A Trait-Based Clustering for Phytoplankton Biomass Modeling and Prediction
Abstract
1. Introduction
2. Materials and Methods
2.1. Description of Data
2.2. Bayesian Model of Cluster-Level Biomass Dynamics
2.3. Description of the Trait-Based Clustering Method
2.3.1. Analyzing the Environmental Drivers of Species Occurrence
2.3.2. Clustering Using GMM and the E-M Algorithm
2.4. Application to the Station L4 Data
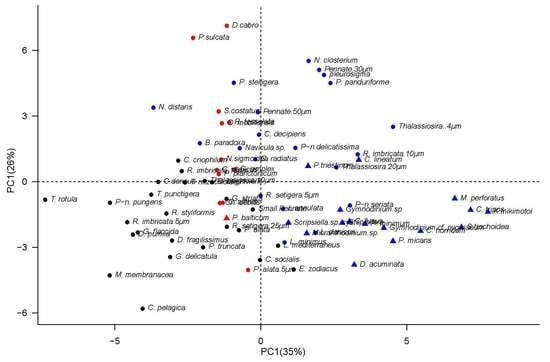
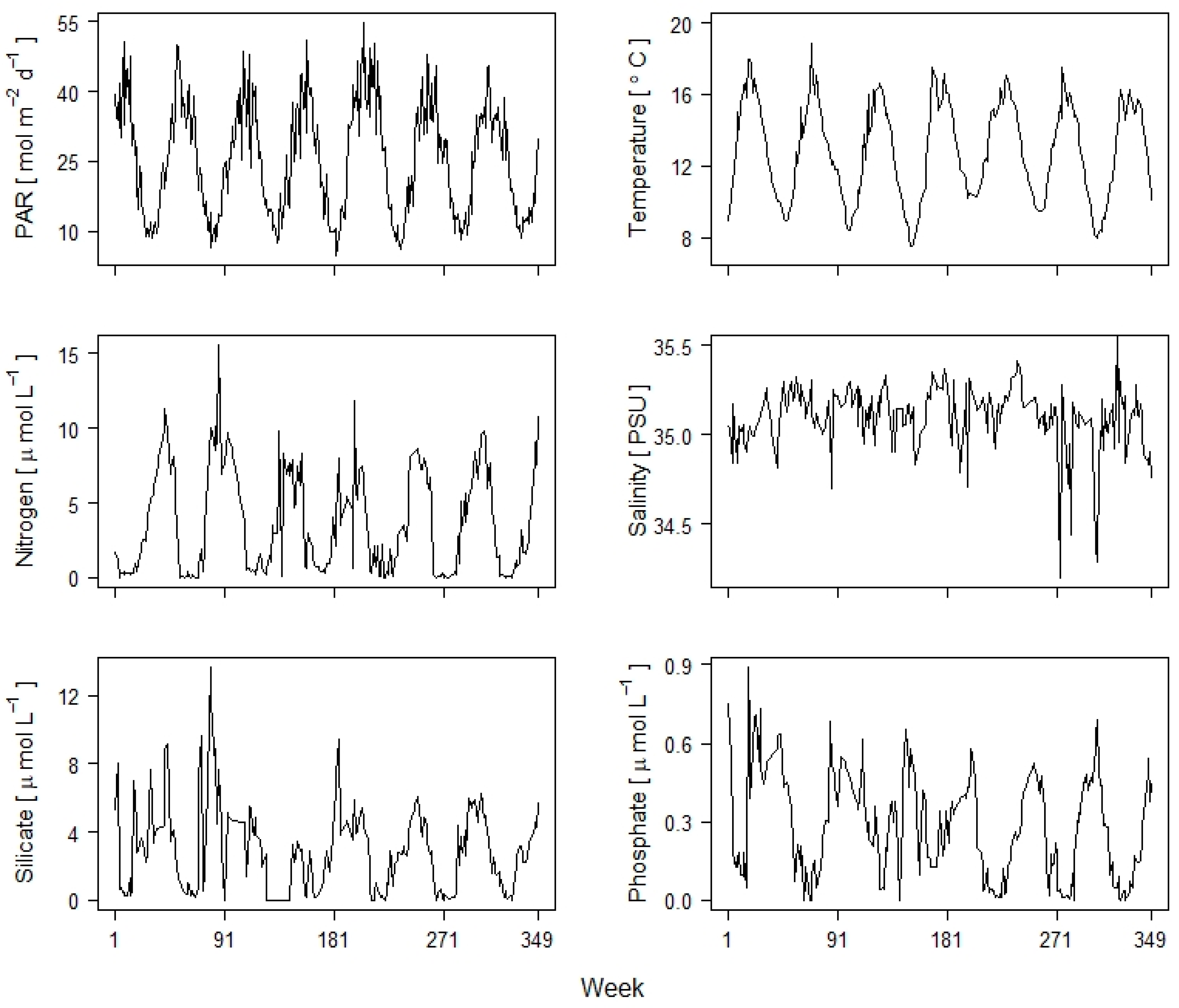
2.4.1. Analyzing the Environmental Controls of Species Occurrence
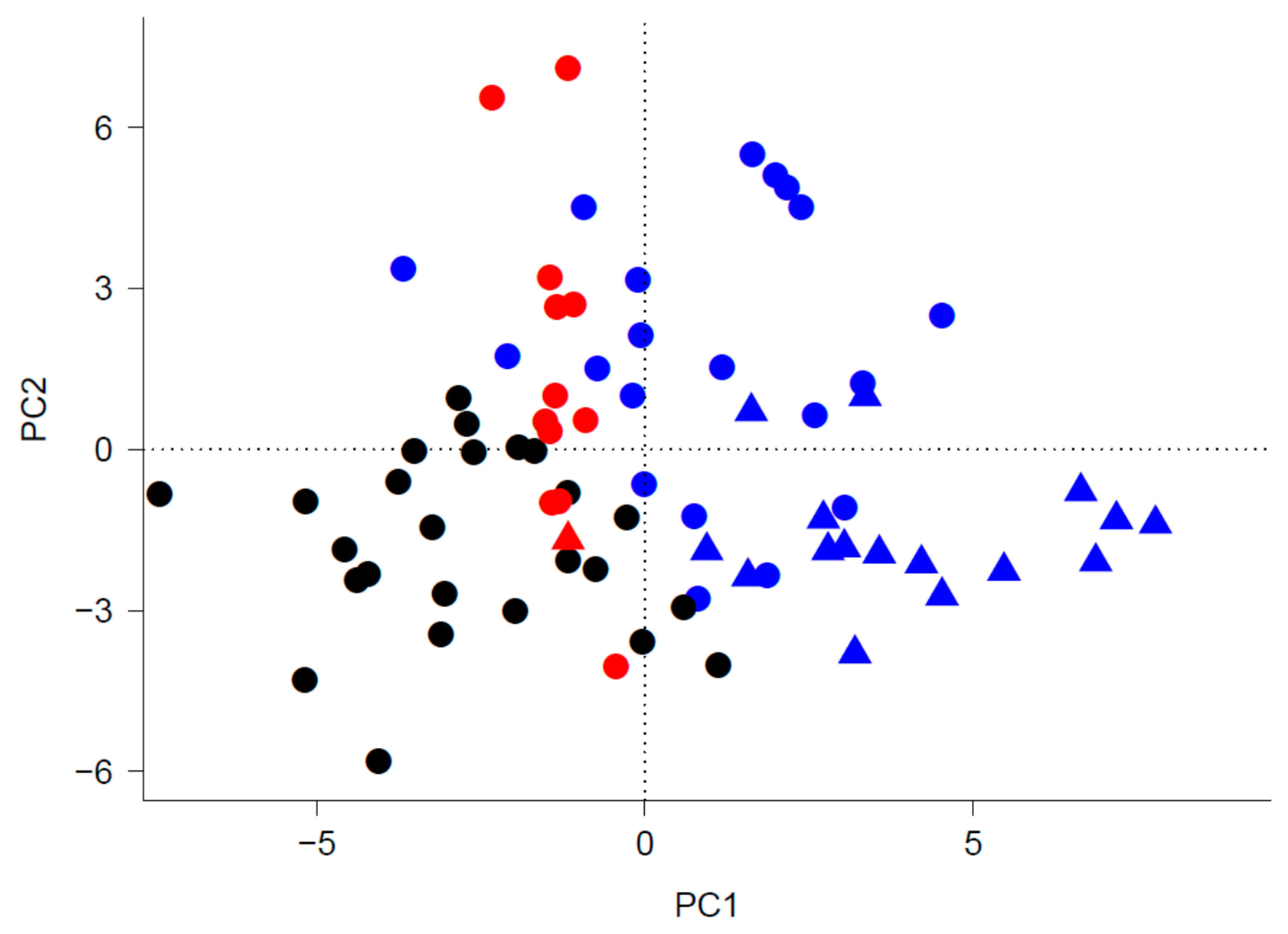
2.4.2. Implementation of the Trait-Based Clustering
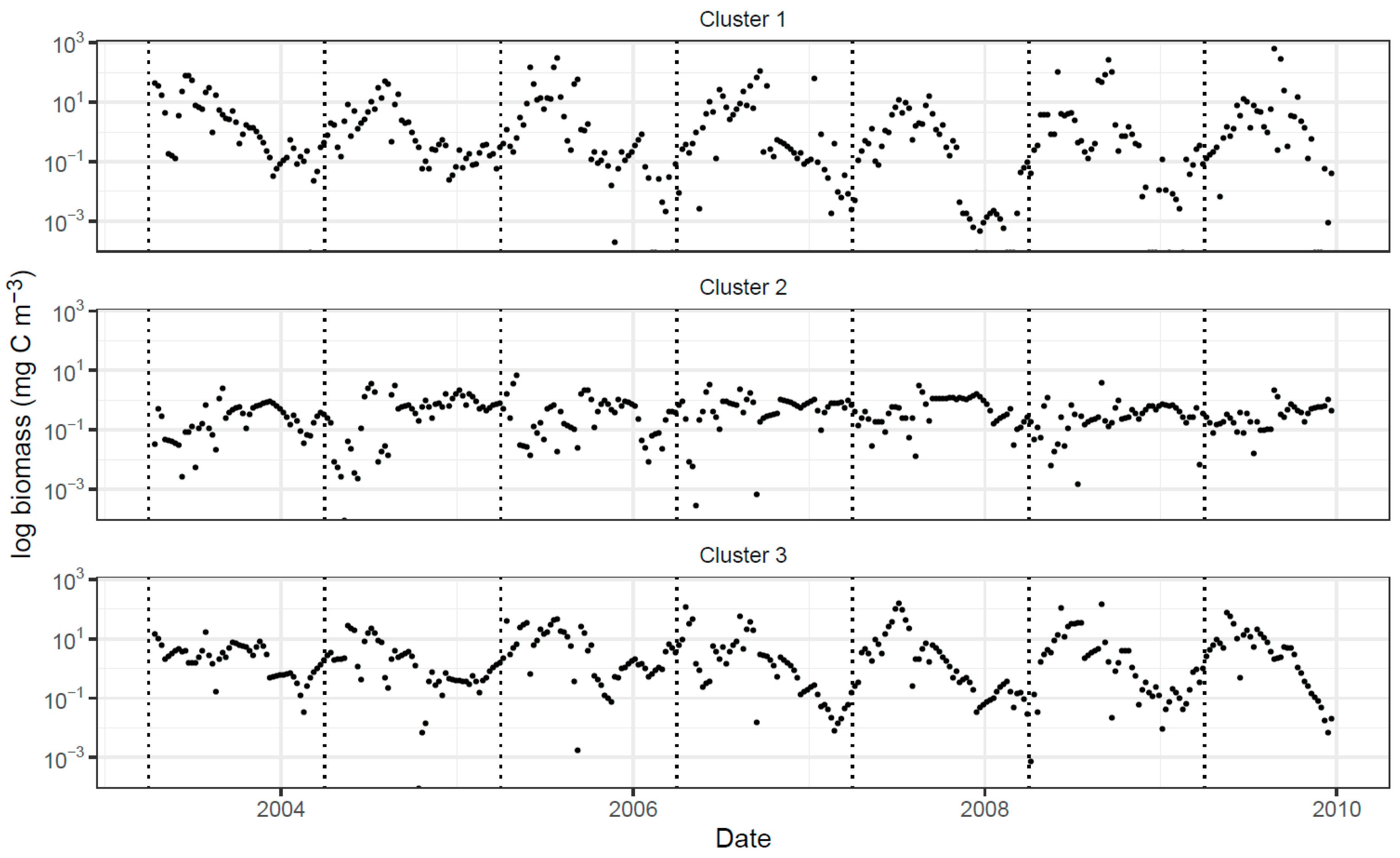
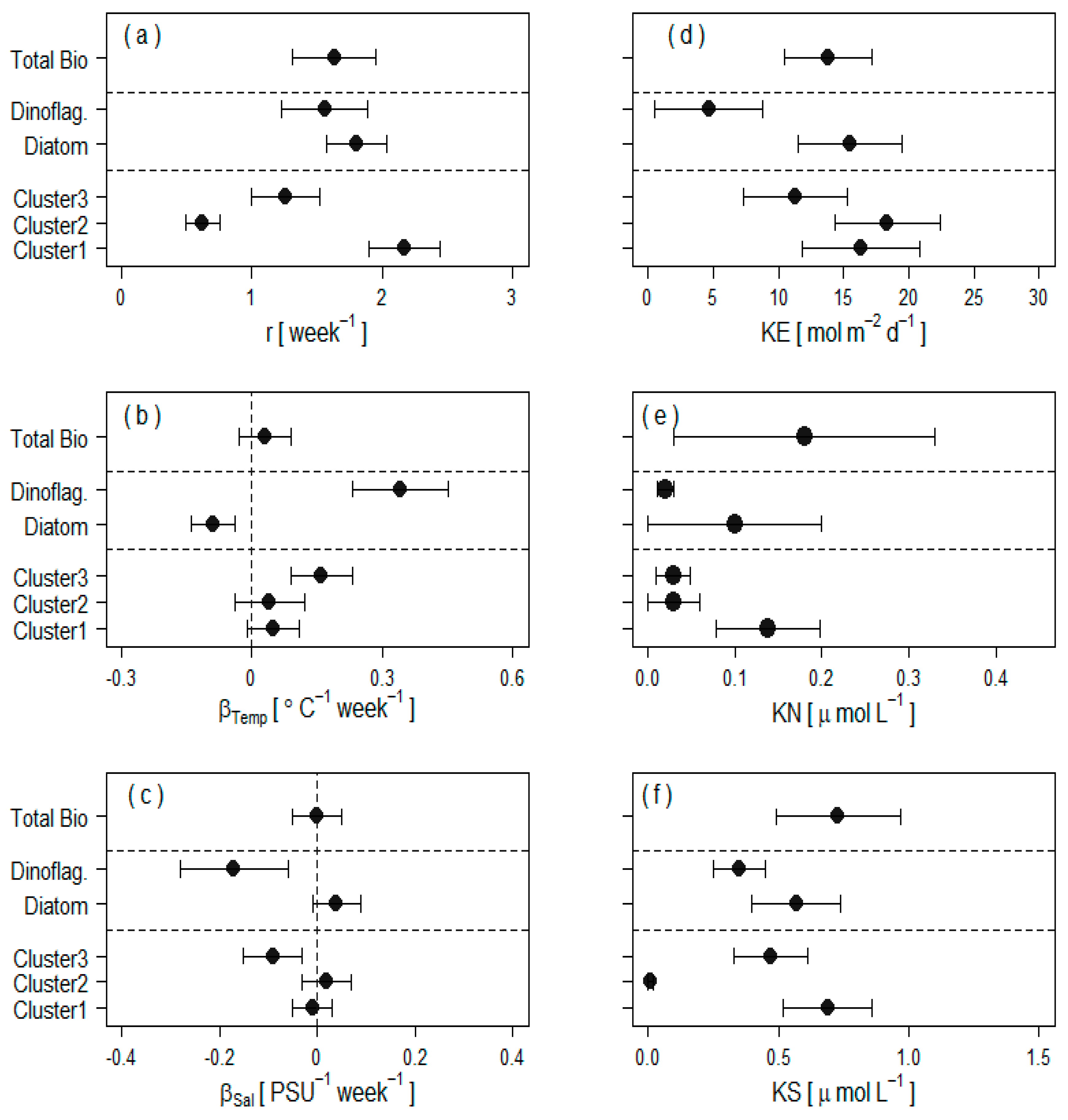
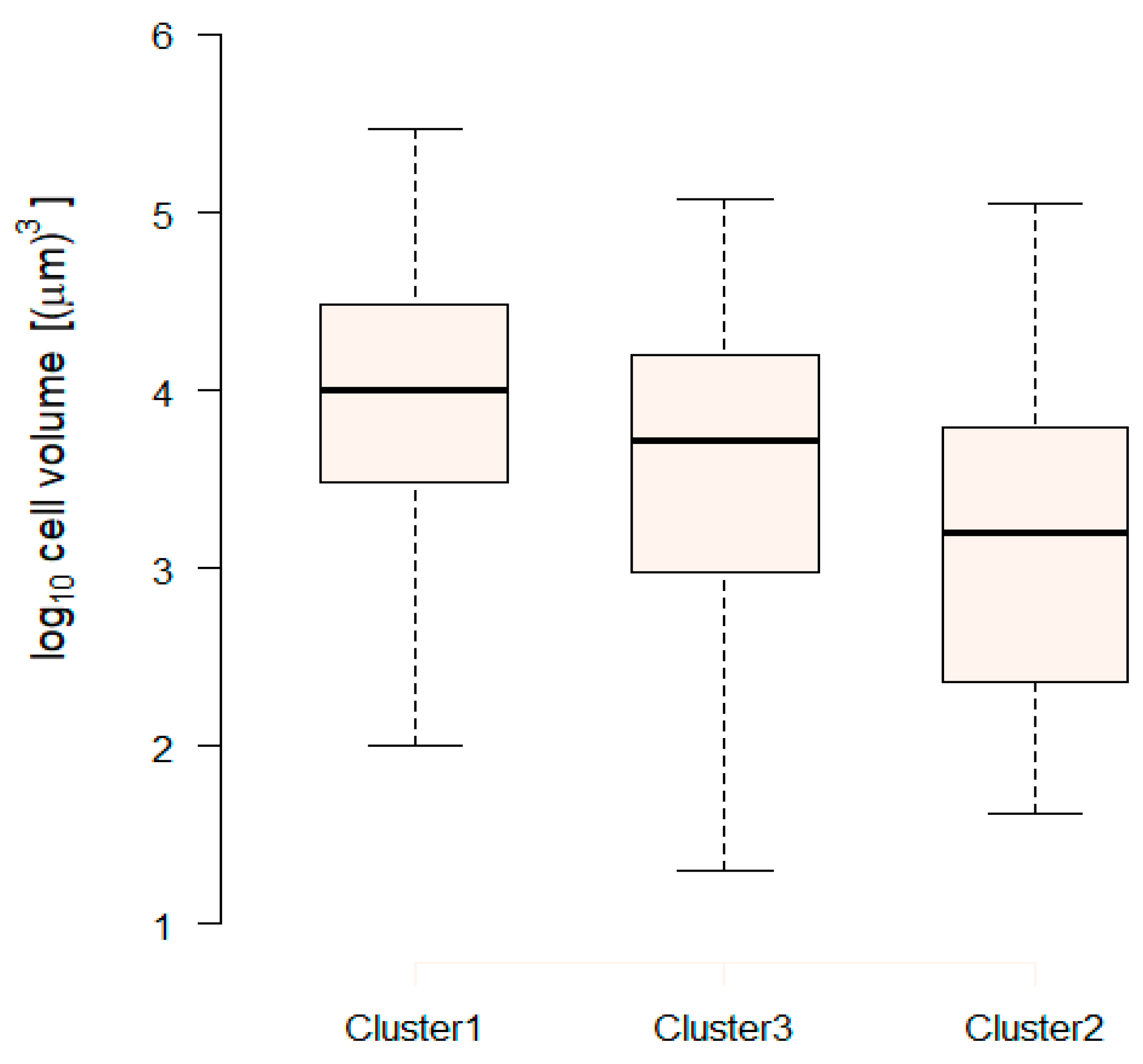
2.4.3. Extraction of Cluster-Specific Trait Values and Total Biomass Prediction
3. Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Species | Functional Type | Cluster Responsibilities | ||
|---|---|---|---|---|
| Cluster 1 | Cluster 2 | Cluster 3 | ||
| Guinardia delicatula | diatom | 0.98 | 0.00 | 0.02 |
| Meuniera membranacea | diatom | 1.00 | 0.00 | 0.00 |
| Cerataulina pelagica | diatom | 1.00 | 0.00 | 0.00 |
| Thalassiosira 10 µm | diatom | 0.70 | 0.18 | 0.12 |
| Eucampia zodiacus | diatom | 1.00 | 0.00 | 0.00 |
| Thalassionema nitzschioides | diatom | 0.98 | 0.00 | 0.02 |
| Guinardia striata | diatom | 0.50 | 0.45 | 0.05 |
| Guinardia flaccida | diatom | 0.98 | 0.00 | 0.02 |
| Dactyliosolen fragilimus | diatom | 0.96 | 0.00 | 0.04 |
| Chaetoceros densus | diatom | 0.86 | 0.00 | 0.14 |
| Corethron criophilum | diatom | 0.59 | 0.00 | 0.41 |
| Ditylum brightwel | diatom | 0.60 | 0.40 | 0.00 |
| Rhizosolenia imbricata 5 µm | diatom | 0.97 | 0.00 | 0.03 |
| Rizosolenia imbricata 15 µm | diatom | 0.68 | 0.00 | 0.32 |
