The Spider Anatomy Ontology (SPD)—A Versatile Tool to Link Anatomy with Cross-Disciplinary Data
Abstract
1. Introduction
Anatomical Ontologies and What They are Used for
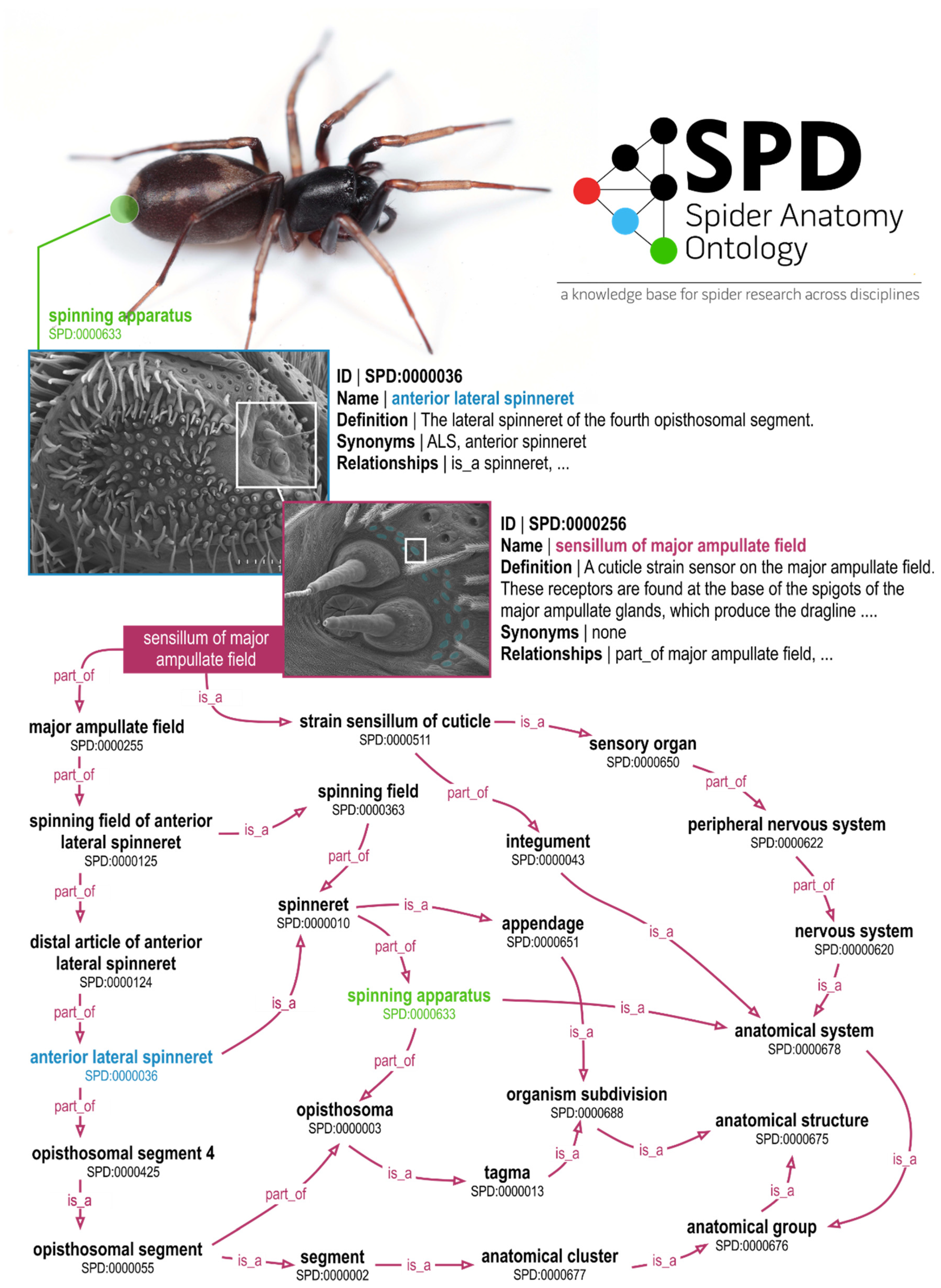
2. The Spider Anatomy Ontology (SPD)
Application of the Spider Anatomy Ontology (SPD)—A Few Use Cases
3. How to Contribute
4. Limitations and Future Developments
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Nyffeler, M.; Birkhofer, K. An estimated 400–800 million tons of prey are annually killed by the global spider community. Sci. Nat. 2017, 104, 30. [Google Scholar] [CrossRef] [PubMed]
- Yoder, M.J.; Miko, I.; Seltmann, K.C.; Bertone, M.A.; Deans, A.R. A gross anatomy ontology for Hymenoptera. PLoS ONE 2010, 5, 12. [Google Scholar] [CrossRef] [PubMed]
- Westring, N. Araneae svecicae. Göteborgs Kungliga Vetenskaps och Vitterhets Samhälles Handlingar 1861, 7, 1–615. [Google Scholar]
- Simon, E. Histoire Naturelle Des Araignées; Roret: Paris, France, 1892; Volume 1, pp. 1–256. [Google Scholar]
- Ubick, D.; Paquin, P.; Cushing, P.E.; Roth, V. Spiders of North America—An Identification Manual, 2nd ed.; American Arachnological Society: Keene, NH, USA, 2017; p. 425. [Google Scholar]
- Bennett, R.G. The spermathecal pores of spiders with special reference to dictynoids and amaurobioids (Araneae, Arneomorphae, Araneoclada). Proc. Entomol. Soc. Ont. 1992, 123, 1–21. [Google Scholar]
- Forster, R.R. The spiders of New Zealand. Part III. Otago Mus. Bull. 1970, 3, 1–184. [Google Scholar]
- Wheeler, W.; Coddington, J.A.; Crowley, L.M.; Dimitrov, D.; Goloboff, P.A.; Griswold, C.E.; Hormiga, G.; Prendini, L.; Ramirez, M.J.; Sierwald, P.; et al. The spider tree of life: Phylogeny of Araneae based on target-gene analyses from an extensive taxon sampling. Cladistics 2017, 33, 574–616. [Google Scholar] [CrossRef]
- Ramirez, M.J. The morphology and phylogeny of dionychan spiders (Araneae: Araneomorphae). Bull. Am. Nat. Hist. 2014, 390, 1–374. [Google Scholar] [CrossRef]
- Huber, B.A. Evolutionary transformation from muscular to hydraulic movements in spider (Arachnida, Araneae) genitalia: A study based on histological serial sections. J. Morphol. 2004, 261, 364–376. [Google Scholar] [CrossRef]
- Labarque, F.; Wolff, J.O.; Michalik, P.; Griswold, C.E.; Ramirez, M.J. The evolution and function of the pretarsus of spiders (Araneae, Arachnida). Zool. J. Linn. Soc. 2017, 181, 308–341. [Google Scholar]
- Coddington, J.A. Ontogeny and homology in the male palpus of orb-weaving spiders and their relatives, with comments on phylogeny (Araneoclada: Araneoidea, Deinopoidea). Smithson. Cont. Zool. 1990, 496, 1–52. [Google Scholar] [CrossRef]
