Evolution and Biogeographic History of Rubyspot Damselflies (Hetaerininae: Calopterygidae: Odonata)
Abstract
1. Introduction
2. Materials and Methods
2.1. Taxon Sampling
2.2. DNA Extraction, Amplification and Sequencing
2.3. Phylogenetic Analyses
2.4. Divergence Dating Analysis
2.5. Ancestral Character State Reconstruction
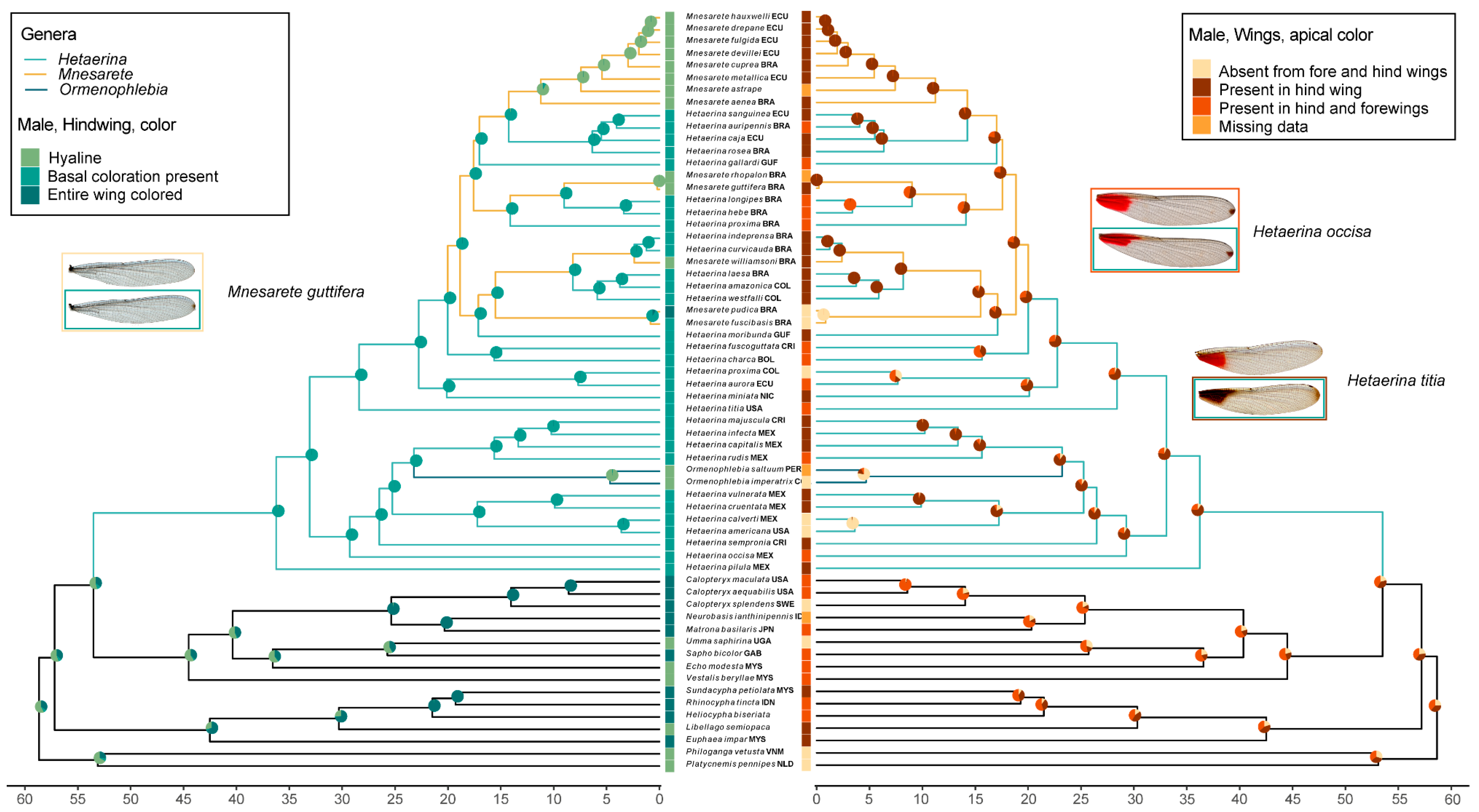
- Male, Hindwing, color: hyaline wings (0), color only basally on wing (1), entire wing colored (2)
- Male, Hindwing, apical color: absent (0), apical melanization only (1), multiple cells colored black or red (2)
- Male, Forewing, apical color: absent (0), apical melanization only (1), multiple cells colored black or red (2)
- Male, apical color: absent from forewings and hindwings (0), present in hindwing (1), present in hindwings and forewings (2). Presence was indicated by any color present, melanization or multiple cells.
