Fungal Diversity in the Phyllosphere of Pinus heldreichii H. Christ—An Endemic and High-Altitude Pine of the Mediterranean Region
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Sites
2.2. Experimental Design and Sampling
2.3. DNA Isolation, Amplificationand Sequencing
2.4. Bioinformatics
2.5. Statistical Analyses
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Janković, M.M. Betrachtungen uber gegenseitigen Bezichungen der Molika (Pinus peuce) und Panzerkiefer (Pinus heldreichii) sowie auch Uber ihre okologishen Eigenschaften, besonders in Bezug auf ihre geologishe Grundlage. Bull. L’Inst. Jard. Bot. L’univ. Beogr. 1960, 1, 141–181. [Google Scholar]
- Jovanović, B. Dendrology, 6th ed.; Faculty of Forestry, University of Belgrade: Belgrade, Serbia, 2007; p. 536. [Google Scholar]
- Blečić, V.; Lakušić, R. Forests of Pinius heldreichii Christ at Štitovo and Bjelasica in Montenegro. Bull. Repub. Inst. Prot. Nat. Mus. Nat. Hist. Titogr. 1969, 4, 5–10. [Google Scholar]
- Stevanović, V.; Jovanović, S.; Lakušić, D. Diversity of vegetation of Yugoslavia. In Biodiversity of Yugoslavia with List of Species of Special Importance; Radović, I., Angelus, J., Eds.; Yugoslavia: Ecolibri, Beograd, 1995; pp. 219–241. [Google Scholar]
- Lazarević, J.; Menkis, A. Fungi inhabiting fine roots of Pinus heldreichii in the Montenegrin montane forests. Symbiosis 2018, 74, 189–197. [Google Scholar] [CrossRef]
- Vendramin, G.G.; Fineschi, S.; Fady, B. EUFORGEN Technical Guidelines for Genetic Conservation and Use for Bosnian Pine (Pinus heldreichii); Bioversity International: Rome, Italy, 2008; ISBN 978-92-9043-789-5. [Google Scholar]
- Baldrian, P. Forest microbiome: Diversity, complexity and dynamics. Fems Mycrobiol. Rev. 2017, 41, 109–130. [Google Scholar] [CrossRef]
- Stewart, J.E.; Kim, M.S.; Klopfenstein, N.B. Molecular genetic approaches toward understanding forest-associated fungi and their interactive roles within forest ecosystems. Curr. For. Rep. 2018, 4, 72–84. [Google Scholar] [CrossRef]
- Zanne, A.E.; Abarenkov, K.; Afkhami, M.E.; Aguilar-Trigueros, C.A.; Bates, S.; Bhatnagar, J.M.; Busby, P.E.; Christian, N.; Cornwell, W.K.; Crowther, T.W.; et al. Fungal functional ecology: Bringing a trait-based approach to plant-associated fungi. Biol. Rev. 2020, 95, 409–433. [Google Scholar] [CrossRef]
- Drenkhan, R.; Tomešová-Haataja, V.; Fraser, S.; Bradshaw, R.; Vahalik, P.; Mullett, M.; Martín-García, J.; Bulman, L.; Wingfield, M.; Kirisits, T.; et al. Global geographic distribution and host range of Dothistroma species: A comprehensive review. For. Pathol. 2016, 46, 408–442. [Google Scholar] [CrossRef]
- Elvira-Recuenco, M.; Cacciola, S.O.; Sanz-Ros, A.V.; Garbelotto, M.; Aguayo, J.; Solla, A.; Mullett, M.; Drenkhan, T.; Oskay, F.; Aday Kaya, A.G.; et al. Potential Interactions between Invasive Fusarium circinatum and Other Pine Pathogens in Europe. Forests 2020, 11, 7. [Google Scholar] [CrossRef]
- Rodriguez, R.J.; Redman, R.S. More than 400 million years of evolution and some plants still can’t make it on their own: Plant stress tolerance via fungal symbiosis. J. Exp. Bot. 2008, 59, 1109–1114. [Google Scholar] [CrossRef]
- Terhonen, E.; Blumenstein, K.; Kovalchuk, A.; Asiegbu, F.O. Forest tree microbiomes and associated fungal endophytes: Functional roles and impact on forest health. Forests 2019, 10, 42. [Google Scholar] [CrossRef]
- Lazarević, J. Contribution to the study of physiological characteristics of the pathogenic fungus Herpotrichia juniperi. Agric. For. 2003, 49, 95–109. [Google Scholar]
- Lazarević, J. Pinus heldreichii as the host of brown felt blight (Herpotrichia juniperi) in Montenegro. Mycol. Monten. 2004, 7, 77–89. [Google Scholar]
- Lazarević, J. Contribution to the study of te morphological characteristics of the pathogenic fungus Herpotrichia juniperi. Mycol. Monten. 2005, 8, 137–147. [Google Scholar]
- Lazarević, J.; Davydenko, K.; Millberg, H. Dothistroma Needle Blight on High Altitude Pine Forests in Montenegro. Balt. For. 2017, 23, 294–302. [Google Scholar]
- Caudullo, G.; Welk, E.; San-Miguel-Ayanz, J. Chorological maps for the main European woody species. Data Brief 2017, 12, 662–666. [Google Scholar] [CrossRef]
- Dinaric_Alps_map-fr.svg: Sémhur, Relief Map of Montenegro svg. Available online: https://commons.wikimedia.org/wiki/File:Relief_Map_of_Montenegro.svg (accessed on 10 January 2020).
