Phylogenetic Characterization of Botryosphaeria Strains Associated with Asphondylia Galls on Species of Lamiaceae
Abstract
1. Introduction
| Species | Main Hosts | Country | Reference |
|---|---|---|---|
| B. auasmontanum | Acacia | Namibia | [14] |
| B. kawatsukai | Malus, Pyrus | China | [15] |
| B. minutispermatia | dead wood | China | [16] |
| B. pseudoramosa | Eucalyptus, Melastoma | China | [17] |
| B. qinlingensis | Quercus | China | [18] |
| B. qingyuanensis | Eucalyptus | China | [17] |
| B. rosaceae | Amygdalus, Malus, Pyrus | China | [19] |
| B. sinensia | Juglans, Malus, Morus, Populus | China | [20] |
| B. wangensis | Cedrus | China | [17] |
2. Materials and Methods
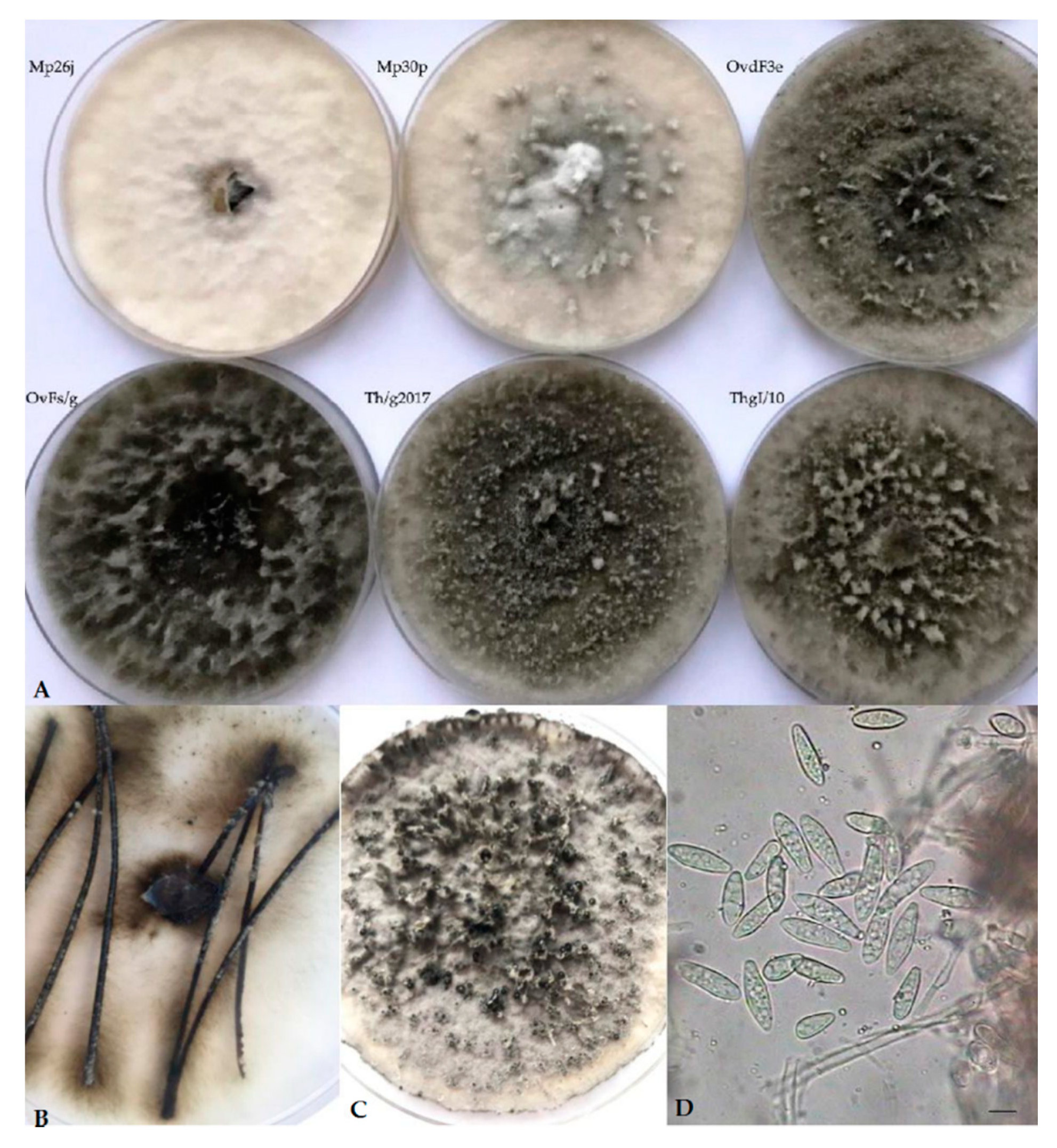
2.1. Isolation and Morphological Observations
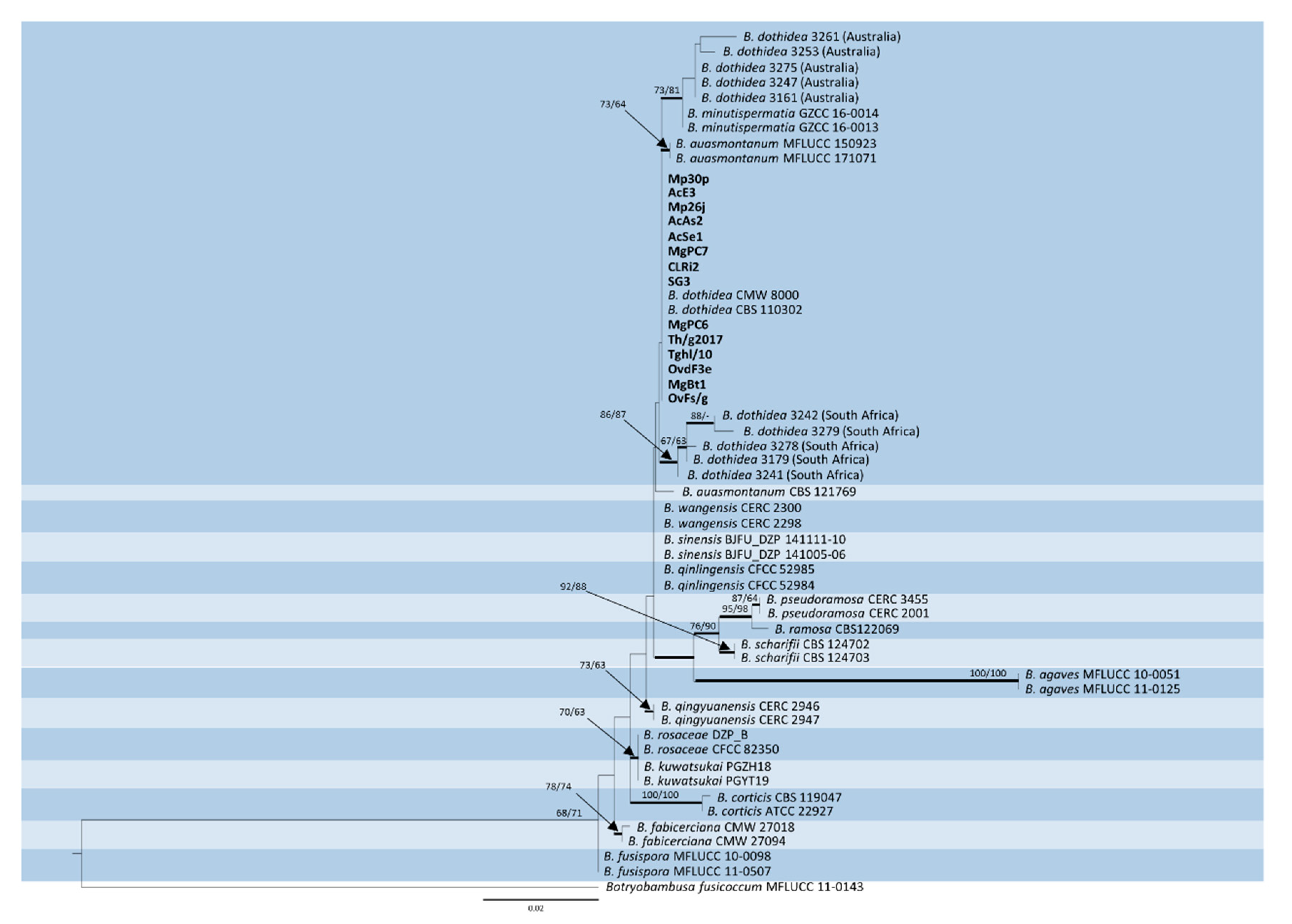
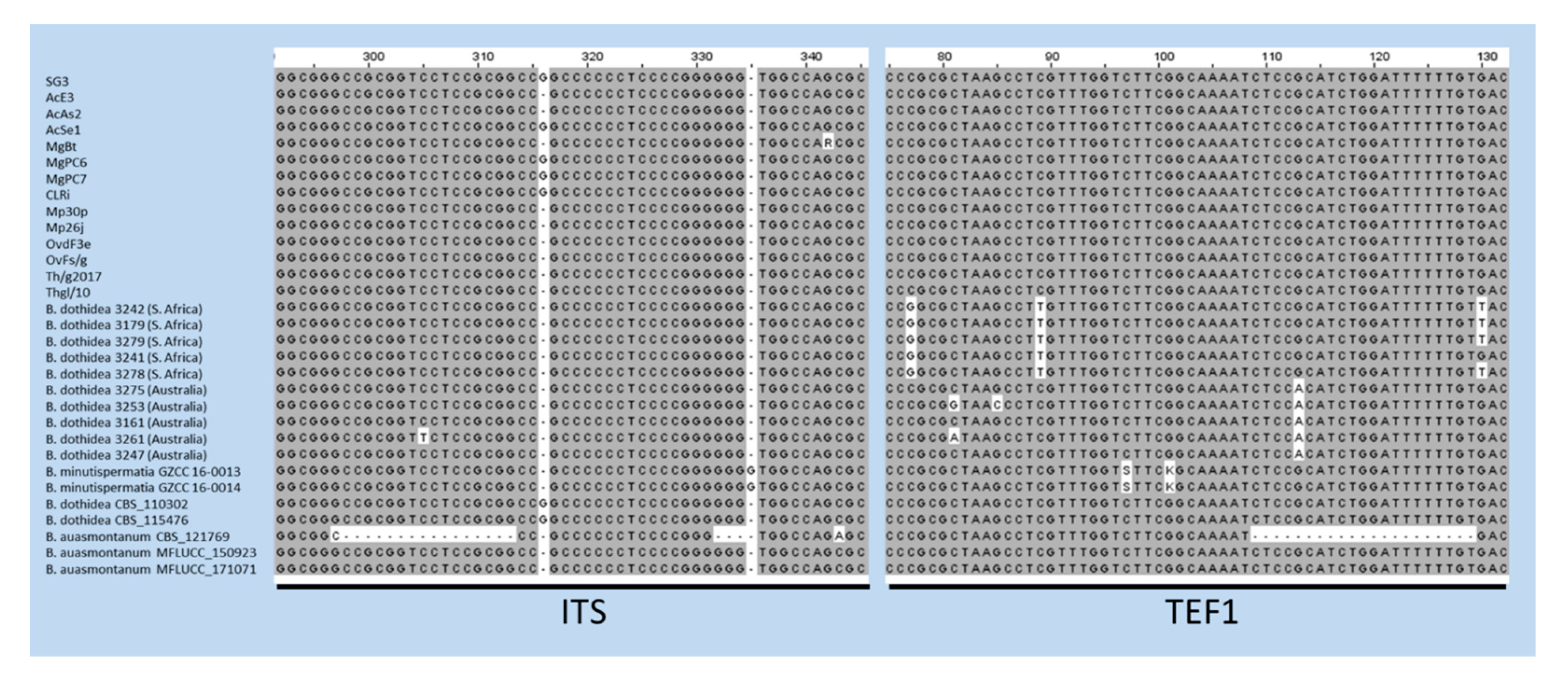
2.2. DNA Sequencing and Phylogenetic Analysis
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Adair, R.J.; Burgess, T.; Serdani, M.; Barber, P. Fungal associations in Asphondylia (Diptera: Cecidomyiidae) galls from Australia and South Africa: Implications for biological control of invasive acacias. Fungal Ecol. 2009, 2, 121–134. [Google Scholar] [CrossRef]
- Lebel, T.; Peele, C.; Veenstra, A. Fungi associated with Asphondylia (Diptera: Cecidomyiidae) galls on Sarcocornia quinqueflora and Tecticornia arbuscula (Chenopodiaceae). Fungal Divers. 2012, 55, 143–154. [Google Scholar] [CrossRef]
