Phylogeny and Biogeography of Branchipolynoe (Polynoidae, Phyllodocida, Aciculata, Annelida), with Descriptions of Five New Species from Methane Seeps and Hydrothermal Vents
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling
2.2. Morphology
2.3. DNA Extraction, Amplification, and Sequencing
2.4. Phylogenetic Analysis and Species Delimitation
2.5. Haplotype Networks and Population Genetics
2.6. Character Transformations
3. Results
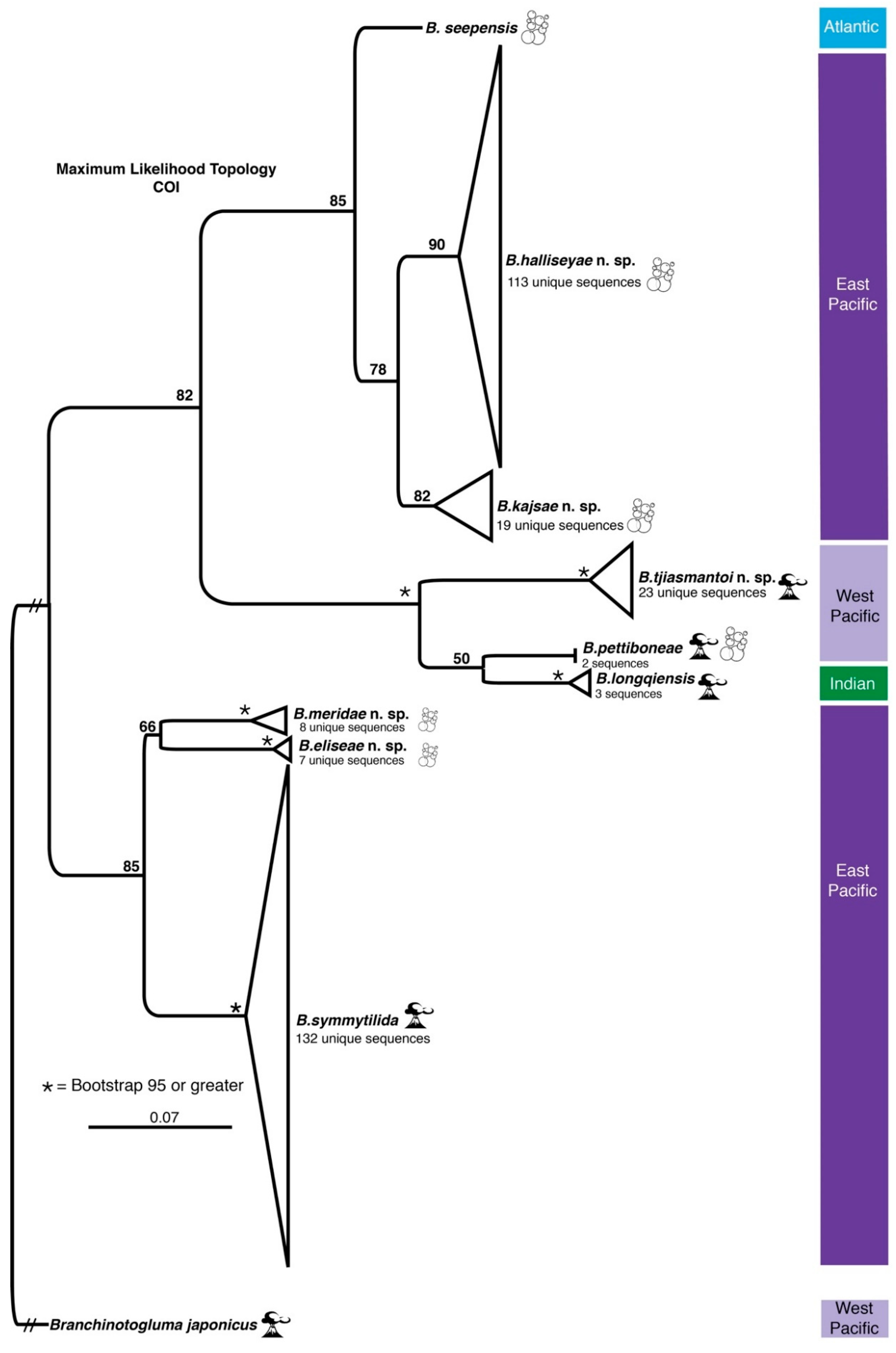