- Fuštić, B.; Đuretić, G. Soils of Montenegro; Biotechnical Institute, University of Montenegro: Podgorica, Montenegro, 2000; pp. 156–172. [Google Scholar]
- IUSS Working Group WRB. Descriptions, distribution, use and management of Reference Soil Groups. In World Reference Base for Soil Resources. International Soil Classification System for Naming Soils and Creating Legends for Soil Maps, Udate 2015; World Soil Resource Reports No. 106; Food and Agriculture Organization of the United Nations: Rome, Italy, 2015; pp. 144–180. [Google Scholar]
- Burić, M.; Micev, B.; Mitrović, L. Atlas of Climate of Montenegro; Montenegrin Academy of Science and Arts: Podgorica, Montenegro, 2012; pp. 18–62, 86–102, 126–129. [Google Scholar]
- Burić, D.; Ducić, V.; Mihajlović, J. The climate of Montenegro: Modificators and types-part two. Bull. Serb. Geogr. Soc. 2014, 94, 73–90. [Google Scholar] [CrossRef]
- Janković, M.M.; Stefanović, K. Ecological relations between relict (sub)endemic pine species Pinus heldreichii and the character of geological substrate and soil in Jugoslavia. Ekologija 1971, 6, 49–61. [Google Scholar]
- Menkis, A.; Ihrmark, K.; Stenlid, J.; Vasaitis, R. Root-associated fungi of Rosa rugosa grown on the frontal dunes of the Baltic Sea coast in Lithuania. Microb. Ecol. 2014, 67, 769–774. [Google Scholar] [CrossRef]
- Ihrmark, K.; Bodeker, I.T.M.; Cruz-Martinez, K.; Friberg, H.; Kubartova, A.; Schenck, J.; Strid, Y.; Stenlid, J.; Brandstrom-Durling, M.; Clemmensen, K.E.; et al. New primers to amplify the fungal ITS2 region—Evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol. Ecol. 2012, 82, 666–677. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press, Inc.: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Menkis, A.; Marčiulynas, A.; Gedminas, A.; Lynikienė, J.; Povilaitienė, A. High-throughput sequencing reveals drastic changes in fungal communities in the phyllosphere of Norway spruce (Picea abies) following invasion of the spruce bud scale (Physokermes piceae). Microb. Ecol. 2015, 70, 904–911. [Google Scholar] [CrossRef]
- Lynikienė, J.; Marčiulynienė, D.; Marčiulynas, A.; Gedminas, A.; Vaičiukynė, M.; Menkis, A. Managed and unmanaged Pinus sylvestris forest stands harbour similar diversity and composition of the phyllosphere and soil fungi. Microorganisms 2020, 8, 259. [Google Scholar] [CrossRef]
- Brandström Durling, M.; Clemmensen, K.; Stenlid, J.; Lindahl, B. SCATA—An Efficient Bioinformatic Pipeline for Species Identification and Quantification after High-Throughput Sequencing of Tagged Amplicons (Submitted). 2011. Available online: https://www.scata.mykopat.slu.se (accessed on 12 November 2019).
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [PubMed]
- Analytic Rarefaction, UGA Stratigraphy Lab. Available online: http://www.uga.edu/strata/software/index.html (accessed on 10 January 2020).
- Magurran, A.E. Ecological Diversity and Its Measurement; Princeton University Press: Princeton, NJ, USA, 1988; p. 192. [Google Scholar]
- Sokal, R.R.; Rohlf, F.J. Biometry: The Principles and Practice of Statistics in Biological Research, 4th ed.; H. Freeman & Company: New York, NY, USA, 2005; p. 896. [Google Scholar]
- Shannon, C.E. A mathematical theory of communication. Bell Syst. Tech. J. 1948, 27, 379–423. [Google Scholar]
- Ter Braak, C.J.F.; Smilauer, P. Canoco Reference Manual and User’s Guide to Canoco for Windows: Software for Canonical Community Ordination, Version 4; Microcomputer Power: Ithaca, NY, USA, 1998; p. 351. [Google Scholar]
- Bringel, F.; Couee, I. Pivotal roles of phyllosphere microorganisms at the interface between plant functioning and atmospheric trace gas dynamics. Front. Microbiol. 2015, 6, 486. [Google Scholar] [CrossRef] [PubMed]
- Porazinska, D.L.; Sung, W.; Giblin-Davis, R.M.; Thomas, W.K. Reproducibility of read numbers in high-throughput sequencing analysis of nematode community composition and structure. Mol. Ecol. Resour. 2010, 10, 666–676. [Google Scholar] [CrossRef] [PubMed]
