Muddy Boots Beget Wisdom: Implications for Rare or Endangered Plant Species Distribution Models
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Species
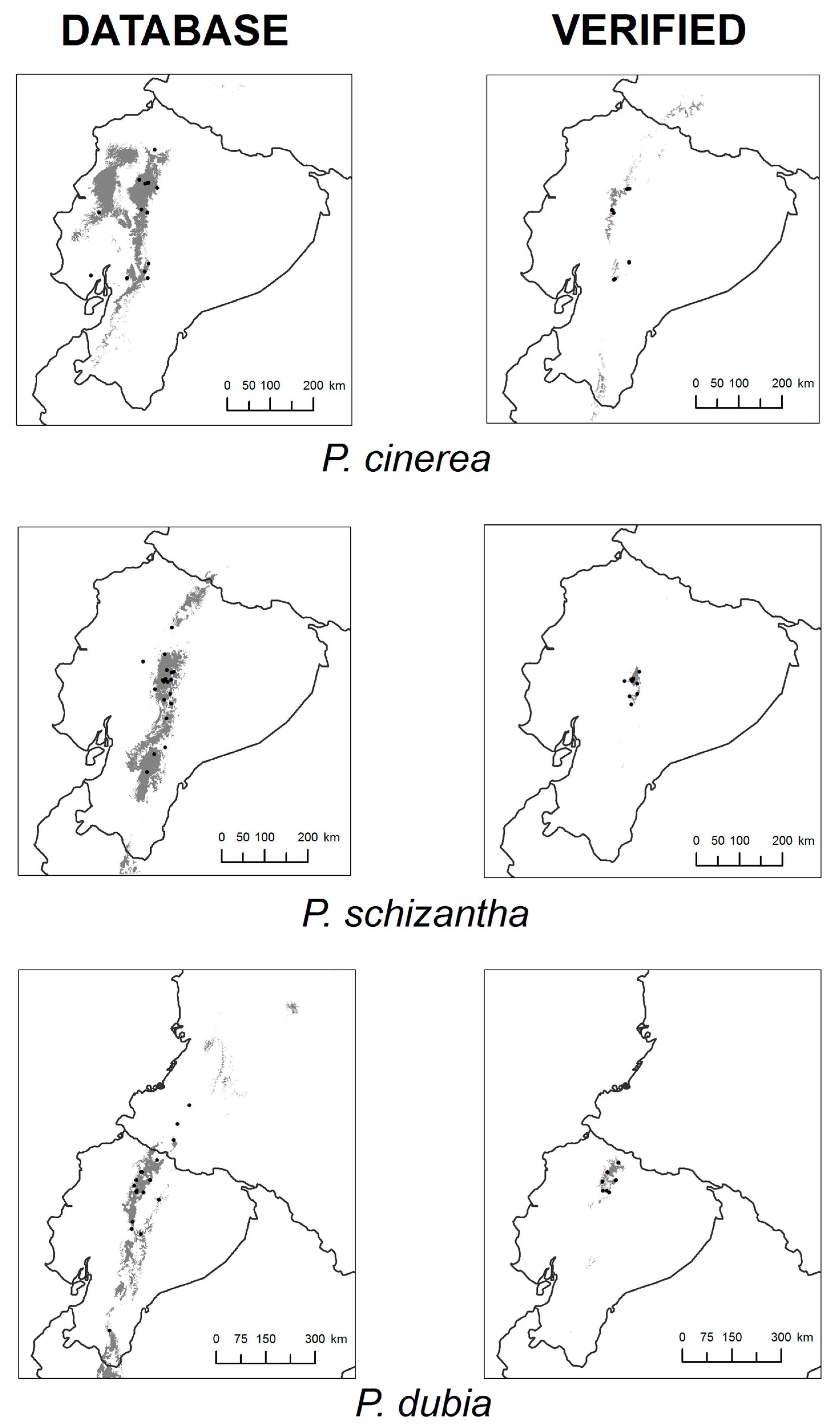
2.2. Collection of Occurrence Data
2.3. Model Building
2.4. Model Evaluation
2.5. Conservation Assessment
3. Results
4. Discussion
Our Recommendation List Summary
- Be critical of each occurrence record. For the conservation of rare and endangered species, each point should be revised.
- Check the taxonomy identification of the same collector’s number voucher. Sometimes specialists curate one of those specimens but not all and because GBIF combines data from multiple herbaria, the same collection can be identified as different species.
- Plot the coordinates into a map. It is an easy way to identify outliers.
- Use filters such as altitude or habitat when that kind of information is known for the species.