| Thalassiosira rotula | diatom | 0.99 | 0.00 | 0.01 |
| Thalassiosira 20 µm | diatom | 0.76 | 0.00 | 0.24 |
| Rhizoselenia styliformis | diatom | 0.88 | 0.00 | 0.12 |
| Rhizosolenia setigera 25 µm | diatom | 0.77 | 0.23 | 0.00 |
| Pseudo-nitzschia pungens | diatom | 0.99 | 000 | 0.01 |
| Chaetoceros socialis | diatom | 0.64 | 0.22 | 0.14 |
| Thalassiosira punctigera | diatom | 1.00 | 0.00 | 0.00 |
| Small pennate | diatom | 0.87 | 0.11 | 0.02 |
| Proboscia truncata | diatom | 0.99 | 0.01 | 0.00 |
| Leptocylindrus mediterraneus | diatom | 1.00 | 0.00 | 0.00 |
| Proboscia alata | diatom | 0.47 | 0.45 | 0.08 |
| Detonula pumila | diatom | 1.00 | 0.00 | 0.00 |
| Species | Functional Type | Cluster Responsibilities | ||
|---|---|---|---|---|
| Cluster 1 | Cluster 2 | Cluster 3 | ||
| Paralia sulcata | diatom | 0.00 | 0.79 | 0.21 |
| Diplomesis cabro | diatom | 0.00 | 0.78 | 0.22 |
| Chaetoceros debilis | diatom | 0.20 | 0.54 | 0.26 |
| Proboscia alata 5µm | diatom | 0.00 | 0.84 | 0.16 |
| Chaetoceros danicus | diatom | 0.24 | 0.48 | 0.28 |
| Nitzschia sigmoidea | diatom | 0.20 | 0.64 | 0.17 |
| Roperia tesselata | diatom | 0.04 | 0.79 | 0.17 |
| Skeletonema costatum | diatom | 0.03 | 0.95 | 0.02 |
| Chaetoceros affinis | diatom | 0.41 | 0.44 | 0.15 |
| Odontella mobiliensis | diatom | 0.02 | 0.97 | 0.01 |
| Pleurosigma planctonicum | diatom | 0.33 | 0.52 | 0.15 |
| Chaetoceros simplex | diatom | 0.17 | 0.49 | 0.34 |
| Prorocentrum balticum | dinoflagellate | 0.34 | 0.48 | 0.18 |
| Species | Functional Type | Cluster Responsibilities | ||
|---|---|---|---|---|
| Cluster 1 | Cluster 2 | Cluster 3 | ||
| Nitzschia closterium | diatom | 0.00 | 0.00 | 1.00 |
| Pseudo-nitzschia delicatissima | diatom | 0.04 | 0.04 | 0.96 |