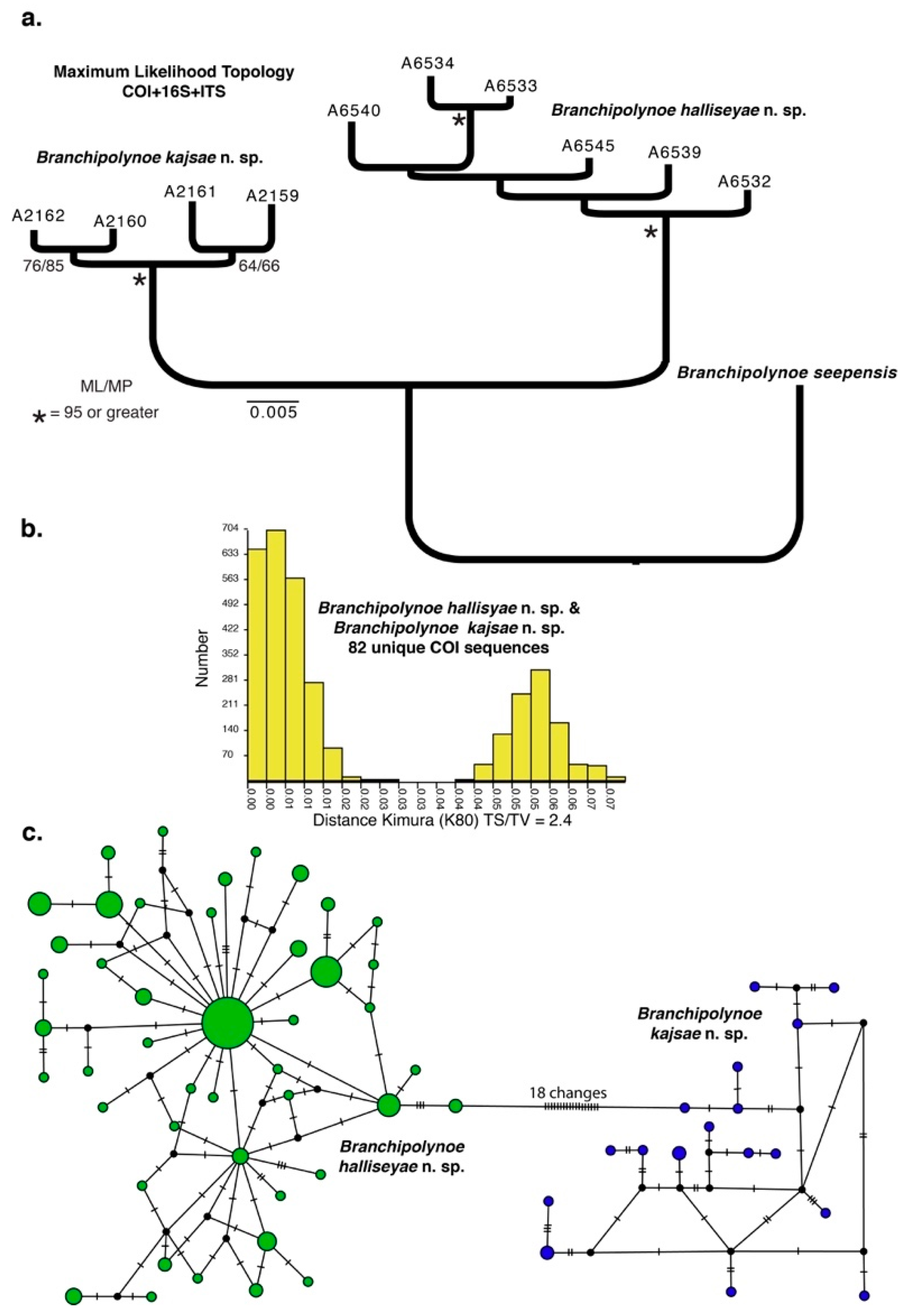
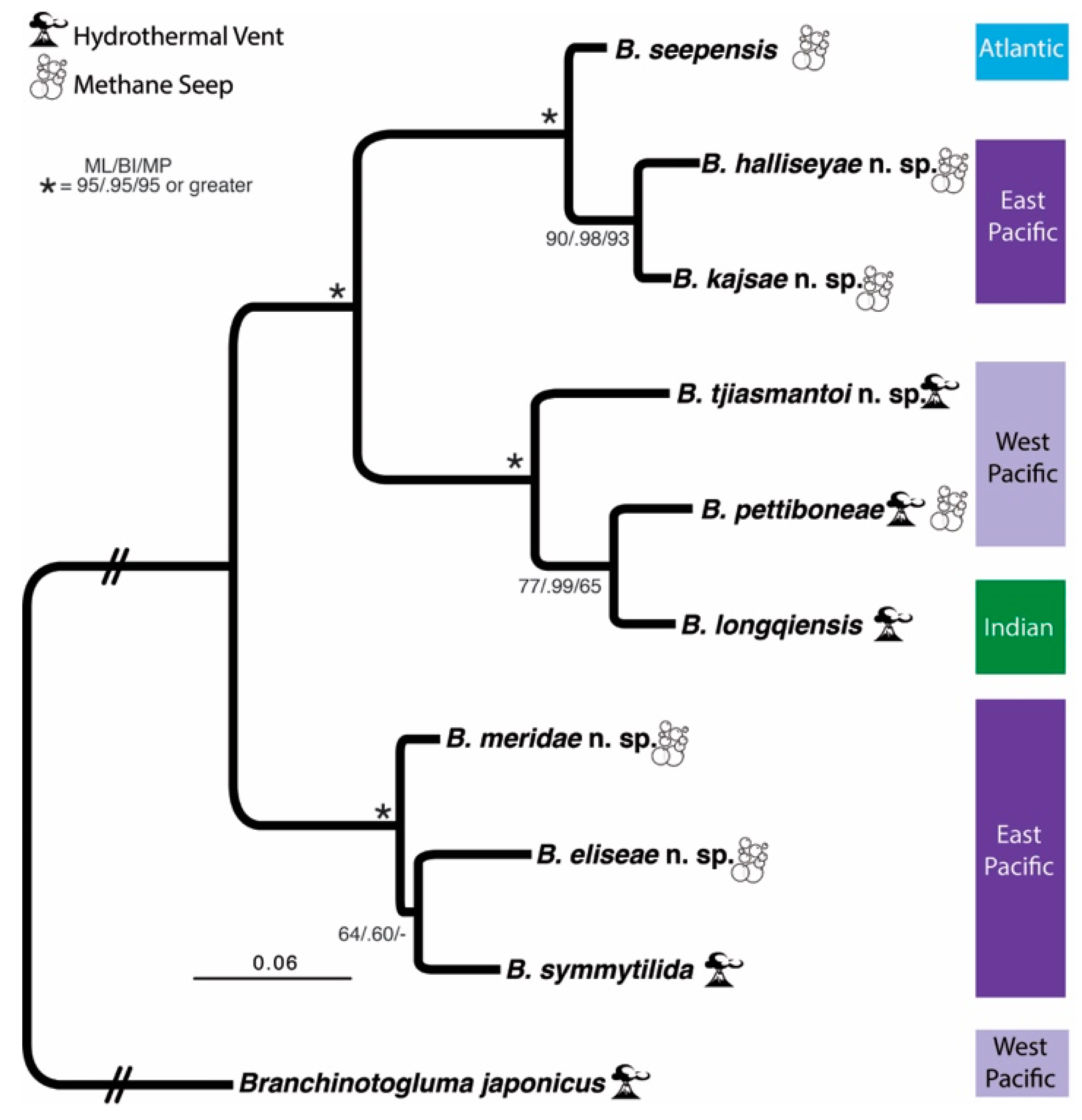
3.1. Species Delimitation and Phylogeny
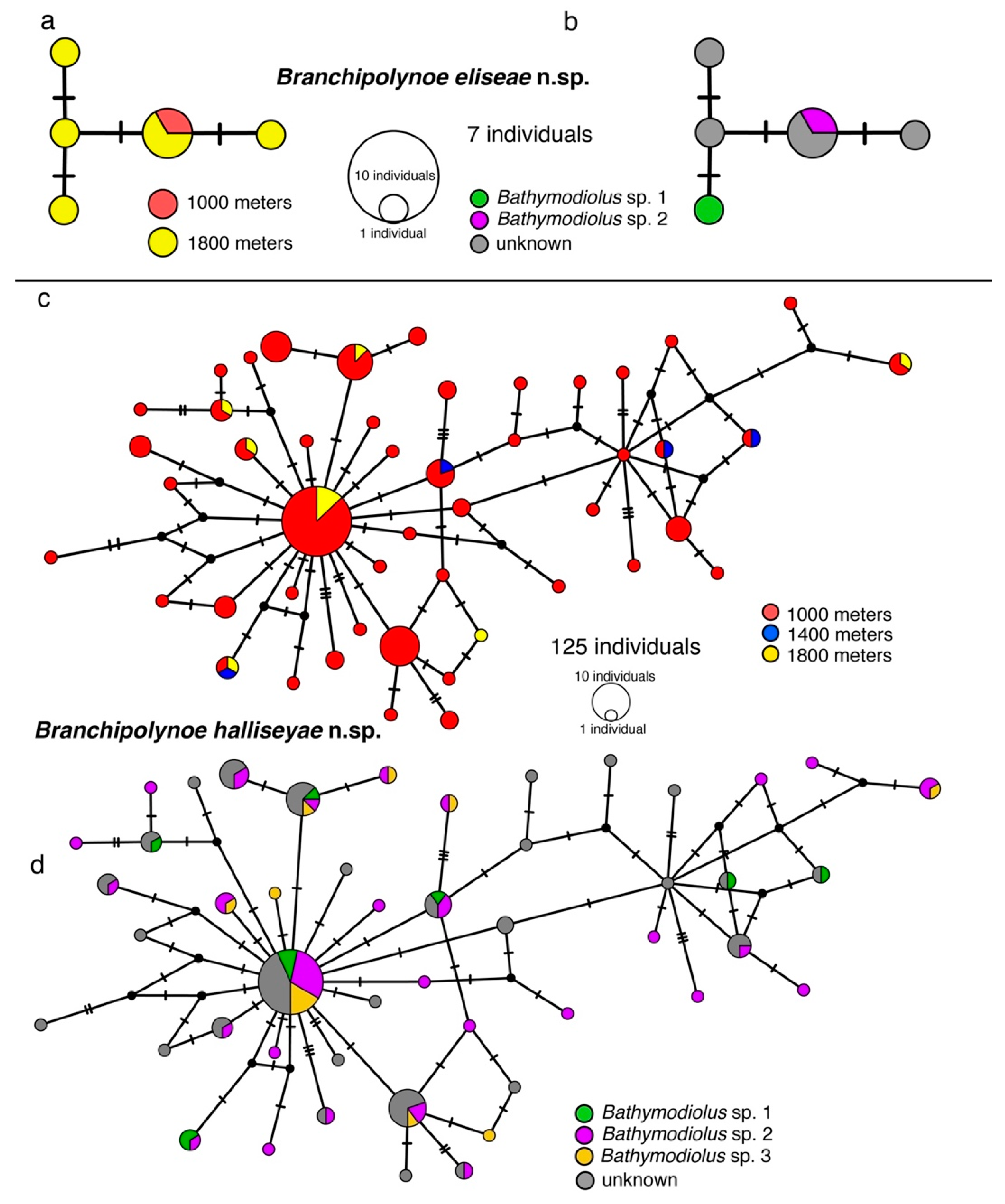
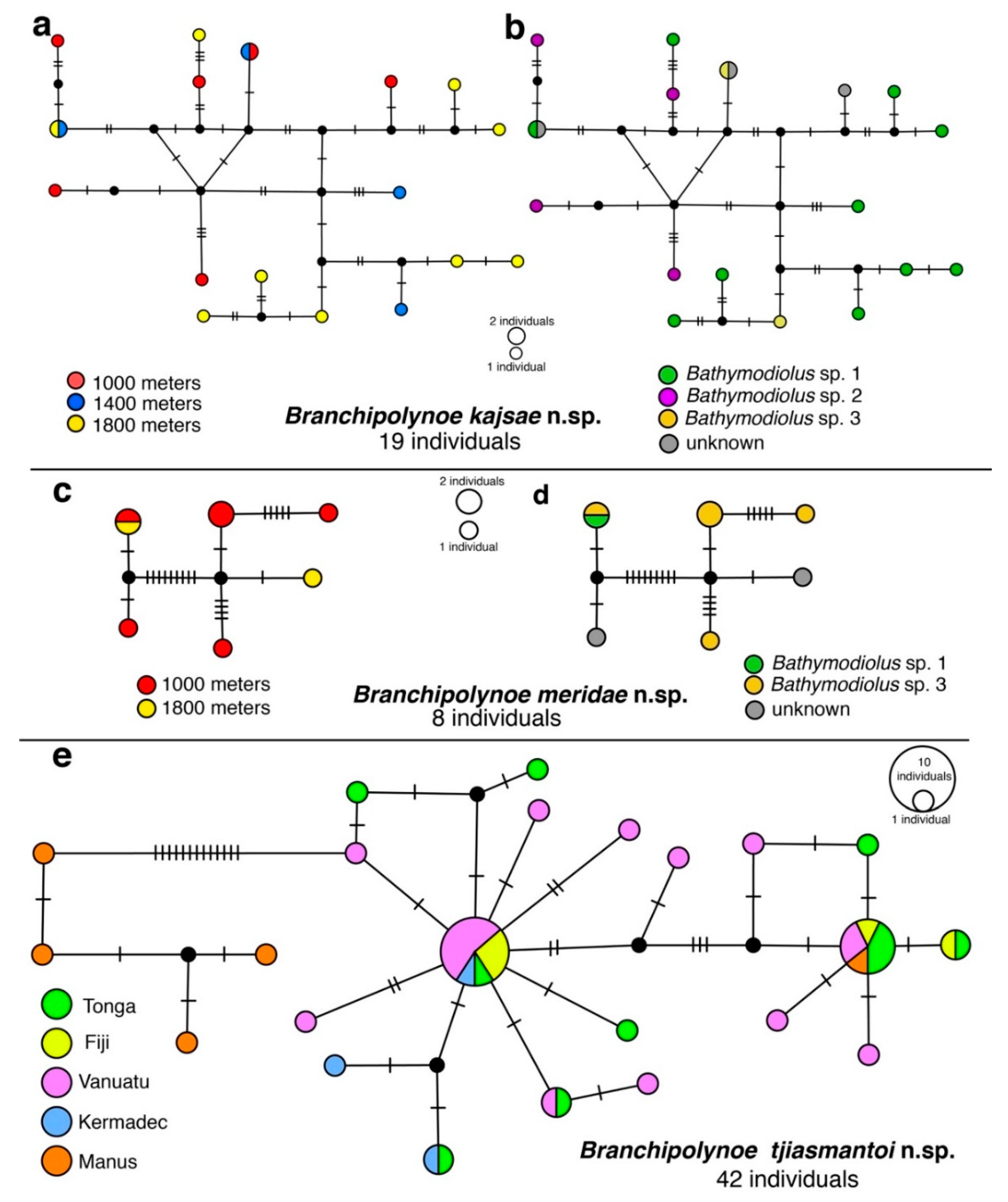
3.2. Haplotype Networks, Connectivity Across Hosts and Depths
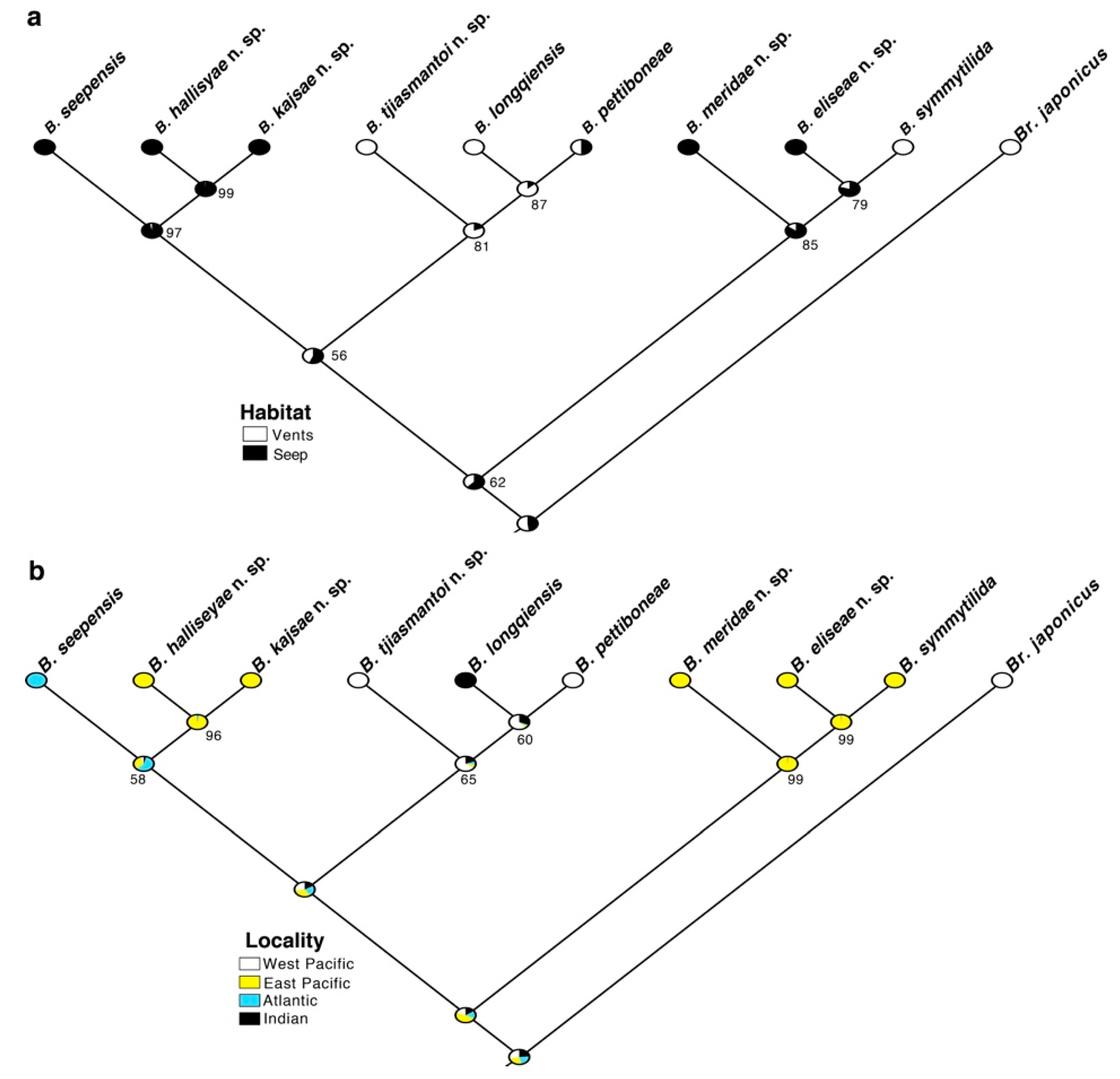
3.3. Character Transformations
4. Discussion
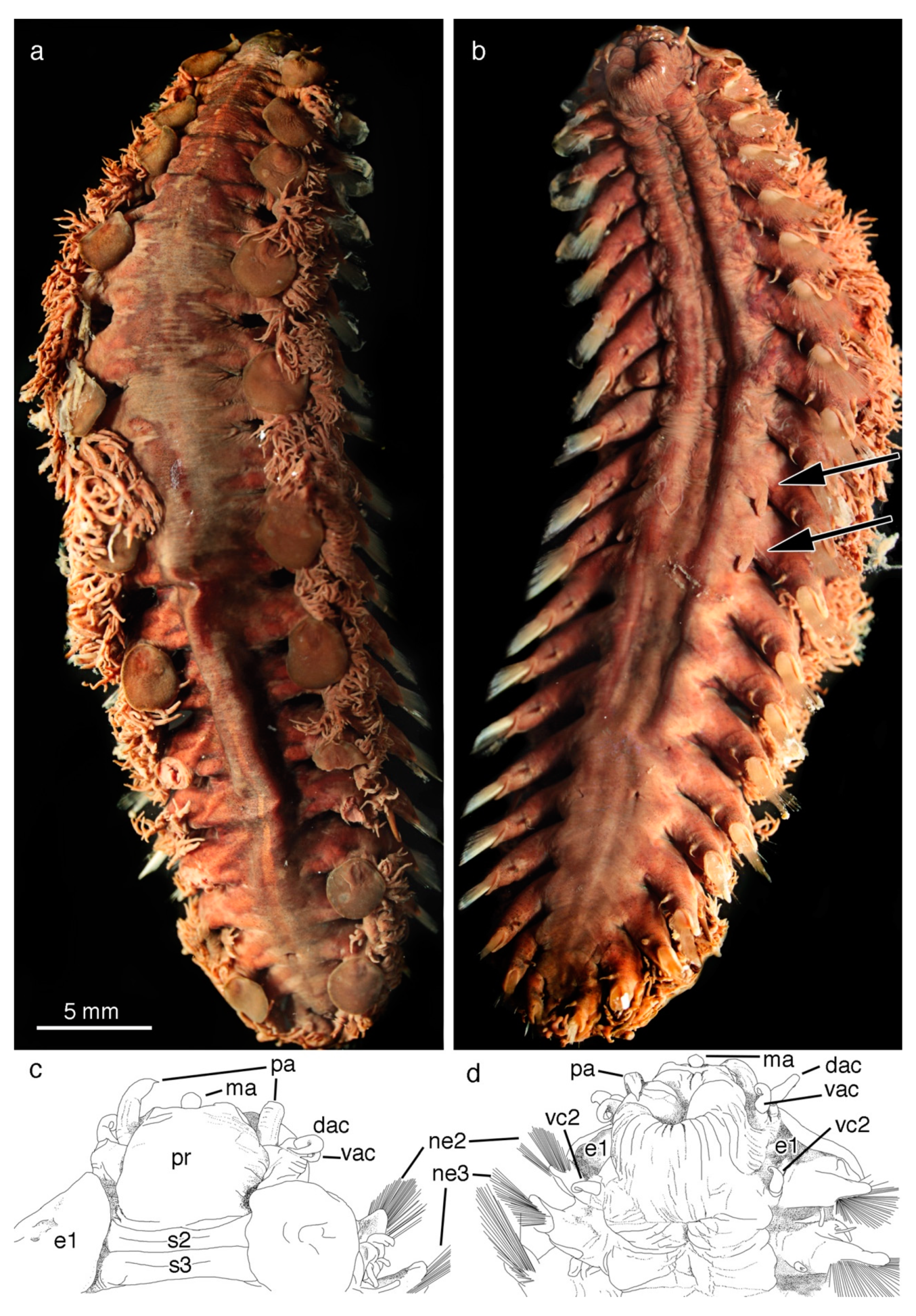
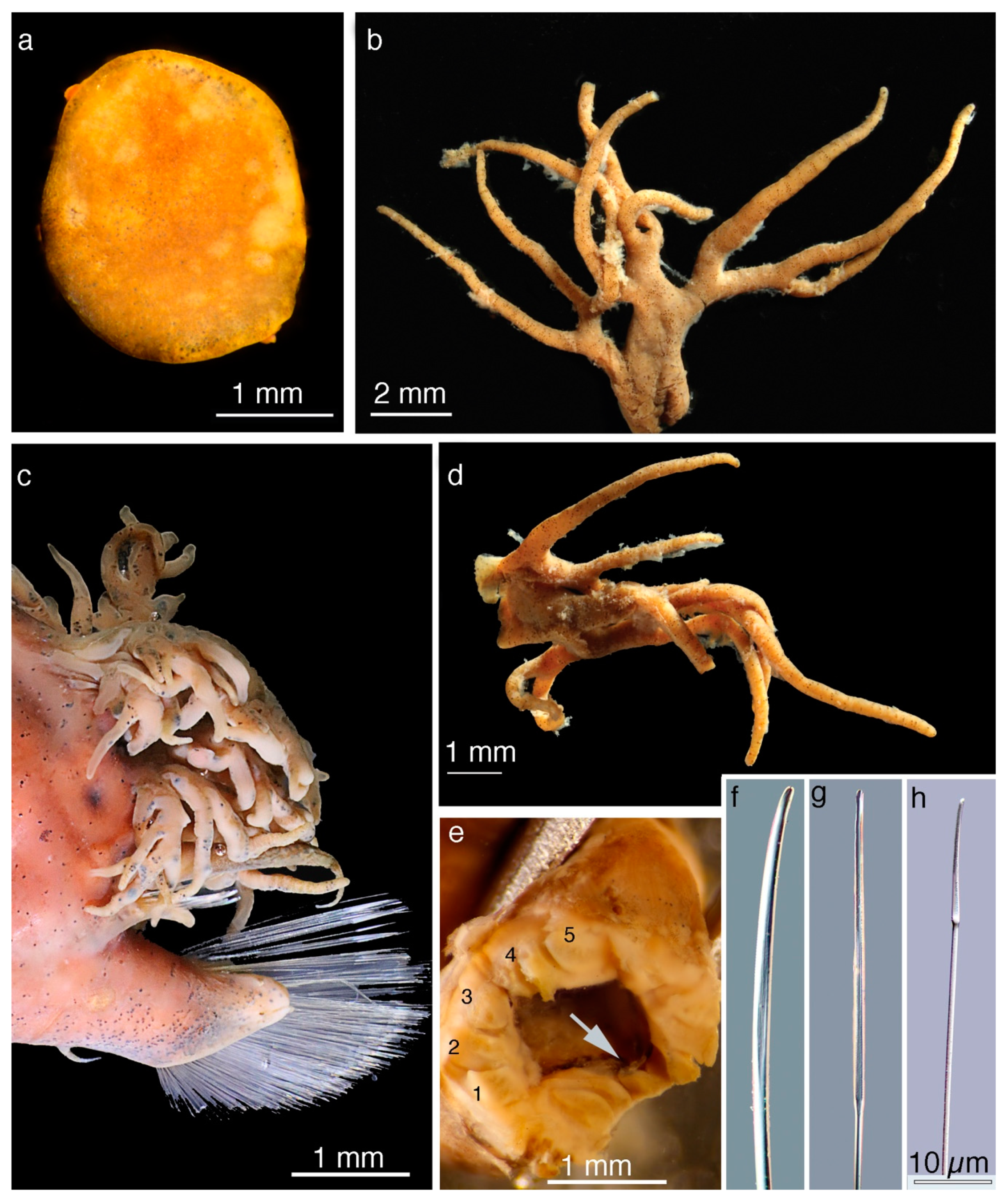
5. Taxonomy
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gonzalez, B.C.; Martínez, A.; Borda, E.; Iliffe, T.M.; Eibye-Jacobsen, D.; Worsaae, K. Phylogeny and systematics of Aphroditiformia. Cladistics 2017, 34, 225–259. [Google Scholar] [CrossRef]
- Norlinder, E.; Nygren, A.; Wiklund, H.; Pleijel, F. Phylogeny of scale-worms (Aphroditiformia, Annelida), assessed from 18SrRNA, 28SrRNA, 16SrRNA, mitochondrial cytochrome c oxidase subunit I (COI), and morphology. Mol. Phylogenet. Evol. 2012, 65, 490–500. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Sun, J.; Rouse, G.W.; Wiklund, H.; Pleijel, F.; Watanabe, H.K.; Cheng, C.; Qian, P.Y.; Qiu, J.W. Phylogeny, evolution and mitochondrial gene order rearrangement in scale worms (Aphroditiformia, Annelida): Insights from low-coverage genome sequencing. Mol. Phylogenet. Evol. 2018, 125, 220–231. [Google Scholar] [CrossRef] [PubMed]
- WoRMS Editorial Board. World Register of Marine Species. 2018. Available online: http://www.marinespecies.org at VLIZ (accessed on 8 June 2018).