- Zimowska, B.; Viggiani, G.; Nicoletti, R.; Furmańczyk, A.; Becchimanzi, A.; Kot, I. First report of the gall midge Asphondylia serpylli on thyme (Thymus vulgaris), and identification of the associated fungal symbiont. Ann. Appl. Biol. 2017, 171, 89–94. [Google Scholar] [CrossRef]
- Bernardo, U.; Nugnes, F.; Gualtieri, L.; Nicoletti, R.; Varricchio, P.; Sasso, R.; Viggiani, G. A new gall midge species of Asphondylia (Diptera: Cecidomyiidae) inducing flower galls on Clinopodium nepeta (Lamiaceae) from Europe, its phenology, and associated fungi. Environ. Entomol. 2018, 47, 609–622. [Google Scholar] [CrossRef] [PubMed]
- Slippers, B.; Crous, P.W.; Denman, S.; Coutinho, T.A.; Wingfield, B.D.; Wingfield, M.J. Combined multiple gene genealogies and phenotypic characters differentiate several species previously identified as Botryosphaeria dothidea. Mycologia 2004, 96, 83–101. [Google Scholar] [CrossRef]
- Denman, S.; Crous, P.W.; Taylor, J.E.; Kang, J.-C.; Pascoe, I.; Wingfield, M.J. An overview of the taxonomic history of Botryosphaeria, and a re-evaluation of its anamorphs based on morphology and ITS rDNA phylogeny. Stud. Mycol. 2000, 45, 129–140. [Google Scholar]
- Park, I.; Sanogo, S.; Hanson, S.F.; Thompson, D.C. Molecular identification of Botryosphaeria dothidea as a fungal associate of the gall midge Asphondylia prosopidis on mesquite in the United States. BioControl 2019, 64, 209–219. [Google Scholar] [CrossRef]