| Pleurosigma | diatom | 0.00 | 0.00 | 1.00 |
| Pseudo-nitzchia seriata | diatom | 0.00 | 0.00 | 1.00 |
| Lauderia annulata | diatom | 0.22 | 0.00 | 0.78 |
| Navicula distans | diatom | 0.05 | 0.00 | 0.95 |
| Leptocylindrus danicus | diatom | 0.03 | 0.00 | 0.97 |
| Rhizosolenia setigera 5µm | diatom | 0.17 | 0.04 | 0.79 |
| Navicula sp. | diatom | 0.12 | 0.37 | 0.51 |
| Leptocylindrus minimus | diatom | 0.01 | 0.01 | 0.98 |
| Chaetoceros decipiens | diatom | 0.08 | 0.02 | 0.90 |
| Pennate 50µm | diatom | 0.04 | 0.01 | 0.95 |
| Rhizosolenia imbricata 10µm | diatom | 0.03 | 0.00 | 0.97 |
| Podosira stelligera | diatom | 0.00 | 0.48 | 0.52 |
| Thalassiosira 4µm | diatom | 0.00 | 0.00 | 1.00 |
| Bacillaria paradoxa | diatom | 0.25 | 0.23 | 0.52 |
| Pennate 30µm | diatom | 0.00 | 0.00 | 1.00 |
| Coscinodiscus radiatus | diatom | 0.25 | 0.05 | 0.70 |
| Psammodictyon panduriforme | diatom | 0.00 | 0.00 | 1.00 |
| Ceratium fusus | dinoflagellate | 0.01 | 0.00 | 0.99 |
| Ceratium horridum | dinoflagellate | 0.00 | 0.00 | 1.00 |
| Ceratium lineatum | dinoflagellate | 0.01 | 0.00 | 0.99 |
| Ceratium tripos | dinoflagellate | 0.00 | 0.00 | 1.00 |
| Dinophysis acuminata | dinoflagellate | 0.00 | 0.00 | 1.00 |
| Karenia mikimotoi | dinoflagellate | 0.00 | 0.00 | 1.00 |
| Gonyaulax spinifera | dinoflagellate | 0.00 | 0.00 | 1.00 |
| Gymnodium sp. | dinoflagellate | 0.02 | 0.00 | 0.98 |
| Gymnodium cf. pygmaeum | dinoflagellate | 0.00 | 0.00 | 1.00 |
| Mesoporos perforatus | dinoflagellate | 0.00 | 0.00 | 1.00 |
| Micranthodinium sp. | dinoflagellate | 0.00 | 0.00 | 1.00 |
| Prorocentrum micans | dinoflagellate | 0.00 | 0.00 | 1.00 |
| Prorocentrum minimum | dinoflagellate | 0.16 | 0.00 | 0.84 |
| Prorocentrum triestinum | dinoflagellate | 0.07 | 0.00 | 0.93 |