- Behnke-Borowczyk, J.; Kwaśna, H.; Kulawinek, B. Fungi associated with Cyclaneusma needle cast in Scots pine in the west of Poland. For. Pathol. 2019, 49, e12487. [Google Scholar] [CrossRef]
- Lindahl, B.D.; Nilsson, R.H.; Tedersoo, L.; Abarenkov, K.; Carlsen, T.; Kjøller, R.; Kõljalg, U.; Pennanen, T.; Rosendahl, S.; Stenlid, J.; et al. Fungal community analysis by high-throughput sequencing of amplified markers—A user’s guide. New Phytol. 2013, 199, 288–299. [Google Scholar]
- Tedersoo, L.; Tooming-Klunderud, A.; Anslan, S. PacBio metabarcoding of Fungi and other eukaryotes: Errors, biases and perspectives. New Phytol. 2018, 217, 1370–1385. [Google Scholar]
- Millberg, H.; Boberg, J.; Stenid, J. Changes in fungal community of Scots pine (Pinus sylvestris) needles along a latitudinal gradient in Sweden. Fungal Ecol. 2015, 17, 126–139. [Google Scholar] [CrossRef]
- Oono, R.; Lefèvre, E.; Simha, A.; Lutzoni, F. A comparison of the community diversity of foliar fungal endophytes between seedling and adult loblolly pines (Pinus taeda). Fungal Biol. 2015, 119, 917–928. [Google Scholar] [CrossRef]
- Botella, L.; Santamaria, O.; Diez, J.J. Fungi associated with decline of Pinus halepensis in Spain. Fungal Divers. 2010, 40, 1–11. [Google Scholar] [CrossRef]
- Sanz-Ros, A.V.; Muller, M.M.; San Martin, R.; Diez, J.J. Fungal endophytic communities on twigs of fast and slow growing Scots pine (Pinus sylvestris L.) in northern Spain. Fungal Biol. 2015, 119, 870–883. [Google Scholar] [CrossRef]
- Parfitt, D.; Hunt, J.; Dockrell, D.; Rogers, H.J.; Boddy, L. Do all trees carry the seeds of their own destruction? PCR reveals numerous wood decay fungi latently present in sapwood of a wide range of angiosperm trees. Fungal Ecol. 2010, 3, 338–346. [Google Scholar] [CrossRef]
- Bowman, E.A.; Arnold, A.E. Distribution of ectomicorrhizal and foliar endophytic fungal communities associated with Pinus ponderosa along a spatially constrained elevation gradient. Am. J. Bot. 2018, 105, 687–699. [Google Scholar] [CrossRef]
- Botella, L.; Diez, J.J. Phylogenetic diversity of fungal endophytes in Spanish stands of Pinus halepensis. Fungal Divers. 2011, 47, 9–18. [Google Scholar]
- Talgo, V.; Chastanger, G.; Thomsen, I.M.; Cech, T.; Riley, K.; Lang, K.; Klemsdal, S.S. Sydowia polyspora associated with current season needle necrosis (CSNN) on true fir (Abies spp.). Fungal Biol. 2010, 114, 545. [Google Scholar] [CrossRef] [PubMed]
- Karadžić, D.; Milijašević, T. The most important parasitic and saprophytic fungi in Austrian pine and Scots pine plantations in Serbia. Bull. Fac. For. 2008, 97, 147–170. [Google Scholar]
- Tinivella, F.; Dani, E.; Minuto, G.; Minuto, A. First Report of Sydowia polyspora on Aleppo Pine (Pinus halepensis) in Italy. Plant Dis. 2014, 98, 2. [Google Scholar] [CrossRef]
- Ridout, M.; Newcombe, G. Sydowia polyspora is both a Foliar Endophyte and a Preemergent Seed Pathogen in Pinus Ponderosa. Plant Dis. 2018, 118, 3. [Google Scholar] [CrossRef]
- Muñoz-Adalia, E.J.; Sanz-Ros, A.V.; Flores-Pacheco, J.A.; Hantula, J.; Diez, J.J.; Vainio, E.J.; Fernández, M. Sydowia polyspora Dominates Fungal Communities Carried by Two Tomicus Species in Pine Plantations Threatened by Fusarium circinatum. Forests 2017, 8, 127. [Google Scholar] [CrossRef]
- Pan, Y.; Ye, H.; Lu, J.; Chen, P.; Zhou, X.D.; Qiao, M.; Yu, Z.-F. Isolation and identification of Sydowia polyspora and its pathogenicity on Pinus yunnanensis in Southwestern China. J. Phytopatol. 2018, 166, 386–395. [Google Scholar] [CrossRef]
- Dobreva, M.; Georgieva, M.; Dermedzhiev, P.; Nachev, R.; Velinov, V.; Terziev, P.; Georgiev, G. Fungal pathogens associated with Pinus species in the region of forest protection station Plovdiv in the period 2013–2016. For. Sci. 2016, 1–2. [Google Scholar]
- Minter, D.W.; Dudka, I.O. Fungi of Ukraine. A Preliminary Checklist; International Mycological Institute: Egham, Surrey, UK; M.G. Kholodny Institute of Botany: Kiev, Ukraine, 1996; pp. 1–361.
