Soil Organic Carbon Shapes AMF Communities in Soils and Roots of Cynodon dactylon under Anti-Seasonal Drying-Wetting Cycles
Abstract
1. Introduction
2. Materials and Methods
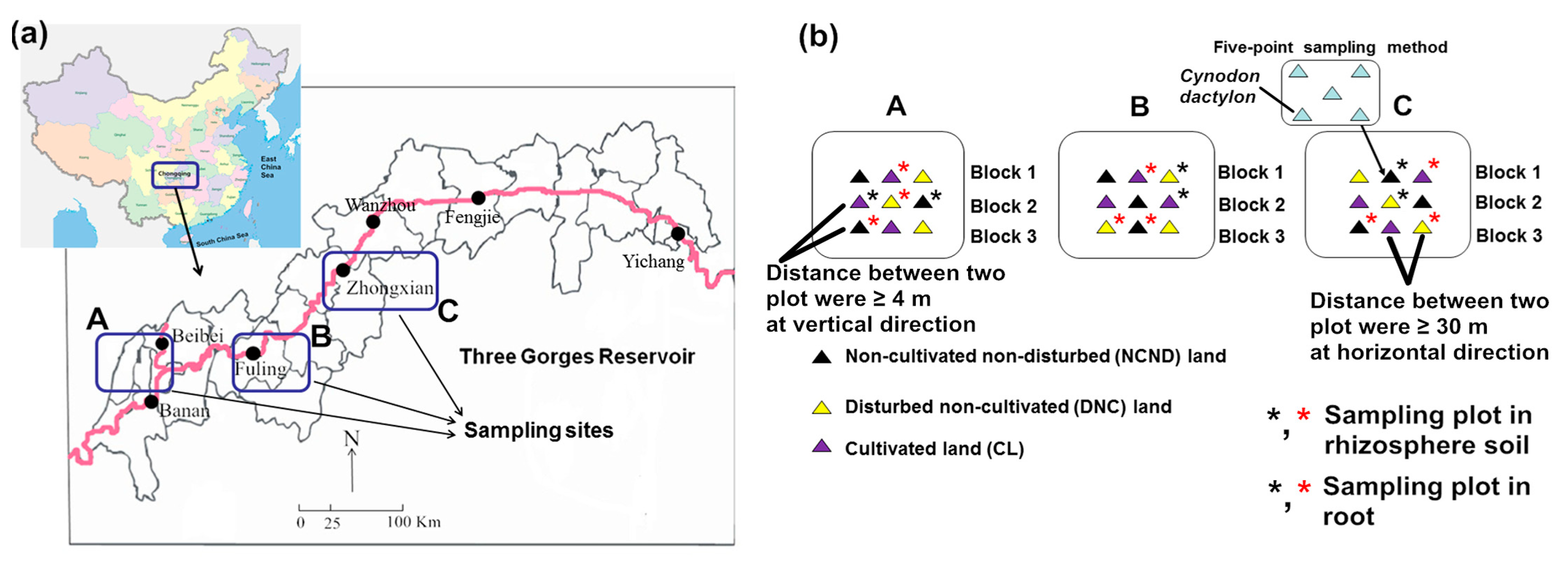
2.1. Site Description and Sampling
2.2. Soil Physical and Chemical Properties
2.3. Determination of AM Colonization
2.4. Extraction of Soil and Root DNA and Amplicon Generation
2.5. Bioinformatics Analysis
2.6. Data and Statistical Analyses
2.7. Accession Number
3. Results
3.1. Differences in Soil Properties between Three Contrasting Land-Use Types
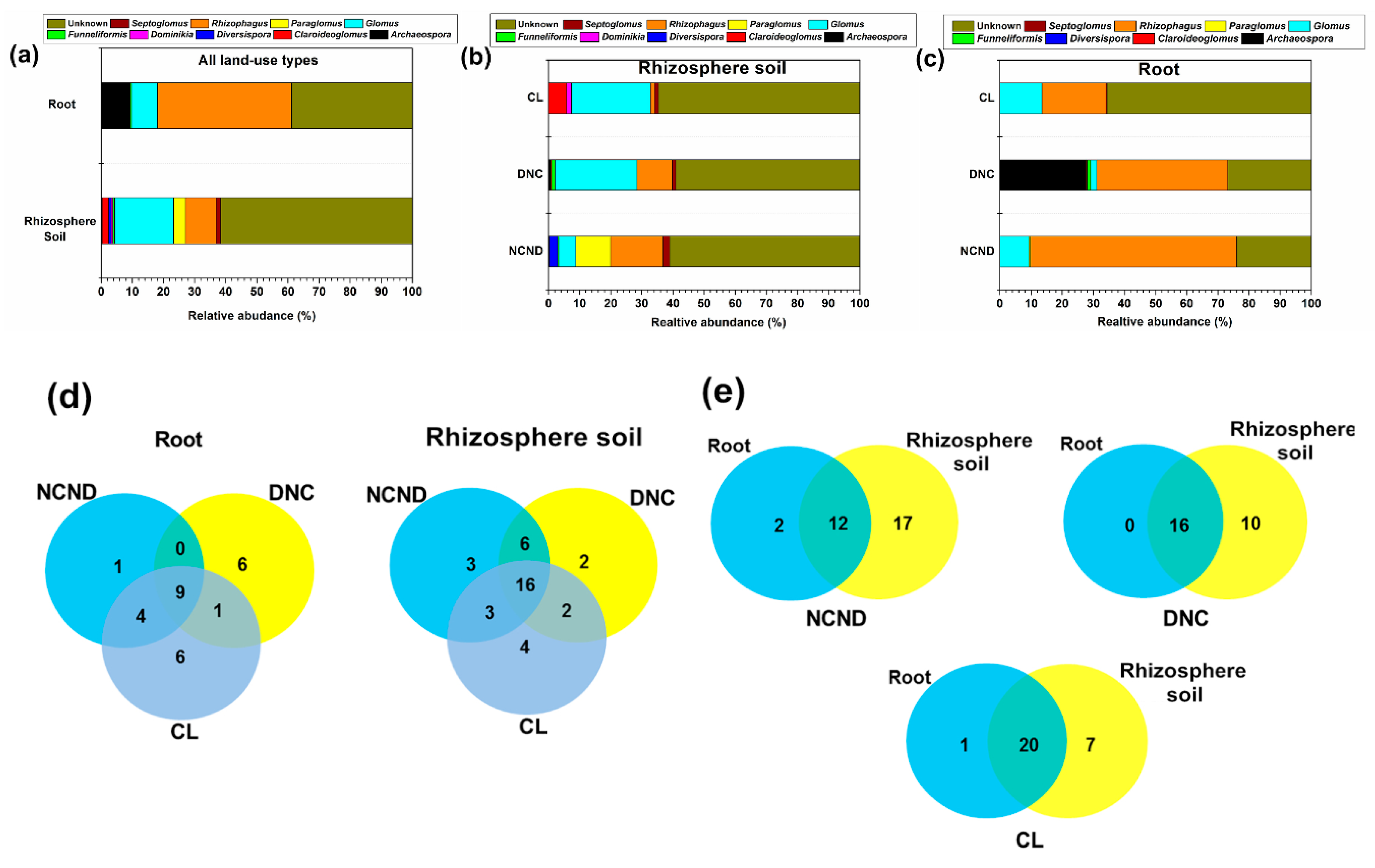
3.2. Differences between Soil and Root DNA Sequences
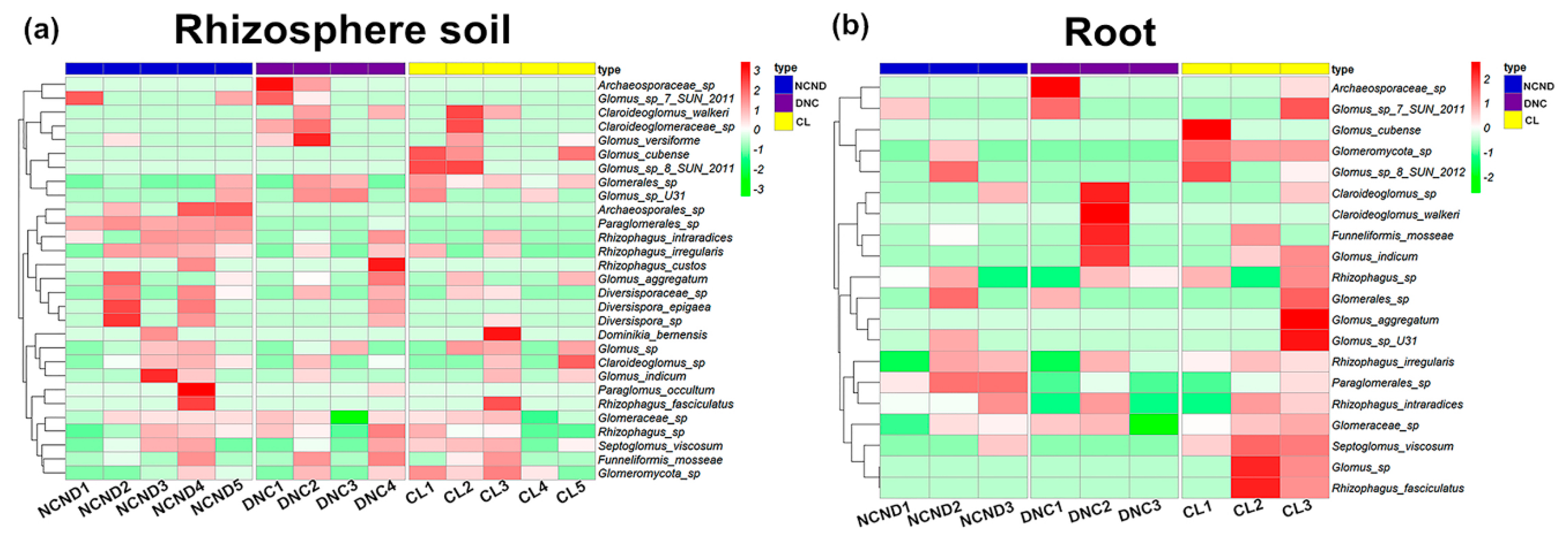
3.3. AMF Community Structure in Rhizosphere Soils under Three Contrasting Land-Use Types
3.4. AMF Community Structure in Roots under Three Contrasting Land-Use Types
3.5. Defferences of AMF Community Structure between Rhizosphere Soils and Roots under Three Contrasting Land-Use Types
3.6. AMF α-Diversity, Colonization and Community under Three Contrasting Land-Use Types Lands
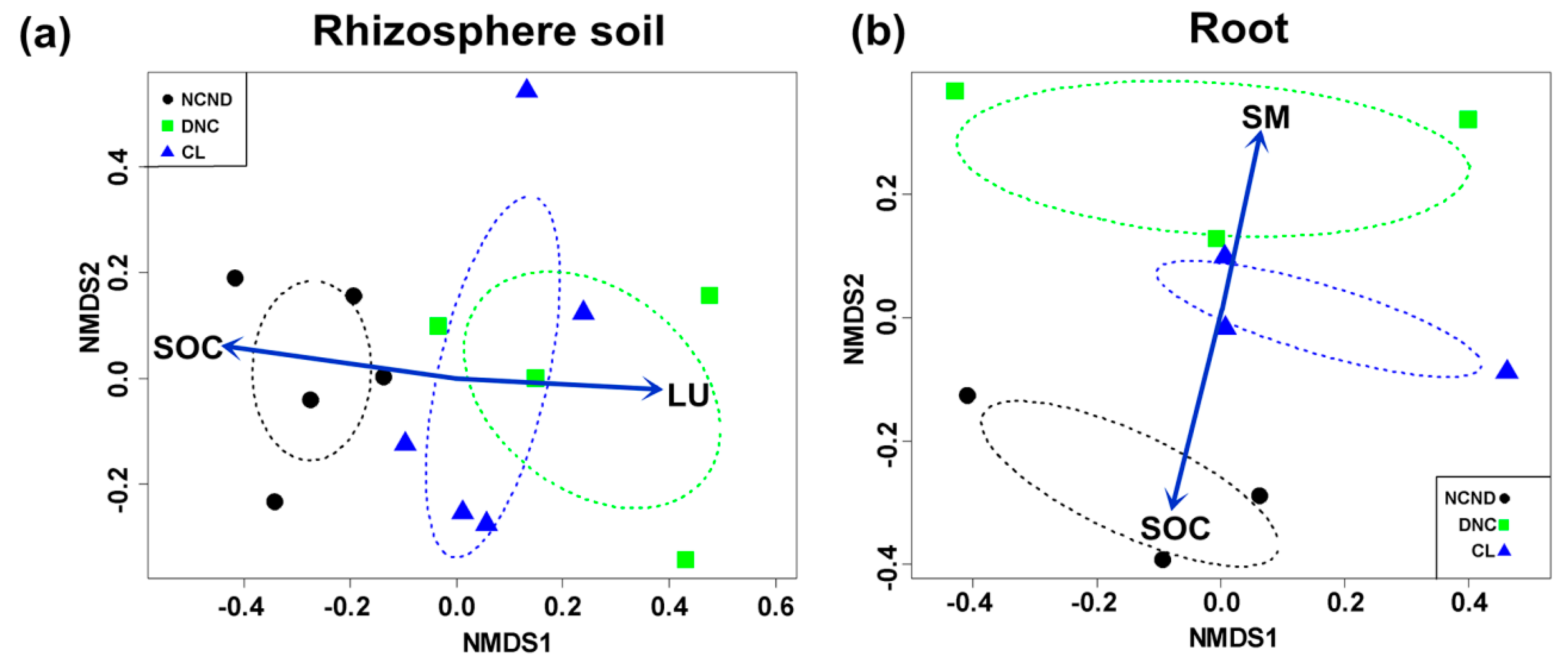
3.7. Effects of Abiotic Factors on the AMF Community between Rhizosphere Soils and Roots
4. Discussions
4.1. Differences in AMF Communities in Rhizosphere Soils and Roots
4.2. Differences in AMF Colonization, Diversity and Community in the Rhizosphere Soils and Roots under Three Contrasting Land-Use Types
4.3. Correlations of Differences between AMF Community and Relevant Abiotic Factors
4.4. Primer Systems for Evaluating AMF Community
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wang, Q.; Yuan, X.; Willison, J.H.; Zhang, Y.; Liu, H. Diversity and above-ground biomass patterns of vascular flora induced by flooding in the drawdown area of China’s Three Gorges Reservoir. PLoS ONE 2014, 9, e100889. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yan, T.; Yang, L.; Campbell, C.D. Microbial biomass and metabolic quotient of soils under different land use in the Three Gorges Reservoir area. Geoderma 2003, 115, 129–138. [Google Scholar] [CrossRef]
- Osborne, L.L.; Kovacic, D.A. Riparian vegetated buffer strips in water-quality restoration and stream management. Freshw. Biol. 1993, 29, 243–258. [Google Scholar] [CrossRef]
- Richardson, D.; Holmes, P.; Esler, K.; Galatowitsch, S.; Stromberg, J.; Kirkman, S.; Pyšek, P.; Hobbs, R. Riparian vegetation: Degradation, alien plant invasions, and restoration prospects. Divers. Distrib. 2007, 13, 126–139. [Google Scholar] [CrossRef]