| Scripsiella trochoidea | dinoflagellate | 0.00 | 0.00 | 1.00 |
| Scripsiella sp. cyst | dinoflagellate | 0.08 | 0.00 | 0.92 |
References
- Sournia, A.; Chretiennot-Dinet, M.-J.; Ricard, M. Marine phytoplankton: How many species in the world ocean? J. Plankton Res. 1991, 13, 1093–1099. [Google Scholar] [CrossRef]
- Tuljapurkar, S.; Caswell, H. Structured Population Models in Marine, Terrestrial and Freshwater Systems; Chapman & Hall: New York, NY, USA, 1997. [Google Scholar]
- Falkowski, P.G.; Katz, M.E.; Knoll, A.H.; Quigg, A.; Raven, J.A.; Schofield, O.; Taylor, F.J.R. The evolution of modern eukaryotic phytoplankton. Science 2004, 305, 354–360. [Google Scholar] [CrossRef]
- Blaum, N.; Mosner, E.; Schwager, M.; Jeltsch, F. How functional is functional? Ecological groupings in terrestrial animal ecology: Towards an animal functional type approach. Biodivers. Conserv. 2011, 20, 2333–2345. [Google Scholar] [CrossRef]
- Yoshio, M.; Yasuhiro, Y.; Takafumi, H.; Hideyuki, N. Competition and community assemblage dynamics within a phytoplankton functional group: Simulation using an eddy-resolving model to disentangle deterministic and random effects. Ecol. Model. 2017, 343, 1–14. [Google Scholar]
- Le Quéré, C.; Harrison, S.P.; Prentice, I.C.; Buitenhuis, E.T.; Aumont, O.; Bopp, L.; Claustre, H.; Cunha, L.C.D.; Geider, R.; Giraud, X.; et al. Ecosystem dynamics based on phytoplankton functional types for global ocean bio-geochemistry models. Glob. Chang. Biol. 2005, 11, 2016–2040. [Google Scholar]
- Irwin, A.J.; Finkel, Z.V. Phytoplankton functional types: A functional trait perspective. In Microbial Ecology of the Ocean; Kirchman, D.M., Gasol, J.M., Eds.; Wiley: Hoboken, NJ, USA, 2016. [Google Scholar]