- Read the original specimen label. Important annotations might not be available in the online database.
- Besides online data, complement the number of records with information from recent taxonomy monographs, databases from permanent plots, and local herbaria. Collaborate with taxonomic specialists! Their understanding of the species is invaluable.
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Canhos, V.P.; Souza, S.; Giovanni, R.; Canhos, D.A.L. Global biodiversity informatics: Setting the scene for a “New World” of ecological modeling. Biodivers Inform. 2004. [Google Scholar] [CrossRef]
- Soberón, J.; Peterson, A.T. Biodiversity informatics: Managing and applying primary biodiversity data. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 2004, 359, 689–698. [Google Scholar] [CrossRef] [PubMed]
- Jiménez-Valverde, A.; Lobo, J.M.; Hortal, J. Not as good as they seem: The importance of concepts in species distribution modelling. Divers. Distrib. 2008, 14, 885–890. [Google Scholar] [CrossRef]
- Feeley, K.J.; Silman, M.R. The data void in modeling current and future distributions of tropical species. Glob. Chang. Biol. 2011, 17, 626–630. [Google Scholar] [CrossRef]
- Global Biodiversity Information Facility. Available online: www.gbif.org/ (accessed on 5 November 2018).
- O Projeto Species Link. Available online: Splink.cria.org.br/ (accessed on 5 November 2018).
- Franklin, J. Mapping Species Distributions: Spatial Inference and Prediction; Cambridge University Press: Cambridge, UK, 2009; 320p, ISBN 9780521700023. [Google Scholar]
- Peterson, A.T. Predicting the geography of species’ invasions via ecological niche modeling. Quat. Rev. Biol. 2003, 78, 419–433. [Google Scholar] [CrossRef]
- Martinez-Meyer, E. Climate change and biodiversity: Some considerations in forecasting shifts in species’ potential distributions. Biodivers. Inform. 2005, 2, 42–55. [Google Scholar] [CrossRef]
- Powell, M.; Accad, A.; Shapcott, A. Geographic Information System (GIS) predictions of past, present habitat distribution and areas for re-introduction of the endangered subtropical rainforest shrub Triunia robusta (Proteaceae) from south-east Queensland Australia. Biol. Conserv. 2005, 123, 165–175. [Google Scholar] [CrossRef]
- Guisan, A.; Broennimann, O.; Engler, R.; Vust, M.; Yoccoz, N.G.; Lehmann, A.; Zimmermann, N.E. Using niche-based models to improve the sampling of rare species. Conserv. Biol. 2006, 20, 501–511. [Google Scholar] [CrossRef]
- Peterson, A.T. Uses and requirements of ecological niche models and related distributional models. Biodivers. Inform. 2006, 3, 59–72. [Google Scholar] [CrossRef]
- Thuiller, W.; Lavorel, S.; Sykes, M.T.; Araújo, M.B. Using niche-based modelling to assess the impact of climate change on tree functional diversity in Europe. Divers. Distrib. 2006, 12, 49–60. [Google Scholar] [CrossRef]
- Peterson, A.T.; Nakazawa, Y. Environmental data sets matter in ecological niche modelling: An example with Solenopsis invicta and Solenopsis richteri. Glob. Ecol. Biogeogr. 2008, 17, 135–144. [Google Scholar] [CrossRef]
- Feeley, K.J.; Silman, M.R. Modelling the responses of Andean and Amazonian plant species to climate change: The effects of georeferencing errors and the importance of data filtering. J. Biogeogr. 2010, 37, 733–740. [Google Scholar] [CrossRef]
- Araújo, M.B.; Peterson, A.T. Uses and misuses of bioclimatic envelope modeling. Ecology 2012, 93, 1527–1539. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, P.A.; Graham, C.H.; Master, L.L.; Albert, D.L. The effect of sample size and species characteristics on performance of different species distribution modeling methods. Ecography 2006, 29, 773–785. [Google Scholar] [CrossRef]
- Wisz, M.S.; Hijmans, R.J.; Li, J.; Peterson, A.T.; Graham, C.H.; Guisan, A.; NCEAS Predicting Species Distributions Working Group. Effects of sample size on the performance of species distribution models. Divers. Distrib. 2008, 14, 763–773. [Google Scholar] [CrossRef]