- Phillips, A.J.L.; Alves, A.; Abdollahzadeh, J.; Slippers, B.; Wingfield, M.J.; Groenewald, J.Z.; Crous, P.W. The Botryosphaeriaceae: genera and species known from culture. Stud. Mycol. 2013, 76, 51–167. [Google Scholar] [CrossRef]
- Marsberg, A.; Kemler, M.; Jami, F.; Nagel, J.H.; Postma-Smidt, A.; Naidoo, S.; Wingfield, M.J.; Crous, P.W.; Spatafora, J.W.; Hesse, C.N.; et al. Botryosphaeria dothidea: a latent pathogen of global importance to woody plant health. Mol. Plant Pathol. 2017, 18, 477–488. [Google Scholar] [CrossRef]
- Slippers, B.; Wingfield, M.J. Botryosphaeriaceae as endophytes and latent pathogens of woody plants: diversity, ecology and impact. Fungal Biol. Rev. 2007, 21, 90–106. [Google Scholar] [CrossRef]
- Janson, E.M.; Peeden, E.R.; Stireman, J.O.; Abbot, P. Symbiont-mediated phenotypic variation without co-evolution in an insect–fungus association. J. Evol. Biol. 2010, 23, 2212–2228. [Google Scholar] [CrossRef] [PubMed]
- Malagaris, P. Biology and ecology of Asphondylia coridothymi (Diptera: Cecidomyiidae) inducing galls on Coridothymus capitatus on the island of Samos, Greece. Acta Soc. Zool. Bohemicae 2011, 75, 239–251. [Google Scholar]
- Viggiani, G.; Stinca, A. Infestations on oregano by the gall midge Asphondylia hornigi Wachtl. Prot. Colture 2018, 4, 29–31. [Google Scholar]