- Mutshinda, C.M.; Finkel, Z.V.; Widdicombe, C.E.; Irwin, A.J. Phytoplankton traits from long-term oceanographic time-series. Mar. Ecol. Prog. Ser. 2017, 576, 11–25. [Google Scholar] [CrossRef]
- Mutshinda, C.M.; Finkel, Z.V.; Widdicombe, C.E.; Irwin, A.J. Bayesian inference to partition determinants of community dynamics from observational time series. Community Ecol. 2019, 20, 238–251. [Google Scholar] [CrossRef]
- Nogueira, E.; Ibanez, F.; Figueiras, F.G. Effect of meteorological and hydrographic disturbances on the microplankton community structure in the Ría de Vigo (NW Spain). Mar. Ecol. Prog. Ser. 2000, 203, 23–45. [Google Scholar] [CrossRef]
- Bode, A.; Estevez, M.G.; Varela, M.; Vilar, J.A. Annual trend patterns of phytoplankton species abundance belie homogeneous taxonomical group responses to climate in the NE Atlantic upwelling. Mar. Environ. Res. 2015, 110, 81–91. [Google Scholar] [CrossRef]
- Mutshinda, C.M.; Finkel, Z.V.; Irwin, A.J. Which environmental factors control phytoplankton populations? A Bayesian variable selection approach. Ecol. Model. 2013, 269, 1–8. [Google Scholar] [CrossRef]
- Shimoda, Y.; Arhonditsis, G.B. Phytoplankton functional type modelling: Running before we can walk? A critical evaluation of the current state of knowledge. Ecol. Model. 2016, 320, 29–43. [Google Scholar] [CrossRef]
- McGill, B.; Enquist, B.; Weiher, E.; Westoby, M. Rebuilding community ecology from functional traits. Trends Ecol. Evol. 2006, 21, 178–185. [Google Scholar] [CrossRef]
- Litchman, E.; Klausmeier, C.A. Trait-based community ecology of phytoplankton. Ann. Rev. Ecol. Evol. Syst. 2008, 39, 615–639. [Google Scholar] [CrossRef]
- Pomati, F.; Nizzetto, L. Assessing triclosan-induced ecological and trans-generational effects in natural phytoplankton communities: A trait-based field method. Ecotoxicology 2013, 22, 779–794. [Google Scholar] [CrossRef] [PubMed]