- Wulff, A.S.; Hollingsworth, P.M.; Ahrends, A.; Jaffré, T.; Veillon, J.M.; L’Huillier, L.; Fogliani, B. Conservation priorities in a biodiversity hotspot: Analysis of narrow endemic plant species in New Caledonia. PLoS ONE 2013, 8, e73371. [Google Scholar] [CrossRef] [PubMed]
- Breiner, F.T.; Guisan, A.; Bergamini, A.; Nobis, M.P. Overcoming limitations of modelling rare species by using ensembles of small models. Methods Ecol. Evol. 2015, 6, 1210–1218. [Google Scholar] [CrossRef]
- Fois, M.; Fenu, G.; Lombrana, A.C.; Cogoni, D.; Bacchetta, G. A practical method to speed up the discovery of unknown populations using Species Distribution Models. J. Nat. Conserv. 2015, 24, 42–48. [Google Scholar] [CrossRef]
- Oleas, N.H.; Meerow, A.W.; Feeley, K.J.; Gebelein, J.; Francisco-Ortega, J. Using species distribution models as a tool to discover new populations of Phaedranassa brevifolia Meerow, 1987 (Liliopsida: Amaryllidaceae) in Northern Ecuador. Check List 2014, 10, 689–691. [Google Scholar] [CrossRef]
- Hortal, J.; Jiménez-Valverde, A.; Gómez, J.F.; Lobo, J.M.; Baselga, A. Historical bias in biodiversity inventories affects the observed environmental niche of the species. Oikos 2008, 117, 847–858. [Google Scholar] [CrossRef]
- Newbold, T. Applications and limitations of museum data for conservation and ecology, with particular attention to species distribution models. Prog. Phys. Geogr. 2010, 34, 3–22. [Google Scholar] [CrossRef]
- Beaman, R.S.; Conn, B.J. Automated geoparsing and georeferencing of Malesian collection locality data. Telopea 2003, 10, 43–52. [Google Scholar] [CrossRef]
- Murphey, P.C.; Guralnick, R.P.; Glaubitz, R.; Neufeld, D.; Ryan, J.A. Georeferencing of museum collections: A review of the problems and automated tools, and the methodology developed by the Mountain and Plains Spatial-Temporal Database-informatics initiative (MaPSTeDI). PhyloInformatics 2004. [Google Scholar] [CrossRef]
- Guralnick, R.P.; Wieczorek, J.; Beaman, R.; Hijmans, R.J. BioGeomancer: Automated georeferencing to map the world’s biodiversity data. PLoS Biol. 2006, 4, 1908–1909. [Google Scholar] [CrossRef]
- Hijmans, R.J.; Schreuder, M.; Cruz, J.D.; Guarino, L. Using GIS to check co-ordinates of genebank accessions. Genet. Resour. Crop Evol. 1999, 46, 291–296. [Google Scholar] [CrossRef]
- Guralnick, R.P.; Hill, A.W.; Lane, M. Towards a collaborative, global infrastructure for biodiversity assessment. Ecol. Lett. 2007, 10, 663–672. [Google Scholar] [CrossRef] [PubMed]
- Lozier, J.D.; Aniello, P.; Hickerson, M.J. Predicting the distribution of Sasquatch in western North America: Anything goes with ecological niche modelling. J. Biogeogr. 2009, 36, 1623–1627. [Google Scholar] [CrossRef]
- Rocchini, D.; Hortal, J.; Lengyel, S.; Lobo, J.M.; Jiménez-Valverde, A.; Ricotta, C.; Bacaro, G.; Chiarucci, A. Accounting for uncertainty when mapping species distributions: The need for maps of ignorance. Prog. Phys. Geogr. 2011, 35, 211–226. [Google Scholar] [CrossRef]
- Meerow, A.W. Amaryllidaceae. In Flora of Ecuador; Harling, G., Andersson, L., Eds.; University of Göteborg and Pontificia Universidad Católica del Ecuador: Göteborg, Sweden; Quito, Ecuador, 1990; Volume 41, pp. 1–52. ISBN 9788788702460. [Google Scholar]
- Minga, D.; Ulloa, C.U.; Oleas, N.; Verdugo, A. A new species of Phaedranassa (Amaryllidaceae) from Ecuador. Phytotaxa 2015, 192, 50–53. [Google Scholar] [CrossRef]
- Oleas, N.H. Amaryllidaceae. In Libro Rojo de las Plantas Endémicas del Ecuador, 2nd ed.; León-Yánes, S., Valencia, R., Pitman, N., Endara, L., Ulloa Ulloa, C., Navarrete, H., Eds.; Pontificia Universidad Católica del Ecuador: Quito, Ecuador, 2011; pp. 87–90. ISBN 9789942033932. [Google Scholar]
- Tropicos. The Missouri Botanical Garden. Available online: Tropicos.org/ (accessed on 5 November 2018).