- Yang, Y.; Li, C. Photosynthesis and growth adaptation of Pterocarya stenoptera and Pinus elliottii seedlings to submergence and drought. Photosynthetica 2016, 54, 120–129. [Google Scholar] [CrossRef]
- Zhang, Q.; Lou, Z. The environmental changes and mitigation actions in the Three Gorges Reservoir region, China. Environ. Sci. Policy 2011, 14, 1132–1138. [Google Scholar] [CrossRef]
- Veach, A.M.; Dodds, W.K.; Jumpponen, A. Woody plant encroachment, and its removal, impact bacterial and fungal communities across stream and terrestrial habitats in a tallgrass prairie ecosystem. FEMS Microbiol. Ecol. 2015, 91, 1–11. [Google Scholar] [CrossRef]
- Tenten, N.; Zeng, B.; Kazda, M. Soil stabilizing capability of three plant species growing on the Three Gorges Reservoir riverside. J. Earth. Sci. 2010, 21, 888–896. [Google Scholar] [CrossRef]
- Willison, J.H.; Li, R.; Yuan, X. Conservation and ecofriendly utilization of wetlands associated with the Three Gorges Reservoir. Environ. Sci. Pollut. Res. 2013, 20, 6907–6916. [Google Scholar] [CrossRef]
- Wardle, D.A.; Bardgett, R.D.; Klironomos, J.N.; Setälä, H.; van der Putten, W.H.; Wall, D.H. Ecological linkages between aboveground and belowground biota. Science 2004, 304, 1629–1633. [Google Scholar] [CrossRef]
- Dam, N.M.V.; Heil, M. Multitrophic interactions below and above ground: En route to the next level. J. Ecol. 2011, 99, 77–88. [Google Scholar]
- Smith, S.E.; Read, D.J. Mycorrhizal Symbiosis; Academic Press: Cambridge, UK, 2008; pp. 1–187. [Google Scholar]
- Bainard, L.D.; Bainard, J.D.; Hamel, C.; Gan, Y. Spatial and temporal structuring of arbuscular mycorrhizal communities is differentially influenced by abiotic factors and host crop in a semi-arid prairie agroecosystem. FEMS Microbiol. Ecol. 2014, 88, 333–344. [Google Scholar] [CrossRef] [PubMed]
- Rillig, M.C.; Mummey, D.L. Mycorrhizas and soil structure. New Phytol. 2006, 171, 41–53. [Google Scholar] [CrossRef] [PubMed]
- Wilson, G.W.; Rice, C.W.; Rillig, M.C.; Springer, A.; Hartnett, D.C. Soil aggregation and carbon sequestration are tightly correlated with the abundance of arbuscular mycorrhizal fungi: Results from long-term field experiments. Ecol. Lett. 2009, 12, 452–461. [Google Scholar] [CrossRef] [PubMed]
- van der Heijden, M.G.A.; Horton, T.R. Socialism in soil? The importance of mycorrhizal fungal networks for facilitation in natural ecosystems. J. Ecol. 2009, 97, 1139–1150. [Google Scholar] [CrossRef]