- Krause, S.; Le Roux, X.; Niklaus, P.; Van Bodegom, P.M.; Lennon, J.T.; Bertilsson, S.; Grossart, H.-P.; Philippot, L.; Bodelier, P.L.E. Trait-based approaches for understanding microbial biodiversity and ecosystem functioning. Front. Microbiol. 2014, 5, 251. [Google Scholar] [PubMed]
- Kruk, C.; Devercelli, M.; Huszar, V.L.M.; Hernández, E.; Beamud, G.; Diaz, M. Classification of Reynolds phytoplankton functional groups using individual traits and machine learning techniques. Freshw. Biol. 2017, 62, 1681–1692. [Google Scholar] [CrossRef]
- Salguero-Gómez, R.; Violle, C.; Gimenez, O.; Childs, D. Delivering the promises of trait-based approaches to the needs of demographic approaches, and vice versa. Funct. Ecol. 2018, 32, 1424–1435. [Google Scholar] [CrossRef]
- Weithoff, G.; Beisner, B.E. Measures and Approaches in Trait-Based Phytoplankton Community Ecology—From Freshwater to Marine Ecosystems. Front. Mar. Sci. 2019, 6, 40. [Google Scholar] [CrossRef]
- Follows, M.J.; Dutkiewicz, S.; Grant, S.; Chisholm, S.W. Emergent Biogeography of Microbial Communities in a Model Ocean. Science 2007, 315, 1843–1846. [Google Scholar] [CrossRef]
- Widdicombe, C.; Eloire, D.; Harbour, D.; Harris, R.; Somerfield, P. Long-term phytoplankton community dynamics in the Western English Channel. J. Plankton Res. 2010, 32, 643–655. [Google Scholar] [CrossRef]
- Menden-Deuer, S.; Lessard, E.J. Carbon to volume relationships for dinoflagellates, diatoms, and other protest plankton. Limnol. Oceanogr. 2000, 45, 569–579. [Google Scholar] [CrossRef]
- Mutshinda, C.M.; Finkel, Z.V.; Widdicombe, C.E.; Irwin, A.J. Ecological equivalence of species within phytoplankton functional groups. Funct. Ecol. 2016, 30, 1714–1722. [Google Scholar] [CrossRef]