- Pettibone, M.H. Additional branchiate scale-worms (Polychaeta: Polynoidae) from Galapagos hydrothermal vent and rift- area off western Mexico at 21 N. Proc. Biol. Soc. Wash. 1985, 98, 447–469. [Google Scholar]
- Pettibone, M.H. An additional new scale worm (Polychaeta: Polynoidae) from the hydrothermal rift area off western Mexico at 21 N. Proc. Biol. Soc. Wash. 1985, 98, 150–157. [Google Scholar]
- Pettibone, M.H. A new scale-worm commensal with deep-sea mussels on the Galapagos hydrothermal vent (Polychaeta: Polynoidae). Proc. Biol. Soc. Wash. 1984, 97, 226–239. [Google Scholar]
- Desbruyères, D.; Laubier, L. Exploitation of a concentrated organic matter source in the deep sea: Role of a new polychaetous annelid C. r. hebd. Séanc. Acad. Sci. Paris 1988, 307, 329–336. [Google Scholar]
- Miura, T. Two new scale-worms (Polynoidae: Polychaeta) from the Lau back-arc and north Fiji basins, South Pacific Ocean. Proc. Biol. Soc. Wash. 1994, 107, 532–543. [Google Scholar]
- Hourdez, S.; Jouin-Toulmond, C. Functional anatomy of the respiratory system of Branchipolynoe species (Polychaeta, Polynoidae), commensal with Bathymodiolus species (Bivalvia, Mytilidae) from deep-sea hydrothermal vents. Zoomorphology 1998, 118, 225–233. [Google Scholar] [CrossRef]
- Desbruyères, D.; Gaill, F.; Laubier, L.; Fouquet, Y. Polychaetous annelids from hydrothermal vent ecosystems: An ecological overview. Bull. Biol. Soc. Wash. 1985, 6, 103–116. [Google Scholar]
- Colaço, A.; Dehairs, F.; Desbruyères, D. Nutritional relations of deep-sea hydrothermal fields at the Mid-Atlantic Ridge: A stable isotope approach. Deep Sea Res. Part I 2002, 49, 395–412. [Google Scholar] [CrossRef]
- Britayev, T.A.; Krylova, E.M.; Martin, D.; Cosel, R.; Aksiuk, T.S. Symbiont-host interraction in the association of the scaleworm Branchipolynoe aff. seepensis (Polychaeta: Polynoidae) with the hydrothermal mussel Bathymodiolus spp. (Bivalvia: Mytilidae). InterRidge News 2003, 12, 13–16. [Google Scholar]
- Britayev, T.A.; Martin, D.; Krylova, E.M.; von Cosel, R.; Aksuik, T.S. Life-history traits of the symbiotic scale-worm Branchipolynoe seepensis and its relationships with host mussels of the genus Bathymodiolus Hydrothermal Vents. Mar. Ecol. 2007, 28, 36–48. [Google Scholar] [CrossRef]
- Ward, M.; Shields, J.; Van Dover, C. Parasitism in species of Bathymodiolus (Bivalvia: Mytilidae) mussels from deep-sea seep and hydrothermal vents. Dis. Aquat. Org. 2004, 62, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Van Dover, C.; Trask, J.; Gross, J.; Knowlton, A. Reproductive biology of free-living and commensal polynoid polychaetes at the Lucky Strike hydrothermal vent field (Mid-Atlantic Ridge). Mar. Ecol. Prog. Ser. 1999, 181, 201–214. [Google Scholar] [CrossRef]
- Jollivet, D.; Empis, A.; Baker, M.C.; Hourdez, S.; Comtet, T.; Jouin-Toulmond, C.; Desbruyères, D.; Tyler, P.A. Reproductive biology, sexual dimorphism, and population structure of the deep sea hydrothermal vent scale-worm, Branchipolynoe seepensis (Polychaeta: Polynoidae). J. Mar. Biol. Assoc. UK 2000, 80, 55–68. [Google Scholar] [CrossRef]
- Gaudron, S.M.; Hourdez, S.; Olu, K. Aspects on gametogenesis, fertilization and embryogenesis of two deep-sea polychaetes from Eastern Atlantic cold seeps. Deep Sea Res. Part I 2017, 129, 59–68. [Google Scholar] [CrossRef]
- Pettibone, M.H. A new scale-worm commensal with deep-sea mussels in the seep-sites at the Florida escarpment in the eastern Gulf of Mexico (Polychaeta: Polynoidae: Branchipolynoinae). Proc. Biol. Soc. Wash. 1986, 99, 444–451. [Google Scholar]
- Miura, T.; Hashimoto, J. Two new branchiate scale-worms (Polynoidae: Polychaeta) from the hydrothermal vent of the Okinawa Trough and the volcanic seamount off Chichijima Island. Proc. Biol. Soc. Wash. 1991, 104, 166–174. [Google Scholar]
- Zhou, Y.; Zhang, D.; Lu, B.; Wang, C. Description of a new branchiate scale-worm (Polychaeta: Polynoidae) from the hydrothermal vent on Southwest Indian Ocean Ridge. Zootaxa 2017, 4282, 123–134. [Google Scholar] [CrossRef]
- Fisher, C.; Childress, J.; Arp, A.; Brooks, J.; Distel, D.; Favuzzi, J.; Felbeck, H.; Hessler, R.; Johnson, K.; Kennicutt, M.; et al. Microhabitat variation in the hydrothermal vent mussel, Bathymodiolus thermophilus, at the Rose Garden vent on the Galapagos Rift. Deep. Sea Res. Part A. Oceanogr. Res. Pap. 1988, 35, 1769–1791. [Google Scholar] [CrossRef]
- Chevaldonné, P.; Jollivet, D.; Feldman, R.A.; Desbruyères, D.; Lutz, R.A.; Vrijenhoek, R.C. Commensal scale-worms of the genus Branchipolynoe (Polychaeta: Polynoidae) at deep-sea hydrothermal vents and cold seeps. Cah. De Biol. Mar. 1998, 39, 347–350. [Google Scholar]
- Hurtado, L.A.; Lutz, R.A.; Vrijenhoek, R.C. Distinct patterns of genetic differentiation among annelids of eastern Pacific hydrothermal vents. Mol. Ecol. 2004, 13, 2603–2615. [Google Scholar] [CrossRef] [PubMed]
- Miura, T. Branchipolynoe Pettiboneae Miura & Hashimoto, 1991. In Handbook of Deep-Sea Hydrothermal Vent Fauna. Denisia 18; Desbruyères, D., Segonzac, M., Bright, M., Eds.; Biologiezentrum der Oberösterreichischen Landesmuseen: Linz, Austria, 2006; p. 231. [Google Scholar]
- Sui, J.; Li, X. A new species and new record of deep-sea scale-worms (Polynoidae: Polychaeta) from the Okinawa Trough and the South China Sea. Zootaxa 2017, 4238, 562. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chen, C.; Qiu, J.W. Sexually Dimorphic Scale Worms (Annelida: Polynoidae) From Hydrothermal Vents in the Okinawa Trough: Two New Species and Two New Sex Morphs. Front. Mar. Sci. 2018, 5, 112. [Google Scholar] [CrossRef]
- Miura, T. Branchipolynoe Pettiboneae Miura & Hashimoto, 1991. In Handbook of Deep-Sea Hydrothermal Vent Fauna; Desbruyères, D., Segonzac, M., Eds.; IFREMER: Brest, France, 1997; p. 55. [Google Scholar]