- Habitat: generalist (0), forest specialist (1)
2.6. Biogeography Analysis
3. Results
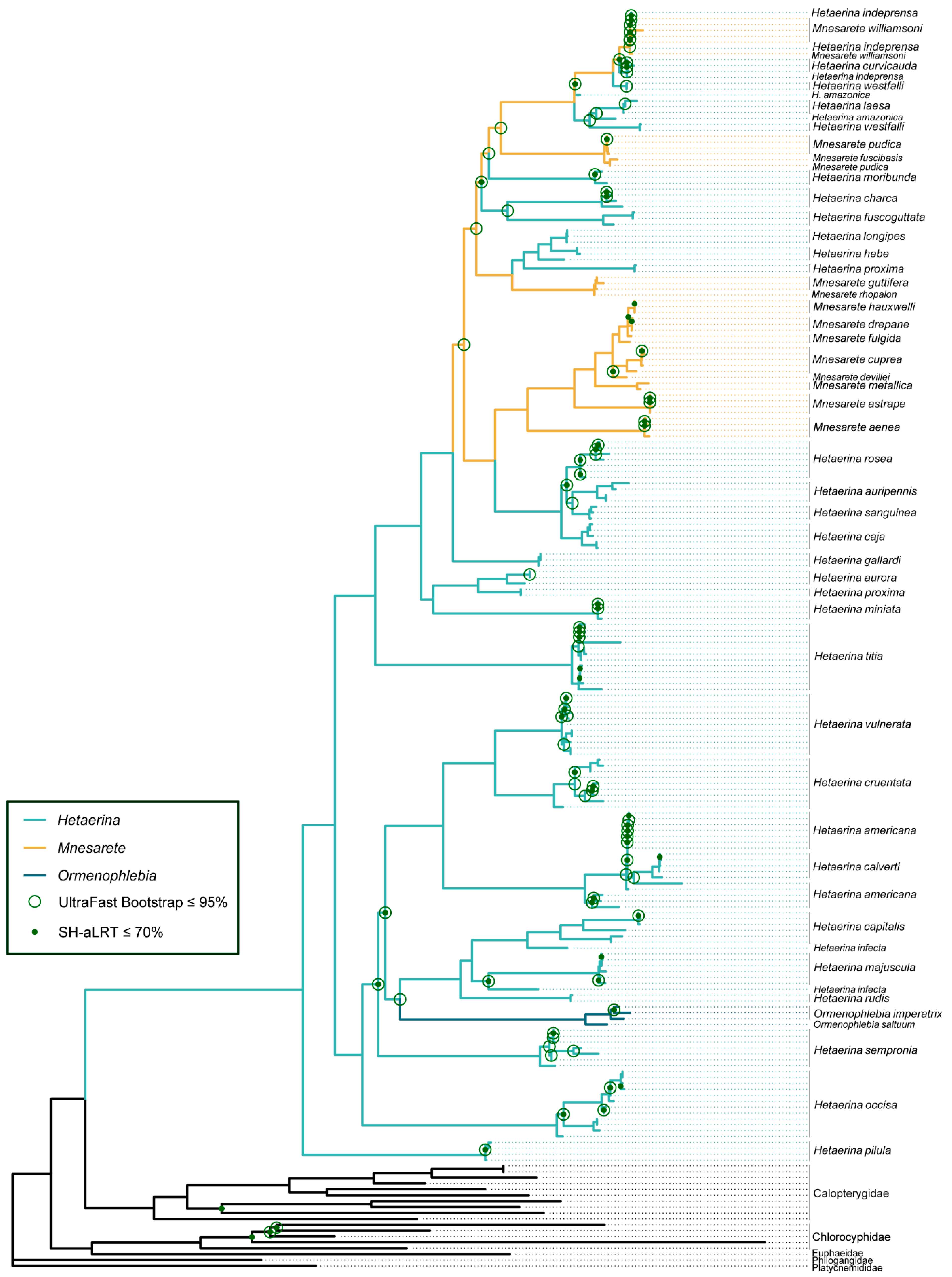
3.1. Phylogenetic Analysis
3.2. Ancestral Character State Reconstruction: Wing Coloration
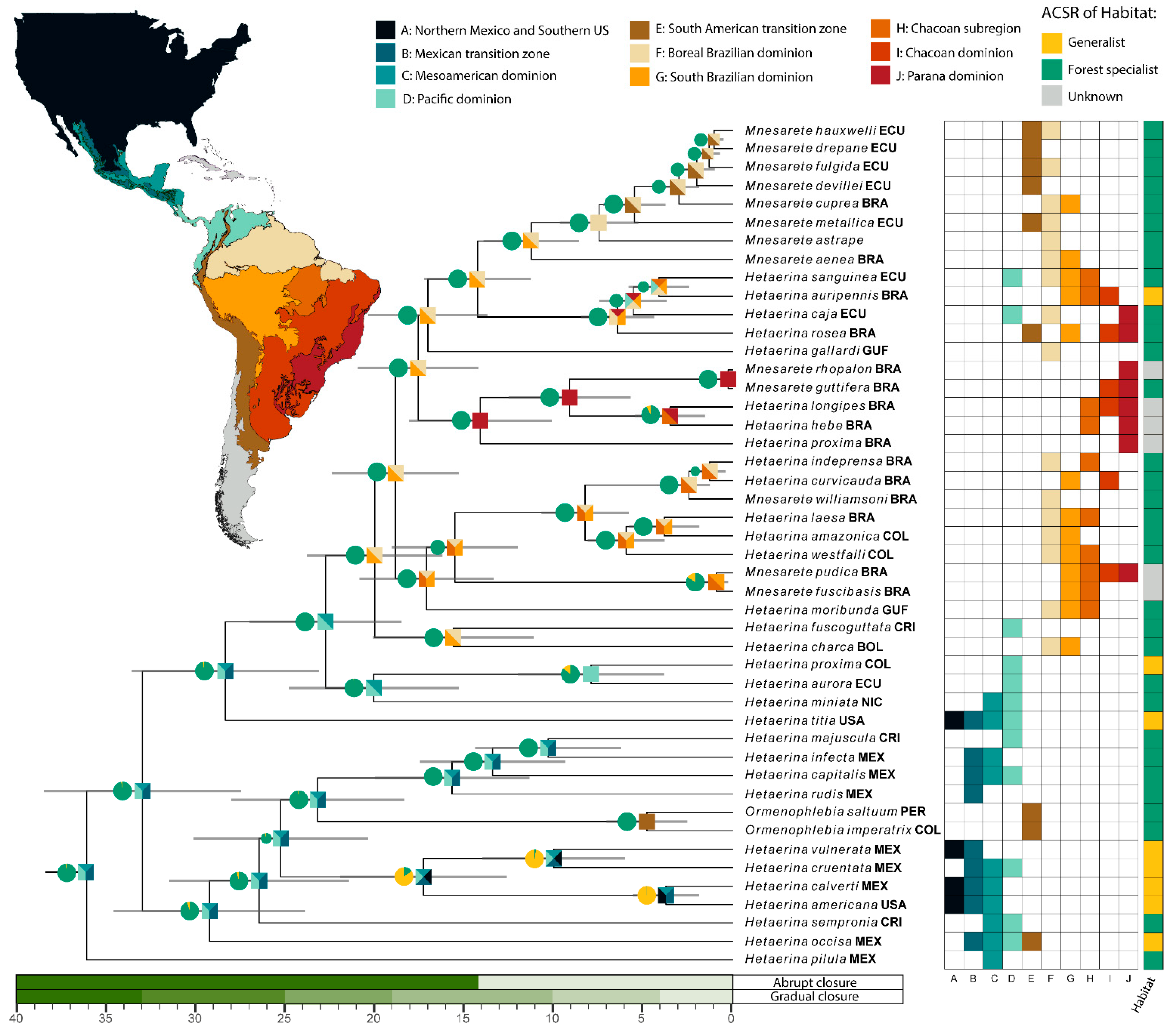
3.3. Ancestral Character State Reconstruction: Habitat
3.4. Divergence Dating and Biogeography
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Misof, B.; Liu, S.; Meusemann, K.; Peters, R.S.; Donath, A.; Mayer, C.; Frandsen, P.B.; Ware, J.; Flouri, T.; Beutel, R.G.; et al. Phylogenomics Resolves the Timing and Pattern of Insect Evolution. Science 2014, 346, 763–767. [Google Scholar] [CrossRef] [PubMed]
- Suárez-Tovar, C.M.; Sarmiento, C.E. Beyond the Wing Planform: Morphological Differentiation between Migratory and Nonmigratory Dragonfly Species. J. Evol. Biol. 2016, 29, 690–703. [Google Scholar] [CrossRef] [PubMed]
- Corbet, P.S. Dragonflies: Behaviour and Ecology of Odonata; Harley Books: Colchester, UK, 1999. [Google Scholar]
- Bybee, S.M.; Kalkman, V.J.; Erickson, R.J.; Frandsen, P.B.; Breinholt, J.W.; Suvorov, A.; Dijkstra, K.D.B.; Cordero-Rivera, A.; Skevington, J.H.; Abbott, J.C.; et al. Phylogeny and Classification of Odonata Using Targeted Genomics. Mol. Phylogenet. Evol. 2021, 160, 107115. [Google Scholar] [CrossRef] [PubMed]
- Garrison, R.W. A Synopsis of the Genus Hetaerina with Descriptions of Four New Species. Trans. Am. Entomol. Soc. 1990, 116, 175–259. [Google Scholar]
- McGirr, R.; Seton, M.; Williams, S. Kinematic and Geodynamic Evolution of the Isthmus of Panama Region: Implications for Central American Seaway Closure. Bull. Geol. Soc. Am. 2021, 133, 867–884. [Google Scholar] [CrossRef]
- Grether, G.F. Sexual Selection and Survival Selection on Wing Coloration and Body Size in the Rubyspot Damselfly Hetaerina Americana. Evolution 1996, 50, 1939–1948. [Google Scholar] [CrossRef]
- Córdoba-Aguilar, A. Adult Survival and Movement in Males of the Damselfly Hetaerina Cruentata (Odonata: Calopterygidae). Fla. Entomol. 1994, 77, 256–264. [Google Scholar] [CrossRef]
- Jaramillo, C.A. Evolution of the Isthmus of Panama: Biological, Palaeoceanographic and Palaeoclimatological Implications. In Mountains, Climate and Biodiversity; Hoorn, C., Perrigo, A., Antonelli, A., Eds.; John Wiley & Sons Ltd.: Hoboken, NJ, USA, 2018; pp. 323–338. [Google Scholar]
