. Species Delimitation of the Dermacentor Ticks Based on Phylogenetic Clustering and Niche Modeling. PeerJ 2019, 7, e6911. [Google Scholar] [CrossRef]
- Malausa, T.; Fenis, A.; Warot, S.; Germain, J.; Ris, N.; Prado, E.; Botton, M.; Vanlerberghe-Masutti, F.; Sforza, R.; Cruaud, C.; et al. DNA Markers to Disentangle Complexes of Cryptic Taxa in Mealybugs (Hemiptera: Pseudococcidae). J. Appl. Entomol. 2011, 135, 142–155. [Google Scholar] [CrossRef]
- Park, D.S.; Suh, S.J.; Hebert, P.D.N.; Oh, H.W.; Hong, K.J. DNA Barcodes for Two Scale Insect Families, Mealybugs (Hemiptera: Pseudococcidae) and Armored Scales (Hemiptera: Diaspididae). Bull. Entomol. Res. 2011, 101, 429–434. [Google Scholar] [CrossRef]
- Amouroux, P.; Crochard, D.; Germain, J.F.; Correa, M.; Ampuero, J.; Groussier, G.; Kreiter, P.; Malausa, T.; Zaviezo, T. Genetic Diversity of Armored Scales (Hemiptera: Diaspididae) and Soft Scales (Hemiptera: Coccidae) in Chile. Sci. Rep. 2017, 7, 2014. [Google Scholar] [CrossRef]
- Oliveira, P.V.; Matos, N.S.; Klippel, A.H.; Oliveira-Costa, J.; Careta, F.D.P.; Paneto, G.G. Using DNA Barcodes to Identify Forensically Important Species of Diptera in Espírito Santo State, Brazil. Braz. Arch. Biol. Technol. 2017, 60, e17160106. [Google Scholar] [CrossRef]
- Pacheco da Silva, V.C.; Kaydan, M.B.; Malausa, T.; Germain, J.-F.; Palero, F.; Botton, M. Integrative Taxonomy Methods Reveal High Mealybug (Hemiptera: Pseudococcidae) Diversity in Southern Brazilian Fruit Crops. Sci. Rep. 2017, 7, 15741. [Google Scholar] [CrossRef]
- Dewer, Y.; Abdel-Fattah, R.S.; Schneider, S.A. Molecular and Morphological Identification of the Mealybug, Phenacoccus solani Ferris (Hemiptera: Pseudococcidae): First Report in Egypt. EPPO Bull. 2018, 48, 155–159. [Google Scholar] [CrossRef]
- Hebert, P.D.N.; Ratnasingham, S.; de Waard, J.R. Barcoding Animal Life: Cytochrome c Oxidase Subunit 1 Divergences among Closely Related Species Barcoding Animal Life: Cytochrome c Oxidase Subunit 1 Divergences among Closely Related Species. Proc. R. Soc. Lond. B 2003, 270, S96–S99. [Google Scholar] [CrossRef] [PubMed]
- Klippel, A.H.; Oliveira, P.V.; Britto, K.B.; Freire, B.F.; Moreno, M.R.; Dos Santos, A.R.; Banhos, A.; Paneto, G.G. Using DNA Barcodes to Identify Road-Killed Animals in Two Atlantic Forest Nature Reserves, Brazil. PLoS ONE 2015, 10, e0134877. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, P.V.; de Almeida, F.A.N.; Lugon, M.D.; Britto, K.B.; Oliveira-Costa, J.; Santos, A.R.; Paneto, G.G. Using High-Resolution Melting to Identify Calliphoridae (Blowflies) Species from Brazil. PeerJ 2020, 8, e9680. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Guo, H.; Han, L.; Chai, J.; Che, X.; Shi, F. Singleton Molecular Species Delimitation Based on COI-5P Barcode Sequences Revealed High Cryptic/Undescribed Diversity for Chinese Katydids (Orthoptera: Tettigoniidae). BMC Evol. Biol. 2019, 19, 79. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Vaamonde, C.; Kirichenko, N.; Cama, A.; Doorenweerd, C.; Godfray, H.C.J.; Guiguet, A.; Gomboc, S.; Huemer, P.; Landry, J.-F.; Laštůvka, A.; et al. Evaluating DNA Barcoding for Species Identification and Discovery in European Gracillariid Moths. Front. Ecol. Evol. 2021, 9, 66. [Google Scholar] [CrossRef]
