Characterization and Phylogenetic Analysis of Chloroplast and Mitochondria Genomes from the Antarctic Polytrichaceae Species Polytrichum juniperinum and Polytrichum strictum
Abstract
:1. Introduction
2. Materials and Methods
3. Results
3.1. Sequence Data
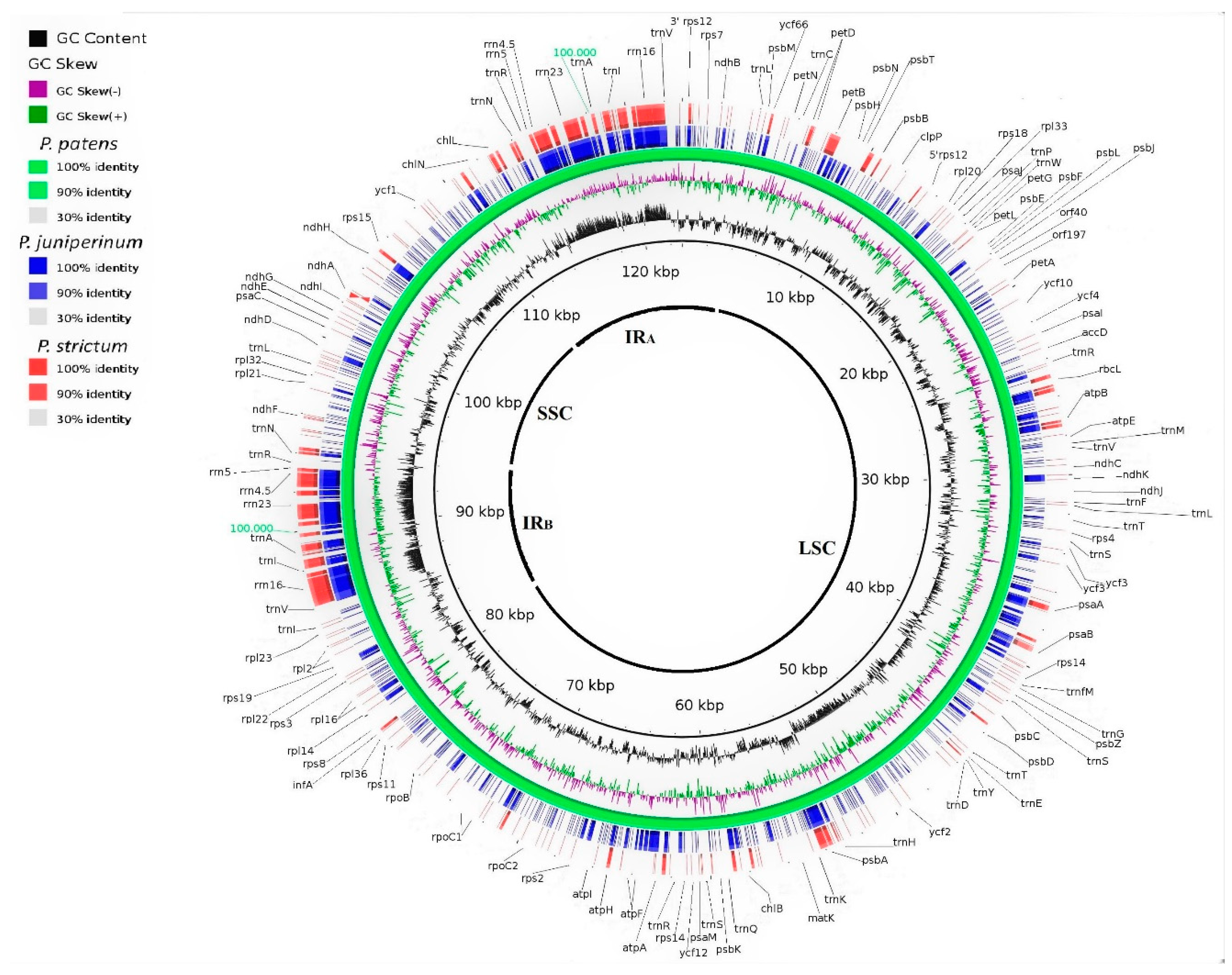
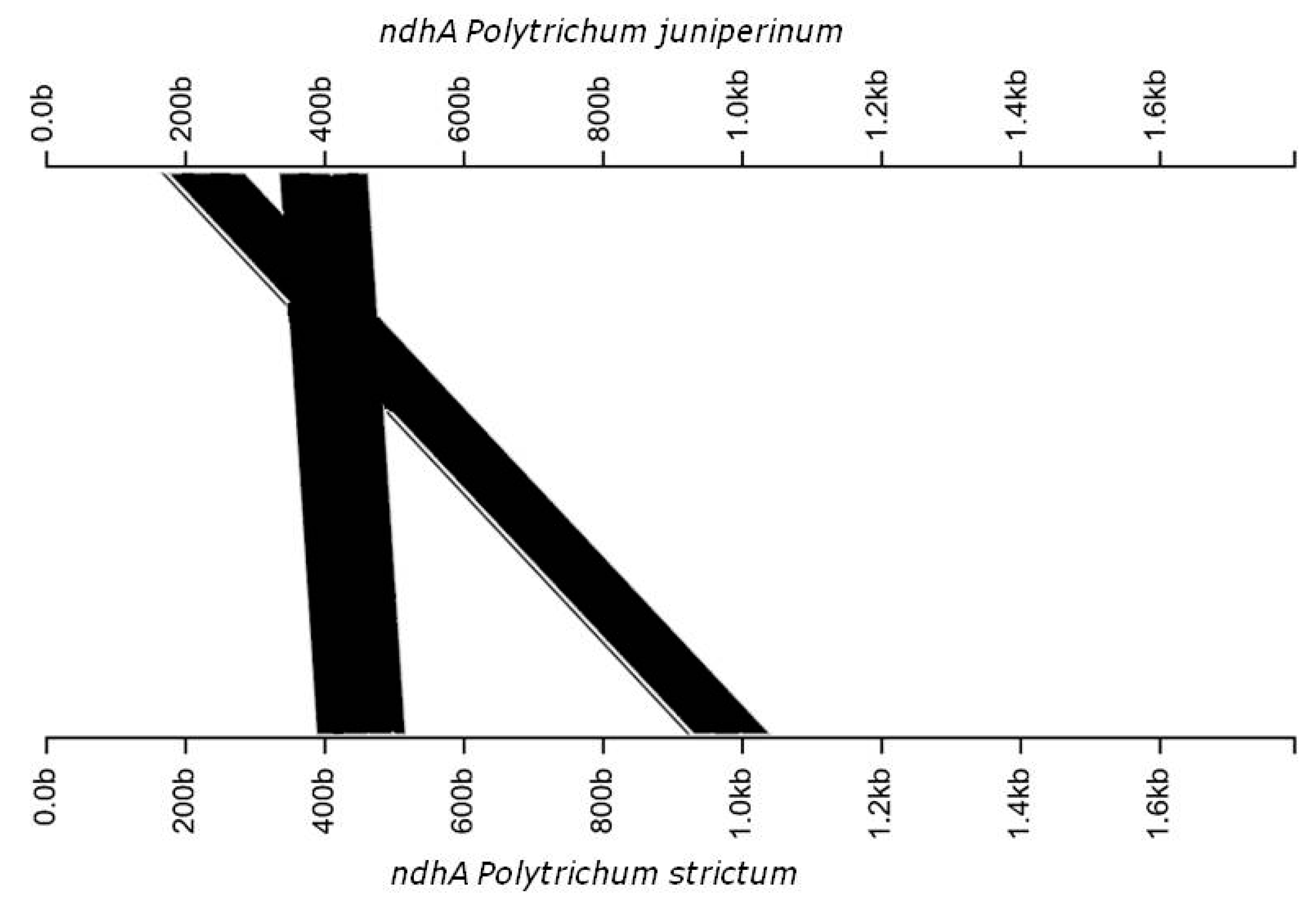
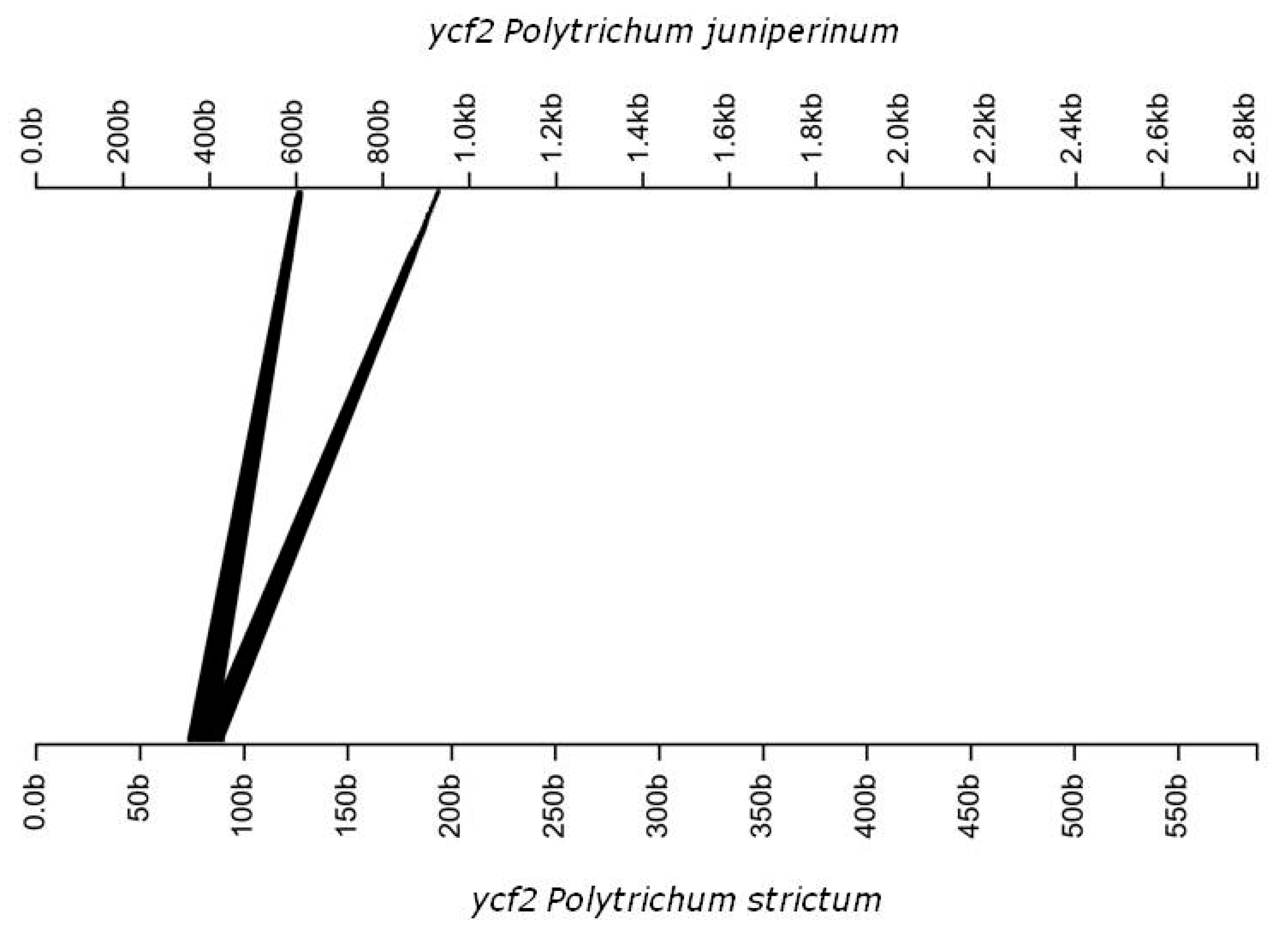
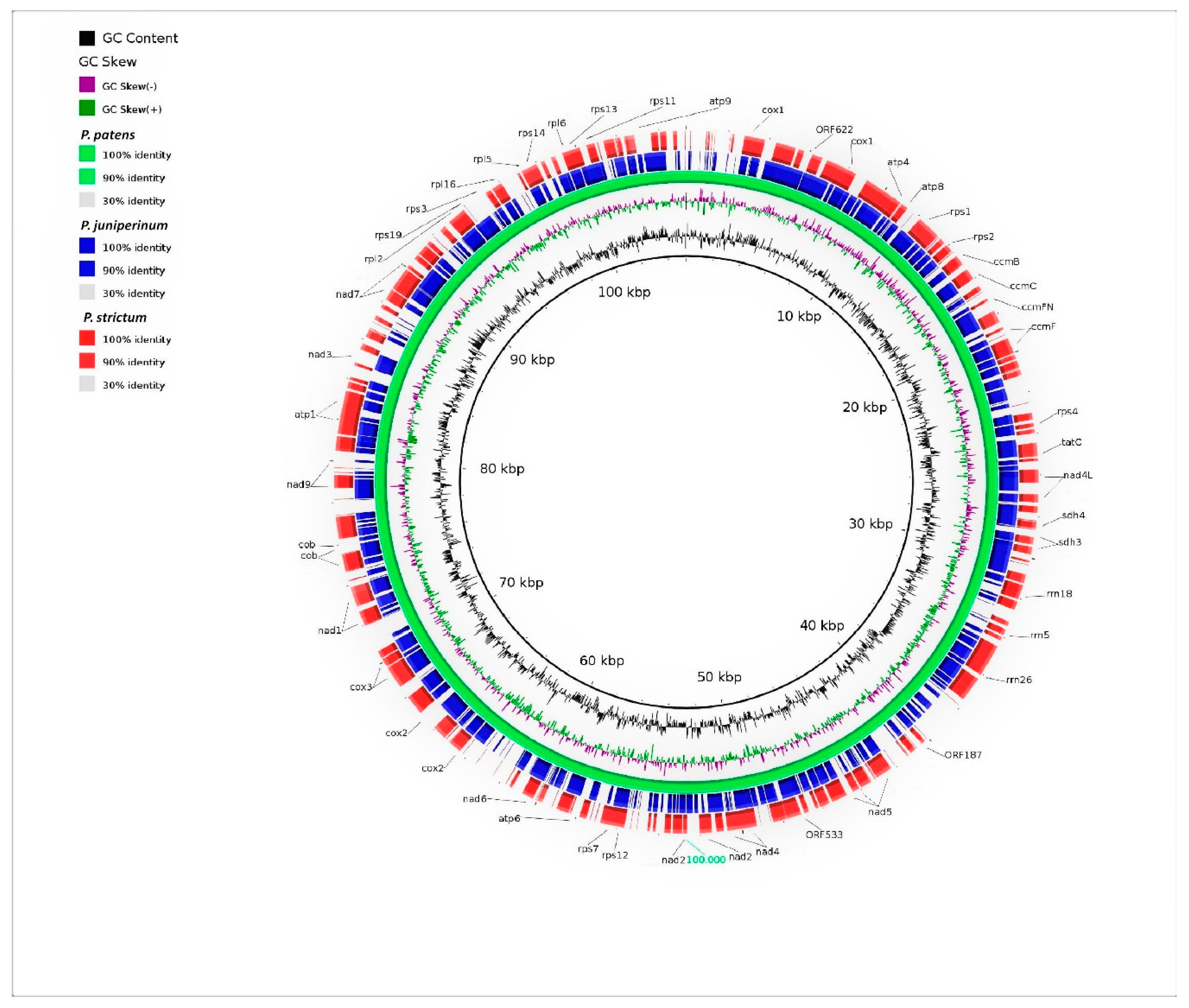
3.2. Genomic Organization and Gene Content
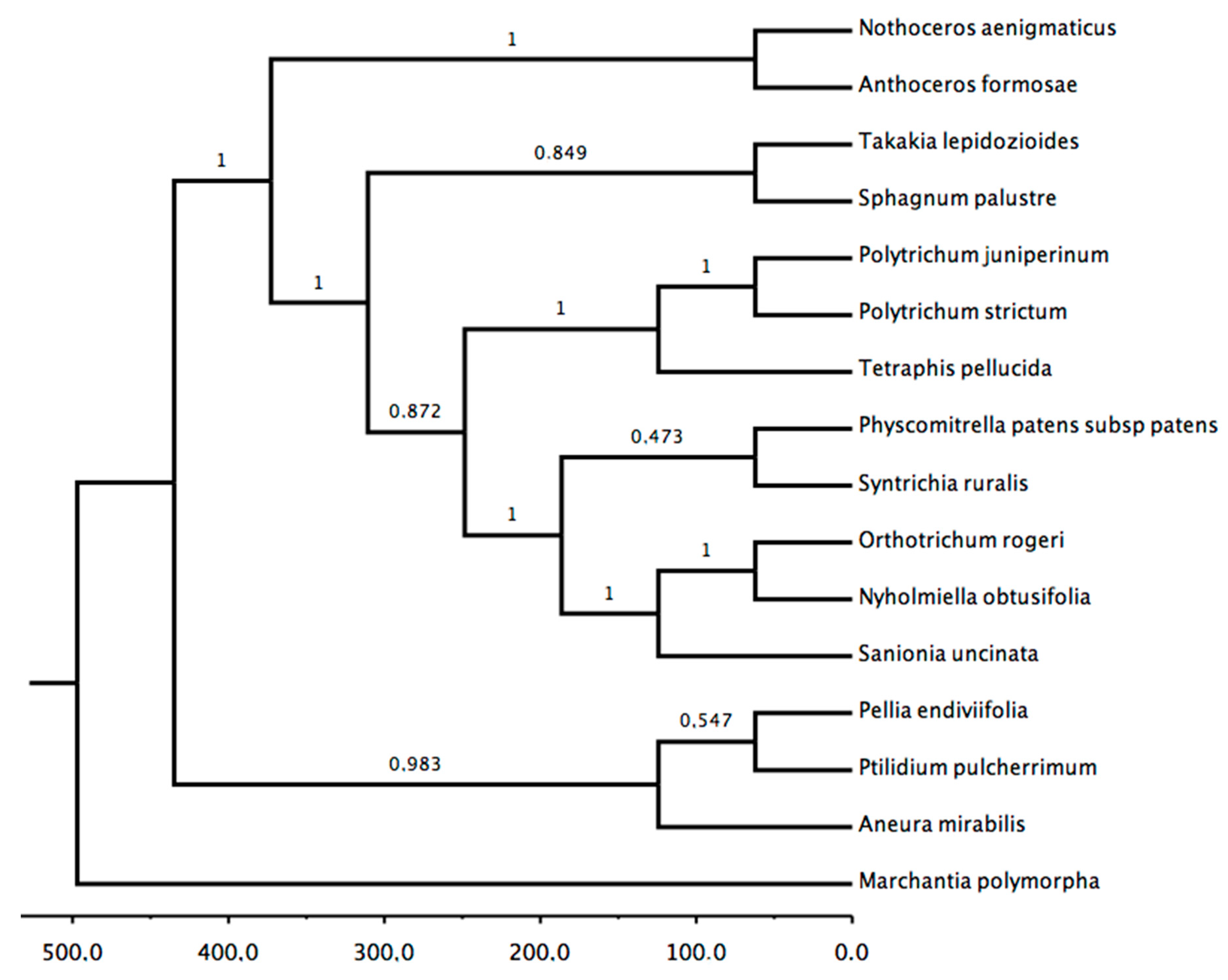
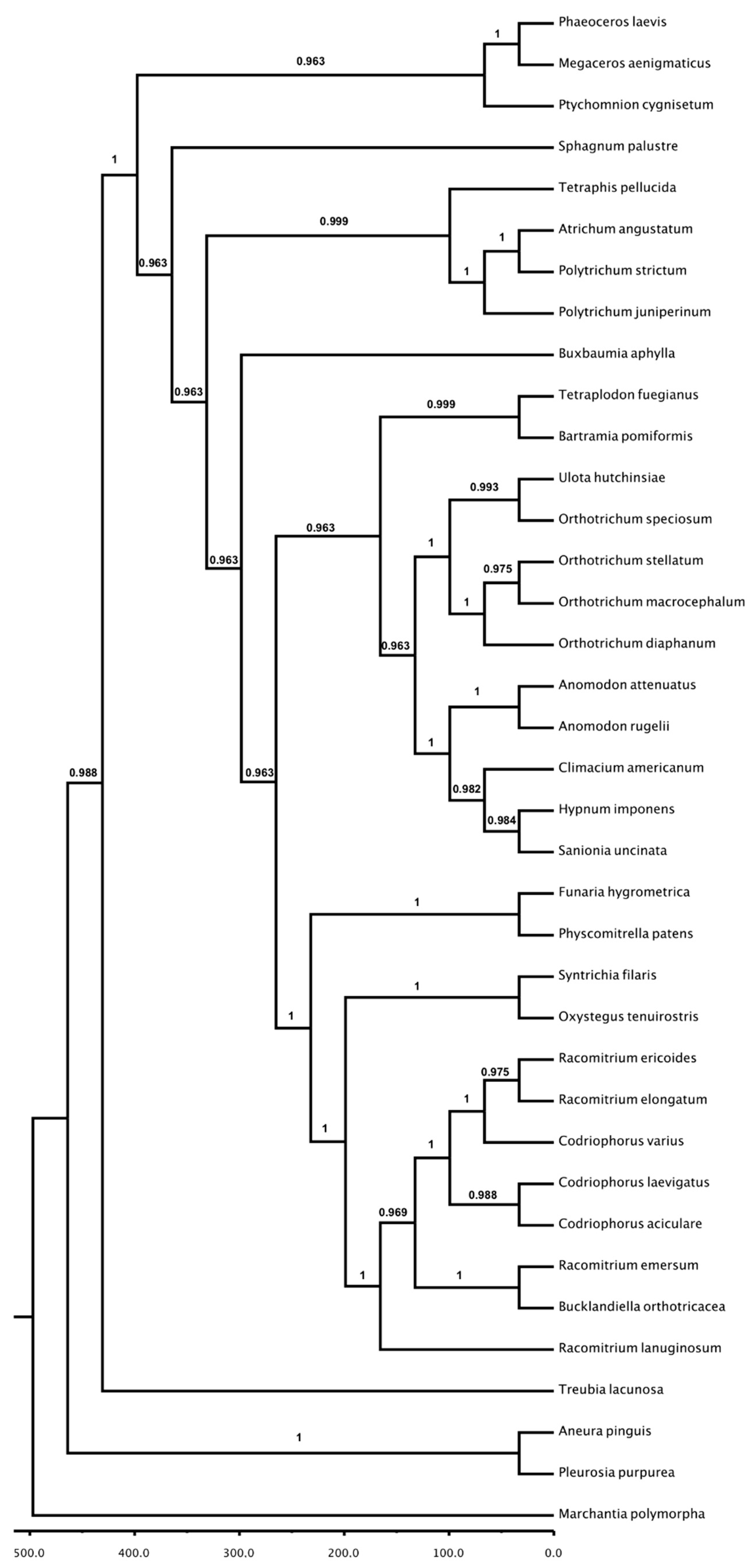
3.3. Phylogenetic Analysis: Chloroplast and Mitochondria
4. Discussion
5. Nucleotide Sequence Accession Numbers
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ochyra, R. The Moss Flora of King George Island Antarctic; Polish Academy of Sciences, W. Szafer Institute of Botany: Cracow, Poland, 1998. [Google Scholar]
- Greene, S.W.; Greene, D.M.; Brown, P.D.; Pacey, J.M. Antarctic Moss Flora: 1. The Genera Andreaea, Pohlia, Polytrichum, Psilopilum and Sarconeurum, 64th ed.; British Antartic Survey: London, UK, 1970. [Google Scholar]
- Mishler, B.D.; Churchill, S.P. A Cladistic Approach to the Phylogeny of the “Bryophytes”. Brittonia 1984, 36, 406–424. [Google Scholar] [CrossRef]
- Bell, N.E.; Hyvönen, J.P. Phylogeny of the moss class Polytrichopsida (BRYOPHYTA): Generic level structure and incongruent gene trees. Mol. Phylogenet. Evol. 2010, 55, 381–398. [Google Scholar] [CrossRef] [PubMed]
- Nock, C.; Waters, D.L.; Edwards, M.A.; Bowen, S.G.; Rice, N.; Cordeiro, G.M.; Henry, R.J. Chloroplast genome sequences from total DNA for exploring plant relationships. Plant Biotechnol. J. 2010, 9, 328–333. [Google Scholar] [CrossRef] [PubMed]
- Atherton, R.A.; McComish, B.J.; Shepherd, L.D.; Berry, L.A.; Albert, N.W.; Lockhart, P.J. Whole genome sequencing of enriched chloroplast DNA using the Illumina GAII platform. Plant Methods 2010, 6, 22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Drummond, A.J.; Suchard, M.A.; Xie, D.; Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 2012, 29, 1969–1973. [Google Scholar] [CrossRef] [PubMed]
- Leggett, R.M.; Ramirez-Gonzalez, R.H.; Clavijo, B.; Waite, D.; Davey, R.P. Sequencing quality assessment tools to enable data-driven informatics for high throughput genomics. Front. Genet. 2013, 4, 288. [Google Scholar] [CrossRef] [PubMed]
- Zerbino, D.R.; Birney, E. Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 2008, 18, 821–829. [Google Scholar] [CrossRef] [PubMed]
- Chikhi, R.; Medvedev, P. Informed and automated k-mer size selection for genome assembly. Bioinformatics 2014, 30, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Benson, D.A.; Karsch-Mizrachi, I.; Lipman, D.J.; Ostell, J.; Wheeler, D.L. GenBank. Nucleic Acids Res. 2005, 33, 34–38. [Google Scholar] [CrossRef] [PubMed]
- Wyman, S.K.; Jansen, R.K.; Boore, J.L. Automatic annotation of organellar genomes with DOGMA. Bioinformatics 2004, 20, 3252–3255. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, C.; Shi, L.; Zhu, Y.; Chen, H.; Zhang, J.; Lin, X.; Guan, X. CpGAVAS, an integrated web server for the annotation, visualization, analysis, and GenBank submission of completely sequenced chloroplast genome sequences. BMC Genom. 2012, 13, 715. [Google Scholar] [CrossRef] [PubMed]