- Montes, C.; Cardona, A.; Jaramillo, C.; Pardo, A.; Silva, J.C.; Valencia, V.; Ayala, C.; Pérez-Angel, L.C.; Rodriguez-Parra, L.A.; Ramirez, V.; et al. Middle Miocene Closure of the Central American Seaway. Science 2015, 348, 226–229. [Google Scholar] [CrossRef]
- Dumont, H.J.; Vanfleteren, J.R.; de Jonckheere, J.F.; Weekers, P.H.H. Phylogenetic Relationships, Divergence Time Estimation, and Global Biogeographic Patterns of Calopterygoid Damselflies (Odonata, Zygoptera) Inferred from Ribosomal DNA Sequences. Syst. Biol. 2005, 54, 347–362. [Google Scholar] [CrossRef]
- Mulcahy, D.G.; Noonan, B.P.; Moss, T.; Townsend, T.M.; Reeder, T.W.; Sites, J.W.; Wiens, J.J. Estimating Divergence Dates and Evaluating Dating Methods Using Phylogenomic and Mitochondrial Data in Squamate Reptiles. Mol. Phylogenet. Evol. 2012, 65, 974–991. [Google Scholar] [CrossRef]
- Suvorov, A.; Scornavacca, C.; Fujimoto, M.S.; Bodily, P.; Clement, M.; Crandall, K.A.; Whiting, M.F.; Schrider, D.R.; Bybee, S.M. Deep Ancestral Introgression Shapes Evolutionary History of Dragonflies and Damselflies. Syst. Biol. 2022, 71, 526–546. [Google Scholar] [CrossRef] [PubMed]
- Kohli, M.; Letsch, H.; Greve, C.; Béthoux, O.; Deregnaucourt, I.; Liu, S.; Zhou, X.; Donath, A.; Mayer, C.; Podsiadlowski, L.; et al. Evolutionary History and Divergence Times of Odonata (Dragonflies and Damselflies) Revealed through Transcriptomics. iScience 2021, 24, 103324. [Google Scholar] [CrossRef] [PubMed]
- Alcock, J. Post-Copulatory Mate Guarding by Males of the Demselfly Hetaerina Vulnerata Selys (Odonata: Calopterygidae). Anim. Behav. 1982, 30, 99–107. [Google Scholar] [CrossRef]
- Grether, G.F.; Grey, R.M. Novel Cost of a Sexually Selected Trait in the Rubyspot Damselfly Hetaerina Americana: Conspicuousness to Prey. Behav. Ecol. 1996, 7, 465–473. [Google Scholar] [CrossRef]
- Córdoba-Aguilar, A.; Serrano-Meneses, M.A.; Contreras-Garduño, J.; Raihani, G. The Lek Mating System of Hetaerina Damselflies (Insecta: Calopterygidae). Behaviour 2009, 146, 189–207. [Google Scholar] [CrossRef]
- Guillermo-Ferreira, R.; Del-Claro, K. Resource Defense Polygyny by Hetaerina Rosea Selys (Odonata: Calopterygidae): Influence of Age and Wing Pigmentation. Neotrop. Entomol. 2011, 40, 78–84. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.N.; Grether, G.F. Multiple Routes to Reduced Interspecific Territorial Fighting in Hetaerina Damselflies. Behav. Ecol. 2011, 22, 527–534. [Google Scholar] [CrossRef]