- Bukowski, B.; Ratnasingham, S.; Hanisch, P.E.; Hebert, P.D.N.; Perez, K.; deWaard, J.; Tubaro, P.L.; Lijtmaer, D.A. DNA Barcodes Reveal Striking Arthropod Diversity and Unveil Seasonal Patterns of Variation in the Southern Atlantic Forest. PLoS ONE 2022, 17, e0267390. [Google Scholar] [CrossRef]
- Martoni, F.; Bulman, S.; Pitman, A.; Taylor, G.; Armstrong, K. DNA Barcoding Highlights Cryptic Diversity in the New Zealand Psylloidea (Hemiptera: Sternorrhyncha). Diversity 2018, 10, 50. [Google Scholar] [CrossRef]
- Dalstein, V.; Eberle, J.; Fabrizi, S.; Etzbauer, C.; Ahrens, D. COI-Based Species Delimitation in Indochinese Tetraserica chafers Reveal Hybridisation despite Strong Divergence in Male Copulation Organs. Org. Divers. Evol. 2019, 19, 277–286. [Google Scholar] [CrossRef]
- Sabadini, C.P.; Machado, C.B.; Vilhena, P.D.S.; Garófalo, C.A.; Del Lama, M.A. Species Delimitation and Phylogenetic Relationships in the Genus Trypoxylon (Hymenoptera: Crabronidae) Using Molecular Markers: An Alternative to Taxonomic Impediment. Syst. Biodivers. 2020, 18, 315–327. [Google Scholar] [CrossRef]
- Martínez-Arce, A.; De Jesús-Navarrete, A.; Leasi, F. DNA Barcoding for Delimitation of Putative Mexican Marine Nematodes Species. Diversity 2020, 12, 107. [Google Scholar] [CrossRef]
- Zhang, H.; Ning, X.; Yu, X.; Bu, W.J. Integrative Species Delimitation Based on COI, ITS, and Morphological Evidence Illustrates a Unique Evolutionary History of the Genus Paracercion (Odonata: Coenagrionidae). PeerJ 2021, 9, e11459. [Google Scholar] [CrossRef] [PubMed]
- Collado, G.A.; Torres-Díaz, C.; Valladares, M.A. Phylogeography and Molecular Species Delimitation Reveal Cryptic Diversity in Potamolithus (Caenogastropoda: Tateidae) of the Southwest Basin of the Andes. Sci. Rep. 2021, 11, 15735. [Google Scholar] [CrossRef] [PubMed]
- Koroiva, R.; Gomes, V.G.N.; Vilela, D.S. DNA Barcoding and New Records of Odonates (Insecta: Odonata) from Paraíba State, Brazil. Diversity 2022, 14, 203. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D.N. BOLD: The Barcode of Life Data System (www.Barcodinglife.Org). Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Sirisena, U.G.A.I.; Watson, G.W.; Hemachandra, K.S.; Wijayagunasekar, H.N.P. A Modified Technique for the Preparation of Specimens of Sternorryncha for Taxonomic Studies. Trop. Agric. Res. 2013, 24, 139–149. [Google Scholar]
- Cox, J.M.; Ben-Dov, Y. Planococcine Mealybugs of Economic Importance from the Mediterranean Basin and Their Distinction from a New African Genus (Hemiptera: Pseudococcidae). Bull. Entomol. Res. 1986, 76, 481–489. [Google Scholar] [CrossRef]
- Miller, D.R.; Gimpel, W.F. Systematic Analysis of the Mealybugs in the Pseudococcus Maritimus Complex (Homoptera: Pseudococcidae); Associated Publishers: Gainesville, FL, USA, 1996; Volume 2. [Google Scholar]
- Granara de Willink, M.C.; Szumik, C. Phenacoccinae de Centro y Sudamérica (Hemiptera:Coccoidea:Pseudococcidae): Sistemática y Filogenia. Rev. Soc. Entomológica Argent. 2007, 66, 29–129. [Google Scholar]
- Granara de Willink, M.C. Dysmicoccus de La Región Neotropical (Hemiptera: Pseudococcidae). Rev. Soc. Entomológica Argent. 2009, 68, 11–95. [Google Scholar]
