Morphological and Genetic Diversity of Sea Buckthorn (Hippophae rhamnoides L.) in the Karakoram Mountains of Northern Pakistan
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material and Sampling Area
2.2. Dendrometric Traits
2.3. Leaf and Fruit Morphometry Traits
2.4. Leaf and Fruit Colors
2.5. DNA Isolation and EST-SSR Analysis
2.6. Data Analysis
3. Results
3.1. Dendrometric Traits
3.2. Leaf and Fruit Morphometry Traits
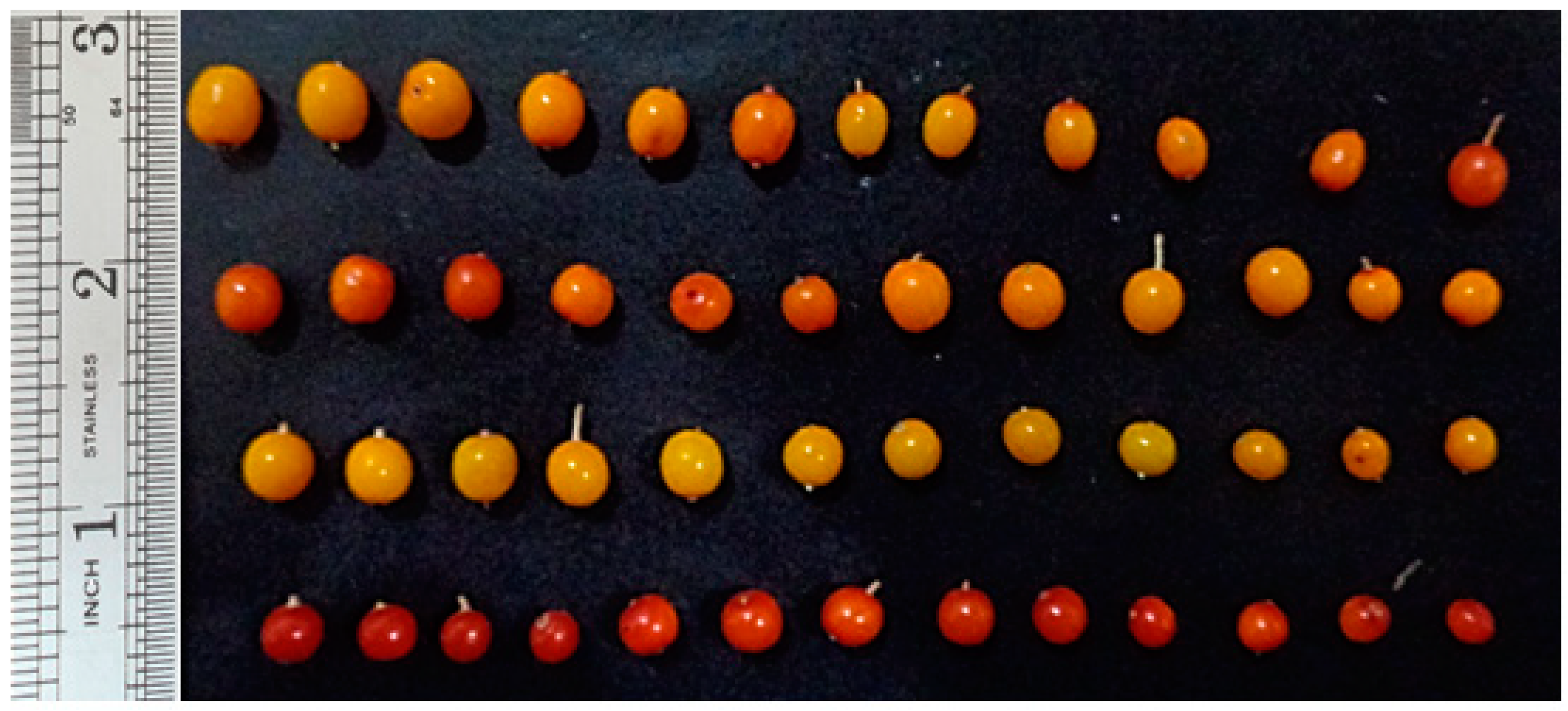
3.3. Leaf and Fruit Color
3.4. Genetic Diversity and Differentiation of Markers
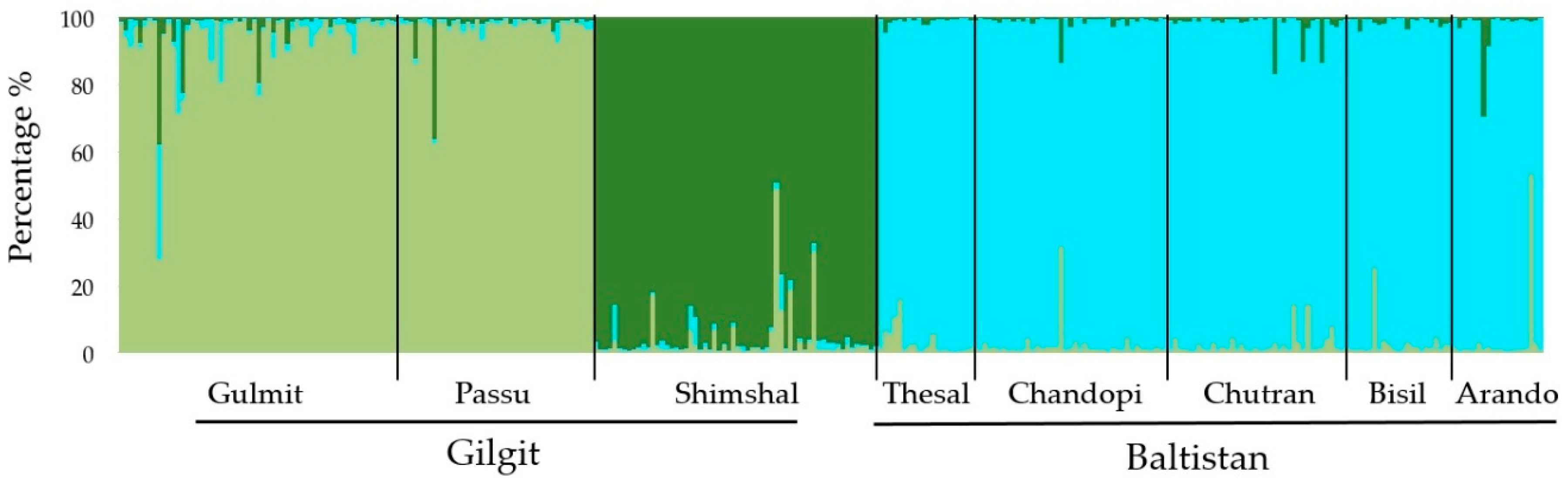
3.5. Genetic Diversity and Differentiation of Populations
4. Discussion
4.1. Morphological Variation
4.2. Genetic Diversity and Differentiation
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Region | Population | Stand | N | Na | PNa | He | Ho | FIS | FIS without HrMS004 Marker |
|---|---|---|---|---|---|---|---|---|---|
| Gilgit | Gulmit | wild | 30 | 8.2 | 3 | 0.704 | 0.683 | 0.047 | 0.035 |
| village | 30 | 8.2 | 4 | 0.699 | 0.653 | 0.083 * | 0.043 | ||
| Passu | wild | 20 | 6.5 | 2 | 0.674 | 0.667 | 0.037 | 0.010 | |
| village | 20 | 6.9 | 1 | 0.685 | 0.674 | 0.041 * | 0.002 | ||
| Shimshal | wild | 30 | 6.8 | 2 | 0.729 | 0.758 | −0.020 *** | −0.058 ** | |
| village | 30 | 6.3 | 2 | 0.686 | 0.703 | −0.01 | −0.026 | ||
| Baltistan | Thesal | wild | 10 | 5.7 | 3 | 0.643 | 0.675 | 0.002 | −0.025 |
| village | 10 | 6.2 | 5 | 0.693 | 0.758 | −0.04 | −0.056 | ||
| Chandopi | wild | 20 | 7.6 | 2 | 0.667 | 0.629 | 0.082 * | 0.064 | |
| village | 20 | 7.9 | 6 | 0.687 | 0.671 | 0.050 * | 0.008 | ||
| Chutran | wild | 20 | 8.5 | 2 | 0.699 | 0.652 | 0.092 ** | 0.061 * | |
| village | 20 | 8.4 | 7 | 0.698 | 0.650 | 0.094 *** | 0.096 ** | ||
| Bisil | wild | 10 | 6.3 | 2 | 0.696 | 0.667 | 0.094 * | 0.034 | |
| village | 10 | 6.2 | 1 | 0.682 | 0.683 | 0.05 | 0.027 | ||
| Arando | wild | 10 | 5.8 | 1 | 0.647 | 0.650 | 0.048 | 0.003 | |
| village | 10 | 4.9 | 0 | 0.672 | 0.683 | 0.035 | −0.018 | ||