- Herbarium (AAU), Institut for Bioscience, Aarhus University. Available online: Bios.au.dk/faciliteter/herbarium/ (accessed on 5 November 2018).
- C. V. Starr Virtual Herbarium, The New York Botanical Garden. Available online: Sweetgum.nybg.org/science/vh/ (accessed on 5 November 2018).
- Kew, Royal Botanic Gardens. Welcome to the Kew Herbarium Catalogue. Available online: Apps.kew.org/herbcat/navigator.do (accessed on 5 November 2018).
- University of Florida Herbarium (FLAS). Florida Museum of Natural History. Available online: www.flmnh.ufl.edu/herbarium/ (accessed on 5 November 2018).
- Colecciones Científicas en Línea. Instituto de Ciencias Naturales, Universidad Nacional de Colombia. Available online: www.biovirtual.unal.edu.co/ICN/ (accessed on 5 November 2018).
- Garcia-Milagros, E.; Funk, V.A. Improving the use of information from museum specimens: Using Google Earth© to georeference Guiana Shield specimens in the US National Herbarium. Front. Biogeogr. 2010, 2, 71–77. [Google Scholar] [CrossRef]
- Oleas, N. Landscape Genetics of Phaedranassa Herb. (Amaryllidaceae) in Ecuador. Ph.D. Thesis, Florida International University, Miami, FL, USA, 2011; 115p. [Google Scholar]
- Araújo, M.B.; New, M. Ensemble forecasting of species distributions. Trends Ecol. Evol. 2007, 22, 42–47. [Google Scholar] [CrossRef] [PubMed]
- Thuiller, W.; Georges, D.; Engler, R.; Breiner, F. biomod2: Ensemble Platform for Species Distribution Modeling. R Package version 3.3-7. 2016. Available online: http://cran.r-project.org/package=biomod2 (accessed on 5 January 2019).
- Phillips, S.J.; Anderson, R.P.; Schapire, R.E. Maximum entropy modeling of species geographic distributions. Ecol. Model. 2006, 190, 231–259. [Google Scholar] [CrossRef]
- McCullagh, P.; Nelder, J.A. Generalized Linear Models, 2nd ed.; Chapman and Hall: London, UK, 1989; 532p, ISBN 9780412317606. [Google Scholar]
- Friedman, J.H. Greedy function approximation: A gradient boosting machine. Ann. Stat. 2001, 29, 1189–1232. [Google Scholar] [CrossRef]
- Friedman, J.H. Multivariate adaptive regression splines. Ann. Stat. 1991, 19, 123–141. [Google Scholar] [CrossRef]
- Elith, J.; Graham, C.H.; Anderson, R.P.; Dudík, M.; Ferrier, S.; Guisan, A.; Hijmans, R.J.; Huettmann, F.; Leathwick, J.R.; Lehmann, A.; et al. Novel methods improve prediction of species’ distributions from occurrence data. Ecography 2006, 29, 129–151. [Google Scholar] [CrossRef]
- Mateo, R.G.; de la Estrella, M.; Felicísimo, A.M.; Muñoz, J.; Guisan, A. A new spin on a compositionalist predictiv modelling framework of conservtion planning: A tropical case study in Ecuador. Biol. Conserv. 2013, 160, 150–161. [Google Scholar] [CrossRef]
- WorldClim—Global Climate Data. Available online: www.worldclim.org/ (accessed on 5 November 2018).
- Hijmans, R.J.; Cameron, J.D.; Parra, J.L.; Jones, P.; Jarvis, A. Very high resolution interpolated climate surface for global land areas. Int. J. Clim. 2005, 25, 1965–1978. [Google Scholar] [CrossRef]
- Allouche, O.; Tsoar, A.; Kadmon, R. Assessing the accuracy of species distribution models: Prevalence, kappa and the true skill statistics (TSS). J. Appl. Ecol. 2006. [Google Scholar] [CrossRef]
- Hirzel, A.H.; Le Lay, G.; Helfer, V.; Randin, C.; Guisan, A. Evaluating the ability of habitat suitability models to predict species presences. Ecol. Model. 2006, 199, 142–152. [Google Scholar] [CrossRef]
- Pearson, R.G.; Raxworthy, C.J.; Nakamura, M.; Peterson, A.T. Predicting species distributions from small numbers of occurrence records: A test case using cryptic geckos in Madagascar. J. Biogeogr. 2007, 34, 102–117. [Google Scholar] [CrossRef]
- Mouton, A.M.; De Baets, B.; Goethals, P.L.M. Ecological relevance of performance criteria for species distribution models. Ecol. Model. 2010, 221, 1995–2002. [Google Scholar] [CrossRef]
- Broennimann, O.; Di Cola, V.; Guisan, A. Ecospat: Spatial Ecology Miscellaneous Methods. R Package Version 3.0. 2018. Available online: https://CRAN.R-project.org/package=ecospat (accessed on 5 January 2019).