- Quade, F.S.C.; Holtzheimer, J.; Frohn, J.; Topperwien, M.; Salditt, T.; Prpic, N.M. Formation and development of the male copulatory organ in the spider Parasteatoda tepidariorum involves a metamorphosis-like process. Sci. Rep. 2019, 9, 1. [Google Scholar] [CrossRef] [PubMed]
- Griswold, C.E.; Ramírez, M.J.; Coddington, J.A.; Platnick, N.I. Atlas of phylogenetic data for entelegyne spiders (Araneae: Araneomorphae: Entelegynae) with comments on their phylogeny. Proc. Calif. Acad. Sci. 2005, 56, 1–325. [Google Scholar]
- Vogt, L.; Bartolomaeus, T.; Giribet, G. The linguistic problem of morphology: Structure versus homology and the standardization of morphological data. Cladistics 2010, 26, 301–325. [Google Scholar] [CrossRef]
- Vogt, L.; Nickel, M.; Jenner, R.A.; Deans, A.R. The need for data standards in zoomorphology. J. Morphol. 2013, 274, 793–808. [Google Scholar] [CrossRef] [PubMed]
- Jocqué, R.; Dippenaar-Schoeman, A.S. Spider Families of the World; Musée Royal de l’Afrique Central: Tervuren, Belgium, 2006; p. 336. [Google Scholar]
- Richter, S.; Loesel, R.; Purschke, G.; Schmidt-Rhaesa, A.; Scholtz, G.; Stach, T.; Vogt, L.; Wanninger, A.; Brenneis, G.; Doring, C.; et al. Invertebrate neurophylogeny: Suggested terms and definitions for a neuroanatomical glossary. Front. Zool. 2010, 7, 1. [Google Scholar] [CrossRef] [PubMed]
- Gorb, S.N.; Barth, F.G. A new mechanosensory organ on the anterior spinnerets of the spider cupiennus salei (araneae, ctenidae). Zoomorphology 1996, 116, 7–14. [Google Scholar] [CrossRef]
- Hilbrant, M.; Damen, W.G.M. The embryonic origin of the ampullate silk glands of the spider Cupiennius salei. Arthropod Str. Dev. 2015, 44, 280–288. [Google Scholar] [CrossRef]
- Mabee, P.M.; Ashburner, M.; Cronk, Q.; Gkoutos, G.V.; Haendel, M.; Segerdell, E.; Mungall, C.; Westerfield, M. Phenotype ontologies: The bridge between genomics and evolution. Trends Eco. Evol. 2007, 22, 345–350. [Google Scholar] [CrossRef]
- Edmunds, R.C.; Su, B.F.; Balhoff, J.P.; Eames, B.F.; Dahdul, W.M.; Lapp, H.; Lundberg, J.G.; Vision, T.J.; Dunham, R.A.; Mabee, P.M.; et al. Phenoscape: Identifying candidate genes for evolutionary phenotypes. Mol. Biol. Evol. 2016, 33, 13–24. [Google Scholar] [CrossRef]
- Walls, R.L.; Athreya, B.; Cooper, L.; Elser, J.; Gandolfo, M.A.; Jaiswal, P.; Mungall, C.J.; Preece, J.; Rensing, S.; Smith, B.; et al. Ontologies as integrative tools for plant science. Am. J. Bot. 2012, 99, 1263–1275. [Google Scholar] [CrossRef]
- Deans, A.R.; Lewis, S.E.; Huala, E.; Anzaldo, S.S.; Ashburner, M.; Balhoff, J.P.; Blackburn, D.C.; Blake, J.A.; Burleigh, J.G.; Chanet, B.; et al. Finding our way through phenotypes. PLoS Boil. 2015, 13, 1. [Google Scholar]
- Dececchi, T.A.; Balhoff, J.P.; Lapp, H.; Mabee, P.M. Toward synthesizing our knowledge of morphology: Using ontologies and machine reasoning to extract presence/absence evolutionary phenotypes across studies. Syst. Biol. 2015, 64, 936–952. [Google Scholar] [CrossRef] [PubMed]
