Genotyping-By-Sequencing Reveals Population Structure and Genetic Diversity of a Buffelgrass (Cenchrus ciliaris L.) Collection
Abstract
1. Introduction
2. Materials and Methods
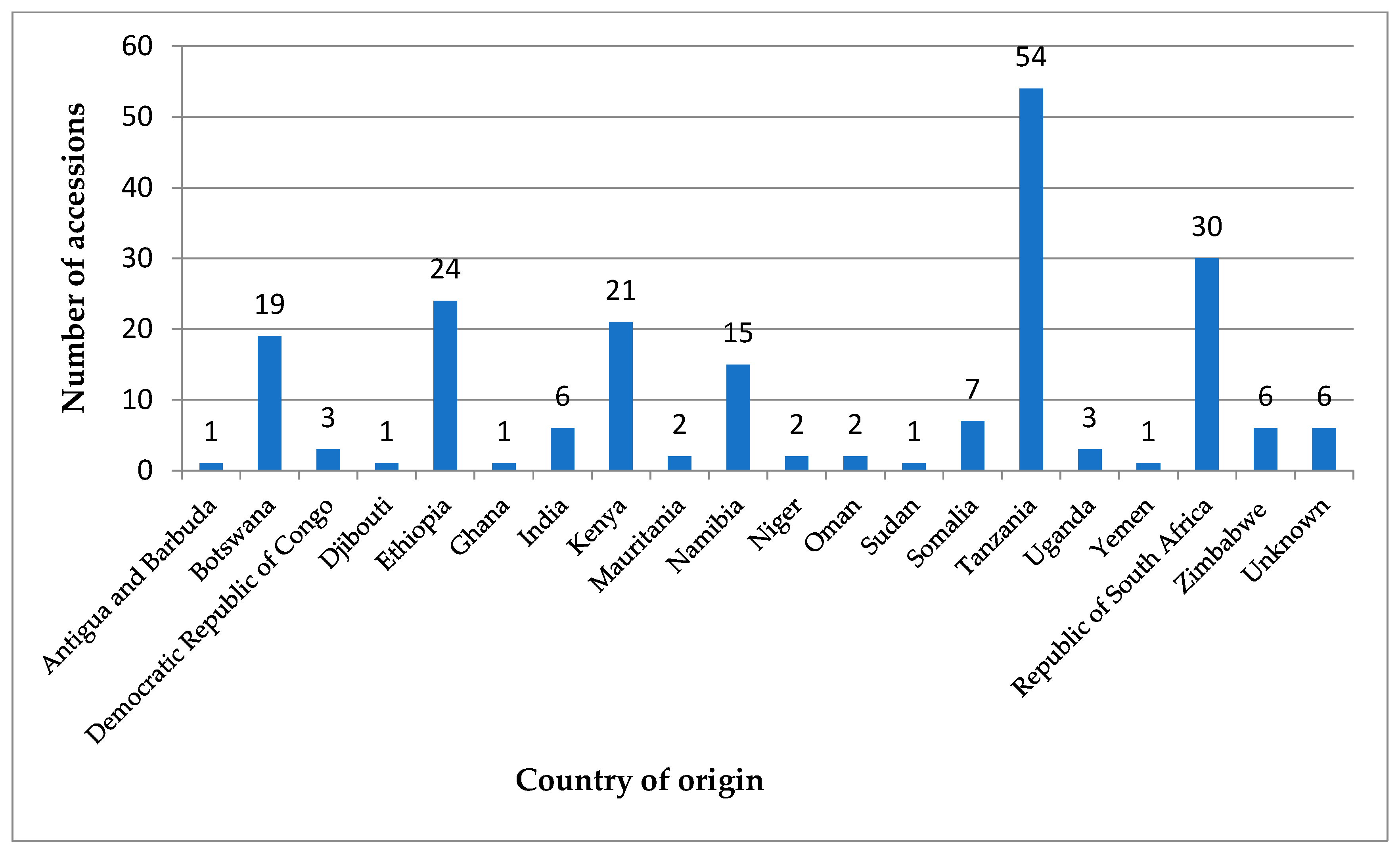
2.1. Materials
2.2. DNA Extraction
2.3. Genotyping
2.4. Data Analysis
2.5. Core Collection Development
3. Results
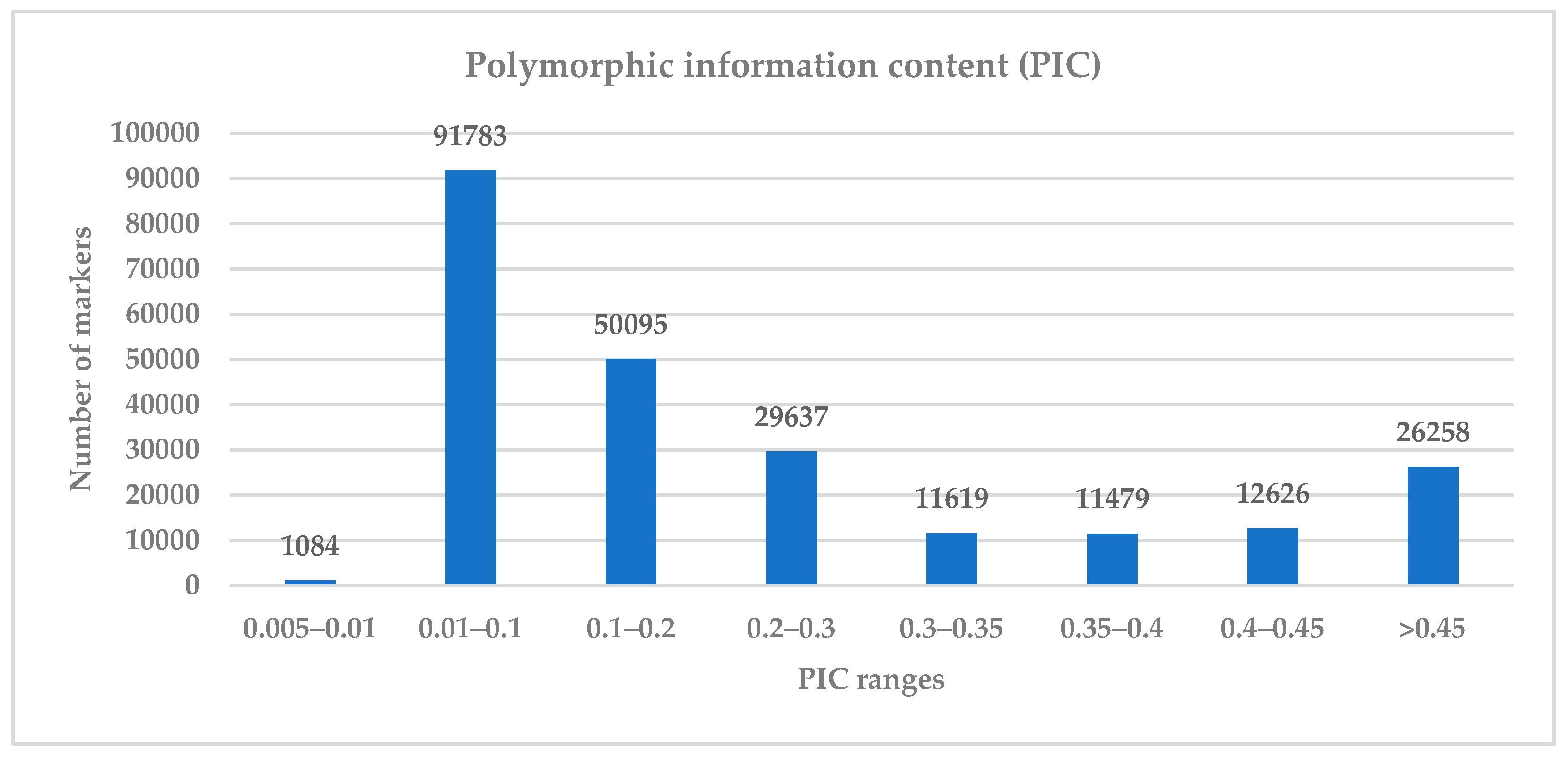
3.1. Informativeness and Diversity of the SNP Markers
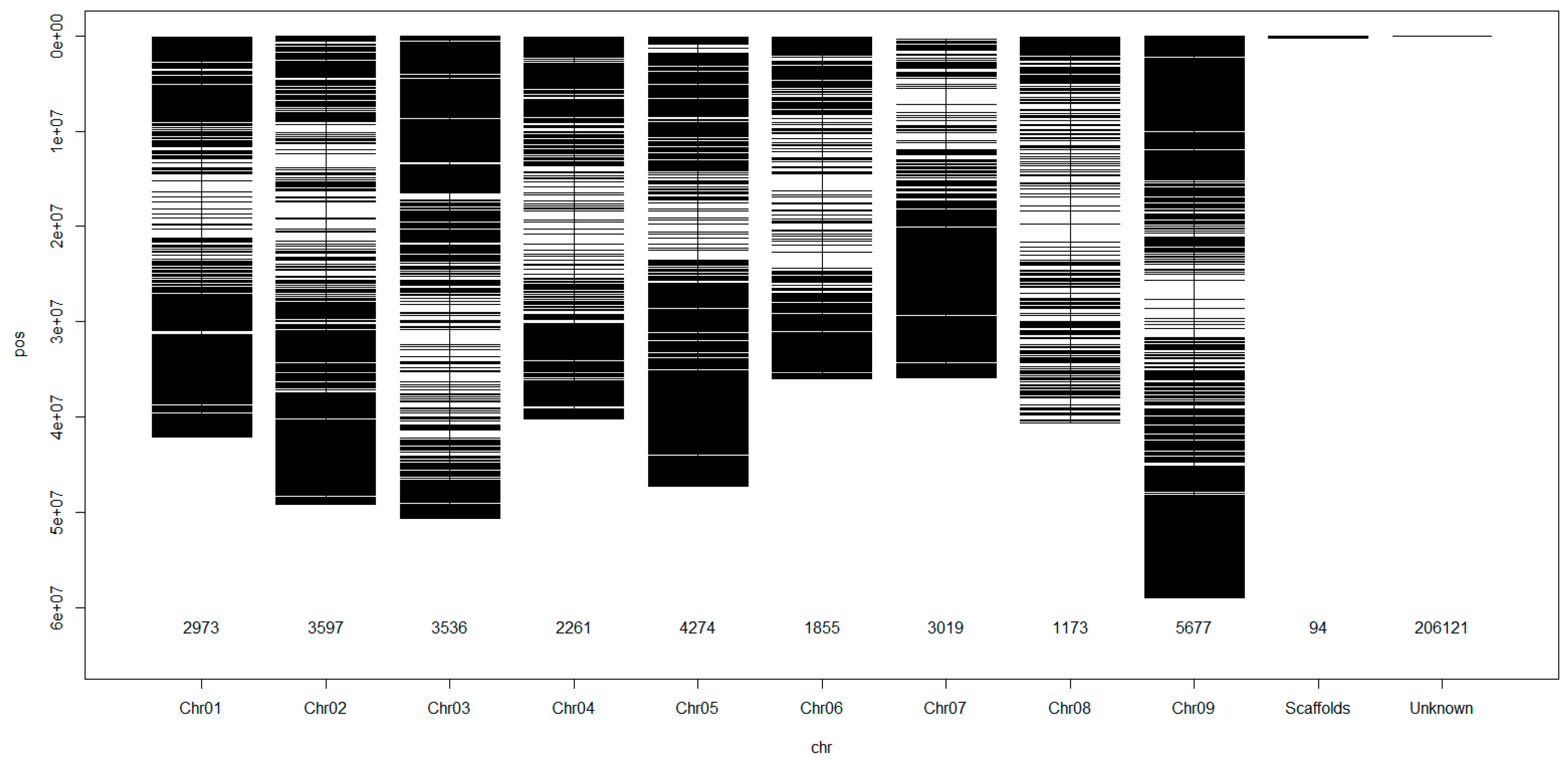
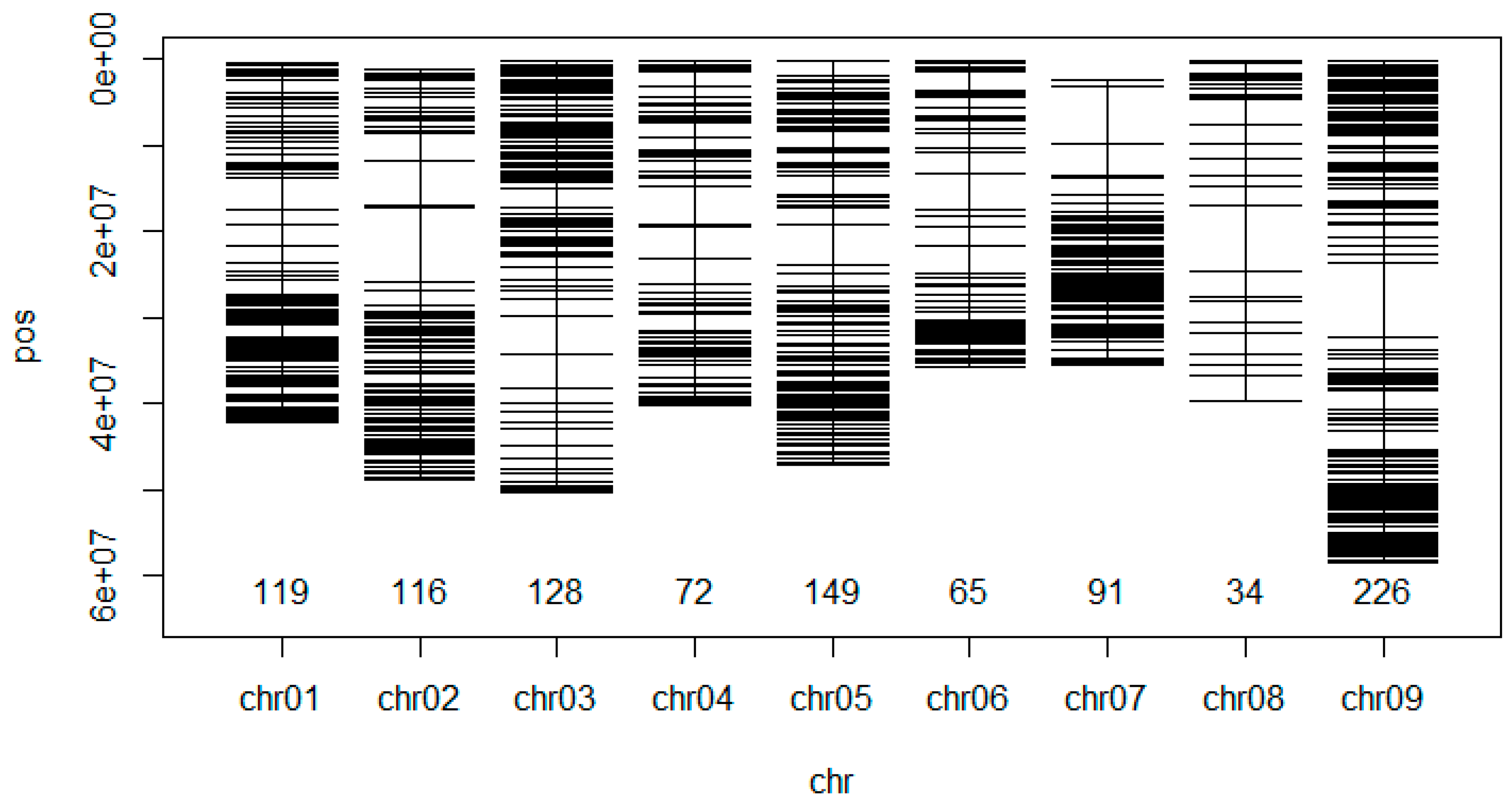
3.2. Mapping and Genome Wide Distribution of the SNP Markers
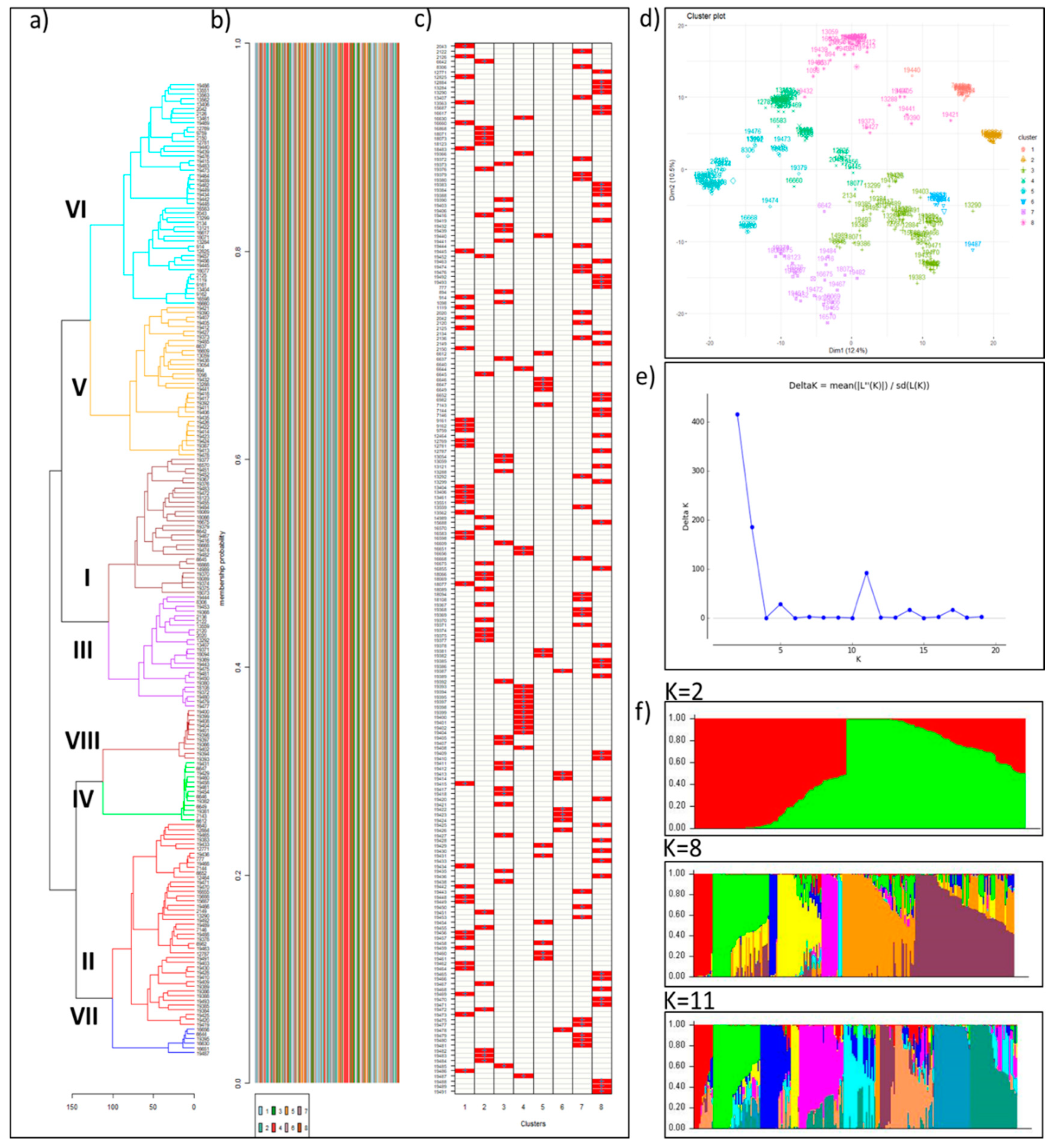
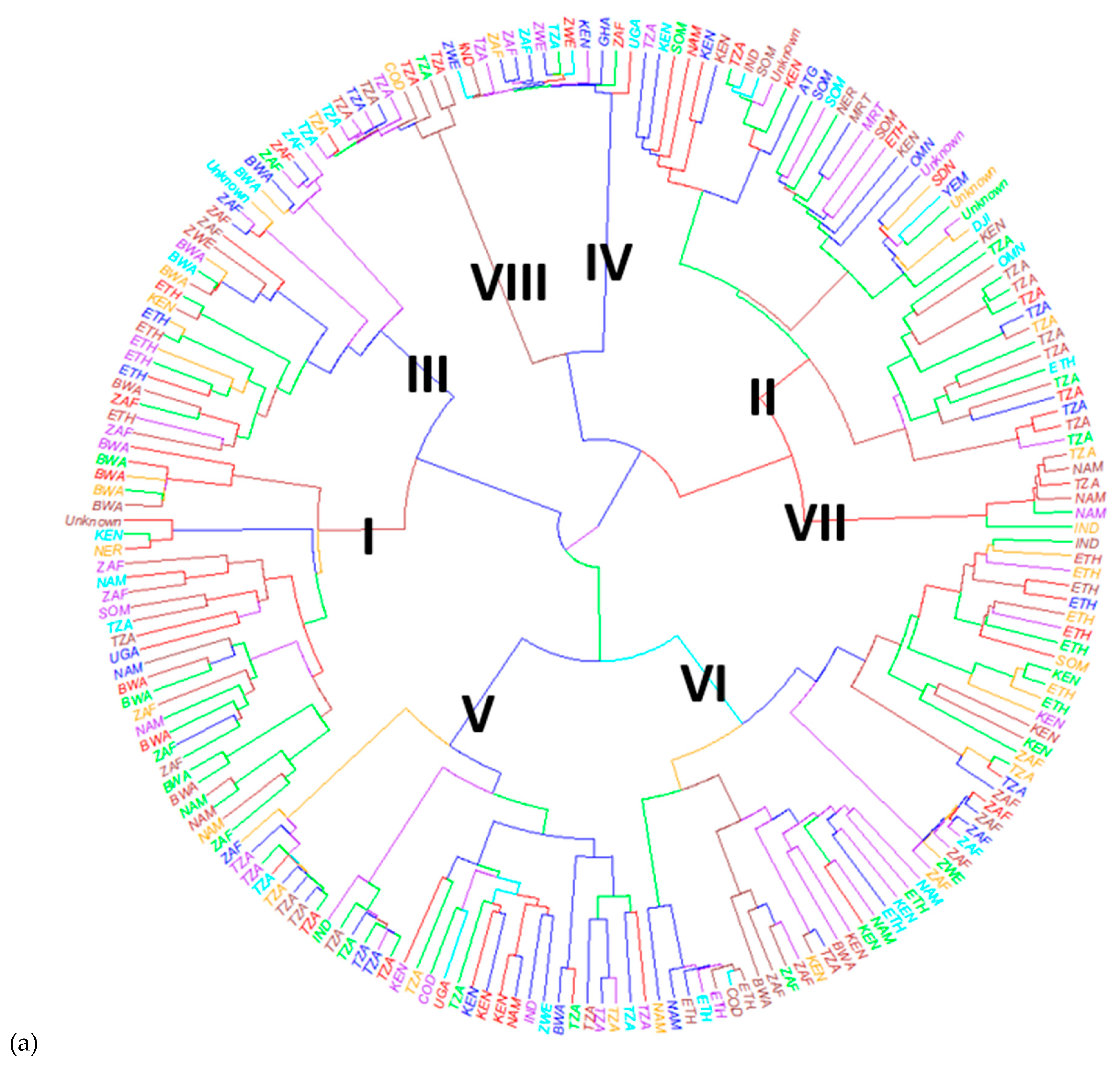
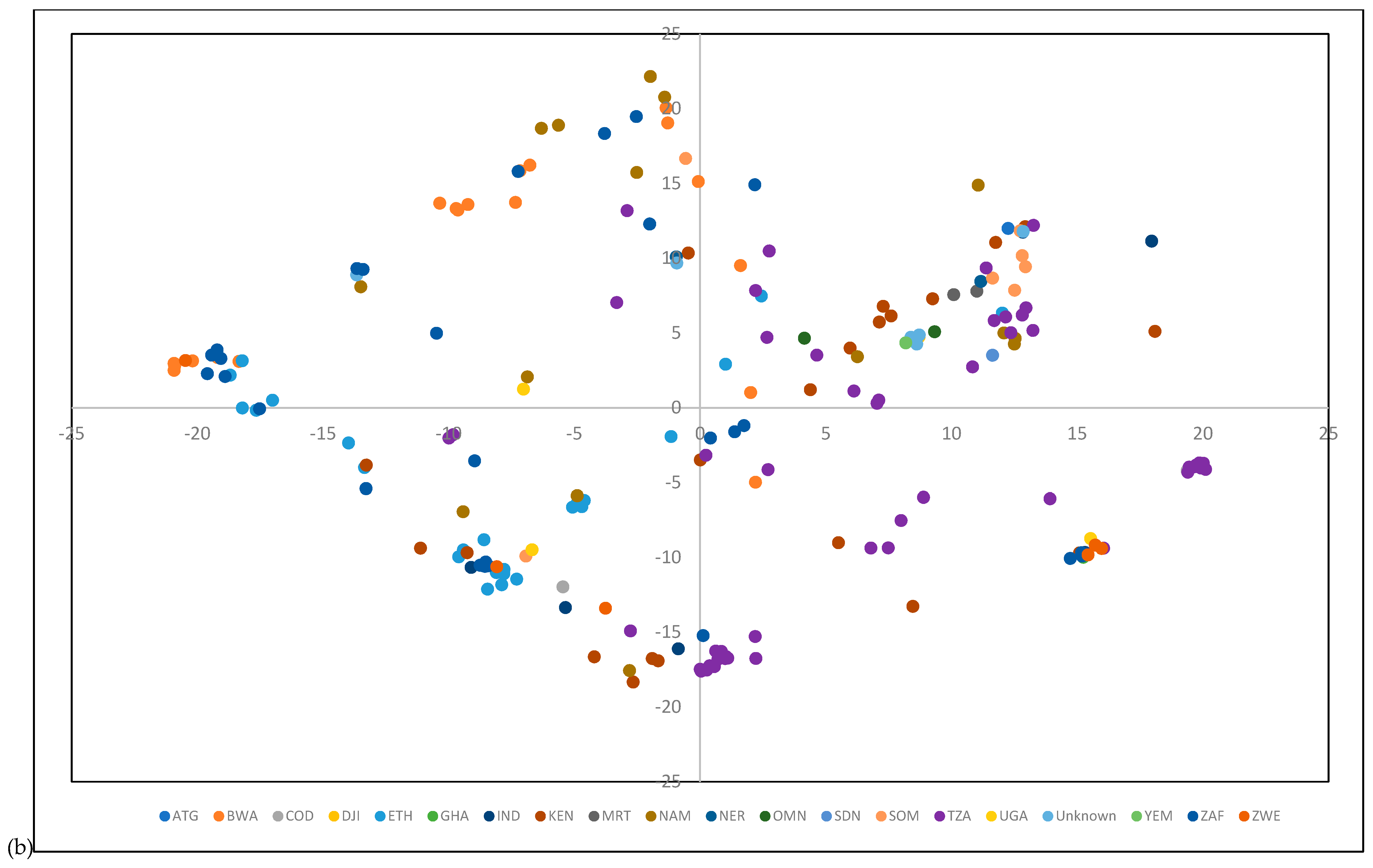