- Alverson, A.J.; Wei, X.; Rice, D.W.; Stern, D.B.; Barry, K.; Palmer, J.D. Insights into the evolution of mitochondrial genome size from complete sequences of Citrullus lanatus and Cucurbita pepo (Cucurbitaceae). Mol. Biol. Evol. 2010, 27, 1436–1448. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acid Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Alikhan, N.F.; Petty, N.K.; Ben Zakour, N.L.; Beatson, S.A. BLAST Ring Image Generator (BRIG): Simple prokaryote genome comparisons. BMC Genom. 2011, 12, 402. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular Evolutionary Genetic Analysis using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Posada, D. JModelTest: Phylogenetic model averaging. Mol. Biol. Evol. 2008, 25, 1253–1256. [Google Scholar] [CrossRef] [PubMed]
- Magallón, S.; Hilu, K.W. The Timetree of Life; Hedges, S.B., Kumar, S., Eds.; Oxford University Press: Oxford, UK, 2009. [Google Scholar]
- Oliver, M.J.; Murdock, A.G.; Mishler, B.D.; Kuehl, J.V.; Boore, J.L.; Mandoli, D.F.; Everett, K.D.; Wolf, P.G.; Duffy, A.M.; Karol, K.G. Chloroplast genome sequence of the moss Tortula ruralis: Gene content, polymorphism, and structural arrangement relative to other green plant chloroplast genomes. BMC Genom. 2010, 11, 143. [Google Scholar] [CrossRef] [PubMed]
- Bell, N.E.; Boore, J.L.; Mishler, B.D.; Hyvönen, J. Organellar genomes of the four-toothed moss. Tetraphis pellucida. BMC Genom. 2014, 15, 383. [Google Scholar]
- Sugiura, C.; Kobayashi, Y.; Aoki, S.; Sugita, C.; Sugita, M. Complete chloroplast DNA sequence of the moss Physcomitrella patens: Evidence for the loss and relocation of rpoA from the chloroplast to the nucleus. Nucleic Acids Res. 2003, 31, 5324–5331. [Google Scholar] [CrossRef] [PubMed]
- Silva, G.G.; Dutilh, B.E.; Matthews, T.D.; Elkins, K.; Schmieder, R.; Dinsdale, E.A.; Edwards, R.A. Combining de novo and reference-guided assembly with scaffold builder. Source Code Biol. Med. 2013, 8, 23. [Google Scholar] [CrossRef] [PubMed]
- Terasawa, K.; Odahara, M.; Kabeya, Y.; Kikugawa, T.; Sekine, Y.; Fujiwara, M.; Sato, N. The mitochondrial genome of the moss Physcomitrella patens sheds new light on mitochondrial evolution in land plants. Mol. Biol. Evol. 2007, 24, 699–709. [Google Scholar] [CrossRef] [PubMed]
- Oda, K.; Yamato, K.; Ohta, E.; Nakamura, Y.; Takemura, M.; Nozato, N.; Akashi, K.; Kanegae, T.; Ogura, Y.; Kohchi, T.; et al. Gene organization deduced from the complete sequence of liverwort Marchantia polymorpha mitochondrial DNA: A primitive form of plant mitochondrial genome. J. Mol. Biol. 1992, 223, 1–7. [Google Scholar] [CrossRef]
- Graham, L.E. Green algae to land plants: An evolutionary transition. J. Plant Res. 1996, 109, 241–251. [Google Scholar] [CrossRef]