- Contreras-Garduño, J.; Villanueva, G.; Alonso-Salgado, A. Phenoloxidase Production: The Importance of Time after Juvenile Hormone Analogue Administration in Hetaerina Americana (Fabricius) (Zygoptera: Calopterygidae). Odonatologica 2012, 41, 1–88. [Google Scholar]
- Córdoba-Aguilar, A.; González-Tokman, D.M. The Behavioral and Physiological Ecology of Adult Rubyspot Damselflies (Hetaerina, Calopterygidae, Odonata). Adv. Study Behav. 2014, 46, 311–341. [Google Scholar] [CrossRef]
- Grether, G.F. Intrasexual Competition Alone Favors a Sexually Dimorphic Ornament in the Rubyspot Damselfly Hetaerina Americana. Evolution 1996, 50, 1949–1957. [Google Scholar] [CrossRef]
- Grether, G.F. Survival Cost of an Intrasexually Selected Ornament in a Damselfly. Proc. R. Soc. Lond. 1997, 264, 207–210. [Google Scholar] [CrossRef]
- Garrison, R.W. A Synopsis of the Genera Mnesarete Cowley, Bryoplathanon Gen. Nov., and Ormenophlebia Gen. Nov. (Odonata: Calopterygidae). Contrib. Sci. 2006, 50, 61–84. [Google Scholar]
- Dijkstra, K.D.B.; Kalkman, V.J.; Dow, R.A.; Stokvis, F.R.; van Tol, J. Redefining the Damselfly Families: A Comprehensive Molecular Phylogeny of Zygoptera (Odonata). Syst. Entomol. 2014, 39, 68–96. [Google Scholar] [CrossRef]
- Weekers, P.H.H.; de Jonckheere, J.F.; Dumont, H.J. Phylogenetic Relationships Inferred from Ribosomal ITS Sequences and Biogeographic Patterns in Representatives of the Genus Calopteryx (Insecta: Odonata) of the West Mediterranean and Adjacent West European Zone. Mol. Phylogenet. Evol. 2001, 20, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Misawa, K.; Kuma, K.I.; Miyata, T. MAFFT: A Novel Method for Rapid Multiple Sequence Alignment Based on Fast Fourier Transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast Model Selection for Accurate Phylogenetic Estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef]
- Yu, G. Using Ggtree to Visualize Data on Tree-Like Structures. Curr. Protoc. Bioinform. 2020, 69, e96. [Google Scholar] [CrossRef]
- Paradis, E.; Schliep, K. Ape 5.0: An Environment for Modern Phylogenetics and Evolutionary Analyses in R. Bioinformatics 2019, 35, 526–528. [Google Scholar] [CrossRef] [PubMed]
- Wickham, H.; Hester, J.; Chang, W. Devtools: Tools to Make Developing R Packags Easier. R Package Version 1.1.6. 2018. Available online: https://CRAN.R-project.org/package=devtools (accessed on 12 April 2022).
- Revell, L.J. Phytools: An R Package for Phylogenetic Comparative Biology (and Other Things). Methods Ecol. Evol. 2012, 3, 217–223. [Google Scholar] [CrossRef]