| Mean | 8.5 | 3 | 0.699 | 0.683 | 0.043 | 0.012 | |||
References
- Rousi, A. The genus Hippophae L. A taxonomic study. Ann. Bot. Fenn. 1971, 8, 177–227. [Google Scholar]
- Zeb, A. Chemical and nutritional constituents of sea buckthorn juice. Pak. J. Nutr. 2004, 3, 99–106. [Google Scholar]
- Kato, K.; Kanayama, Y.; Ohkawa, W.; Kanahama, K. Nitrogen fixation in seabuckthorn (Hippophae rhamnoides L.) root nodules and effect of nitrate on nitrogenase activity. J. Jpn. Soc. Hortic. Sci. 2007, 76, 185–190. [Google Scholar] [CrossRef]
- Enescu, C.M. Sea buckthorn: A species with a variety of uses, especially in land reclamation. Dendrobiology 2014, 72, 41–46. [Google Scholar] [CrossRef]
- Ali, A.; Kaul, V.; Tamchos, S. Pollination biology in Hippophae rhamnoides L. Growing in cold desert of Ladakh Himalayas. In Sea Buckthorn: Emerging Technologies for Health Protection and Environmental Conservation; International Sea buckthorn Association: New Dehli, India, 2015. [Google Scholar]
- Mangla, Y.; Chaudhary, M.; Gupta, H.; Thakur, R.; Goel, S.; Raina, S.; Tandon, R. Facultative apomixis and development of fruit in a deciduous shrub with medicinal and nutritional uses. AoB Plants 2015, 7. [Google Scholar] [CrossRef] [PubMed]
- Mangla, Y.; Tandon, R. Pollination ecology of Himalayan sea buckthorn, Hippophae rhamnoides L. (Elaeagnaceae). Curr. Sci. 2014, 106, 1731–1735. [Google Scholar]
- Shah, A.H.; Ahmed, D.; Sabir, M.; Arif, S.; Khaliq, I.; Batool, F. Biochemical and nutritional evaluations of sea buckthorn (Hippophae rhamnoides L. spp. turkestanica) from different locations of Pakistan. Pak. J. Bot. 2007, 39, 2059–2065. [Google Scholar]
- Yang, W.; Laaksonen, O.; Kallio, H.; Yang, B. Proanthocyanidins in sea buckthorn (Hippophaë rhamnoides L.) berries of different origins with special reference to the influence of genetic background and growth location. J. Agric. Food Chem. 2016, 64, 1274–1282. [Google Scholar] [CrossRef] [PubMed]
- Durst, P.B.; Ulrich, W.; Kashio, M. Non-Wood Forest Products in Asia; Rapa Publication (FAO): Rome, Italy, 1994; Volume 28. [Google Scholar]