- Ohyama, K.; Fukuzawa, H.; Kohchi, T.; Shirai, H.; Sano, T.; Sano, S.; Umesono, K.; Shiki, Y.; Takeuchi, M.; Chang, Z.; et al. Chloroplast gene organization deduced from complete sequence of liverwort Marchantia polymorpha chloroplast DNA. Nature 1986, 322, 572–574. [Google Scholar] [CrossRef]
- Ohyama, K. Chloroplast and mitochondrial genomes from a liverwort, Marchantia polymorpha—Gene organization and molecular evolution. Biosci. Biotechnol. Biochem. 1996, 60, 16–24. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Medina, R.; Goffinet, B. 350 My of mitochondrial genomes stasis in mosses, an early land plant lineage. Mol. Biol. Evol. 2014, 31, 2586–2591. [Google Scholar] [CrossRef] [PubMed]
- Kugita, M.; Kaneko, A.; Yamamoto, Y.; Takeya, Y.; Matsumoto, T.; Yoshinaga, K. The complete nucleotide sequence of the hornwort (Anthoceros formosae) chloroplast genome: Insight into the earliest land plants. Nucleic Acids Res. 2003, 31, 716–721. [Google Scholar] [CrossRef] [PubMed]
- Lockhart, J. Plastid genes that were lost along the road to parasitism. Plant Cell 2013, 25, 3636. [Google Scholar] [CrossRef] [PubMed]
- Daniell, H.; Chase, C.D. Molecular Biology and Biotechnology of Plant Organelles: Chloroplasts and Mitochondria; Springer Science & Business Media: Berlin, Germany, 2007. [Google Scholar]
- Knoop, V. Looking for sense in the nonsense: A short review of non-coding organellar DNA elucidating the phylogeny of bryophytes. Trop. Bryol. 2010, 31, 51–60. [Google Scholar]
- Qiu, Y.L.; Li, L.; Wang, B.; Chen, Z.; Knoop, V.; Groth-Malonek, M.; Dombrovska, O.; Lee, J.; Kent, L.; Rest, J.; et al. The deepest divergences in land plants inferred from phylogenomic evidence. Proc. Natl. Acad. Sci. USA 2006, 103, 15511–15516. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Edwards, S.R. New Manual of Bryology; The Hattori Botanical Laboratory: Nichican, Japan, 1984. [Google Scholar]
- Magombo, Z.L. The phylogeny of basal peristomate mosses: Evidence from cpDNA, and implications for peristome evolution. Syst. Bot. 2003, 28, 24–38. [Google Scholar]
- Volkmar, U.; Knoop, V. Introducing intron locus cox1i624 for phylogenetic analysis in bryophytes: On the issue of Takakia as sister genus to all other extant mosses. J. Mol. Evol. 2010, 70, 506–518. [Google Scholar] [CrossRef] [PubMed]
- Ligrone, R.; Ducket, J.G. Morphology versus molecules in moss phylogeny: New insights (or controversies) from placental and vascular anatomy in Oedipodium griffithianum. Plant Syst. Evol. 2011, 296, 275–282. [Google Scholar] [CrossRef]
- Cox, C.J.; Goffinet, B.; Shaw, J.; Boles, S.B. Phylogenetic relationships among the mosses based on heterogeneous bayesian analysis of multiple genes from multiple genomic compartments. Syst. Bot. 2004, 29, 234–250. [Google Scholar] [CrossRef]
- Hyvönen, J.; Koskinen, S.; Smith, M.G.L.; Hedderson, T.A.; Stenroos, S. Phylogeny of the Polytrichales (Bryophyta) based on simultaneous analysis of molecular and morphological data. Mol. Phylogenet. Evol. 2004, 31, 915–928. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Newton, A.E.; Cox, C.J.; Duckett, J.G.; Wheeler, J.A.; Goffinet, B.; Hedderson, T.A.J.; Mishler, B.D. Evolution of the major moss lineages: Phylogenetic analyses based on multiple gene sequences and morphology. Bryologist 2000, 103, 187–211. [Google Scholar] [CrossRef]