- Bouckaert, R.; Vaughan, T.G.; Barido-Sottani, J.; Duchêne, S.; Fourment, M.; Gavryushkina, A.; Heled, J.; Jones, G.; Kühnert, D.; de Maio, N.; et al. BEAST 2.5: An Advanced Software Platform for Bayesian Evolutionary Analysis. PLoS Comput. Biol. 2019, 15, e1006650. [Google Scholar] [CrossRef] [PubMed]
- Ho, S.Y.W.; Lo, N. The Insect Molecular Clock. Aust. J. Entomol. 2013, 52, 101–105. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A.J.; Xie, D.; Baele, G.; Suchard, M.A. Posterior Summarization in Bayesian Phylogenetics Using Tracer 1.7. Syst. Biol. 2018, 67, 901–904. [Google Scholar] [CrossRef]
- Parham, J.F.; Donoghue, P.C.J.; Bell, C.J.; Calway, T.D.; Head, J.J.; Holroyd, P.A.; Inoue, J.G.; Irmis, R.B.; Joyce, W.G.; Ksepka, D.T.; et al. Best Practices for Justifying Fossil Calibrations. Syst. Biol. 2012, 61, 346–359. [Google Scholar] [CrossRef]
- Lin, Q.B.; Petrulevičius, J.F.; Huang, D.Y.; Nel, A.; Engel, M.S. First Fossil Calopterygoidea (Odonata: Zygoptera) from Southeastern Asia: A New Genus and Species from the Paleogene of China. Geobios 2010, 43, 349–353. [Google Scholar] [CrossRef]
- NEL, A.; BRISAC, P. Découverte Du Premier Calopteryx Fossile Du Miocène de l’Ardèche (Odonota, Calopterygidae). Bull. Soc. Entomol. Fr. 1994, 99, 27–30. [Google Scholar] [CrossRef]
- Steininger, F.F.; Berggren, W.A.; Kent, D.V.; Bernor, R.L.; Sen, S.; Agusti, J. Circum-Mediterranean Neogene (Miocene and Pliocene) Marine-Continental Chronologic Correlations of European Mammal Units. In The Evolution of Western Eurasian Neogene Mammal Faunas; Bernor, R.L., Fahlbusch, V., Mitmann, H.-W., Eds.; Columbia University Press: New York, NY, USA, 1996; pp. 7–46. [Google Scholar]
- Nel, A.; Petrulevičius, J.F. Afrotropical and Nearctic Genera of Odonata in the French Oligocene: Biogeographic and Paleoclimatic Implications (Insecta: Calopterygidae, Aeshnidae). Ann. Soc. Entomol. Fr. 2010, 46, 228–236. [Google Scholar] [CrossRef][Green Version]
- Nel, A.; Gross, M.; Engel, M.S. First Fossil Occurrence of the Jewel Damselflies (Odonata: Chlorocyphidae): A New Species from the Late Miocene of Styria, Austria. Ann. Soc. Entomol. Fr. 2017, 53, 280–285. [Google Scholar] [CrossRef]
- Renner, S.; Périco, E.; Ely, G.J.; Sahlén, G. Preliminary Dragonfly (Odonata) Species List from the Pampa Biome in Rio Grande Do Sul, Brazil, with Ecological Notes for 19 New Records for the State. Biota Neotrop. 2017, 17, e20170374. [Google Scholar] [CrossRef]