- Slippers, B.; Roux, J.; Wingfield, M.J.; van der Walt, F.J.J.; Jami, F.; Mehl, J.W.M.; Marais, G.J. Confronting the constraints of morphological taxonomy in the Botryosphaeriales. Persoonia 2014, 33, 155–168. [Google Scholar] [CrossRef]
- Xu, C.; Wang, C.S.; Ju, L.L.; Zhang, R.; Biggs, A.R.; Tanaka, E.; Li, B.; Sun, G.Y. Multiple locus genealogies and phenotypic characters reappraise the causal agent of apple ring rot in China. Fungal Divers. 2015, 71, 215–231. [Google Scholar] [CrossRef]
- Ariyawansa, H.A.; Hyde, K.D.; Liu, J.K.; Wu, S.P.; Liu, Z.Y. Additions to Karst Fungi 1: Botryosphaeria minutispermatia sp. nov., from Guizhou Province, China. Phytotaxa 2016, 275, 35–44. [Google Scholar] [CrossRef]
- Li, G.Q.; Liu, F.F.; Li, J.Q.; Liu, Q.L.; Chen, S.F. Botryosphaeriaceae from Eucalyptus plantations and adjacent plants in China. Persoonia 2018, 40, 63–95. [Google Scholar] [CrossRef]
- Liang, L.Y.; Jiang, N.; Chen, W.Y.; Liang, Y.M.; Tian, C.M. Botryosphaeria qinlingensis sp. nov. causing oak frogeye leaf spot in China. Mycotaxon 2019, 134, 463–473. [Google Scholar] [CrossRef]
- Zhou, Y.P.; Zhang, M.; Dou, Z.P.; Zhang, Y. Botryosphaeria rosaceae sp. nov. and B. ramosa, new botryosphaeriaceous taxa from China. Mycosphere 2017, 8, 162–171. [Google Scholar] [CrossRef]
- Zhou, Y.; Dou, Z.; He, W.; Zhang, X.; Zhang, Y. Botryosphaeria sinensia sp nov., a new species from China. Phytotaxa 2016, 245, 43–50. [Google Scholar] [CrossRef]