- Drenkhan, R.; Hanso, M. Recent invasion of foliage fungi of pines (Pinus spp.) to the Northern Baltics. For. Stud. 2009, 51, 49–64. [Google Scholar] [CrossRef]
- Markovskaja, S.; Kačergius, A.; Davydenko, K.; Fraser, S. First record of Neocatenulostroma germanicum on pines in Lithuania and Ukraine and its co- occurrence with Dothistroma spp. and other pathogens. Path 2016, 46, 522–533. [Google Scholar] [CrossRef]
- Wajihi, A.H.; Lee, S.; Das, K.; Eom, A.; Jung, H. First Report of Allantophomopsiella pseudotsugae Isolated from Soil in Korea. Korean J. Mycol. 2019, 47, 29–34. [Google Scholar] [CrossRef]
- Crous, P.W.; Quaedvlieg, W.; Hansen, K.; Hawksworth, D.L.; Groenewald, J.Z. Phacidium and Ceuthospora (Phacidiaceae) are congeneric: Taxonomic and nomenclatural implications. Ima Fungus 2014, 5, 173–193. [Google Scholar] [CrossRef]
- Sieber, T.N. Endophytic fungi in forest trees: Are they mutualists? Fungal Biol. Rev. 2007, 21, 75–89. [Google Scholar] [CrossRef]
- Kogel, K.-H.; Franken, P.; Huckelhoven, R. Endophyte or parasite-what decides. Curr. Opin. Plant Biol. 2006, 9, 358–369. [Google Scholar] [CrossRef]
- Soltani, J.; Hosseyni Moghaddam, M.S. Fungal Endophyte Diversity and Bioactivity in the Mediterranean Cypress Cupressus Sempervirens. Curr. Microbiol. 2015, 70, 580–586. [Google Scholar]
- Jeewon, R.; Yeung, Q.S.; Wannasinghe, D.N.; Rampadarath, S.; Puchooa, D.; Wang, H.K.; Hyde, K.D. Hidden mycota of pine needles: Molecular signatures from PCRDGGE and Ribosomal DNA phylogenetic characterization of novel phylotypes. Sci. Rep. 2018, 8, 18053. [Google Scholar] [CrossRef]
- Minter, D.W.; Staley, J.M.; Millar, C.S. Four species of Lophodermium on Pinus sylvestris. Trans. Br. Mycol. Soc. 1978, 71, 295–301. [Google Scholar]
- Reignoux, S.A.; Green, S.; Ennos, R.A. Molecular identification and relative abundance of cryptic Lophodermium species in natural populations of Scots pine, Pinus sylvestris L. Fungal Biol. 2014, 118, 835–845. [Google Scholar] [CrossRef] [PubMed]
- Stierle, A.A.; Stierle, D.B. Bioactive secondary metabolites produced by fungal Endophytes of conifers. Nat. Prod. Commun. 2015, 10, 1671–1682. [Google Scholar] [PubMed]
- Wang, L.W.; Xu, B.G.; Wang, J.Y.; Su, Z.Z.; Lin, F.C.; Zhang, C.L.; Kubicek, C.P. Bioactive metabolites from Phoma species, an endophytic fungus from the Chinese medicinal plant Arisaema erubescens. Appl. Microbiol. Biotechnol. 2012, 93, 1231–1239. [Google Scholar] [CrossRef] [PubMed]
- Johne, S.; Watzke, R.; Meusel, W.; Mollmann, V.; Harti, A.; Dahse, H.M.; Matthes, B.; Seifert, K. Biotechnological production and bioactivities of mollisin and two new, structuraly related fungal naphthequinone metabolites. Chem. Biodivers. 2005, 2, 1109–1115. [Google Scholar] [PubMed]
- Bertscha, C.; Ramırez-Sueroa, M.; Magnin-Robertb, M.; Larignonc, P.; Chonga, J.; Abou-Mansourd, E.; Spagnolob, A.; Clementb, C.; Fontaineb, F. Grapevine trunk diseases: Complex and still poorly understood. Plant Pathol. 2013, 62, 243–265. [Google Scholar] [CrossRef]
- Bien, S.; Kraus, C.; Damm, U. Novel Collophorina-like genera and species from Prunus trees and vineyards in Germany. Persoonia 2020, 45, 46–67. [Google Scholar] [CrossRef]
- Piperkova, N.; Yonkova, I. Symptoms, Etiology and Control of Sooty Blotch and Flyspeck in Bulgaria. Turk. J. Agric. Nat. Sci. 2014, 1, 817–822. [Google Scholar]
- Epidemiology of the Fungus Athelia arachnoidea in Epiphytic Communities of Broadleaved Forests under Strong Anthropogenic Impact. Available online: http://www.elibrary.lt/resursai/LMA/Ekologija/0504_07_Eko.pdf (accessed on 5 March 2020).
- Adams, G.C.; Kropp, B.R. Athelia arachnoidea, the sexual state of Rhizoctonia carotae, a pathogen of carrot in cold storage. Mycologia 1996, 88, 459–472. [Google Scholar] [CrossRef]
- Menkis, A.; Vasiliauskas, R.; Taylor, A.F.S.; Stenlid, J.; Finlay, R. Fungal communities in mycorrhizal roots of conifer seedlings in forest nurseries under different cultivation systems, assessed by morphotyping, direct sequencing and mycelial isolation. Mycorrhiza 2005, 16, 33–41. [Google Scholar]
- Gehring, C.A.; Theimer, T.C.; Whitham, G.; Keim, P. Ectomycorrhizal fungal community structure of pinyon pines growing in two environmental extremes. Ecology 1998, 79, 1562–1572. [Google Scholar]
- Rosa, L.H.; Vaz, A.B.M.; Caligiorne, R.B.; Campolina, S.; Rosa, C.A. Endophytic fungi associaed with Antarctic grass Deschampsia antarctica Devs (Poaceae). Polar Biol. 2009, 32, 161–167. [Google Scholar] [CrossRef]
- Perić, B.; Perić, O. Diversity of macromycetes in Montenegro. MASA 1997, 11, 45–142. [Google Scholar]
- Perić, B.; Perić, O. Preliminary red list of Macromycetes of Montenegro 2. Mycol. Monten. 2004, 7, 7–33. [Google Scholar]
- Lazarević, J.; Perić, O.; Perić, B. Ectomycorrhizal fungi in Montenegro—Diversity and distribution. Mycol. Monten. 2011, 14, 85–115. [Google Scholar]
- Lazarević, J. Forests and biodiversity of Kuči mountains. In Katun of the Kuči Mountains; Laković, I., Ed.; University of Montenegro: Podgorica, Montenegro, 2017; pp. 64–78. [Google Scholar]
- Official Gazette of Montenegro, No 054/16, 15.08.2016. Law of Nature Protection of Montenegro. Available online: http://www.sluzbenilist.me/pregled-dokumenta/?id={0C659042-9DD7-43FB-8E8C-B7DF445298A1} (accessed on 23 March 2020).