- Vilela, D.S.; Koroiva, R.; Tosta, T.H.A.; Novaes, M.C.; Guillermo-Ferreira, R. Dragonflies and Damselflies from the West of Minas Gerais, Brazil: Checklist and New Records. Biota Neotrop. 2020, 20, e20190851. [Google Scholar] [CrossRef]
- Bota-Sierra, C.A.; Flórez-V, C.; Escobar, F.; Sandoval-H, J.; Novelo-Gutiérrez, R.; Londoño, G.A.; Cordero-Rivera, A. The Importance of Tropical Mountain Forests for the Conservation of Dragonfly Biodiversity: A Case from the Colombian Western Andes. Int. J. Odonatol. 2021, 24, 233–247. [Google Scholar] [CrossRef]
- Abbot, J.C.; Bota-Sierra, C.A.; Guralnick, R.; Kalkman, V.; González-Soriano, E.; Novelo-Gutiérrez, R.; Bybee, S.M.; Ware, J.L.; Belitz, M.W. Diversity of Nearctic Dragonflies and Damselflies (Odonata). Diversity 2022, 14, 575. [Google Scholar] [CrossRef]
- Matzke, N.J. Probabilistic Historical Biogeography: New Models for Founder-Event Speciation, Imperfect Detection, and Fossils Allow Improved Accuracy and Model-Testing; University of California: Berkeley, CA, USA, 2013. [Google Scholar]
- Morrone, J.J.; Escalante, T.; Rodriguez-Tapia, G.; Carmona, A.; Arana, M.; Mercado-Gomez, J.D. Biogeographic Regionalization of the Neotropical Region: New Map and Shapefile. An. Acad. Bras. Cienc. 2022, 94, e20211167. [Google Scholar] [CrossRef] [PubMed]
- Iturralde-Vinent, M.A.; MacPhee, R.D.E. Paleogeography of the Caribbean Region: Implications for Cenozoic Biogeography. Bull. Am. Mus. Nat. Hist. 1999, 238, 1–72. [Google Scholar]
- Matzke, N.J. Statistical Comparison of DEC and DEC+J Is Identical to Comparison of Two ClaSSE Submodels, and Is Therefore Valid. J. Biogeogr. 2021, 1–26. [Google Scholar] [CrossRef]
- Wesselingh, F.P.; Salo, J.A. A Miocene Perspective on the Evolution of the Amazonian Biota. Scr. Geol. 2006, 133, 439–458. [Google Scholar]
- Hoorn, C.; Wesselingh, F.P.; ter Steege, H.; Bermudez, M.A.; Mora, A.; Sevink, J.; Sanmartín, I.; Sanchez-Meseguer, A.; Anderson, C.L.; Figueiredo, J.P.; et al. Amazonia through Time: Andean Uplift, Climate Change, Landscape Evolution, and Biodiversity. Science 2010, 330, 927–931. [Google Scholar] [CrossRef]
- Kotthoff, U.; Greenwood, D.R.; McCarthy, F.M.G.; Müller-Navarra, K.; Prader, S.; Hesselbo, S.P. Late Eocene to Middle Miocene (33 to 13 Million Years Ago) Vegetation and Climate Development on the North American Atlantic Coastal Plain (IODP Expedition 313, Site M0027). Clim. Past 2014, 10, 1523–1539. [Google Scholar] [CrossRef]