- Van Dover, C.L.; Humphris, S.E.; Fornari, D.; Cavanaugh, C.M.; Collier, R.; Goffredi, S.K.; Hashimoto, J.; Lilley, M.D.; Reysenbach, A.L.; Shank, T.M.; et al. Biogeography and ecological setting of Indian Ocean hydrothermal vents. Science 2001, 294, 818–823. [Google Scholar] [CrossRef] [PubMed]
- Copley, J.T.; Marsh, L.; Glover, A.G.; Hühnerbach, V.; Nye, V.E.; Reid, W.D.K.; Sweeting, C.J.; Wigham, B.D.; Wiklund, H. Ecology and biogeography of megafauna and macrofauna at the first known deep-sea hydrothermal vents on the ultraslow-spreading Southwest Indian Ridge. Sci. Rep. 2016, 6, 39158. [Google Scholar] [CrossRef]
- Faure, B.; Schaeffer, S.W.; Fisher, C.R. Species distribution and population connectivity of deep-sea mussels at hydrocarbon seeps in the Gulf of Mexico. PLoS ONE 2015, 10, e011846019. [Google Scholar] [CrossRef]
- Segonzac, M. Les peuplements associés à l’hydrothermalisme océanique du Snake Pit (dorsale médio-atlantique; 23° N, 3480 m): composition et microdistribution de la mégafaune. Comptes Rendus Acad. Sci. Paris 1992, 314, 593–600. [Google Scholar]
- Desbruyères, D.; Alayse, A.M.; Antoine, E.; Barbier, G.; Barriga, F.; Biscoito, M.; Briand, P.; Brulport, J.P.; Comtet, T.; Cornec, L.; et al. New information on the ecology of deep-sea vent communities in the Azores Triple Junction area: Preliminary results of the Diva 2 cruise (May 31–July 4, 1994). InterRidge News 1994, 3, 18–19. [Google Scholar]
- Plouviez, S.; Daguin-Thiébaut, C.; Hourdez, S.; Jollivet, D. Juvenile and adult scale worms Branchipolynoe seepensis in Lucky Strike hydrothermal vent mussels are genetically unrelated. Aquat. Biol. 2008, 3, 79–87. [Google Scholar] [CrossRef]
- Daguin, C.; Jollivet, D. Development and cross-amplification of nine polymorphic microsatellite markers in the deep-sea hydrothermal vent polychaete Branchipolynoe seepensis. Mol. Ecol. Notes 2005, 5, 780–783. [Google Scholar] [CrossRef]
- Levin, L.A.; Orphan, V.J.; Rouse, G.W.; Rathburn, A.E.; Ussler, W.; Cook, G.S.; Goffredi, S.K.; Perez, E.M.; Waren, A.; Grupe, B.M.; et al. A hydrothermal seep on the Costa Rica margin: middle ground in a continuum of reducing ecosystems. Proc. R. Soc. B Biol. Sci. 2012, 279, 2580–2588. [Google Scholar] [CrossRef] [PubMed]
- Levin, L.A.; Mendoza, G.F.; Grupe, B.; Gonzalez, J.P.; Jellison, B.; Rouse, G.W.; Thurber, A.R.; Waren, A. Biodiversity on the rocks: Macrofauna inhabiting authigenic carbonate at Costa Rica methane seeps. PLoS ONE 2015, 10, e0131080. [Google Scholar] [CrossRef] [PubMed]
- Folmer, O.; Black, M.; Hoeh, W.R.; Lutz, R.A.; Vrijenhoek, R.C. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotech. 1994, 3, 294–299. [Google Scholar]
- Palumbi, S.R. Nucleic Acids II: The Polymerase Chain Reaction. In Molecular Systematics; Hillis, D.M., Moritz, C., Mable, B.K., Eds.; Sinauer Associates: Sunderland, MA, USA, 1996; pp. 205–247. [Google Scholar]
- Nygren, A.; Eklöf, J.; Pleijel, F. Arctic-boreal sibling species of Paranaitis (Polychaeta, Phyllodocidae). Mar. Biol. Res. 2009, 5, 315–327. [Google Scholar] [CrossRef][Green Version]
- Glover, A.G.; Goetze, E.; Dhalgren, T.G.; Smith, C.R. Morphology, reproductive biology and genetic structure of the whale-fall and hydrothermal vent specialist, Bathykurila guaymasensis Pettibone, 1989 (Annelida: Polynoidae). Mar. Ecol. 2005, 26, 223–234. [Google Scholar] [CrossRef]
- Roy, K.O.L.; Von Cosel, R.; Hourdez, S.; Carney, S.; Jollivet, D. Amphi-Atlantic cold-seep Bathymodiolus species complexes across the equatorial belt. Deep. Sea Res. Part I Oceanogr. Res. Pap. 2007, 54, 1890–1911. [Google Scholar]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Plouviez, S.; Shank, T.M.; Faure, B.; Daguin-Thiebaut, C.; Viard, F.; Lallier, F.H.; Jollivet, D. Comparative phylogeography among hydrothermal vent species along the East Pacific Rise reveals vicariant processes and population expansion in the South. Mol. Ecol. 2009, 18, 3903–3917. [Google Scholar] [CrossRef] [PubMed]
- Van Audenhaege, L.; Farinas-Bermejo, A.; Schultz, T.; Van Dover, C.L. An environmental baseline for food webs at deep-sea hydrothermal vents in Manus Basin (Papua New Guinea). Deep Sea Res. Part I 2019, 148, 88–99. [Google Scholar] [CrossRef]
- Vaidya, G.; Lohman, D.J.; Meier, R. Sequence Matrix: Concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 2011, 27, 171–180. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Silvestro, D.; Michalak, I. raxmlGUI: A graphical front-end for RAxML. Org. Divers. Evol. 2012, 12, 335–337. [Google Scholar] [CrossRef]
- Ronquist, F.; Teslenko, M.; Van Der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian Phylogenetic Inference and Model Choice Across a Large Model Space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [PubMed]
- Swofford, D.L. PAUP: Phylogenetic Analysis Using Parsimony (and Other Methods); Version 4; Sinauer Associates: Sunderland, MA, USA, 2002. [Google Scholar]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar]
- Guindon, S.; Gascuel, O. A Simple, Fast, and Accurate Algorithm to Estimate Large Phylogenies by Maximum Likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef]
- Puillandre, N.; Lambert, A.; Brouillet, S.; Achaz, G. ABGD, Automatic Barcode Gap Discovery for primary species delimitation. Mol. Ecol. 2012, 21, 1864–1877. [Google Scholar] [CrossRef]
- Leigh, J.W.; Bryant, D. POPART: Full-feature software for haplotype network construction. Methods Ecol. Evol. 2015, 6, 1110–1116. [Google Scholar]
- Maddison, W.P.; Maddison, D.R. Mesquite: A Modular System for Evolutionary Analysis. Version 3.5. 2018. Available online: http://www.mesquiteproject.org (accessed on 8 June 2018).