- Official Gazette of Montenegro, No 76/06, 12.12.2006, Act on Protection of Certain Plant and Animal Species. 2006. Available online: http://www.sluzbenilist.me/pregled-dokumenta/?id={631C3E5D-4129-4985-B55B-CE4D3703CA2E} (accessed on 23 March 2020).
- Perić, B.; Baral, H.-O. Erioscyphella curvispora, spec. nov. from Montenegro. Mycol. Monten. 2015, 17, 89–104. [Google Scholar]
- Baral, H.-O.; Perić, B. Perzia triseptata Gen. et sp. nov. (Ascomycota, Insertae sedis) from xeric bark in France and Montenegro. Mycol. Monten. 2016, 19, 7–20. [Google Scholar]
- Baral, H.-O.; Perić, B. Velutarina bertiscensis and V. alpestris spp. nov., with a redescription of V. rufoolivacea and discussion of its synonyms. Mycol. Monten. 2015, 17, 17–52. [Google Scholar]
- Perić, B.; Grebenc, T. Une espèce nouvelle du genre Peziza (Pezizales): P. ontirivicola spec. Nov. Ascomycete. Org. 2015, 7, 347–356. [Google Scholar]
- Perić, B.; Baral, H.-O.; Partel, K. Cenangiopsis raghavanii and C. junipericola spp. nov (Cenangiaceae, Helotiales) collected in Montenegro, with redescription of a recent collection of C. quercicola. Mycol. Monten. 2015, 17, 7–40. [Google Scholar]
- Perić, B.; van Vooren, N.; Healy, R.; Lazarević, J. Une Trichophaea rare récoltée en France et au Monténégro: T. Flavobrunnea Comb. Nov. (Pezizales). Mycol. Monten. 2014, 16, 65–87. [Google Scholar]
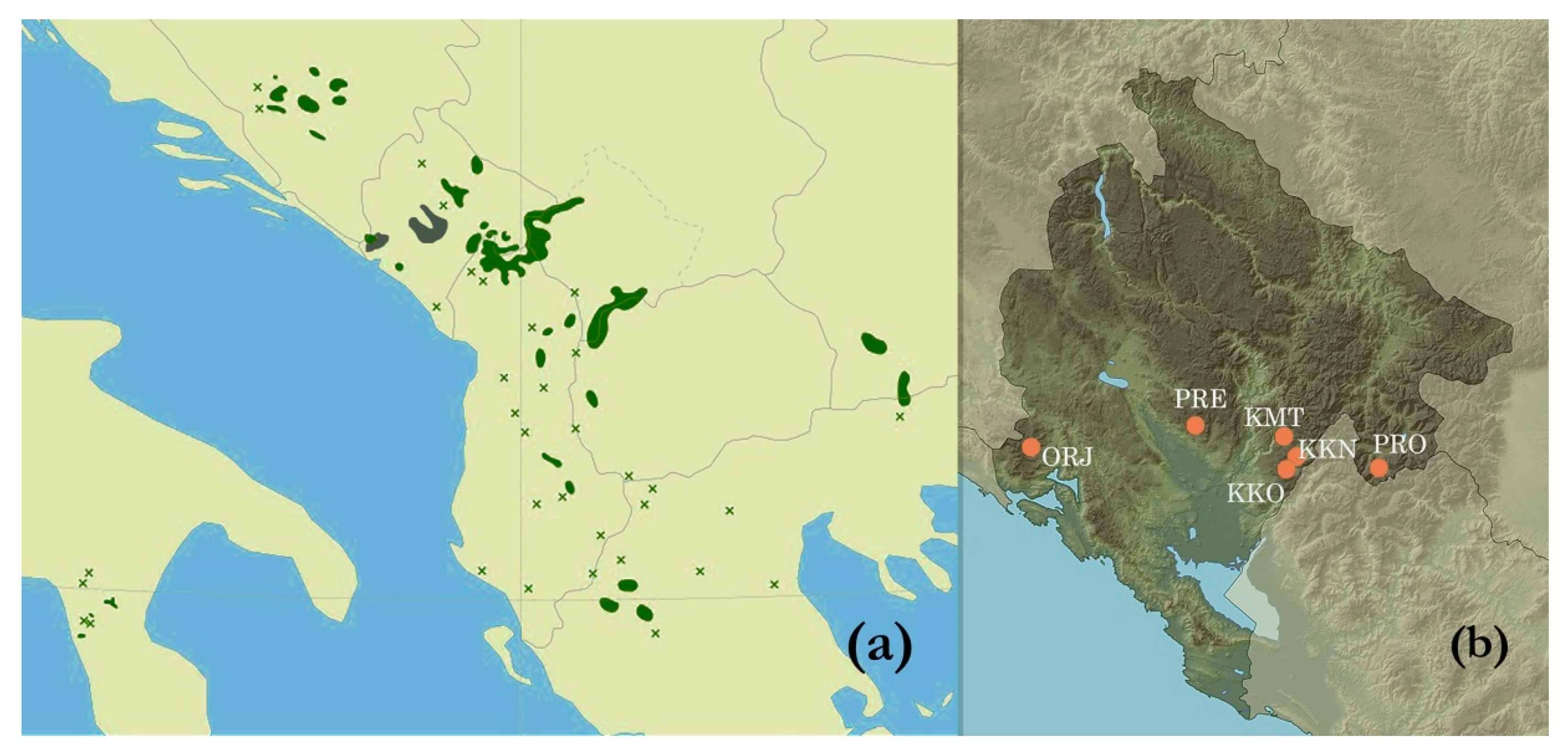

| Site | GPS Coordinates | Altitude (m) | Soil Type a | Climate b | Health c |
|---|---|---|---|---|---|