- Li, L.F.; Li, T.; Zhao, Z.W. Differences of arbuscular mycorrhizal fungal diversity and community between a cultivated land, an old field, and a never-cultivated field in a hot and arid ecosystem of southwest China. Mycorrhiza 2007, 17, 655–665. [Google Scholar] [CrossRef] [PubMed]
- Boddington, C.L.; Dodd, J.C. The effect of agricultural practices on the development of indigenous arbuscular mycorrhizal fungi. I. Field studies in an Indonesian ultisol. Plant. Soil 2000, 218, 137–144. [Google Scholar] [CrossRef]
- Islam, K.R.; Weil, R.R. Soil quality indicator properties in mid-Atlantic soils as influenced by conservation management. J. Soil Water Conserv. 2000, 55, 69–78. [Google Scholar]
- Li, B.; Xiao, H.Y.; Yuan, X.Z.; Willison, J.H.M.; Liu, H.; Chen, Z.L.; Zhang, Y.W.; Deng, W.; Yue, J.S. Analysis of ecological and commercial benefits of a dike-pond project in the drawdown zone of the Three Gorges Reservoir. Ecol. Eng. 2013, 61, 1–11. [Google Scholar] [CrossRef]
- Helgason, T.; Daniell, T.J.; Husband, R.; Fitter, A.H.; Young, J.P.W. Ploughing up the wood-wide web? Nature 1998, 394, 431. [Google Scholar] [CrossRef]
- Muthukumar, T.; Udaiyan, K. Arbuscular mycorrhizas of plants growing in the Western Ghats region, Southern India. Mycorrhiza 2000, 9, 297–313. [Google Scholar] [CrossRef]
- Zhao, X.; Yuan, S.; Song, H.; Su, X.; Mao, H.; Shen, W.; Qu, X.L.; Dong, J.Y. Arbuscular mycorrhizal and dark septate fungal associations in riparian plants of the Three Gorges Reservoir Region, Southwest China. Aquat. Bot. 2016, 133, 28–37. [Google Scholar] [CrossRef]
- Luo, X.; Su, X.; Cui, J.; Lou, Y.; Li, R.; Luo, X.M.; Zeng, Y.Q.; Xu, Y.B.; Dong, J.Y. Biodiversity of arbuscular mycorrhizal fungi in the drawdown zone of the Three Gorges Reservoir under different fertilization histories. Ecol. Res. 2016, 31, 407–416. [Google Scholar] [CrossRef]
- Nielsen, K.B.; Kjøller, R.; Olsson, P.A.; Schweiger, P.F.; Andersen, F.Ø.; Rosendahl, S. Colonisation and molecular diversity of arbuscular mycorrhizal fungi in the aquatic plants Littorella uniflora and Lobelia dortmanna in southern Sweden. Mycol. Res. 2004, 108, 616–625. [Google Scholar] [CrossRef] [PubMed]