- Crous, P.W.; Slippers, B.; Wingfield, M.J.; Rheeder, J.; Marasas, W.F.; Philips, A.J.; Alves, A.; Burgess, T.; Barber, P.; Groenewald, J.Z. Phylogenetic lineages in the Botryosphaeriaceae. Stud. Mycol. 2006, 55, 235–253. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.J.W.T.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetic. In PCR Protocols, A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: New York, NY, USA, 1990; Volume 18, pp. 315–322. [Google Scholar]
- Carbone, I.; Kohn, L.M. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999, 91, 553–556. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Larsson, A. AliView: A fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 2014, 30, 3276–3278. [Google Scholar] [CrossRef]
- Dissanayake, A.J.; Camporesi, E.; Hyde, K.D.; Yan, J.Y.; Li, X.H. Saprobic Botryosphaeriaceae, including Dothiorella italica sp. nov., associated with urban and forest trees in Italy. Mycosphere 2017, 8, 1157–1176. [Google Scholar] [CrossRef]
- Wilgenbusch, J.C.; Swofford, D. Inferring evolutionary trees with PAUP*. Curr. Prot. Bioinformatics 2003, 1, 6.4.1–6.4.28. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 2006, 22, 2688–2690. [Google Scholar] [CrossRef]
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef]
- Molecular Evolution, Phylogenetics and Epidemiology. Available online: http://tree.bio.ed.ac.uk/software/figtree/ (accessed on 25 November 2018).