- Smith, B.; Ashburner, M.; Rosse, C.; Bard, J.; Bug, W.; Ceusters, W.; Goldberg, L.J.; Eilbeck, K.; Ireland, A.; Mungall, C.J.; et al. The OBO Foundry: Coordinated evolution of ontologies to support biomedical data integration. Nat. Biol. 2007, 25, 1251–1255. [Google Scholar]
- Day-Richter, J.; Harris, M.A.; Haendel, M.A.; Lewis, S. Obo-edit—An ontology editor for biologists. Bioinformatics 2007, 23, 2198–2200. [Google Scholar] [CrossRef]
- Ramirez, M.J.; Coddington, J.A.; Maddison, W.P.; Midford, P.E.; Prendini, L.; Miller, J.; Griswold, C.E.; Hormiga, G.; Sierwald, P.; Scharff, N.; et al. Linking of digital images to phylogenetic data matrices using a morphological ontology. Syst. Biol. 2007, 56, 283–294. [Google Scholar] [CrossRef]
- Haendel, M.A.; Neuhaus, F.; Osumi-Sutherland, D.; Mabee, P.M.; Mejino, J.L.; Mungall, C.J.; Smith, B. Caro—The common anatomy reference ontology. In Anatomy Ontologies for Bioinformatics; Burger, A., Davidson, D., Baldock, R., Eds.; Springer: London, UK, 2008; pp. 327–349. [Google Scholar]
- Dahdul, W.M.; Lundberg, J.G.; Midford, P.E.; Balhoff, J.P.; Lapp, H.; Vision, T.J.; Haendel, M.A.; Westerfield, M.; Mabee, P.M. The teleost anatomy ontology: Anatomical representation for the genomics age. Syst. Biol. 2010, 59, 369–383. [Google Scholar] [CrossRef]
- Wirkner, C.S.; Göpel, T.; Runge, J.; Keiler, J.; Klussmann-Fricke, B.J.; Huckstorf, K.; Scholz, S.; Miko, I.; Yoder, M.J.; Richter, S. The first organ-based ontology for arthropods (Ontology of Arthropod Circulatory Systems—OArCS) and its integration into a novel formalization scheme for morphological descriptions. Syst. Biol. 2017, 66, 754–768. [Google Scholar] [CrossRef]
- Costa, M.; Reeve, S.; Grumbling, G.; Osumi-Sutherland, D. The Drosophila anatomy ontology. J. Biomed. Semant. 2013, 4, 1. [Google Scholar] [CrossRef]
- Rosse, C.; Mejino, J.L.V. A reference ontology for biomedical informatics: The foundational model of anatomy. J. Biomed. Inf. 2003, 36, 478–500. [Google Scholar] [CrossRef]
- Smith, B.; Ceusters, W.; Klagges, B.; Kohler, J.; Kumar, A.; Lomax, J.; Mungall, C.; Neuhaus, F.; Rector, A.L.; Rosse, C. Relations in biomedical ontologies. Genome Biol. 2005, 6, 5. [Google Scholar]
- Michalik, P.; Ramirez, M.J. Evolutionary morphology of the male reproductive system, spermatozoa and seminal fluid of spiders (Araneae, Arachnida)—Current knowledge and future directions. Arthropod. Str. Dev. 2014, 43, 291–322. [Google Scholar] [CrossRef] [PubMed]
- Comstock, J.H. The palpi of male spiders. Ann. Entomol. Soc. Am. 1910, 3, 161–185. [Google Scholar] [CrossRef]
- Sierwald, P. Morphology and ontogeny of female copulatory organs in American Pisauridae, with special reference to homologous features (Arachnida: Araneae). Smithson. Cont. Zool. 1989, 484, 1–24. [Google Scholar] [CrossRef]