- Wilde, P.; Manal, A.; Stodden, M.; Sieverding, E.; Hildebrandt, U. Biodiversity of arbuscular mycorrhizal fungi in roots and soils of two salt marshes. Environ. Microbiol. 2009, 11, 1548–1561. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.T.; Huang, Y.L.; Qiu, Q.; Xin, G.R.; Yang, Z.Y.; Shi, S.H. Flooding greatly affects the diversity of arbuscular mycorrhizal fungi (AMF) communities in the roots of wetland plants. PLoS ONE 2011, 6, e24512. [Google Scholar]
- Twanabasu, B.R.; Smith, C.M.; Stevens, K.J.; Venables, B.J.; Sears, W.C. Triclosan inhibits arbuscular mycorrhizal colonization in three wetland plants. Sci. Total Environ. 2013, 447, 450–457. [Google Scholar] [CrossRef]
- Wang, Y.; Li, Y.; Bao, X.; Björn, L.O.; Li, S.; Olsson, P.A. Response differences of arbuscular mycorrhizal fungi communities in the roots of an aquatic and a semiaquatic species to various flooding regimes. Plant Soil 2016, 403, 361–373. [Google Scholar] [CrossRef]
- Ye, C.; Zhang, K.; Deng, Q.; Zhang, Q. Plant communities in relation to flooding and soil characteristics in the water level fluctuation zone of the Three Gorges Reservoir, China. Environ. Sci. Pollut. Res. 2013, 20, 1794–1802. [Google Scholar] [CrossRef]
- Bao, S.D. Soil Agro-Chemistry Analyses; Chinese Agriculture Press: Beijing, China, 2008; pp. 1–100. (In Chinese) [Google Scholar]
- Rhoades, J.D. Salinity: Electrical conductivity and total dissolved solids. In Methods of Soil Analysis, Part 3–Chemical Methods; Sparks, D.L., Ed.; American Society of Agronomy: Madison, WI, USA, 1996; pp. 417–436. [Google Scholar]
- de Souza, F.A.; Dalpé, Y.; Declerck, S.; de la Providencia, I.E.; Séjalon-Delmas, N. Life history strategies in Gigasporaceae: Insight from monoxenic culture. In In Vitro Culture of Mycorrhizas; Declerck, S., Strullu, D.G., Fortin, J.A., Eds.; Springer: Heidelberg, Germany, 2005; pp. 73–91. [Google Scholar]
- Mcgonigle, T.P.; Miller, M.H.; Evans, D.G.; Fairchild, G.L.; Swan, J.A. A new method which gives an objective measure of colonization of roots by vesicular-arbuscular mycorrhizal fungi. New Phytol. 1990, 115, 495–501. [Google Scholar] [CrossRef]
- Krüger, M.; Stockinger, H.; Krüger, C.; Schüssler, A. DNA-based species level detection of Glomeromycota: One PCR primer set for all arbuscular mycorrhizal fungi. New Phytol. 2009, 183, 212–223. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols, a Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Ihrmark, K.; Bödeker, I.T.; Cruz-Martinez, K.; Friberg, H.; Kubartova, A.; Schenck, J.; Strid, Y.; Stenlid, J.; Brandström-Durling, M.; Clemmensen, K.E.; et al. New primers to amplify the fungal ITS2 region–evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol. Ecol. 2012, 82, 666–677. [Google Scholar] [CrossRef] [PubMed]