- Vega-Sánchez, Y.M.; Mendoza-Cuenca, L.F.; Gonzálezrodríguez, A. Hetaerina Calverti (Odonata: Zygoptera: Calopterygidae) Sp. Nov., a New Cryptic Species of the American Rubyspot Complex. Zootaxa 2020, 4766, 485–497. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.N.; Grether, G.F. Character Displacement in the Fighting Colours of Hetaerina Damselflies. Proc. R. Soc. B Biol. Sci. 2010, 277, 3669–3675. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.N.; Grether, G.F. Interspecific Aggression and Character Displacement of Competitor Recognition in Hetaerina Damselflies. Proc. Biol. Sci. R. Soc. 2010, 277, 549–555. [Google Scholar] [CrossRef] [PubMed]
- González-Tokman, D.; Córdoba-Aguilar, A.; González-Santoyo, I.; Lanz-Mendoza, H. Infection Effects on Feeding and Territorial Behaviour in a Predatory Insect in the Wild. Anim. Behav. 2011, 81, 1185–1194. [Google Scholar] [CrossRef]
- Guillermo-Ferreira, R.; Bispo, P.C. Male and Female Interactions during Courtship of the Neotropical Damselfly Mnesarete Pudica (Odonata: Calopterygidae). Acta Ethol. 2012, 15, 173–178. [Google Scholar] [CrossRef]
- Drury, J.P.; Grether, G.F. Interspecific Aggression, Not Interspecific Mating, Drives Character Displacement in the Wing Coloration of Male Rubyspot Damselflies (Hetaerina). Proc. R. Soc. B Biol. Sci. 2014, 281, 20141737. [Google Scholar] [CrossRef]
- Grether, G.F.; Drury, J.P.; Berlin, E.; Anderson, C.N. The Role of Wing Coloration in Sex Recognition and Competitor Recognition in Rubyspot Damselflies (Hetaerina Spp.). Ethology 2015, 121, 674–685. [Google Scholar] [CrossRef]
- Drury, J.P.; Anderson, C.N.; Castillo, M.B.C.; Fisher, J.; McEachin, S.; Grether, G.F. A General Explanation for the Persistence of Reproductive Interference. Am. Nat. 2019, 194, 268–275. [Google Scholar] [CrossRef]
- Grether, G.F.; Drury, J.P.; Okamoto, K.W.; McEachin, S.; Anderson, C.N. Predicting Evolutionary Responses to Interspecific Interference in the Wild. Ecol. Lett. 2020, 23, 221–230. [Google Scholar] [CrossRef]
- Pena-Firme, P.; Guillermo-Ferreira, R. Females of the Red Damselfly Mnesarete Pudica Are Attracted to More Ornamented Males and Attract Rival Males. Sci. Rep. 2020, 10, 14320. [Google Scholar] [CrossRef]

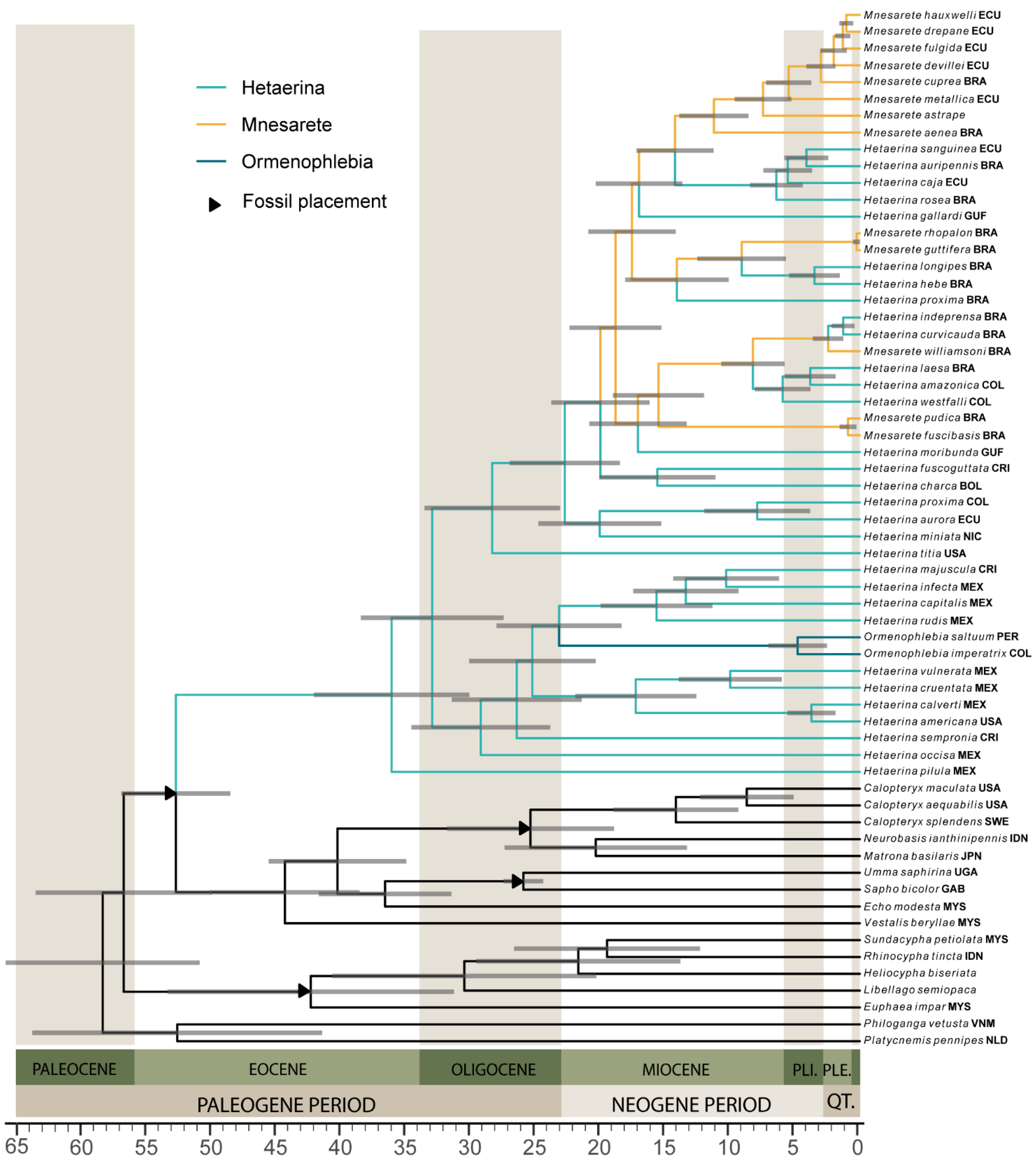
| Fossil | Accession Number | Publication | Fossil Placement | Age and Justification | Prior | Shape | Mean Age | CI |
|---|---|---|---|---|---|---|---|---|