- Chen, S.; Pavlic, D.; Roux, J.; Slippers, B.; Xie, Y.; Wingfield, M.J.; Zhou, X.D. Characterization of Botryosphaeriaceae from plantation-grown Eucalyptus species in South China. Plant Pathol. 2011, 60, 739–751. [Google Scholar] [CrossRef]
- Abdollahzadeh, J.; Zare, R.; Phillips, A.J. Phylogeny and taxonomy of Botryosphaeria and Neofusicoccum species in Iran, with description of Botryosphaeria scharifii sp. nov. Mycologia 2013, 105, 210–220. [Google Scholar] [CrossRef]
| Number | Host | Location | GenBank Accession | |
|---|---|---|---|---|
| ITS | TEF1 | |||
| SG3 | Clinopodium nepeta | San Giorgio a Cremano, Italy | MN731265 | MN737437 |
| AcE3 | C. nepeta | Astroni Nature Reserve, Italy | MN731266 | MN737438 |
| AcAs2 | C. nepeta | Astroni Nature Reserve, Italy | MN731267 | MN737439 |
| AcSe1 | C. nepeta | Serino, Italy | MN731268 | MN737440 |
| CLRi2 | Clinopodium vulgare | Rivello, Italy | MN731272 | MN737444 |
| Mp26j | Mentha piperita | Konopnica, Poland | MN731273 | MN737445 |
| Mp30p | M. piperita | Konopnica, Poland | MN731274 | MN737446 |
| MgBt1 | Micromeria graeca | Boscotrecase, Italy | MN731269 | MN737441 |
| MgPC6 | M. graeca | Palma Campania, Italy | MN731270 | MN737442 |
| MgPC7 | M. graeca | Palma Campania, Italy | MN731271 | MN737443 |
| OvdF3e | Origanum vulgare | Fajsławice, Poland | MN731275 | MN737447 |
| OvFs/g | O. vulgare | Fajsławice, Poland | MN731276 | MN737448 |
| Th/g2017 | Thymus vulgaris | Fajsławice, Poland | MN731277 | MN737449 |
| ThgI/10 | T. vulgaris | Fajsławice, Poland | MN731278 | MN737450 |
| Species | Number | GenBank Accession | |
|---|---|---|---|
| ITS | TEF1 | ||
| Botryobambusa fusicoccum | MFLUCC 11-0143 | JX646792 | JX646857 |
| Botryosphaeria agaves | MFLUCC 10-0051 | JX646790 | JX646855 |
| MFLUCC 11-0125 | JX646791 | JX646856 | |
| Botryosphaeria auasmontanum | CBS 121769 | KF766167 | EU101348 |
| MFLUCC 15-0923 | MF398858 | MF398910 | |
| MFLUCC 17-1071 | MF398863 | MF398915 | |
| Botryosphaeria corticis | CBS 119047 | DQ299245 | EU017539 |
| ATCC 22927 | DQ299247 | EU673291 | |
| Botryosphaeria dothidea | CBS 110302 | AY259092 | AY573218 |
| CBS 115476 | AY236949 | AY236898 | |
| 3161 | EF614924 | EF614940 | |
| 3179 | EF614917 | EF614933 | |
| 3241 | EF614920 | EF614937 | |
| 3242 | EF614916 | EF614936 | |
| 3247 | EF614923 | EF614941 | |
| 3253 | EF614921 | EF614938 | |
| 3261 | EF614926 | EF614943 | |
| 3275 | EF614925 | EF614942 | |
| 3278 | EF614919 | EF614934 | |
| 3279 | EF614918 | EF614935 | |
| Botryosphaeria fabicerciana | CMW 27094 | HQ332197 | HQ332213 |
| CMW 27108 | HQ332200 | HQ332216 | |
| Botryosphaeria fusispora | MFLUCC 10-0098 | JX646789 | JX646854 |
| MFLUCC 11-0507 | JX646788 | JX646853 | |
| Botryosphaeria kawatsukai | PGZH18 | MG637267 | MG637243 |
| PGZH19 | MG637266 | MG637242 | |
| Botryosphaeria minutispermatia | GZCC 16-0013 | KX447675 | KX447678 |
| GZCC 16-0014 | KX447676 | KX447679 | |