- Goffinet, B.; Buck, W.R. Monographs in Systematic Botany from the Missouri Botanical Garden. In Molecular Systematics of Bryophytes; Goffinet, B., Hollowell, V., Magill, R., Eds.; Missouri Botanical Garden Press: St. Louis, MO, USA, 2004; pp. 205–239. [Google Scholar]
- Liu, Y.; Cox, C.J.; Wang, W.; Goffinet, B. Mitochondrial phylogenomics of early land plants: Mitigating the effects of saturation, compositional heterogeneity, and codon-usage bias. Syst. Biol. 2014, 63, 862–878. [Google Scholar] [CrossRef] [PubMed]
- Daniell, H.; Lin, C.S.; Yu, M.; Chang, W.J. Chloroplast genomes: Diversity, evolution, and applications in genetic engineering. Genome Biol. 2016, 17, 134. [Google Scholar] [CrossRef] [PubMed]
- Lane, N.; Martin, W. The energetics of genome complexity. Nature 2010, 467, 929–934. [Google Scholar] [CrossRef] [PubMed]
- Leitch, I.J.; Chase, M.W.; Bennett, M.D. Phylogenetic analysis of DNA c-values provides evidence for a small ancestral genome size in flowering plants. Ann. Bot. 1998, 82, 85–94. [Google Scholar] [CrossRef]
- Vesely, P.; Bures, P.; Smarda, P.; Pavlicek, T. Genome size and DNA base composition of geophytes: The mirror of phenology and ecology? Ann. Bot. 2011, 109, 65–75. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.-M.; Wang, J.; Feng, L.; Liu, S.; Pang, H.; Qi, L.; Li, J.; Sun, W.Y.; Qiao, W.; Zhang, L.; et al. Inferring the evolutionary mechanism of the chloroplast genome size by comparing whole-chloroplast genome sequences in seed plants. Sci. Rep. 2017, 7, 1555. [Google Scholar]
- Jansen, R.K.; Ruhlman, T.A. Genomics of Chloroplasts and Mitochondria; Bock, R., Knoop, V., Eds.; Springer: New York, NY, USA, 2012; pp. 103–126. [Google Scholar]
- Cai, Z.; Guisinger, M.; Kim, H.G.; Ruck, E.; Blazier, J.C.; McMurtry, V.; Kuehl, J.V.; Boore, J.; Jansen, R.K. Extensive reorganization of the plastid genome of Trifolium subterraneum (Fabaceae) is associated with numerous repeated sequences and novel DNA insertions. J. Mol. Evol. 2008, 67, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Xue, J.Y.; Wang, B.; Li, L.; Qiu, Y.L. The mitochondrial genomes of the early land plants Treubia lacunosa and Anomodon rugelii: Dynamic and conservative evolution. PLoS ONE 2011, 6, e25836. [Google Scholar] [CrossRef] [PubMed]
- Goffinet, B.; Cox, C.J. Phylogenetic relationships among basal-most arthrodontous mosses with special emphasis on the evolutionary significance of the Funariineae. Bryologist 2000, 103, 212–223. [Google Scholar] [CrossRef]
- Cox, C.J.; Hedderson, T.A.J. Phylogenetic relationships among the ciliate arthrodontous mosses: Evidence from chloroplast and nuclear DNA sequences. Plant Syst. Evol. 1999, 215, 119–139. [Google Scholar] [CrossRef]
- Goffinet, B.; Wickett, N.J.; Shaw, A.J.; Cox, C.J. Phylogenetic significance of the rpoA loss in the chloroplast genome of mosses. Taxon 2005, 54, 353–360. [Google Scholar] [CrossRef]