| Calopteryx andancensis | PaleoDB collection 113893 | Nel and Brisac [41] | Node 68 | 9.0–5.3 mya; Steininger et al. [42] | Lognormal | Offset = 5.3 Mean = 13.4 | 25.4 | 19.1, 32.0 |
| Sapho legrandi | PaleoDB collection 194946 | Nel and Petrulevičius [43] | Node 73 | 27.82–24.8 mya; Steininger et al. [42] | Lognormal | Offset = 24.8 Mean = 31.4 | 25.7 | 24.8, 27.9 |
| Chlorocypha cordasevae | PaleoDB collection 105962 | Nel et al. [44] | Node 119 | 11.1–9.4 mya; Steininger et al. [42] | Lognormal | Offset = 9.4 Mean = 13.2 | 42.5 | 31.2, 53.3 |
| Sinocalopteryx shangyongensis | PaleoDB collection 194570 | Lin et al. [40] | Node 65 | 56–5.3 mya | Uniform | Min = 5.3 Max = 56 | 53.5 | 47.6, 56.0 |
| MRCA of Hetaerininae | Node 74 | 36.2 | 30.1, 42.1 | |||||
| North & Central American clade | Node 76 | 29.3 | 24.1, 34.8 | |||||
| South American clade | Node 87 | 28.4 | 23.1, 33.6 | |||||
| Ormenophlebia split | Node 82 | 23.2 | 18.5, 28.1 | |||||
| Ormenophlebia diverged | Node 86 | 4.7 | 2.6, 7.1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Standring, S.; Sánchez-Herrera, M.; Guillermo-Ferreira, R.; Ware, J.L.; Vega-Sánchez, Y.M.; Clement, R.; Drury, J.P.; Grether, G.F.; González-Rodríguez, A.; Mendoza-Cuenca, L.; et al. Evolution and Biogeographic History of Rubyspot Damselflies (Hetaerininae: Calopterygidae: Odonata). Diversity 2022, 14, 757. https://doi.org/10.3390/d14090757
Standring S, Sánchez-Herrera M, Guillermo-Ferreira R, Ware JL, Vega-Sánchez YM, Clement R, Drury JP, Grether GF, González-Rodríguez A, Mendoza-Cuenca L, et al. Evolution and Biogeographic History of Rubyspot Damselflies (Hetaerininae: Calopterygidae: Odonata). Diversity. 2022; 14(9):757. https://doi.org/10.3390/d14090757
Chicago/Turabian StyleStandring, Samantha, Melissa Sánchez-Herrera, Rhainer Guillermo-Ferreira, Jessica L. Ware, Yesenia Margarita Vega-Sánchez, Rebecca Clement, Jonathan P. Drury, Gregory F. Grether, Antonio González-Rodríguez, Luis Mendoza-Cuenca, and et al. 2022. "Evolution and Biogeographic History of Rubyspot Damselflies (Hetaerininae: Calopterygidae: Odonata)" Diversity 14, no. 9: 757. https://doi.org/10.3390/d14090757
APA StyleStandring, S., Sánchez-Herrera, M., Guillermo-Ferreira, R., Ware, J. L., Vega-Sánchez, Y. M., Clement, R., Drury, J. P., Grether, G. F., González-Rodríguez, A., Mendoza-Cuenca, L., Bota-Sierra, C. A., & Bybee, S. (2022). Evolution and Biogeographic History of Rubyspot Damselflies (Hetaerininae: Calopterygidae: Odonata). Diversity, 14(9), 757. https://doi.org/10.3390/d14090757