- Kohout, P.; Sudová, R.; Janoušková, M.; Čtvrtlíková, M.; Hejda, M.; Pánková, H.; Slavíková, R.; Stajerová, K.; Vosátka, M.; Sýkorová, Z. Comparison of commonly used primer sets for evaluating arbuscular mycorrhizal fungal communities: Is there a universal solution? Soil Biol. Biochem. 2014, 68, 482–493. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.F.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef]
- Bengtsson-Palme, J.; Ryberg, M.; Hartmann, M.; Branco, S.; Wang, Z.; Godhe, A.; Wit, P.D.; Sánchez-García, M.; Ebersberger, I.; de Sousa, F.; et al. Improved software detection and extraction of ITS1 and ITS2 from ribosomal ITS sequences of fungi and other eukaryotes for analysis of environmental sequencing data. Methods Ecol. Evol. 2013, 4, 914–919. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community–supported software for describing and comparing microbial communities. Appl. Environ. Microb. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Kõljalg, U.; Nilsson, R.H.; Abarenkov, K.; Tedersoo, L.; Taylor, A.F.; Bahram, M.; Bates, S.T.; Bruns, T.D.; Bengtsson-Palme, J.; Callaghan, T.M.; et al. Towards a unified paradigm for sequence-based identification of fungi. Mol. Ecol. 2013, 22, 5271–5277. [Google Scholar]
- Edgar, R.C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996–998. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Abarenkov, K.; Nilsson, R.H.; Larsson, K.H.; Alexander, I.J.; Eberhardt, U.; Erland, S.; Høiland, K.; Kjøller, R.; Larsson, E.; Pennanen, T.; et al. The UNITE database for molecular identification of fungi--recent updates and future perspectives. New Phytol. 2010, 186, 281–285. [Google Scholar] [CrossRef]
- Redecker, D.; Schüssler, A.; Stockinger, H.; Stürmer, S.L.; Morton, J.B.; Walker, C. An evidence-based consensus for the classification of arbuscular mycorrhizal fungi (Glomeromycota). Mycorrhiza 2013, 23, 515–531. [Google Scholar] [CrossRef] [PubMed]
- Anderson, M.J. Distance-based tests for homogeneity of multivariate dispersions. Biometrics 2006, 62, 245–253. [Google Scholar] [CrossRef] [PubMed]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; Stevens, M.H.H.; Wagner, H. Vegan: Community Ecology Package. R Package Version 3.2.1. 2012. Available online: http://vegan.r-forge. r-project.org/ (accessed on 11 July 2016).