- Lewis, P.O. A Likelihood Approach to Estimating Phylogeny from Discrete Morphological Character Data. Syst. Biol. 2001, 50, 913–925. [Google Scholar] [CrossRef] [PubMed]
- Bickford, D.; Lohman, D.J.; Sodhi, N.S.; Ng, P.K.; Meier, R.; Winker, K.; Ingram, K.K.; Das, I. Cryptic species as a window on diversity and conservation. Trends Ecol. Evol. 2007, 22, 148–155. [Google Scholar] [CrossRef] [PubMed]
- Peek, A.S.; Gustafson, R.G.; Lutz, R.A.; Vrijenhoek, R.C. Evolutionary relationships of deep-sea hydrothermal vent and cold-water seep clams (Bivalvia: Vesicomyidae): Results from the mitochondrial cytochrome oxidase subunit I. Mar. Biol. 1997, 130, 151–161. [Google Scholar] [CrossRef]
- Johnson, S.B.; Warén, A.; Tunnicliffe, V.; van Dover, C.L.; Wheat, C.G.; Schultz, T.F.; Vrijenhoek, R.C. Molecular taxonomy and naming of five cryptic species of Alviniconcha snails (Gastropoda: Abyssochrysoidea) from hydrothermal vents. Syst. Biod. 2015, 13, 278–295. [Google Scholar] [CrossRef]
- Borda, E.; Kudenov, J.D.; Chevaldonné, P.; Desbruyères, D.; Blake, J.A.; Fabri, M.C.; Hourdez, S.; Shank, T.M.; Wilson, N.G.; Pleijel, F.; et al. Cryptic species of Archinome (Annelida: Amphinomidae) from hydrothermal vents and cold seeps. Proc. Roy Soc. B 2013, 280, 20131876. [Google Scholar] [CrossRef] [PubMed]
- Stiller, J.; Rousset, V.; Pleijel, F.; Chevaldonné, P.; Vrijenhoek, R.C.; Rouse, G.W. Phylogeny, biogeography and systematics of hydrothermal vent and methane seep Amphisamytha (Ampharetidae, Annelida), with descriptions of three new species. Syst. Biodivers. 2013, 11, 35–65. [Google Scholar] [CrossRef]
- Halt, M.N.; Kupriyanova, E.K.; Cooper, S.J.B.; Rouse, G.W. Naming species with no morphological indicators: Species status of Galeolaria caespitosa (Annelida, Serpulidae) inferred from nuclear and mitochondrial gene sequences and morphology. Invertebr. Syst. 2009, 23, 205–222. [Google Scholar] [CrossRef]
- Nygren, A. Cryptic polychaete diversity: A review. Zool. Scr. 2013, 43, 172–183. [Google Scholar] [CrossRef]
- Berriman, J.S.; Ellingson, R.A.; Awbrey, J.D.; Rico, D.M.; Valdés, Á.A.; Wilson, N.G.; Aguilar, A.; Herbert, D.G.; Hirano, Y.M.; Trowbridge, C.D.; et al. A biting commentary: Integrating tooth characters with molecular data doubles known species diversity in a lineage of sea slugs that consume “killer algae”. Mol. Phylogenet. Evol. 2018, 126, 356–370. [Google Scholar] [CrossRef] [PubMed]
- Meyer, C.P.; Paulay, G. DNA barcoding: Error rates based on comprehensive sampling. PLoS Biol. 2005, 3, 422–431. [Google Scholar] [CrossRef] [PubMed]
- Meier, R.; Zhang, G.; Ali, F. The Use of Mean Instead of Smallest Interspecific Distances Exaggerates the Size of the “Barcoding Gap” and Leads to Misidentification. Syst. Biol. 2008, 57, 809–813. [Google Scholar] [CrossRef] [PubMed]
- Nygren, A.; Norlinder, E.; Panova, M.; Pleijel, F. Colour polymorphism in the polychaete Harmothoe imbricata (Linnaeus, 1767). Mar. Biol. Res. 2011, 7, 54–62. [Google Scholar] [CrossRef]
- O’Dea, A.; Lessios, H.A.; Coates, A.G.; Eytan, R.I.; Restrepo-Moreno, S.A.; Cione, A.L.; Collins, L.S.; de Queiroz, A.; Farris, D.W.; Norris, R.D.; et al. Formation of the Isthmus of Panama. Sci. Adv. 2016, 2, e1600883. [Google Scholar] [CrossRef] [PubMed]
- Farris, D.W.; Jaramillo, C.; Bayona, G.; Restrepo-Moreno, S.A.; Montes, C.; Cardona, A.; Mora, A.; Speakman, R.J.; Glascock, M.D.; Valencia, V. Fracturing of the Panamanian Isthmus during initial collision with South America. Geology 2011, 39, 1007–1010. [Google Scholar] [CrossRef]
- Rouse, G.W.; Pleijel, F. Polychaetes; Oxford University Press: London, UK, 2001. [Google Scholar]




| Scientific Name | Site | COI | 16S | ITS | Specimen: SIO-BIC (A), MZUCR, or MNHN (IA) |
|---|---|---|---|---|---|
| Branchipolynoe symmytilida (type species) | German Flats, EPR | MH369876-77 | MH396821-22 | MH396787-88 | A6554-55 |
| Choo Choo, EPR | MH369875 | MH396823 | - | A6556 | |
| Branchipolynoe seepensis | Florida Escarpment, GM | MH369885 | MH596848 | - | A6553 |
| Branchipolynoe kajsae n. sp. | Jaco Scar, CR | MH369859-62 | MH396799-800, MH396811-12 | MH396774, MH396776, MH396782-83 | A2159-62 |
| MH369872-73 | MH396816, MH396819 | - | A6516, A6518 | ||
| - | MH396817-18, MH396820 | - | A6517, A6519-20 | ||
| MH369906-07, MH370018 | - | - | A6652, A6669, MZUCR 405-01 | ||
| Mound 12, CR | MH369893, MH369899, MH369902, MH369939-41 | - | - | A6574, A6597, A6611, A6626, A6662, A6703 | |
| Quepos Seep, CR | MH369923, MH369961, MH369983-84 | - | - | A6707, A6710, A6712-13 | |
| Branchipolynoe halliseyae n. sp. | Mound 12, CR | MH369853-56, MH369858, MH369870 | MH396795, MH396801, MH396803-04, MH396810, MH396815 | MH396775, MH396777-81 | A6532-34, A6539-40, A6545 |
| MH369845, MH369847-51, MH369857 | MH396794, MH396797-98, MH396805-06, MH396808-09 | - | A2127-A2128, A6531, A6535-38 | ||
| MH369869 | - | MH396786 | A6544 | ||
| MH369841-44, MH369846, MH369852, MH369868, MH369871, MH369887-92, MH369894-98, MH369900-01, MH369903-04, MH369909-13, MH369915-22, MH369927-28, MH369938, MH369960, MH369962-82, MH369985-370015, MH370017, MH370019-22 | - | - | A1322, A2144, A6530, A6541-43, A6546-47, A6558, A6568, A6570, A6575, A6578, A6582-89, A6592-93, A6595-96, A6603-10, A6612, A6620-25, A6627-34, A6636-37, A6640-49, A6656-59, A6661, A6673-85, A6687-702, A6704-05, MZUCR 404-01 | ||
| Jaco Scar, CR | MH369886, MH369905, MH369926, MH369930-35, MH369937 | - | - | A6613, A6617, A6619, A6650-51, A6653-54, A6667, A6670, A6672 | |
| Quepos Seep, CR | MH369914, MH369924-25, MH369929 | - | - | A6706, A6708-09, A6711 | |
| Branchipolynoe meridae n. sp. | Jaco Scar, CR | MH369884 | MH396829 | MH396793 | A2131 |
| MH369936 | - | - | A6616 | ||
| Mound 12, CR | MH369865 | MH396796 | MH396784 | A6529 | |
| MH369866 | MH396807 | - | A6521 | ||
| - | MH396802, MH396813 | - | A6523, A6526 | ||
| MH369863-64, MH369867, MH370016 | - | - | A6524-25, A6527, MZUCR 404-02 | ||
| - | MH396814 | MH396785 | A6528 | ||
| Branchipolynoe eliseae n. sp. | Jaco Scar, CR | MH369878, MH369882-83 | MH396826-28 | MH396789, MH396791-92, | MZUCR 403-01, A6548, A6552 |
| MH369879-80 | MH396824-25 | - | A6549-50 | ||