3.3. Population Structure and Genetic Diversity of the Buffelgrass Collection
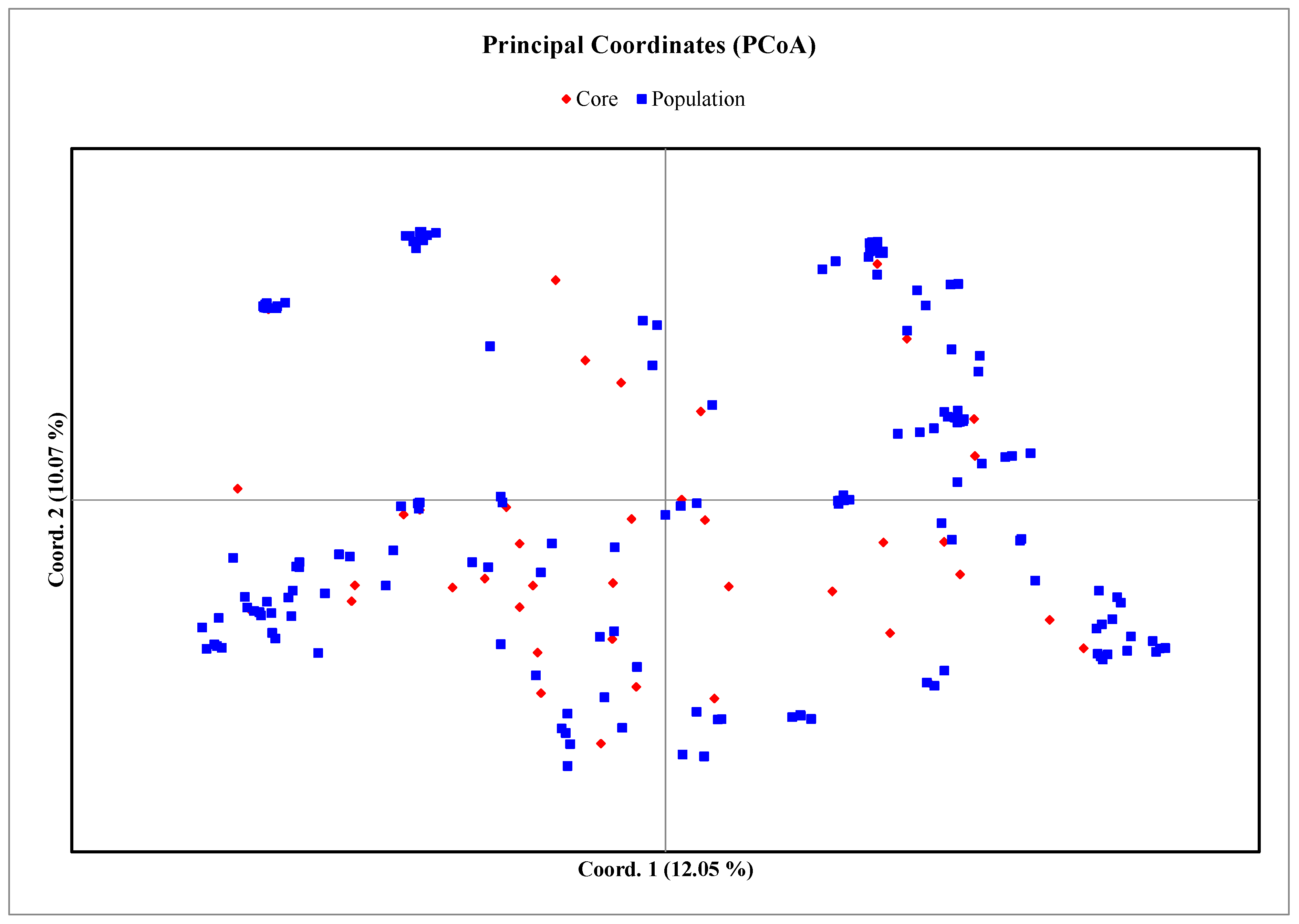
3.4. Core Collection Development
4. Discussion
4.1. Population Structure and Genetic Diversity of the Buffelgrass Collection
4.2. GBS Data Revealed a Lack of Genetic Differentiation among Germplasm from Diverse Origins
4.3. The GBS Data could be Used for Gap Filling in the Collection
4.4. Core Collection Establishment
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Devendra, C.; Swanepoel, F.; Stroebel, A.; Van Rooyen, C. Implications and innovative strategies for enhancing the future contribution of livestock. In The Role of Livestock in Developing Communities; SUN MeDIA Bloemfontein: Bloemfontein, South Africa, 2010. [Google Scholar]