- Oksanen, J. Multivariate Analysis of Ecological Communities in R: Vegan Tutorial. 2011. Available online: http://cc.oulu.fi/~jarioksa /opetus/metodi/vegantutor.pdf (accessed on 22 July 2016).
- Hempel, S.; Renker, C.; Buscot, F. Differences in the species composition of arbuscular mycorrhizal fungi in spore, root and soil communities in a grassland ecosystem. Environ. Microbiol. 2007, 9, 1930–1938. [Google Scholar] [CrossRef] [PubMed]
- Martínez-García, L.B.; Armas, C.; Miranda, J.D.D.; Padilla, F.M.; Pugnaire, F.I. Shrubs influence arbuscular mycorrhizal fungi communities in a semi-arid environment. Soil Biol. Biochem. 2011, 43, 682–689. [Google Scholar] [CrossRef]
- Liu, Y.J.; He, L.; An, L.Z.; Helgason, T.; Feng, H.Y. Arbuscular mycorrhizal dynamics in a chronosequence of Caragana korshinskii plantations. FEMS Microbiol. Ecol. 2009, 67, 81–82. [Google Scholar] [CrossRef]
- Hontoria, C.; García-González, I.; Quemada, M.; Roldán, A.; Alguacil, M.M. The cover crop determines the AMF community composition in soil and in roots of maize after a ten-year continuous crop rotation. Sci. Total Environ. 2019, 660, 913–922. [Google Scholar] [CrossRef]
- Tamayo, E.; Gómez-Gallego, T.; Azcón-Aguilar, C.; Ferrol, N. Genome-wide analysis of copper, iron and zinc transporters in the arbuscular mycorrhizal fungus Rhizophagus irregularis. Front. Plant. Sci. 2014, 5, 547. [Google Scholar] [CrossRef]
- Garcia, K.; Doidy, J.; Zimmermann, S.D.; Wipf, D.; Courty, P.-E. Take a trip through the plant and fungal transportome of mycorrhiza. Trends Plant. Sci. 2016, 21, 937–950. [Google Scholar] [CrossRef]
- Chen, Y.L.; Zhang, X.; Ye, J.S.; Han, H.Y.; Wan, S.Q.; Chen, B.D. Six-year fertilization modifies the biodiversity of arbuscular mycorrhizal fungi in a temperate steppe in Inner Mongolia. Soil Biol. Biochem. 2014, 69, 371–381. [Google Scholar] [CrossRef]
- Bailey-Serres, J.; Voesenek, L.A.C.J. Flooding stress: Acclimations and genetic diversity. Annu. Rev. Plant. Biol. 2008, 59, 313–339. [Google Scholar] [CrossRef]
- Šraj-Kržič, N.; Pongrac, P.; Klemenc, M.; Kladnik, A.; Regvar, M.; Gaberščik, A. Mycorrhizal colonisation in plants from intermittent aquatic habitats. Aquat. Bot. 2006, 85, 331–336. [Google Scholar] [CrossRef]
- Chen, F.Q.; Xie, Z.Q. Ecophysiological response of two herbaceous species to flooding. IEEE 2011, 4260–4263. [Google Scholar]
- Alguacil, M.M.; Torres, M.P.; Torrecillas, E.; Diaz, G.; Roldán, A. Plant type differently promote the arbuscular mycorrhizal fungi biodiversity in the rhizosphere after revegetation of a degraded, semiarid land. Soil Biol. Biochem. 2011, 43, 167–173. [Google Scholar] [CrossRef]
- Torrecillas, E.; Alguacil, M.M.; Roldán, A. Host preferences of arbuscular mycorrhizal fungi colonizing annual herbaceous plant species in semiarid Mediterranean prairies. Appl. Environ. Microb. 2012, 78, 6180–6186. [Google Scholar] [CrossRef] [PubMed]
- Alguacil, M.M.; Lumini, E.; Roldán, A.; Salinas-García, J.R.; Bonfante, P.; Bianciotto, V. The impact of tillage practices on arbuscular mycorrhizal fungal diversity in subtropical crops. Ecol. Appl. 2008, 18, 527–536. [Google Scholar] [CrossRef] [PubMed]
- Oehl, F.; Laczko, E.; Bogenrieder, A.; Stahr, K.; Bösch, R.; van der Heijden, M. Soil type and land use intensity determine the composition of arbuscular mycorrhizal fungal communities. Soil Biol. Biochem. 2010, 42, 724–738. [Google Scholar] [CrossRef]
- Stürmer, S.L.; Siqueira, J.O. Species richness and spore abundance of arbuscular mycorrhizal fungi across distinct land uses in Western Brazilian Amazon. Mycorrhiza 2011, 21, 255–267. [Google Scholar] [CrossRef]
- González-Cortés, J.C.; Vega-Fraga, M.; Varela-Fregoso, L.; Martínez-Trujillo, M.; Carreón-Abud, Y.; Gavito, M.E. Arbuscular mycorrhizal fungal (AMF) communities and land use change: The conversion of temperate forests to avocado plantations and maize fields in central Mexico. Fungal Ecol. 2012, 5, 16–23. [Google Scholar] [CrossRef]