- Mutshinda, C.M.; O’Hara, R.B.; Woiwod, I.P. What drives community dynamics? Proc. R. Soc. Lond. B 2009, 276, 2923–2929. [Google Scholar] [CrossRef]
- Mutshinda, C.M.; O’Hara, R.B.; Woiwod, I.P. A multispecies perspective on ecological impacts of climatic forcing. J. Anim. Ecol. 2011, 80, 101–107. [Google Scholar] [CrossRef] [PubMed]
- Liebig, J. Organic Chemistry in Its Applications to Agriculture and Physiology; Taylor and Walton: London, UK, 1840. [Google Scholar]
- van der Ploeg, R.R.; Kirkham, M. On the origin of the theory of mineral nutrition of plants and the law of the minimum. Soil Sci. Soc. Am. J. 1999, 63, 1055–1062. [Google Scholar] [CrossRef]
- McCarthy, M. Bayesian Methods in Ecology; Cambridge University Press: New York, NY, USA, 2007. [Google Scholar]
- Gelman, A.; Carlin, J.B.; Stern, H.S.; Dunson, D.B.; Vehtari, A.; Rubin, D.B. Bayesian Data Analysis, 3rd ed.; Chapman & Hall: London, UK, 2013. [Google Scholar]
- Gilks, W.R.; Richardson, S.; Spiegelhalter, D.J. Markov Chain Monte Carlo in Practice; Chapman & Hall: London, UK, 1996. [Google Scholar]
- Thomas, A.; O’Hara, R.B.; Ligges, U.; Sturtz, S. Making BUGS open. R News 2006, 6, 12–17. [Google Scholar]
- Mutshinda, C.M. Markov chain Monte Carlo-based Bayesian analysis of binary response regression, with illustration in dose-response assessment. Mod. Appl. Sci. 2009, 3, 19–29. [Google Scholar] [CrossRef][Green Version]
- Dempster, A.P.; Laird, N.M.; Rubin, D.B. Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. B 1977, 39, 1–38. [Google Scholar]
- Redner, R.A.; Walker, H.F. Mixture Densities, Maximum Likelihood and the EM Algorithm. SIAM Rev. 1984, 26, 195–239. [Google Scholar] [CrossRef]