- Granara de Willink, M.C.; González, P. Revisión Taxonómica de Pseudococcus Westwood (Hemiptera: Pseudococcidae) de Centro y Sud América Con Descripciones de Especies Nuevas. Insecta Mundi 2018, 1775, 1–117. [Google Scholar]
- Kaydan, M.B.; Gullan, P.J. A Taxonomic Revision of the Mealybug Genus Ferrisia Fullaway (Hemiptera: Pseudococcidae), with Descriptions of Eight New Species and a New Genus. Zootaxa 2012, 3543. [Google Scholar] [CrossRef]
- Arseneau, J.R.; Steeves, R.; Laflamme, M. Modified Low-Salt CTAB Extraction of High-Quality DNA from Contaminant-Rich Tissues. Mol. Ecol. Resour. 2017, 17, 686–693. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: A Multiple Sequence Alignment Method with Reduced Time and Space Complexity. BMC Bioinform. 2004, 5, 113. [Google Scholar] [CrossRef] [PubMed]
- Song, H.; Buhay, J.E.; Whiting, M.F.; Crandall, K.A. Many Species in One: DNA Barcoding Overestimates the Number of Species When Nuclear Mitochondrial Pseudogenes Are Coamplified. Proc. Natl. Acad. Sci. USA 2008, 105, 13486–13491. [Google Scholar] [CrossRef]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA Sequence Polymorphism Analysis of Large Data Sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. JModelTest 2: More Models, New Heuristics and Parallel Computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML Version 8: A Tool for Phylogenetic Analysis and Post-Analysis of Large Phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Choi, J.; Lee, S. Higher Classification of Mealybugs (Hemiptera: Coccomorpha) Inferred from Molecular Phylogeny and Their Endosymbionts. Syst. Entomol. 2022, 47, 354–370. [Google Scholar] [CrossRef]
- Drummond, A.J.; Suchard, M.A.; Xie, D.; Rambaut, A. Bayesian Phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 2012, 29, 1969–1973. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A.J.; Xie, D.; Baele, G.; Suchard, M.A. Posterior Summarization in Bayesian Phylogenetics Using Tracer 1.7. Syst. Biol. 2018, 67, 901–904. [Google Scholar] [CrossRef]
- Bouckaert, R.; Heled, J.; Kühnert, D.; Vaughan, T.; Wu, C.H.; Xie, D.; Suchard, M.A.; Rambaut, A.; Drummond, A.J. BEAST 2: A Software Platform for Bayesian Evolutionary Analysis. PLOS Comput. Biol. 2014, 10, e1003537. [Google Scholar] [CrossRef] [PubMed]
- Puillandre, N.; Brouillet, S.; Achaz, G. ASAP: Assemble Species by Automatic Partitioning. Mol. Ecol. Resour. 2021, 21, 609–620. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Kapli, P.; Pavlidis, P.; Stamatakis, A. A General Species Delimitation Method with Applications to Phylogenetic Placements. Bioinformatics 2013, 29, 2869–2876. [Google Scholar] [CrossRef]
- Kapli, P.; Lutteropp, S.; Zhang, J.; Kobert, K.; Pavlidis, P.; Stamatakis, A.; Flouri, T. Multi-Rate Poisson Tree Processes for Single-Locus Species Delimitation under Maximum Likelihood and Markov Chain Monte Carlo. Bioinformatics 2017, 33, 1630–1638. [Google Scholar] [CrossRef]
- Fujisawa, T.; Barraclough, T.G. Delimiting Species Using Single-Locus Data and the Generalized Mixed Yule Coalescent Approach: A Revised Method and Evaluation on Simulated Data Sets. Syst. Biol. 2013, 62, 707–724. [Google Scholar] [CrossRef]