- Sheveleva, E.V.; Giordani, N.V.; Hallick, R.B. Identification and comparative analysis of the chloroplast α-subunit gene of DNA-dependent RNA polymerase from seven Euglena species. Nucleic Acids Res. 2002, 30, 1247–1254. [Google Scholar] [CrossRef] [PubMed]
- Sato, S.; Nakamura, Y.; Kaneko, T.; Asamizu, E.; Tabata, S. Complete structure of the chloroplast genome of Arabidopsis thaliana. DNA Res. 1999, 6, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Wakasugi, T.; Nagai, T.; Kapoor, M.; Sugita, M.; Ito, M.; Ito, S.; Tsudzuki, J.; Nakashima, K.; Tsudzuki, T.; Suzuki, Y.; et al. Complete nucleotide sequence of the chloroplast genome from the green alga Chlorella vulgaris: The existence of genes possibly involved in chloroplast division. Proc. Natl. Acad. Sci. USA 1997, 94, 5967–5972. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Zhou, Y.; Wang, Z.W.; Su, Y.J.; Wang, T. Evolution of the rpoB-psbZ region in fern plastid genomes: Notable structural rearrangements and highly variable intergenic spacers. BMC Plant Biol. 2011, 11, 64. [Google Scholar] [CrossRef] [PubMed]
- Adams, K.L.; Daley, D.O.; Qiu, Y.L.; Whelan, J.; Palmer, J.D. Repeated, recent and diverse transfers of a mitochondrial gene to the nucleus in flowering plants. Nature 2000, 408, 354–357. [Google Scholar] [CrossRef] [PubMed]
- Lelandais, C.; Albert, B.; Gutierres, S.; De Paepe, R.; Godelle, B.; Vedel, F.; Chétrit, P. Organization and expression of the mitochondrial genome in the Nicotiana sylvestris CMSII mutant. Genetics 1998, 150, 873–882. [Google Scholar] [PubMed]
- Gao, L.; Yi, X.; Yang, Y.-X.; Su, Y.-J.; Wang, T. Complete chloroplast genome sequence of a tree fern Alsophila spinulosa: Insights into evolutionary changes in fern chloroplast genomes. BMC Evol. Biol. 2009, 9, 130. [Google Scholar] [CrossRef] [PubMed]
- Doolittle, W.E. You are what you eat: A gene transfer ratchet could account for bacterial genes in eukaryotic nuclear genomes. Trends Genet. 1998, 14, 307–311. [Google Scholar] [CrossRef]
- Palmer, J.D. The Molecular Biology of Plastids; Hermann, R.G., Ed.; Springer: Berlin, Germany, 1991; pp. 5–53. [Google Scholar]
- Downie, S.R.; Palmer, J.D. Molecular Systematics of Plants; Soltis, P.S., Soltis, D.E., Doyle, J.J., Eds.; Chapman and Hall: New York, NY, USA; London, UK, 1992; pp. 14–35. [Google Scholar]
- Quandt, D.; Müller, K.; Huttunen, S. Characterisation of the chloroplast DNA psbT-H region and the influence of dyad symmetrical elements on phylogenetic reconstructions. Plant Biol. 2003, 5, 400–410. [Google Scholar] [CrossRef]
- Huttunen, S.; Ignatov, M.S. Phylogeny of the Brachytheciaceae (Bryophyta) based on morphology and sequence level data. Cladistics 2004, 20, 151–183. [Google Scholar] [CrossRef]
- Hernandez-Maqueda, R.; Quandt, D.; Werner, O.; Munoz, J. Phylogeny and classification of the Grimmiaceae/Ptychomitriaceae complex (Bryophyta) inferred from cpDNA. Mol. Phylogenet. Evol. 2008, 46, 863–877. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, B.; Li, L.; Qiu, Y.-L.; Xue, J. Conservative and dynamic evolution of mitochondrial genomes in early land plants. In Genomics of Chloroplasts and Mitochondria; Springer: Dordrecht, The Netherlands, 2012; Volume 7, pp. 159–174. [Google Scholar]