- Kharrat-Souissi, A.; Siljak-Yakovlev, S.; Brown, S.C.; Chaieb, M. Cytogeography of Cenchrus ciliaris (Poaceae) in Tunisia. Folia. Geobot. 2013, 48, 95–113. [Google Scholar] [CrossRef]
- Marshall, V.M.; Lewis, M.M.; Ostendorf, B. Buffelgrass (Cenchrus ciliaris) as an invader and threat to biodiversity in arid environments: A review. J. Arid Environ. 2012, 78, 1–12. [Google Scholar] [CrossRef]
- Cook, B.G.; Pengelly, B.C.; Brown, S.D.; Donnelly, J.L.; Eagles, D.A.; Franco, M.A.; Hanson, J.; Mullen, B.F.; Partridge, I.J.; Peters, M.; et al. Tropical Forages: An Interactive Selection Tool [CD-ROM]; CSIRO, DPI&F(Qld), CIAT and ILRI: Brisbane, Australia, 2005. [Google Scholar]
- Kharrat-Souissi, A.; Siljak-Yakovlev, S.; Brown, S.C.; Baumel, A.; Torre, F.; Chaieb, M. The polyploid nature of Cenchrus ciliaris L. (Poaceae) has been overlooked: New insights for the conservation and invasion biology of this species—A review. Rangel. J. 2014, 36, 11–23. [Google Scholar] [CrossRef]
- Jessup, R.W.; Burson, B.L.; Burow, G.; Wang, Y.W.; Chang, C.; Li, Z.; Paterson, A.H.; Hussey, M.A. Segmental allotetraploidy and allelic interactions in buffelgrass (Pennisetum ciliare (L.) Link syn. Cenchrus ciliaris L.) as revealed by genome mapping. Genome 2003, 46, 304–313. [Google Scholar] [CrossRef][Green Version]
- Bray, R.A. Evidence for Facultative Apomixis in Cenchrus ciliaris. Euphytica 1978, 27, 801–804. [Google Scholar] [CrossRef]