- Kimura, M. A Simple Method for Estimating Evolutionary Rates of Base Substitutions through Comparative Studies of Nucleotide Sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Carstens, B.C.; Pelletier, T.A.; Reid, N.M.; Satler, J.D. How to Fail at Species Delimitation. Mol. Ecol. 2013, 22, 4369–4383. [Google Scholar] [CrossRef] [PubMed]
- Kol-Maimon, H.; Ghanim, M.; Franco, J.C.; Mendel, Z. Evidence for Gene Flow between Two Sympatric Mealybug Species (Insecta; Coccoidea; Pseudococcidae). PLoS ONE 2014, 9, e88433. [Google Scholar] [CrossRef]
- Wang, X.B.; Zhang, J.T.; Deng, J.; Zhou, Q.S.; Zhang, Y.Z.; Wu, S.A. DNA Barcoding of Mealybugs (Hemiptera: Coccoidea: Pseudococcidae) from Mainland China. Ann. Entomol. Soc. Am. 2016, 109, 438–446. [Google Scholar] [CrossRef]
- Rung, A.; Scheffer, S.J.; Evans, G.; Miller, D. Molecular Identification of Two Closely Related Species of Mealybugs of the Genus Planococcus (Homoptera: Pseudococcidae). Entomol. Soc. Am 2008, 101, 525–532. [Google Scholar] [CrossRef]
- Talavera, G.; Dincă, V.; Vila, R. Factors Affecting Species Delimitations with the GMYC Model: Insights from a Butterfly Survey. Methods Ecol. Evol. 2013, 4, 1101–1110. [Google Scholar] [CrossRef]
- Tan, D.S.H.; Ang, Y.; Lim, G.S.; Bin Ismail, M.R.; Meier, R. From “cryptic Species” to Integrative Taxonomy: An Iterative Process Involving DNA Sequences, Morphology, and Behaviour Leads to the Resurrection of Sepsis Pyrrhosoma (Sepsidae: Diptera). Zool. Scr. 2010, 39, 51–61. [Google Scholar] [CrossRef]
- Delrieu-Trottin, E.; Durand, J.; Limmon, G.; Sukmono, T.; Kadarusman; Sugeha, H.Y.; Chen, W.; Busson, F.; Borsa, P.; Dahruddin, H.; et al. Biodiversity Inventory of the Grey Mullets (Actinopterygii: Mugilidae) of the Indo-Australian Archipelago through the Iterative Use of DNA-based Species Delimitation and Specimen Assignment Methods. Evol. Appl. 2020, 13, 1451–1467. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Xie, X.; Huo, L.; Liang, X.; Wang, X.; Chen, X. An Integrative DNA Barcoding Framework of Ladybird Beetles (Coleoptera: Coccinellidae). Sci. Rep. 2020, 10, 10063. [Google Scholar] [CrossRef]
- Song, C.; Lin, X.L.; Wang, Q.; Wang, X.H. DNA Barcodes Successfully Delimit Morphospecies in a Superdiverse Insect Genus. Zool. Scr. 2018, 47, 311–324. [Google Scholar] [CrossRef]
- Koroiva, R.; Rodrigues, L.R.R.; Santana, D.J. DNA Barcoding for Identification of Anuran Species in the Central Region of South America. PeerJ 2020, 8, e10189. [Google Scholar] [CrossRef]
- Bergsten, J.; Bilton, D.T.; Fujisawa, T.; Elliott, M.; Monaghan, M.T.; Balke, M.; Hendrich, L.; Geijer, J.; Herrmann, J.; Foster, G.N.; et al. The Effect of Geographical Scale of Sampling on DNA Barcoding. Syst. Biol. 2012, 61, 851–869. [Google Scholar] [CrossRef]
- Meiklejohn, K.A.; Damaso, N.; Robertson, J.M. Assessment of BOLD and GenBank—Their Accuracy and Reliability for the Identification of Biological Materials. PLoS ONE 2019, 14, e0217084. [Google Scholar] [CrossRef]
- Guimarães, K.L.A.; Lima, M.P.; Santana, D.J.; de Souza, M.F.B.; Barbosa, R.S.; Rodrigues, L.R.R. DNA Barcoding and Phylogeography of the Hoplias Malabaricus Species Complex. Sci. Rep. 2022, 12, 5288. [Google Scholar] [CrossRef]
- Puillandre, N.; Lambert, A.; Brouillet, S.; Achaz, G. ABGD, Automatic Barcode Gap Discovery for Primary Species Delimitation. Mol. Ecol. 2012, 21, 1864–1877. [Google Scholar] [CrossRef]