- Donoghue, M.J.; Cracraft, J. Assembling the Tree of Life; Cracraft, J., Donoghue, M.J., Eds.; Oxford University Press: New York, NY, USA, 2004; pp. 1–4. [Google Scholar]
- Cronn, R.; Liston, A.; Parks, M.; Gernandt, D.S.; Shen, R.; Mockler, T. Multiplex sequencing of plant chloroplast genomes using Solexa sequencing-by synthesis technology. Nucleic Acids Res. 2008, 36, e122. [Google Scholar] [CrossRef] [PubMed]
- Wolf, P.G.; Der, J.P.; Duffy, A.M.; Davidson, J.B.; Grusz, A.L.; Pryer, K.M. The evolution of chloroplast genes and genomes in ferns. Plant Mol. Biol. 2011, 76, 251–261. [Google Scholar] [CrossRef] [PubMed]
- Henson, J.; Tischler, G.; Ning, Z. Next-generation sequencing and large genome assemblies. Pharmacogenomics 2012, 13, 901–915. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lawton, E. Moss Flora of the Pacific Northwest, 1st ed.; The Hattori Botanical Laboratory: Nichican, Japan, 1971; 389p. [Google Scholar]
- Steere, W.C.; Brassard, G.R. The Mosses of Arctic Alaska, 1st ed.; Cramer Publisher: Lehre, Germany, 1978; 508p. [Google Scholar]
- Koponen, T.; Isoviita, P.; Lammes, T. The Bryophytes of Finland: An Annotated Checklist, 1st ed.; Societas pro fauna et flora Fennica: Helsinki, Finland, 1977; 77p. [Google Scholar]
- Anderson, L.E.; Crum, H.A.; Buck, W.R. List of the mosses of North America and north of Mexico. Bryologist 1990, 93, 448–499. [Google Scholar] [CrossRef]
- Derda, G.S.; Wyatt, R. Genetic variation and population structure in Polytrichum juniperinum and Polytricum strictum (Polytrichaceae). Lindbergia 2003, 28, 23–40. [Google Scholar]
- Cox, C.J.; Goffinet, B.; Wickett, N.J.; Boles, S.B.; Shaw, A.J. Moss diversity: A molecular phylogenetic analysis of genera. Phytotaxa 2010, 9, 175–195. [Google Scholar] [CrossRef]
- La Farge, C.; Mishler, B.D.; Wheeler, J.A.; Wall, D.P.; Johannes, K.; Schaffer, S.; Shaw, A.J. Phylogenetic relationships within the haplolepideous mosses. Bryologist 2000, 103, 257–276. [Google Scholar] [CrossRef]
- De Luna, E.; Newton, A.E.; Withey, A.; Gonzalez, D.; Mishler, B.D. The transition to pleurocarpy: A phylogenetic analysis of the main Diplolepidous lineages based on rbcL sequences and morphology. Bryologist 1999, 102, 634–650. [Google Scholar] [CrossRef]
- De Luna, E.; Buck, W.R.; Akiyama, H.; Arikawa, T.; Tsubota, H.; Gonzalez, D.; Newton, A.E.; Shaw, A.J. Ordinal phylogeny within the Hypnobryalean pleurocarpous mosses inferred from cladistic analyses of three chloroplast DNA sequence data sets: trnL-F, rps4, and rbcL. Bryologist 2000, 103, 242–256. [Google Scholar] [CrossRef]
- Vaidya, G.; Lohman, D.L.; Meier, R. SequenceMatrix: Concatenation software for the fast assembly of multigene datasets with character set and codon information. Cladistics 2010, 27, 171–180. [Google Scholar] [CrossRef]