- Shafer, G.S.; Burson, B.L.; Hussey, M.A. Stigma receptivity and seed set in protogynous buffelgrass. Crop Sci. 2000, 40, 391–397. [Google Scholar] [CrossRef]
- Visser, N.C.; Spies, J.J.; Venter, H.J.T. Aneuploidy in Cenchrus cilliaris (Poaceae, Panicoideae, Paniceae): Truth or fiction? S. Afr. J. Bot. 1998, 64, 337–345. [Google Scholar] [CrossRef]
- Buffelgrass (Cenchrus ciliaris). Feedipedia, A Programme by INRA, CIRAD, AFZ and FAO. Available online: https://www.feedipedia.org/node/482 (accessed on 4 October 2019).
- Burson, B.L.; Actkinson, J.M.; Hussey, M.A.; Jessup, R.W. Ploidy determination of buffelgrass accessions in the USDA National Plant Germplasm System collection by flow cytometry. S. Afr. J. Bot. 2012, 79, 91–95. [Google Scholar] [CrossRef][Green Version]
- Velázquez, S.G.; Herrera, R.R.; Carrillo, A.R.Q.; Quiroz, J.F.E.; Garay, A.H.; Hernández, A.P. Evaluación morfológica, citológica y valor nutritivo de siete nuevos genotipos y un cultivar de pasto Cenchrus ciliaris L., tolerantes a frío. Rev. Mex. Cienc. Agrícolas 2015, 6, 1679–1687. [Google Scholar] [CrossRef][Green Version]
- Al-Dakheel, A.J.; Hussain, M.I. Genotypic variation for salinity tolerance in Cenchrus ciliaris L. Front. Plant Sci. 2016, 7, 1090. [Google Scholar] [CrossRef]
- Genesys PGR. Available online: https://www.genesys-pgr.org/ (accessed on 30 April 2019).