| MH369881 | - | MH396790 | A6551 | ||
| Mound 12, CR | MH369908 | - | - | A6660 | |
| Branchipolynoe tjiasmantoi n. sp. | White Lady, Fiji | MH369874, MH369943-46 | - | - | A4009-10, A4011A-B, A5460 |
| Lau Basin, Tonga | MH369947, MH982238-41 | MH396830 | - | A8511 | |
| Niuatahi Seamount, Tonga | MH982230-35 | A9392 | |||
| Nifonea, Vanuatu | MH369942, MH369948-59, MH607403, MH982243-46 | - | - | A7303A, A7309A-K, A7310B-C, IA-2013-406-08 | |
| Haungaroa, NZ | MH982242 | - | - | IA-2013-340 | |
| Brothers Caldera, NZ | MH982236-37 | IA-2013-338-39 |
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | |
|---|---|---|---|---|---|---|---|---|---|
| 1.B. symmytilida | 0.022 | 0.307 | 0.317 | 0.267 | 0.101 | 0.118 | 0.356 | 0.412 | 0.419 |
| 2.B. seepensis | 0.136 | - | 0.072 | 0.080 | 0.306 | 0.272 | 0.396 | 0.368 | 0.375 |
| 3.B. kajsae n. sp. | 0.132 | 0.053 | 0.030 | 0.045 | 0.247 | 0.301 | 0.335 | 0.359 | 0.347 |
| 4.B. halliseyae n. sp. | 0.139 | 0.057 | 0.037 | 0.019 | 0.325 | 0.310 | 0.404 | 0.338 | 0.437 |
| 5.B. meridae n. sp. | 0.075 | 0.151 | 0.128 | 0.141 | 0.023 | 0.085 | 0.387 | 0.382 | 0.376 |
| 6.B. eliseae n. sp. | 0.082 | 0.131 | 0.139 | 0.137 | 0.065 | 0.006 | 0.413 | 0.302 | 0.302 |
| 7.B. tjiasmantoi n. sp. | 0.151 | 0.141 | 0.136 | 0.141 | 0.154 | 0.157 | 0.017 | 0.155 | 0.149 |
| 8.B. pettiboneae | 0.155 | 0.133 | 0.132 | 0.129 | 0.147 | 0.140 | 0.096 | 0.032 | 0.089 |
| 9.B. longqiensis | 0.156 | 0.140 | 0.135 | 0.142 | 0.154 | 0.157 | 0.091 | 0.064 | 0.011 |
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | |
|---|---|---|---|---|---|---|---|
| 1.B. symmytilida | - | 0.169 | 0.147 | 0.016 | 0.048 | 0.151 | 0.173 |
| 2.B. kajsae n. sp. | 0.126 | - | 0.027 | 0.215 | 0.201 | 0.260 | 0.255 |
| 3.B. halliseyae n. sp. | 0.113 | 0.026 | - | 0.188 | 0.195 | 0.229 | 0.215 |
| 4.B. meridae n. sp. | 0.016 | 0.149 | 0.137 | - | 0.055 | 0.249 | 0.246 |
| 5.B. eliseae n. sp. | 0.044 | 0.141 | 0.139 | 0.049 | - | 0.271 | 0.337 |
| 6.B. pettiboneae | 0.115 | 0.171 | 0.157 | 0.168 | 0.175 | - | 0.081 |
| 7.B. longqiensis | 0.127 | 0.169 | 0.150 | 0.167 | 0.200 | 0.069 | - |
| Characters | B. symmytilida | B. seepensis | B. pettiboneae | B. longqiensis | B. eliseae n. sp. | B. halliseyae n. sp. | B. kajsae n. sp. | B. meridae n. sp. | B. tjiasmantoi n. sp. |
|---|---|---|---|---|---|---|---|---|---|
| Elytra | Partially covering parapodia | Covering parapodia, dorsum exposed | Covering parapodia, dorsum exposed | Covering parapodia, dorsum exposed | Partially covering parapodia | Fully or partially covering parapodia | Fully or partially covering parapodia | Minute, most of parapodia exposed | Covering parapodia, dorsum exposed |
| Frontal filaments | Minute | Absent | Absent | Absent | Absent | Absent | Absent | Absent | Absent |
| First branchiae | Segment 2 | Segment 3 | Segment 3 | Segment 3 | Segment 2 | Segment 2 | Segment 2 | Segment 2 | Segment 3 |
| Branchiae | Long | Short, dense | Short, dense | Short, dense | Long | Short, dense | Short, dense | Long | Short, dense |
| Dorsal cirri | Short | Short | Short | Short | Short | Long | Long | Short | Short |
| Parapodium | Subbiramous | Biramous | Subbiramous | Subbiramous | Subbiramous | Biramous | Biramous | Subbiramous | Subbiramous |
| Notopodial acicular lobe | Rounded, very short | Triangular, ½ notochaetal length | Digitiform, very short | Straight, reaching tip of notochaetae | Digitiform, ½ notochaetal length | Straight, reaching tip of notochaetae | Straight, ½ notochaetal length | Rounded, very short | Digitiform, longer than notochaetae |
| Notochaetae | Few, very short | Several, long | Few, very short | Few, very short | Few, very short | Several, long | Several, long | Few, very short | Few, very short |
| Supra-acicular neurochaetae | Slightly hooked | Tapered, blunt tip | Slender, slightly hooked | Stout, slightly hooked | Slender, tapered into rounded tip | Slender, tapered tip, subdistal swelling | Slender, tapered tip, subdistal swelling | Stout, straight, subdistal swelling | Slender, rounded tip, subdistal swelling |
| Subacicular neurochaetae | Slender, slightly hooked | Slender, slightly hooked | Slightly hooked | Slender or stout, slightly hooked | Similar to supra-acicular, slightly hooked | Similar to supra-acicular, shorter subdistal area | Similar to supra-acicular, shorter subdistal area | Slender, rounded tip, longer subdistal area | Stout, shorter subdistal area |
| Habitat | Vents | Seeps | Vents & Seeps | Vents | Seeps | Seeps | Seeps | Seeps | Vents |
| Host Bathymodiolus mussel | B. thermophilus & B. antarcticus | Unknown B. heckerae? | B. brevior, B. japonicas, & Gigantidas platifrons | B. marisindicus | B. n. sp. 1 and 2 | B. n. sp. 1–3 | B. n. sp. 1–3 | B. n. sp. 1 and 3 | Bathymodiolus spp. including B. brevior |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lindgren, J.; Hatch, A.S.; Hourdez, S.; Seid, C.A.; Rouse, G.W. Phylogeny and Biogeography of Branchipolynoe (Polynoidae, Phyllodocida, Aciculata, Annelida), with Descriptions of Five New Species from Methane Seeps and Hydrothermal Vents. Diversity 2019, 11, 153. https://doi.org/10.3390/d11090153
Lindgren J, Hatch AS, Hourdez S, Seid CA, Rouse GW. Phylogeny and Biogeography of Branchipolynoe (Polynoidae, Phyllodocida, Aciculata, Annelida), with Descriptions of Five New Species from Methane Seeps and Hydrothermal Vents. Diversity. 2019; 11(9):153. https://doi.org/10.3390/d11090153
Chicago/Turabian StyleLindgren, Johanna, Avery S. Hatch, Stephané Hourdez, Charlotte A. Seid, and Greg W. Rouse. 2019. "Phylogeny and Biogeography of Branchipolynoe (Polynoidae, Phyllodocida, Aciculata, Annelida), with Descriptions of Five New Species from Methane Seeps and Hydrothermal Vents" Diversity 11, no. 9: 153. https://doi.org/10.3390/d11090153
APA StyleLindgren, J., Hatch, A. S., Hourdez, S., Seid, C. A., & Rouse, G. W. (2019). Phylogeny and Biogeography of Branchipolynoe (Polynoidae, Phyllodocida, Aciculata, Annelida), with Descriptions of Five New Species from Methane Seeps and Hydrothermal Vents. Diversity, 11(9), 153. https://doi.org/10.3390/d11090153