- Nishiyama, T.; Wolf, P.G.; Kugita, M.; Sinclair, R.B.; Sugita, M.; Sugiura, C.; Wakasugi, T.; Yamada, K.; Yoshinaga, K.; Yamaguchi, K.; et al. Chloroplast phylogeny indicates that bryophytes are monophyletic. Mol. Biol. Evol. 2004, 21, 1813–1819. [Google Scholar] [CrossRef] [PubMed]
- Wahrmund, U.; Rein, T.; Müller, K.F.; Groth-Malonek, M.; Knoop, V. Fifty mosses on five trees: Comparing phylonetic information in three types of non-coding mitochondrial DNA and two chloroplast loci. Plant Syst. Evol. 2009, 282, 241–255. [Google Scholar] [CrossRef]
- Shaw, A.J.; Allen, B.H. Phylogenetic relationships, morphological incongruence, and geographic speciation in the Fontinalaceae (Bryophyta). Mol. Syst. Evol. 2000, 16, 225–237. [Google Scholar] [CrossRef] [PubMed]
- Sawicki, J.; Szczecińska, M.; Bednarek-Ochyra, H.; Ochyra, R. Mitochondrial phylogenomics supports splitting the traditionally conceived genus Racomitrium (Bryophyta: Grimmiaceae). Nova Hedwig. 2015, 100, 293–317. [Google Scholar] [CrossRef]
- Molina, J.; Hazzouri, K.M.; Nickrent, D.; Geisler, M.; Meyer, R.S.; Pentony, M.M.; Flowers, J.M.; Pelser, P.; Barcelona, J.; Inovejas, S.A.; et al. Possible loss of the chloroplast genome in the parasitic flowering plant Rafflesia lagascae (Rafflesiaceae). Mol. Biol. Evol. 2014, 31, 793–803. [Google Scholar] [CrossRef] [PubMed]
- Tsujimura, M.; Mori, N.; Yamagishi, H.; Terachi, T. A possible breakage of linkage disequilibrium between mitochondrial and chloroplast genomes during Emmer and Dinkel wheat evolution. Genome 2013, 56, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Jankowiak, K.; Rybarczyk, A.; Wyatt, R.; Odrzykoski, I.; Pacak, A.; Szweykowska-Kulinska, Z. Organellar inheritance in the allopolyploid moss Rhizomnium pseudopunctatum. Taxon 2005, 54, 383–388. [Google Scholar] [CrossRef]
- Guillon, J.M.; Raquin, C. Maternal inheritance of chloroplasts in the horsetail Equisetum variegatum (Schleich). Curr. Genet. 2000, 37, 53–56. [Google Scholar] [CrossRef] [PubMed]
- Natcheva, R.; Cronberg, N. Maternal transmission of cytoplasmic DNA in interspecific hybrids of peat moss, Sphagnum (Bryophyta). J. Evol. Biol. 2007, 20, 1613–1616. [Google Scholar] [CrossRef] [PubMed]
- Mahoney, L.L.; Quimby, M.L.; Shields, M.E.; Davis, T.M. Mitochondrial DNA transmission, ancestry, and sequences in Fragaria. Acta Hortic. 2010, 859, 301–308. [Google Scholar] [CrossRef]
- Njuguna, W.; Liston, A.; Cronn, R.; Ashman, T.L.; Bassil, N. Insights into phylogeny, sex function and age of Fragaria based on whole chloroplast genome sequencing. Mol. Phylogenet. Evol. 2013, 66, 17–29. [Google Scholar] [CrossRef] [PubMed]
- Govindarajulu, R.; Parks, M.; Tennessen, J.A.; Liston, A.; Ashman, T.L. Comparison of nuclear, plastid, and mitochondrial phylogenies and the origin of wild octaploid strawberry species. Am. J. Bot. 2015, 102, 544–554. [Google Scholar] [CrossRef] [PubMed]
| Reads Count | Polytrichum juniperinum | Polytrichum strictum | ||
|---|---|---|---|---|
| Total reads library I | 4,215,441 | 4,992,524 | ||