- Jorge, M.A.B.; Van De Wouw, M.; Hanson, J.; Mohammed, J. Characterisation of a collection of buffelgrass (Cenchrus ciliaris). Trop. Grassl. 2008, 42, 27–39. [Google Scholar]
- SávGutiéc, R.A.; Morales Nieto, C.R.; Hanson, J.; Santellano Estrada, E.; Jurado Guerra, P.; Villanueva Avalos, J.F.; Melgoza Castillo, A. Caracterización forrajera de ecotipos de zacate buffel en condiciones de temporal en Debre Zeit, Etiopía. Rev. Mex. Cienc. Agrícolas 2017, 8, 13–26. [Google Scholar]
- Hignight, K.W.; Bashaw, E.C.; Hussey, M.A. Cytological and morphological diversity of native apomictic buffelgrass, Pennisetum ciliare (L) Link. Bot. Gaz. 1991, 152, 214–218. [Google Scholar] [CrossRef]
- Arshad, M.; Ashraf, M.Y.; Ahamad, M.; Zaman, F. Morpho-genetic variability potential of Cenchrus ciliaris L., from Cholistan Desert, Pakistan. Pak. J. Bot. 2007, 39, 1481–1488. [Google Scholar]
- Kharrat-Souissi, A.; Baumel, A.; Mseddi, K.; Torre, F.; Chaieb, M. Polymorphism of Cenchrus ciliaris L. a perennial grass of arid zones. Afr. J. Ecol. 2011, 49, 209–220. [Google Scholar] [CrossRef]
- Bruno, L.R.G.P.; Antonio, R.P.; Assis, J.G.D.; Moreira, J.N.; Lira, I.C.D.A. Buffelgrass morphoagronomic characterization from Cenchrus germplasm active bank. Rev. Caatinga. 2017, 30, 487–495. [Google Scholar] [CrossRef]
- Burson, B.L.; Renganayaki, K.; Dowling, C.D.; Hinze, L.L.; Jessup, R.W. Genetic diversity among pentaploid Buffelgrass accessions. Crop Sci. 2015, 55, 1637–1645. [Google Scholar] [CrossRef]
- Kharrat-Souissi, A.; Baumel, A.; Torre, F.; Juin, M.; Siljak-Yakovlev, S.; Roig, A.; Chaieb, M. New insights into the polyploid complex Cenchrus ciliaris L. (Poaceae) show its capacity for gene flow and recombination processes despite its apomictic nature. Aust. J. Bot. 2011, 59, 543–553. [Google Scholar] [CrossRef]
- Kilian, A.; Wenzl, P.; Huttner, E.; Carling, J.; Xia, L.; Blois, H.; Caig, V.; Heller-Uszynska, K.; Jaccoud, D.; Hopper, C.; et al. Diversity arrays technology: A generic genome profiling technology on open platforms. Meth. Mol. Biol. 2012, 888, 67–89. [Google Scholar]
- Reference Genome of Setaria Italica (Foxtail Millet). Available online: https://www.ncbi.nlm.nih.gov/genome/?term=foxtail+millet (accessed on 22 May 2019).
- Wimmer, V.; Albrecht, T.; Auinger, H.-J.; Schon, C.-C. Synbreed: A framework for the analysis of genomic prediction data using R. Bioinformatics 2012, 28, 2086–2087. [Google Scholar] [CrossRef]
- Michael, T.P.; Jackson, S. The First 50 Plant Genomes. Plant Genom. 2013, 6, 2. [Google Scholar] [CrossRef]
- Nei, M. Analysis of gene diversity in subdivided populations. Proc. Natl. Acad. Sci. USA 1973, 70, 3321–3323. [Google Scholar] [CrossRef] [PubMed]
- Jombart, T. Adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 2008, 24, 1403–1405. [Google Scholar] [CrossRef] [PubMed]
- The R Project for Statistical Computing. Available online: https://www.r-project.org/ (accessed on 16 January 2019).
- Galili, T. Dendextend: An R package for visualizing, adjusting and comparing trees of hierarchical clustering. Bioinformatics 2015, 31, 3718–3720. [Google Scholar] [CrossRef]
- Extract and Visualize the Results of Multivariate Data Analyses. Available online: https://cran.r-project.org/web/packages/factoextra/factoextra.pdf (accessed on 15 August 2019).
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar]
- Falush, D.; Stephens, M.; Pritchard, J.K. Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 2003, 164, 1567–1587. [Google Scholar]
- Muktar, M.S.; Teshome, A.; Hanson, J.; Negawo, A.T.; Habte, E.; Entfellner, J.-B.D.; Lee, K.-W.; Jones, C.S. Genotyping by sequencing provides new insights into the diversity of Napier grass (Cenchrus purpureus) and reveals variation in genome-wide LD patterns between collections. Sci. Rep. 2019, 9, 6936. [Google Scholar] [CrossRef]
- Earl, D.A.; VonHoldt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- de Beukelaer, H.; Davenport, G.F.; Fack, V. Core Hunter 3: Flexible core subset selection. BMC Bioinform. 2018, 19, 203. [Google Scholar] [CrossRef] [PubMed]
- Nybom, H.; Weising, K.; Rotter, B. DNA fingerprinting in botany: Past, present, future. Investig. Genet. 2014, 5, 1. [Google Scholar] [CrossRef] [PubMed]
- Using Molecular Marker Technology in Studies on Plant Genetic Diversity. Available online: https://www.bioversityinternational.org/fileadmin/user_upload/online_library/publications/pdfs/Molecular_Markers_Volume_1_en.pdf (accessed on 11 November 2019).
- Gutierrez-Ozuna, R.; Eguiarte, L.E.; Molina-Freaner, F. Genotypic diversity among pasture and roadside populations of the invasive buffelgrass (Pennisetum ciliare L. Link) in north-western Mexico. J. Arid Environ. 2009, 73, 26–32. [Google Scholar] [CrossRef]
- Negawo, A.T.; Jorge, A.; Hanson, J.; Teshome, A.; Muktar, M.S.; Azevedo, A.L.S.; Ledo, F.J.S.; Machado, J.C.; Jones, C.S. Molecular markers as a tool for germplasm acquisition to enhance the genetic diversity of a Napier grass (Cenchrus purpureus syn. Pennisetum purpureum) collection. Trop. Grassl. 2018, 6, 58–69. [Google Scholar]
- Singh, N.; Wu, S.Y.; Raupp, W.J.; Sehgal, S.; Arora, S.; Tiwari, V.; Vikram, P.; Singh, S.; Chhuneja, P.; Gill, B.S.; et al. Efficient curation of genebanks using next generation sequencing reveals substantial duplication of germplasm accessions. Sci. Rep. 2019, 9, 1–10. [Google Scholar] [CrossRef]
- The Core Collection at the Crossroads. Available online: https://www.researchgate.net/publication/285536959 (accessed on 10 February 2020).