| Orjen, Reovačka greda (ORJ) | 42°35′ N 18°35′ E | 1700 | Lithosol | Cfsb | Healthy-looking |
| Kuči Mt., Momonjevo (KMT) | 42°36′ N 19°32′ E | 1800 | Lithosol | Cfb | Healthy-looking |
| Prekornica, Studeno (PRE) | 42°38′ N 19°12′ E | 1200 | Leptic cambisol | Cfsb | Healthy-looking |
| Prokletije, Ropojana (PRO) | 42°29′ N 19°48′ E | 1250 | Leptic cambisol | Cfb | Healthy-looking |
| Kučka korita (KKO) | 42°29′ N 19°30′ E | 1300 | Leptic cambisol | Cfs’’b | Healthy-looking |
| Kučka korita North (KKN) | 42°30′ N 19°32′ E | 1450 | Molic leptosol | Cfs’’b | Attacked by insects |
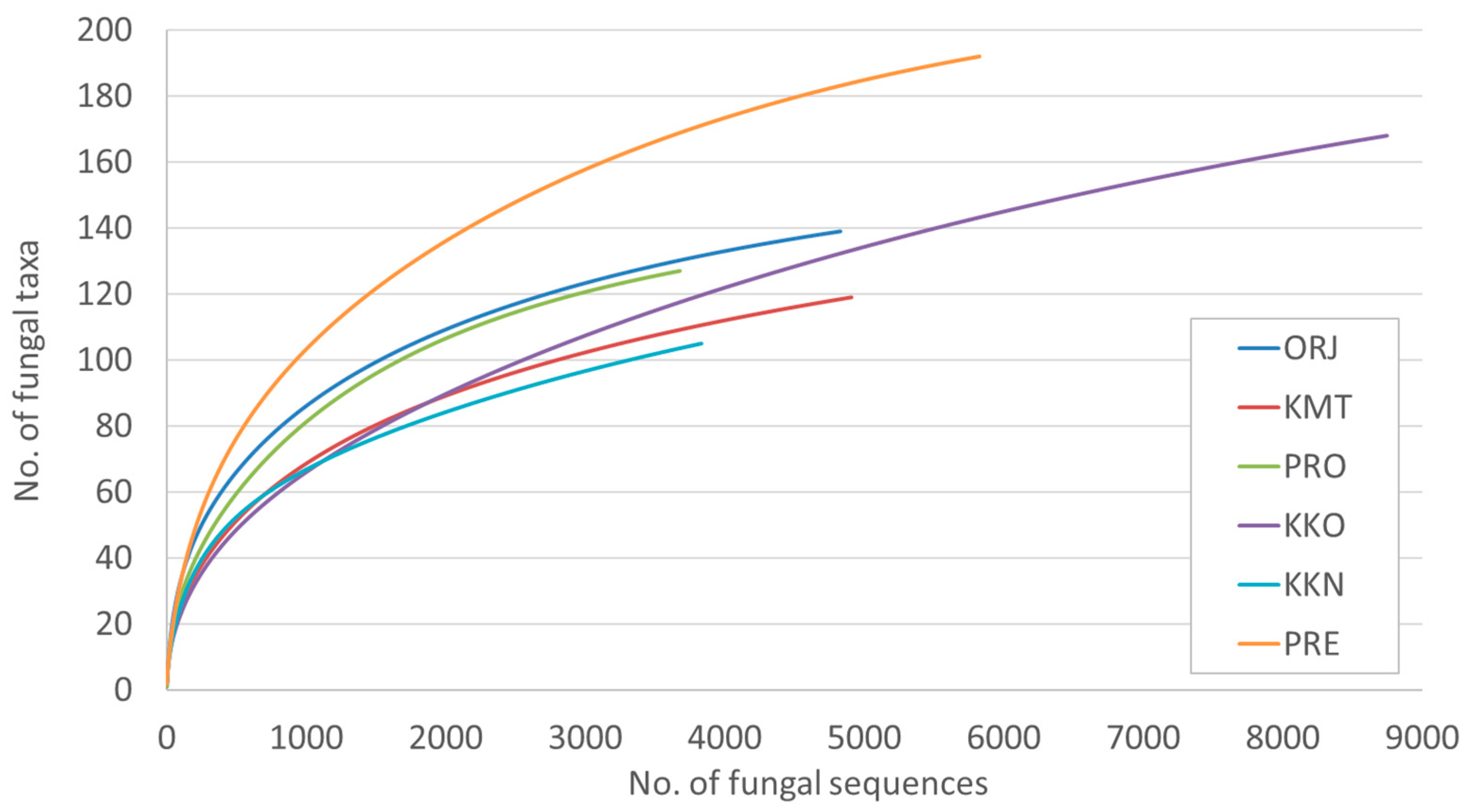
| Site | No. of Fungal Sequences | No. of Fungal Taxa | Shannon Diversity Index |
|---|---|---|---|
| Orjen –ORJ | 4830 | 139 a | 3.4 |
| Kuči Mt.-KMT | 4910 | 119 ab | 2.9 |
| Prekornica-PRE | 5828 | 192 a | 3.1 |
| Prokletije -PRO | 3679 | 127 a | 3.2 |
| Kučka korita -KKO | 8748 | 167 b | 2.9 |
| Kučka korita North –KKN | 3834 | 104 ab | 2.7 |
| All | 31,829 | 375 | - |
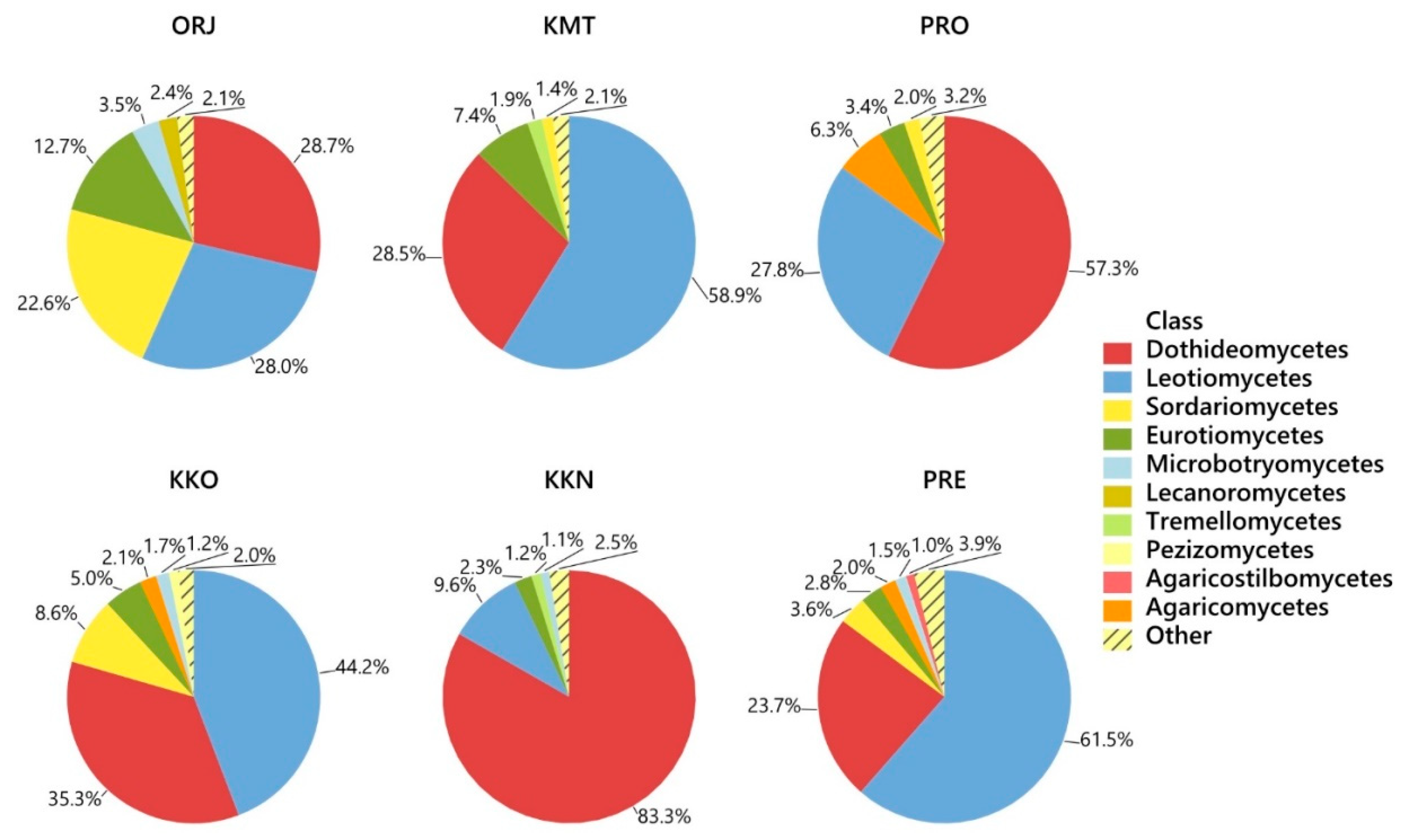
| Fungal Taxon | Phylum * | GenBank Accession No. | Compared (bp) | Similarity (%) | Sites | All | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| ORJ | KMT | PRE | PRO | KKO | KKN | ||||||
| Lophodermium pinastri | A | KC608049 | 361/361 | 100 | 4.7 | 22.1 | 14.3 | 14.2 | 14.9 | 0.4 | 12.5 |
| Lophodermium conigenum | A | HM060650 | 453/453 | 100 | - | 0.1 | 32.4 | 5.5 | 14.5 | 2.8 | 10.9 |
| Sydowia polyspora | A | KU516591 | 473/473 | 100 | 6.8 | 16.0 | 2.1 | 8.2 | 0.4 | 32.2 | 8.8 |
| Cyclaneusma niveum | A | AF013223 | 442/442 | 99 | 1.3 | 1.0 | 4.5 | 5.7 | 12.9 | 0.9 | 5.5 |