| Total reads library II | 8,693,811 | 8,443,552 | ||
| Total reads library III | 3,424,244 | 3,243,657 | ||
| Total reads in concatenate library | 16,333,496 | 16,679,733 | ||
| Sequence length | 25–383 | 25–389 | ||
| High quality reads | 9,164,328 | 9,563,865 | ||
| Chloroplast | Mitochondria | Chloroplast | Mitochondria | |
| Mapped reads | 103.776 (1.13%) | 32.931 (0.35%) | 58.624 (0.61%) | 29.620 (0.30%) |
| N50 | 10,083 | 2988 | 1890 | 26,564 |
| Species | CpDNA Ac. Num. | Chloroplast Genes | MtDNA Ac. Num. | Mitochondrial Genes | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| rpoA | ycf66 | petN | matK | rps15 | rps10 | nad7 | ORF187 | |||
| Chlorella sp. | (NC_001865.1) (NC_023835.1) | + | - | - | - | - | (NC_024626.1) (NC_025413.1) | + | + | - |
| Chaetosphaeridium globosum | NC_004115.1 | + | + | + | + | + | NC_004118.1 | + | + | - |
| Marchantia polymorpha | NC_001319.1 | + | + | - | + | + | NC_001660.1 | + | Ψ | - |
| Anthoceros angustus | NC_004543.1 | + | - | + | Ψ | Ψ | 0 | 0 | 0 | 0 |
| Physcomitrella patens | NC_005087.1 | - | + | + | + | + | NC_007945.1 | - | + | + |
| Polytrichum juniperinum | KY795004 | - | + | - | - | - | KY795005 | - | + | + |
| Polytrichum strictum | KY795006 | - | + | - | - | - | KY795007 | - | + | + |
| Tetraphis pellucida | NC_024291.1 | - | + | - | + | + | NC_024290.1 | - | - | - |
| Arabidopsis thaliana | NC_000932.1 | + | - | - | + | + | NC_001284.2 | - | + | + |
| Oryza sativa Indica Group | NC_027678.1 | + | - | + | + | + | NC_007886.1 | - | + | + |
| Triticum aestivum | NC_002762.1 | + | - | - | + | + | NC_007579.1 | - | + | - |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Freitas, K.E.J.; Metz, G.F.; Cañon, E.R.P.; Roesch, L.F.W.; Pereira, A.B.; Victoria, F.C. Characterization and Phylogenetic Analysis of Chloroplast and Mitochondria Genomes from the Antarctic Polytrichaceae Species Polytrichum juniperinum and Polytrichum strictum. Diversity 2018, 10, 89. https://doi.org/10.3390/d10030089
De Freitas KEJ, Metz GF, Cañon ERP, Roesch LFW, Pereira AB, Victoria FC. Characterization and Phylogenetic Analysis of Chloroplast and Mitochondria Genomes from the Antarctic Polytrichaceae Species Polytrichum juniperinum and Polytrichum strictum. Diversity. 2018; 10(3):89. https://doi.org/10.3390/d10030089
Chicago/Turabian StyleDe Freitas, Karine Elise Janner, Geferson Fernando Metz, Ehidy Rocio Peña Cañon, Luiz Fernando Wurdig Roesch, Antonio Batista Pereira, and Filipe Carvalho Victoria. 2018. "Characterization and Phylogenetic Analysis of Chloroplast and Mitochondria Genomes from the Antarctic Polytrichaceae Species Polytrichum juniperinum and Polytrichum strictum" Diversity 10, no. 3: 89. https://doi.org/10.3390/d10030089
APA StyleDe Freitas, K. E. J., Metz, G. F., Cañon, E. R. P., Roesch, L. F. W., Pereira, A. B., & Victoria, F. C. (2018). Characterization and Phylogenetic Analysis of Chloroplast and Mitochondria Genomes from the Antarctic Polytrichaceae Species Polytrichum juniperinum and Polytrichum strictum. Diversity, 10(3), 89. https://doi.org/10.3390/d10030089