- Martínez, I.; González-Taboada, F.; Wiegand, T.; Camarero, J.J.; Gutiérrez, E. Dispersal limitation and spatial affect model based projections of Pinus uncinata response to climate change in the Pyrenees. Glob. Chang. Biol. 2012, 18, 1714–1724. [Google Scholar] [CrossRef]
- Vieilledent, G.; Cornu, C.; Sanchez, C.A.; Pock-Tsy, J.-M.L.; Danthu, P. Vulnerability of baobab species to climate change and effectiveness of the protected area network in Madagascar: Towards new conservation priorities. Biol. Conserv. 2013, 166, 11–22. [Google Scholar] [CrossRef]
- IUCN. IUCN Red List Categories and Criteria; Version 3.1; IUCN Species Survival Commission: Gland, Switzerland, 2001; 30p, ISBN 2831706335. [Google Scholar]
- Dirnböck, T.; Essl, F.; Rabitsch, W. Disproportional risk for habitat loss of high altitude endemic species under climate change. Glob. Chang. Biol. 2011, 17, 990–996. [Google Scholar] [CrossRef]
- Dullinger, S.; Gattringer, A.; Thuiller, W.; Moser, D.; Zimmermann, N.E.; Guisan, A.; Willner, W.; Plutzar, C.; Leitner, M.; Mang, T.; et al. Extinction debt of high-mountain plants under twenty-first-century climate change. Nat. Clim. Chang. 2012, 2, 619–622. [Google Scholar] [CrossRef]
- Urrutia, R.; Vuille, M. Climate change projections for the tropical Andes using a regional climate model: Temperature and precipitation simulations for the end of the 21st century. J. Geophys. Res. 2009, 114, D00G17. [Google Scholar] [CrossRef]
| Species | N | TSS | AUC | Boyce Index | p-Value * | Altitude (Min) | Altitude (Max) | Distribution Range Area |
|---|---|---|---|---|---|---|---|---|
| P. cinerea 1 | 13 | 0.89 | 0.99 | 0.85 | 0.00 | 25 | 3964 | 20536 |
| P. cinerea 2 | 10 | 0.90 | 1.00 | 0.65 | 0.03 | 1010 | 3252 | 2584 |
| P. dubia 1 | 27 | 0.86 | 0.98 | 0.73 | 0.00 | 1623 | 6169 | 20416 |
| P. dubia 2 | 14 | 0.88 | 1.00 | 1.00 | 0.00 | 1566 | 3844 | 2508 |
| P. schizantha 1 | 20 | 0.91 | 0.99 | 0.90 | 0.00 | 1623 | 6169 | 17481 |
| P. schizantha 2 | 10 | 0.92 | 1.00 | 0.90 | 0.00 | 1730 | 3826 | 859 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oleas, N.H.; Feeley, K.J.; Fajardo, J.; Meerow, A.W.; Gebelein, J.; Francisco-Ortega, J. Muddy Boots Beget Wisdom: Implications for Rare or Endangered Plant Species Distribution Models. Diversity 2019, 11, 10. https://doi.org/10.3390/d11010010
Oleas NH, Feeley KJ, Fajardo J, Meerow AW, Gebelein J, Francisco-Ortega J. Muddy Boots Beget Wisdom: Implications for Rare or Endangered Plant Species Distribution Models. Diversity. 2019; 11(1):10. https://doi.org/10.3390/d11010010
Chicago/Turabian StyleOleas, Nora H., Kenneth J. Feeley, Javier Fajardo, Alan W. Meerow, Jennifer Gebelein, and Javier Francisco-Ortega. 2019. "Muddy Boots Beget Wisdom: Implications for Rare or Endangered Plant Species Distribution Models" Diversity 11, no. 1: 10. https://doi.org/10.3390/d11010010
APA StyleOleas, N. H., Feeley, K. J., Fajardo, J., Meerow, A. W., Gebelein, J., & Francisco-Ortega, J. (2019). Muddy Boots Beget Wisdom: Implications for Rare or Endangered Plant Species Distribution Models. Diversity, 11(1), 10. https://doi.org/10.3390/d11010010