| Unidentified sp. 2814_1 | A | MF976656 | 238/245 | 97 | 20.2 | - | 0.5 | - | 8.2 | - | 5.4 |
| Phaeosphaeria pontiformis | A | KT000144 | 442/442 | 98 | - | - | 3.7 | 11.1 | 8.9 | - | 4.4 |
| Unidentified sp. 2814_11 | A | KU062806 | 203/252 | 81 | - | - | 1.3 | 9.0 | 10.3 | - | 4.1 |
| Unidentified sp. 2814_10 | A | KX220267 | 257/257 | 100 | 3.3 | - | 1.5 | 4.1 | 5.4 | 5.4 | 3.4 |
| Phaeomoniella sp. 2814_15 | A | GQ153187 | 257/259 | 99 | 7.2 | 5.9 | 0.2 | 1.2 | 2.9 | 0.5 | 3.0 |
| Neocatenulostroma germanicum | A | KR995100 | 242/242 | 100 | 1.3 | - | 0.4 | 1.0 | 3.4 | 7.9 | 2.3 |
| Microsphaeropsis olivacea | A | MH871969 | 249/249 | 100 | 0.7 | 0.1 | 0.3 | 0.1 | - | 16.8 | 2.2 |
| Allantophomopsiella pseudotsugae | A | MH857222 | 240/240 | 100 | 6.4 | 7.1 | - | - | - | - | 2.1 |
| Lachnellula calyciformis | A | MH858771 | 239/239 | 100 | 4.7 | 8.1 | - | - | - | - | 2.0 |
| Unidentified sp. 2814_23 | A | KF983527 | 209/244 | 86 | 1.4 | - | 1.5 | 3.7 | 0.3 | 4.3 | 1.5 |
| Ramoconidiophora euphorbiae | A | MG592740 | 232/239 | 97 | 3.2 | 6.0 | - | - | - | 0.6 | 1.5 |
| Geastrumia sp. 2814_26 | A | FJ438389 | 220/235 | 94 | 2.7 | 2.6 | 0.4 | 0.6 | 0.5 | 2.3 | 1.4 |
| Unidentified sp. 2814_16 | A | KP892077 | 240/242 | 99 | 4.8 | 0.1 | 0.6 | 1.9 | 0.3 | 0.5 | 1.2 |
| Unidentified sp. 2814_27 | A | KT244857 | 209/248 | 84 | - | - | 3.5 | 5.0 | - | - | 1.2 |
| Collophora sp. 2814_30 | A | NR_137726 | 229/241 | 95 | 3.2 | 2.2 | 0.4 | - | - | 0.3 | 0.9 |