- Ren, J.M.; Ashfaq, M.; Hu, X.N.; Ma, J.; Liang, F.; Hebert, P.D.N.; Lin, L.; Germain, J.F.; Ahmed, M.Z. Barcode Index Numbers Expedite Quarantine Inspections and Aid the Interception of Nonindigenous Mealybugs (Pseudococcidae). Biol. Invasions 2018, 20, 449–460. [Google Scholar] [CrossRef]
- Wakgari, W.M.; Giliomee, J.H. Natural Enemies of Three Mealybug Species (Hemiptera: Pseudococcidae) Found on Citrus and Effects of Some Insecticides on the Mealybug Parasitoid Coccidoxenoides peregrinus (Hymenoptera: Encyrtidae) in South Africa. Bull. Entomol. Res. 2003, 93, 243–254. [Google Scholar] [CrossRef]
- Lopes, F.S.C.; de Oliveira, J.V.; de Morais Oliveira, J.E.; de Oliveira, M.D.; de Souza, A.M. Host Plants for Mealybugs (Hemiptera: Pseudococcidae) in Grapevine Crops. Pesq. Agropec. Trop. 2019, 49, e54421. [Google Scholar] [CrossRef]
- Ahmed, N.; Abd-Rabou, S. Host Plants, Geographical Distribution, Natural Enemies and Biological Studies of the Citrus Mealybug, Planococcus citri (Risso) (Hemiptera: Pseudococcidae). Egypt. Acad. J. Biol. Sci. A Entomol. 2010, 3, 39–47. [Google Scholar] [CrossRef]
- Fernandes, M.H.D.A.; Oliveira, J.E.D.M.; Costa, V.A.; de Menezes, K.O. Coccidoxenoides perminutus Parasitizing Planococcus citri on Vine in Brazil. Ciência Rural 2016, 46, 1130–1133. [Google Scholar] [CrossRef]
- Mansour, R.; Belzunces, L.P.; Suma, P.; Zappalà, L.; Mazzeo, G.; Grissa-Lebdi, K.; Russo, A.; Biondi, A. Vine and Citrus Mealybug Pest Control Based on Synthetic Chemicals. A Review. Agron. Sustain. Dev. 2018, 38, 37. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oliveira, P.V.; dos Santos, A.R.; Olive, E.L.; Britto, K.B.; de Almeida, F.A.N.; Pacheco da Silva, V.C.; Machado, C.B.; Fornazier, M.J.; Ventura, J.A.; Culik, M.P.; et al. Molecular Species Delimitation Using COI Barcodes of Mealybugs (Hemiptera: Pseudococcidae) from Coffee Plants in Espírito Santo, Brazil. Diversity 2023, 15, 305. https://doi.org/10.3390/d15020305
Oliveira PV, dos Santos AR, Olive EL, Britto KB, de Almeida FAN, Pacheco da Silva VC, Machado CB, Fornazier MJ, Ventura JA, Culik MP, et al. Molecular Species Delimitation Using COI Barcodes of Mealybugs (Hemiptera: Pseudococcidae) from Coffee Plants in Espírito Santo, Brazil. Diversity. 2023; 15(2):305. https://doi.org/10.3390/d15020305
Chicago/Turabian StyleOliveira, Pablo Viana, Alexandre Rosa dos Santos, Emily Lopes Olive, Karolinni Bianchi Britto, Francine Alves Nogueira de Almeida, Vitor Cezar Pacheco da Silva, Carolina Barros Machado, Maurício José Fornazier, José Aires Ventura, Mark Paul Culik, and et al. 2023. "Molecular Species Delimitation Using COI Barcodes of Mealybugs (Hemiptera: Pseudococcidae) from Coffee Plants in Espírito Santo, Brazil" Diversity 15, no. 2: 305. https://doi.org/10.3390/d15020305
APA StyleOliveira, P. V., dos Santos, A. R., Olive, E. L., Britto, K. B., de Almeida, F. A. N., Pacheco da Silva, V. C., Machado, C. B., Fornazier, M. J., Ventura, J. A., Culik, M. P., & Paneto, G. G. (2023). Molecular Species Delimitation Using COI Barcodes of Mealybugs (Hemiptera: Pseudococcidae) from Coffee Plants in Espírito Santo, Brazil. Diversity, 15(2), 305. https://doi.org/10.3390/d15020305