- Jefwa, J.M.; Okoth, S.; Wachira, P.; Karanja, N.; Kahindi, J.; Njuguini, S.; Ichami, S.; Mung’atu, J.; Okoth, P.; Huising, J. Impact of land use types and farming practices on occurrence of arbuscular mycorrhizal fungi (AMF) Taita-Taveta district in Kenya. Agr. Ecosyst. Environ. 2012, 157, 32–39. [Google Scholar] [CrossRef]
- Moora, M.; Davison, J.; Öpik, M.; Metsis, M.; Saks, Ü.; Jairus, T.; Vasar, M.; Zobel, M. Anthropogenic land use shapes the com position and phylogenetic structure of soil arbuscular mycorrhizal fungal communities. FEMS Microbiol. Ecol. 2014, 903, 609–621. [Google Scholar] [CrossRef]
- Ciccolini, V.; Bonari, E.; Pellegrino, E. Land-use intensity and soil properties shape the composition of fungal communities in Mediterranean peaty soils drained for agricultural purposes. Biol. Fertil. Soils 2015, 51, 719–731. [Google Scholar] [CrossRef]
- Jansa, J.; Erb, A.; Oberholzer, H.R.; Smilauer, P.; Egli, S. Soil and geography are more important determinants of indigenous arbuscular mycorrhizal communities than management practices in Swiss agricultural soils. Mol. Ecol. 2014, 23, 2118–2135. [Google Scholar] [CrossRef] [PubMed]
- Pellegrino, E.; Bosco, S.; Ciccolini, V.; Pistocchi, C.; Sabbatini, T.; Silvestri, N.; Bonari, E. Agricultural abandonment in Mediterranean reclaimed peaty soils: Long-term effects on soil chemical properties, arbuscular mycorrhizas and CO2 flux. Agric. Ecosyst. Environ. 2014, 199, 164–175. [Google Scholar] [CrossRef]
- Titus, J.H.; Lepš, J. The response of arbuscular mycorrhizae to fertilization, mowing, and removal of dominant species in a diverse oligotrophic wet meadow. Am. J. Bot. 2000, 87, 392–401. [Google Scholar] [CrossRef] [PubMed]
- Barto, E.K.; Rillig, M.C. Does herbivory really suppress mycorrhiza? A meta-analysis. J. Ecol. 2010, 98, 745–753. [Google Scholar] [CrossRef]
- Jansa, J.; Mozafar, A.; Anken, T.; Ruh, R.; Sanders, I.; Frossard, E. Diversity and structure of AMF communities as affected by tillage in a temperate soil. Mycorrhiza 2002, 12, 225–234. [Google Scholar]
- Öpik, M.; Moora, M.; Liira, J.; Zobel, M. Composition of root–colonizing arbuscular mycorrhizal fungal communities in different ecosystems around the globe. J. Ecol. 2006, 94, 778–790. [Google Scholar] [CrossRef]
- Rosendahl, S.; Matzen, H.B. Genetic structure of arbuscular mycorrhizal populations in fallow and cultivated soils. New Phytol. 2008, 179, 1154–1161. [Google Scholar] [CrossRef]
- Landis, F.C.; Gargas, A.; Givnish, T.J. Relationships among arbuscular mycorrhizal fungi, vascular plants and environmental conditions in oak savannas. New Phytol. 2004, 164, 493–504. [Google Scholar] [CrossRef]
- Tan, S.; Zhu, M.; Zhang, K.; Xu, H.; Zhang, Q. Effects of submergence on morpho-physiological characteristics and recovery of bermudagrass (Cynodon dactylon). Fresen. Environ. Bull. 2013, 22, 2533–2541. [Google Scholar]
- Yang, H.; Yuan, Y.; Zhang, Q.; Tang, J.; Liu, Y.; Chen, X. Changes in soil organic carbon, total nitrogen, and abundance of arbuscular mycorrhizal fungi along a large-scale aridity gradient. Catena 2011, 87, 70–77. [Google Scholar] [CrossRef]
- Stürmer, S.L.; Kemmelmeier, K.; Moreira, B.C.; Kasuya, M.C.M.; Pereira, G.M.D.; da Silva, K. Arbuscular mycorrhizal fungi (Glomeromycota) communities in tropical savannas of Roraima, Brazil. Mycol. Prog. 2018, 17, 1149–1159. [Google Scholar] [CrossRef]
- Tchabi, A.; Coyne, D.; Hountondji, F.; Lawouin, L.; Wiemken, A.; Oehl, F. Arbuscular mycorrhizal fungal communities in sub-Saharan savannas of Benin, West Africa, as affected by agricultural land use intensity and ecological zone. Mycorrhiza 2008, 18, 181–195. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Lu, X.; Liao, Y. Effect of tillage treatment on the diversity of soil arbuscular mycorrhizal fungal and soil aggregate-associated carbon content. Front. Microbiol. 2018, 9, 2986. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.T.; Li, T.; Wen, X.X.; Liu, Y.; Han, J.; Liao, Y.C.; DeBruyn, J.M. Fungal communities in rhizosphere soil under conservation tillage shift in response to plant growth. Front. Microbiol. 2017, 8, 1301. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.J.; He, J.X.; Shi, G.X.; An, L.Z.; Öpik, M.; Feng, H.Y. Diverse communities of arbuscular mycorrhizal fungi inhabit sites with very high altitude in Tibet Plateau. FEMS Microbiol. Ecol. 2011, 78, 355–365. [Google Scholar] [CrossRef]