| Unidentified sp. 2814_29 | A | MF976139 | 209/240 | 87 | 1.4 | - | 0.4 | 4.1 | 0.4 | - | 0.9 |
| Rhytismataceae sp. 2814_22 | A | KR266446 | 234/237 | 99 | 0.4 | - | 1.0 | 1.4 | 1.6 | - | 0.8 |
| Dothideomycetes sp. 2814_13 | A | KP991484 | 253/257 | 98 | 1.1 | 3.5 | - | 0.1 | - | 0.4 | 0.8 |
| Mollisia ligni | A | MF161301 | 237/241 | 98 | - | 4.7 | - | - | - | - | 0.7 |
| Cenangium acuum | A | MG597445 | 239/239 | 100 | - | - | 3.9 | - | - | - | 0.7 |
| Athelia acrospora | B | KP814375 | 296/299 | 99 | - | - | 1.8 | 0.1 | 1.3 | - | 0.7 |
| All of 25 taxa | 74.9 | 79.9 | 74.6 | 76.6 | 86.4 | 75.4 | 79.0 | ||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lazarević, J.; Menkis, A. Fungal Diversity in the Phyllosphere of Pinus heldreichii H. Christ—An Endemic and High-Altitude Pine of the Mediterranean Region. Diversity 2020, 12, 172. https://doi.org/10.3390/d12050172
Lazarević J, Menkis A. Fungal Diversity in the Phyllosphere of Pinus heldreichii H. Christ—An Endemic and High-Altitude Pine of the Mediterranean Region. Diversity. 2020; 12(5):172. https://doi.org/10.3390/d12050172
Chicago/Turabian StyleLazarević, Jelena, and Audrius Menkis. 2020. "Fungal Diversity in the Phyllosphere of Pinus heldreichii H. Christ—An Endemic and High-Altitude Pine of the Mediterranean Region" Diversity 12, no. 5: 172. https://doi.org/10.3390/d12050172
APA StyleLazarević, J., & Menkis, A. (2020). Fungal Diversity in the Phyllosphere of Pinus heldreichii H. Christ—An Endemic and High-Altitude Pine of the Mediterranean Region. Diversity, 12(5), 172. https://doi.org/10.3390/d12050172