- Eberhard, W.G.; Pereira, F. Ultrastructure of cribellate silk of nine species in eight families and possible taxonomic implications (Araneae: Amaurobiidae, Deinopidae, Desidae, Dictynidae, Filistatidae, Hypochilidae, Stiphidiidae, Tengellidae). J. Arachnol. 1993, 21, 161–174. [Google Scholar]
- Foelix, R.F. Biology of Spiders, 3rd ed.; Oxford University Press: Oxford, UK, 2011; p. 419. [Google Scholar]
- Barth, F.G. A Spider’s World-Senses and Behavior; Spinger: Berlin/Heidelberg, Germany; New York, NY, USA, 2002; p. 394. [Google Scholar]
- Millot, J. Ordre. aranéides (Araneae). In Traité De Zoologie-Anatomie, Systématique, Biologie; Grassé, P.-P., Ed.; Masson: Paris, France, 1949; Volume Tome VI, pp. 589–743. [Google Scholar]
- Maddison, W.P.; Ramirez, M.J. Silk: Simple Image Linking Package for Mesquite. 2016. Available online: http://mesquiteproject.org/SILK/ (accessed on 21 October 2019).
- Maddison, W.P.; Maddison, D.R. Mesquite. version 3.51. 2018. Available online: http://www.mesquiteproject.org (accessed on 21 October 2019).
- Ramirez, M.J.; Michalik, P. Calculating structural complexity in phylogenies using ancestral ontologies. Cladistics 2014, 30, 635–649. [Google Scholar] [CrossRef]
- Sereno, P.C. Logical basis for morphological characters in phylogenetics. Cladistics 2007, 23, 565–587. [Google Scholar] [CrossRef]
- Cui, H.; Xu, D.F.; Chong, S.S.; Ramirez, M.; Rodenhausen, T.; Macklin, J.A.; Ludascher, B.; Morris, R.A.; Soto, E.M.; Koch, N.M. Introducing Explorer of Taxon Concepts with a case study on spider measurement matrix building. BMC Bioinfor. 2016, 17, 1. [Google Scholar] [CrossRef]
- Bertone, M.A.; Miko, I.; Yoder, M.J.; Seltmann, K.C.; Balhoff, J.P.; Deans, A.R. Matching arthropod anatomy ontologies to the Hymenoptera Anatomy Ontology: Results from a manual alignment. Database 2013, 2013, bas057. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramírez, M.J.; Michalik, P. The Spider Anatomy Ontology (SPD)—A Versatile Tool to Link Anatomy with Cross-Disciplinary Data. Diversity 2019, 11, 202. https://doi.org/10.3390/d11100202
Ramírez MJ, Michalik P. The Spider Anatomy Ontology (SPD)—A Versatile Tool to Link Anatomy with Cross-Disciplinary Data. Diversity. 2019; 11(10):202. https://doi.org/10.3390/d11100202
Chicago/Turabian StyleRamírez, Martín J., and Peter Michalik. 2019. "The Spider Anatomy Ontology (SPD)—A Versatile Tool to Link Anatomy with Cross-Disciplinary Data" Diversity 11, no. 10: 202. https://doi.org/10.3390/d11100202
APA StyleRamírez, M. J., & Michalik, P. (2019). The Spider Anatomy Ontology (SPD)—A Versatile Tool to Link Anatomy with Cross-Disciplinary Data. Diversity, 11(10), 202. https://doi.org/10.3390/d11100202