| Botryosphaeria pseudoramosa | CERC 2001 | KX277989 | KX278094 |
| CERC 3455 | KX277997 | KX278102 | |
| Botryosphaeria qinlingensis | CFCC 52984 | MK434301 | MK425020 |
| CFCC 52985 | MK434302 | MK425021 | |
| Botryosphaeria qingyuanensis | CERC 2946 | KX278000 | KX278105 |
| CERC 2947 | KX278001 | KX278106 | |
| Botryosphaeria ramosa | CBS 122069 | EU144055 | EU144070 |
| Botryosphaeria rosaceae | CFCC 82350 | KX197079 | KX197097 |
| DZP B | KX197076 | KX197096 | |
| Botryosphaeria scharifii | IRAN1529C | JQ772020 | JQ772057 |
| IRAN1543C | JQ772019 | JQ772056 | |
| Botryosphaeria sinensia | BJFU DZP141005-06 | KT343254 | KU221233 |
| BJFU DZP141111-10 | KT343256 | KU221234 | |
| Botryosphaeria wangensis | CERC 2298 | KX278002 | KX278107 |
| CERC 2300 | KX278004 | KX278109 | |
| Strain Number | Conidial Dimensions (Length × Width, µm) | ||
|---|---|---|---|
| Range (50 Conidia) | Mean | Length/Width Ratio | |
| SG3 | (14–)16.8–21.8 × (3.5–)4.2–5.6 | 18.9 × 4.9 | 3.2–5.2 |
| AcAs2 | 15.9–21 × (3.9–)4.3-5.8(–6.6) | 18.7 × 5.2 | 2.6–4.7 |
| AcE3 | (14.2–)16.4–21.6 × (3.4–)4.6–5.4 | 19.4 × 5.1 | 3.1–5.4 |
| AcSe1 | (13.4–)15.3–21(–22.9) × 3.8–5.7 | 18.2 × 5.1 | 2.3–5.1 |
| CLRi2 | 14.9–21.8 × (3.9–)4.3–5.8(–6.6) | 18.7 × 5.2 | 2.6–5.4 |
| Mp26j | (17.5–)18.5–29.7 × 3.7–7(–7.5) | 22.3 × 5.1 | 3.5–6.1 |
| Mp30p | (16.5–)19.0–28.7 × (3.2–)4.0–7.3 | 22.3 × 5.1 | 3.7–6.1 |
| MgBt1 | (13–)16.8-21.2 × 4.6–5.5(–6.2) | 17.3 × 5.2 | 2.4–4.6 |
| MgPC6 | (12.8–)15.3–21 × 3.8–5.7 | 15.6 × 4.9 | 2.1–5.5 |
| MgPC7 | (13.4–)15.5–21(–22.9) × 3.3–5.7 | 18.9 × 4.4 | 2.3–6.0 |
| OvdF3e | (17–)18.5–25.9 × (3.0–)3.5–7.4 | 24.3 × 5.3 | 3.0–6.1 |
| OvFs/g | (17.5–)18.5–28.7 × (2.8–)3.7–7.4 | 23.5 × 5.1 | 3.5–6.0 |
| Th/g2017 | (15.2–)16.5–25.7 × (2.7–)3.9–6.8 | 20.3 × 4.8 | 2.9–5.5 |
| ThgI/10 | (16.0–)18.5–28.4 × (3.1–)3.5–7.0 | 22.0 × 5.1 | 3.4–6.0 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zimowska, B.; Okoń, S.; Becchimanzi, A.; Krol, E.D.; Nicoletti, R. Phylogenetic Characterization of Botryosphaeria Strains Associated with Asphondylia Galls on Species of Lamiaceae. Diversity 2020, 12, 41. https://doi.org/10.3390/d12020041
Zimowska B, Okoń S, Becchimanzi A, Krol ED, Nicoletti R. Phylogenetic Characterization of Botryosphaeria Strains Associated with Asphondylia Galls on Species of Lamiaceae. Diversity. 2020; 12(2):41. https://doi.org/10.3390/d12020041
Chicago/Turabian StyleZimowska, Beata, Sylwia Okoń, Andrea Becchimanzi, Ewa Dorota Krol, and Rosario Nicoletti. 2020. "Phylogenetic Characterization of Botryosphaeria Strains Associated with Asphondylia Galls on Species of Lamiaceae" Diversity 12, no. 2: 41. https://doi.org/10.3390/d12020041
APA StyleZimowska, B., Okoń, S., Becchimanzi, A., Krol, E. D., & Nicoletti, R. (2020). Phylogenetic Characterization of Botryosphaeria Strains Associated with Asphondylia Galls on Species of Lamiaceae. Diversity, 12(2), 41. https://doi.org/10.3390/d12020041