- Dumbrell, A.J.; Ashton, P.D.; Aziz, N.; Feng, G.; Nelson, M.; Dytham, C.; Fitter, A.H.; Helgason, T. Distinct seasonal assemblages of arbuscular mycorrhizal fungi revealed by massively parallel pyrosequencing. New Phytol. 2011, 190, 794–804. [Google Scholar] [CrossRef]
- Redecker, D. Specific PCR primers to identify arbuscular mycorrhizal fungi within colonized roots. Mycorrhiza 2000, 10, 73–80. [Google Scholar] [CrossRef]
- Stockinger, H.; Krüger, M.; Schüssler, A. DNA barcoding of arbuscular mycorrhizal fungi. New Phytol. 2010, 187, 461–474. [Google Scholar] [CrossRef]
- Gamper, H.A.; Walker, C.; Schussler, A. Diversispora celata sp. nov: Molecular ecology and phylotaxonomy of an inconspicuous arbuscular mycorrhizal fungus. New Phytol. 2009, 182, 495–506. [Google Scholar] [CrossRef]
- Sýkorová, Z.; Ineichen, K.; Wiemken, A.; Redecker, D. The cultivation bias: Different communities of arbuscular mycorrhizal fungi detected in roots from the field, from bait plants transplanted to the field, and from a greenhouse trap experiment. Mycorrhiza 2007, 18, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Stockinger, H.; Walker, C.; Schüssler, A. ‘Glomus intraradices DAOM197198′ a model fungus in arbuscular mycorrhiza research, is not Glomus intraradices. New Phytol. 2009, 183, 1176–1187. [Google Scholar] [CrossRef] [PubMed]
- Glenn, T.C. Field guide to next-generation DNA seqeuencers. Mol. Ecol. Resour. 2011, 11, 759–769. [Google Scholar] [CrossRef] [PubMed]
| Land-Use Type | EC | pH | SM (%) | SOC (g/kg) | TN (g/kg) | C/N ratio | AK (mg/kg) | AP (mg/kg) |
|---|---|---|---|---|---|---|---|---|
| NCND | 2588 ± 10.0a | 8.18 ± 0.01b | 27.58 ± 1.17a | 9.50 ± 0.02a | 0.69 ± 0.01b | 13.71 ± 0.19a | 119.56 ± 1.40b | 12.09 ± 0.19a |
| DNC | 2600 ± 2.00a | 8.28 ± 0.00a | 16.76 ± 2.10b | 6.77 ± 0.10c | 0.59 ± 0.01c | 11.47 ± 0.33c | 40.56 ± 0.92c | 9.95 ± 0.28b |
| CL | 2140 ± 6.00b | 8.27 ± 0.01a | 17.95 ± 1.00b | 8.96 ± 0.03b | 0.92 ±0.02a | 9.70 ± 0.22c | 148.84 ± 4.56a | 9.58 ± 0.09b |
| MC (%) | OTUs | Chao 1 | Shannon-Wiener | |
|---|---|---|---|---|
| Rhizosphere soil | ||||
| NCND | – | 71 ± 21.40 (a, x) | 80.19 ± 21.64 (a, x) | 4.05 ± 0.47 (a, x) |
| DNC | – | 40 ± 20.66 (a, x) | 42.38 ± 20.93 (a, x) | 2.99 ± 0.72 (a, x) |
| CL | – | 63 ± 12.99 (a, x) | 64.40 ± 13.68 (a, x) | 4.16 ± 0.45 (a, x) |
| Root | ||||
| NCND | 20.15 ± 2.45 (c, x) | 47 ± 24.00 (ab, x) | 49.33 ± 27.00 (ab, x) | 3.32 ± 0.20 (ab, x) |
| DNC | 31.85 ± 4.76 (b, x) | 23 ± 1.86 (b, x) | 24.00 ± 1.86 (b, x) | 2.42 ± 0.12 (b, x) |
| CL | 29.00 ± 2.17 (a, x) | 76 ± 20.90 (a, x) | 82.50 ± 22.18 (a, x) | 3.92 ± 0.59 (a, x) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luo, X.; He, X.; Luo, X.; Liu, Y.; Wang, J.; Dong, J. Soil Organic Carbon Shapes AMF Communities in Soils and Roots of Cynodon dactylon under Anti-Seasonal Drying-Wetting Cycles. Diversity 2019, 11, 197. https://doi.org/10.3390/d11100197
Luo X, He X, Luo X, Liu Y, Wang J, Dong J. Soil Organic Carbon Shapes AMF Communities in Soils and Roots of Cynodon dactylon under Anti-Seasonal Drying-Wetting Cycles. Diversity. 2019; 11(10):197. https://doi.org/10.3390/d11100197
Chicago/Turabian StyleLuo, Xie, Xinhua He, Xiumei Luo, Yining Liu, Junqi Wang, and Jinyan Dong. 2019. "Soil Organic Carbon Shapes AMF Communities in Soils and Roots of Cynodon dactylon under Anti-Seasonal Drying-Wetting Cycles" Diversity 11, no. 10: 197. https://doi.org/10.3390/d11100197
APA StyleLuo, X., He, X., Luo, X., Liu, Y., Wang, J., & Dong, J. (2019). Soil Organic Carbon Shapes AMF Communities in Soils and Roots of Cynodon dactylon under Anti-Seasonal Drying-Wetting Cycles. Diversity, 11(10), 197. https://doi.org/10.3390/d11100197