- Frankel, O.H. Genetic perspectives of germplasm conservation. In Genetic Manipulation: Impact on Man and Society; Arber, W., Illmensee, K., Peacock, W.J., Starlinger, P., Eds.; Cambridge University Press: Cambridge, UK, 1984; pp. 161–170. [Google Scholar]
| Source of Variation | Degree of Freedom | Sum of Squares | Mean Sum of Squares | Estimation Variation | Percentage of Variation |
|---|---|---|---|---|---|
| Among clusters | 7 | 18445.55 | 2635.08 | 100.65 | 38% |
| Within clusters | 197 | 32813.78 | 166.57 | 166.57 | 62% |
| Total | 204 | 51259.33 | 267.22 | 100% |
| Accession No. | DOI | Origin | Region | Elevation (m a.s.l.) | Cluster+ | Accession No. | DOI | Origin | Region | Elevation | Cluster + |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 2043 | 10.18730/FZ78M | ETH | Africa | 1670 | 6 | 19388 | 10.18730/FY563 | TZA | Africa | 1117 | 2 |
| 2126 | 10.18730/G03P5 | ETH | Africa | 1600 | 6 | 19390 | 10.18730/FY585 | TZA | Africa | 1117 | 5 |
| 2134 | 10.18730/G06FM | ETH | Africa | 1600 | 2 | 19403 | 10.18730/FY5PK | TZA | Africa | 1140 | 2 |
| 6642 | 10.18730/G5X2H | TZA | Africa | 1 | 19408 | 10.18730/FY5VR | TZA | Africa | 1140 | 8 | |
| 12771 | 10.18730/FRMSK | KEN | Africa | 5 | 2 | 19416 | 10.18730/FY63* | TZA | Africa | 1229 | 1 |
| 12825 | 10.18730/FRPB* | KEN | Africa | 11 | 6 | 19419 | 10.18730/FY66 = | TZA | Africa | 1229 | 2 |
| 13284 | 10.18730/FS158 | KEN | Africa | 945 | 2 | 19427 | 10.18730/FY6E6 | TZA | Africa | 1207 | 5 |
| 13290 | 10.18730/FS1BE | KEN | Africa | 670 | 2 | 19432 | 10.18730/FY6KB | UGA | Africa | 1000 | 5 |
| 13563 | 10.18730/FS8JQ | ETH | Africa | 1400 | 6 | 19439 | 10.18730/FY6TJ | KEN | Africa | 1667 | 5 |
| 16630 | 10.18730/FVPF0 | NAM | Africa | 200 | 7 | 19440 | 10.18730/FY6VK | KEN | Africa | 15 | 4 |
| 16660 | 10.18730/FVQ7R | NAM | Africa | 1200 | 6 | 19441 | 10.18730/FY6WM | TZA | Africa | 909 | 5 |
| 16868 | 10.18730/FVXK6 | NER | Africa | 370 | 2 | 19444 | 10.18730/FY6ZQ | ZAF | Africa | 758 | 3 |
| 18071 | 10.18730/FWX8M | BWA | Africa | 900 | 2 | 19445 | 10.18730/FY70R | ZAF | Africa | 1288 | 6 |
| 18073 | 10.18730/FWXAP | BWA | Africa | 900 | 1 | 19463 | 10.18730/FY7J5 | DJI | Africa | 11 | 2 |
| 18108 | 10.18730/FWYAH | BWA | Africa | 900 | 3 | 19465 | 10.18730/FY7M7 | SOM | Africa | 20 | 2 |
| 18123 | 10.18730/FWYNW | BWA | Africa | 800 | 1 | 19474 | 10.18730/FY7XG | ZAF | Africa | 439 | 3 |
| 18483 | 10.18730/FX9AU | TZA | Africa | 1400 | 3 | 19476 | 10.18730/FY7ZJ | ZAF | Africa | 314 | 3 |
| 19377 | 10.18730/FY4VX | ZAF | Africa | 1 | 19478 | 10.18730/FY81M | ZAF | Africa | 900 | 5 | |
| 19379 | 10.18730/FY4XZ | UGA | Africa | 3 | 19492 | 10.18730/FY8F$ | OMN | Middle East | 2500 | 2 | |
| 19380 | 10.18730/FY4Y* | Unknown | 1 | 19493 | 10.18730/FY8G= | ETH | Africa | 325 | 2 | ||
| 19384 | 10.18730/FY52U | TZA | Africa | 1241 | 2 |
| Source of Variation | Degree of Freedom | Sum of Squares | Mean Sum of Squares | Estimation Variation | Percentage of Variation |
|---|---|---|---|---|---|
| Variation among groups | 1 | 286.89 | 286.89 | 0.55 | 0.22% |
| Variation within groups | 203 | 50972.44 | 251.20 | 251.20 | 99.78% |
| Total | 204 | 51259.33 | 251.65 | 100.00% |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Negawo, A.T.; Assefa, Y.; Hanson, J.; Abdena, A.; Muktar, M.S.; Habte, E.; Sartie, A.M.; Jones, C.S. Genotyping-By-Sequencing Reveals Population Structure and Genetic Diversity of a Buffelgrass (Cenchrus ciliaris L.) Collection. Diversity 2020, 12, 88. https://doi.org/10.3390/d12030088
Negawo AT, Assefa Y, Hanson J, Abdena A, Muktar MS, Habte E, Sartie AM, Jones CS. Genotyping-By-Sequencing Reveals Population Structure and Genetic Diversity of a Buffelgrass (Cenchrus ciliaris L.) Collection. Diversity. 2020; 12(3):88. https://doi.org/10.3390/d12030088
Chicago/Turabian StyleNegawo, Alemayehu Teressa, Yilikal Assefa, Jean Hanson, Asebe Abdena, Meki S. Muktar, Ermias Habte, Alieu M. Sartie, and Chris S. Jones. 2020. "Genotyping-By-Sequencing Reveals Population Structure and Genetic Diversity of a Buffelgrass (Cenchrus ciliaris L.) Collection" Diversity 12, no. 3: 88. https://doi.org/10.3390/d12030088
APA StyleNegawo, A. T., Assefa, Y., Hanson, J., Abdena, A., Muktar, M. S., Habte, E., Sartie, A. M., & Jones, C. S. (2020). Genotyping-By-Sequencing Reveals Population Structure and Genetic Diversity of a Buffelgrass (Cenchrus ciliaris L.) Collection. Diversity, 12(3), 88. https://doi.org/10.3390/d12030088