- R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Benaglia, T.; Chauveau, D.; Hunter, D.R.; Young, D. Mixtools: An R Package for Analyzing Finite Mixture Models. J. Stat. Softw. 2009, 32, 1–29. [Google Scholar] [CrossRef]
- Armbrust, E.V. The life of diatoms in the world’s oceans. Nature 2009, 459, 185–192. [Google Scholar] [CrossRef] [PubMed]
- Dodge, J.D. The Prorocentrales (Dinophyceae). II. Revision of the taxonomy within the genus Prorocentrum. Bot. J. Linn. Soc. 1975, 71, 103–125. [Google Scholar] [CrossRef]
- Bailey, R.C.; Norris, R.H.; Reynoldson, T.B. Taxonomic Resolution of Benthic Macroinvertebrate Communities in Bioassessments. J. N. Am. Benthol. Soc. 2001, 20, 280–286. [Google Scholar] [CrossRef]
| Variable | PC1 | PC2 | PC3 |
|---|---|---|---|
| Irradiance (PAR) | 0.66 | −0.19 | 0.53 |
| Temperature | 0.67 | −0.06 | −0.25 |
| Salinity | 0.02 | 0.14 | 0.14 |
| Nitrogen | 0.23 | 0.94 | −0.07 |
| Silicate | 0.11 | −0.19 | −0.02 |
| Phosphate | 0.20 | −0.16 | −0.79 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mutshinda, C.M.; Finkel, Z.V.; Widdicombe, C.E.; Irwin, A.J. A Trait-Based Clustering for Phytoplankton Biomass Modeling and Prediction. Diversity 2020, 12, 295. https://doi.org/10.3390/d12080295
Mutshinda CM, Finkel ZV, Widdicombe CE, Irwin AJ. A Trait-Based Clustering for Phytoplankton Biomass Modeling and Prediction. Diversity. 2020; 12(8):295. https://doi.org/10.3390/d12080295
Chicago/Turabian StyleMutshinda, Crispin M., Zoe V. Finkel, Claire E. Widdicombe, and Andrew J. Irwin. 2020. "A Trait-Based Clustering for Phytoplankton Biomass Modeling and Prediction" Diversity 12, no. 8: 295. https://doi.org/10.3390/d12080295
APA StyleMutshinda, C. M., Finkel, Z. V., Widdicombe, C. E., & Irwin, A. J. (2020). A Trait-Based Clustering for Phytoplankton Biomass Modeling and Prediction. Diversity, 12(8), 295. https://doi.org/10.3390/d12080295