- Lecoent, A.; Vandecandelaere, E.; Cadilhon, J.-J. Quality linked to the geographical origin and geographical indications: Lessons learned from six case studies in Asia. RAP Publ. 2010, 4, 85–112. [Google Scholar]
- Rongsen, L. Seabuckthorn: A Multipurpose Plant Species for Fragile Mountains, 20th ed.; International Centre for Integrated Mountain Development: Kathmandu, Nepal, 1992; pp. 18–20. [Google Scholar]
- Stobdan, T.; Angchuk, D.; Singh, S.B. Seabuckthorn: An emerging storehouse for researchers in India. Curr. Sci. 2008, 94, 1236–1237. [Google Scholar]
- Yadav, V.; Sharma, S.; Rao, V.; Yadav, R.; Radhakrishna, A. Assessment of morphological and biochemical diversity in sea buckthorn (Hippophae salicifolia D. Don.) populations of Indian Central Himalaya. Proc. Natl. Acad. Sci. India Sect. B Biol. Sci. 2016, 86, 351–357. [Google Scholar] [CrossRef]
- Waehling, A. In Assessment report on the sea buckthorn market in Europe, Russia, NIS-Countries and China Results of a market investigation in 2005. In Proceedings of the 3rd International Sea buckthorn Association Conference, Quebec City, QC, Canada, 12–16 August 2007. [Google Scholar]
- Fan, J.; Ding, X.; Gu, W. Radical-scavenging proanthocyanidins from sea buckthorn seed. Food Chem. 2007, 102, 168–177. [Google Scholar] [CrossRef]
- Hussain, A.; Abid, H.; Ali, S. Physicochemical characteristics and fatty acid composition of sea buckthorn (Hippophae rhamnoides L.) oil. J. Chem. Soc. Pak. 2007, 29, 256. [Google Scholar]
- Li, C.; Xu, G.; Zang, R.; Korpelainen, H.; Berninger, F. Sex-related differences in leaf morphological and physiological responses in Hippophae rhamnoides along an altitudinal gradient. Tree Physiol. 2007, 27, 399–406. [Google Scholar] [CrossRef] [PubMed]
- Ansari, A.S. In Sea Buckthorn (Hippophae L. spp.)—A Potential Resource for Biodiversity Conservation in Nepal Himalayas. In Proceedings of the International Workshop on Underutilized Plant Species, Leipzig, Germany, 6–8 May 2003. [Google Scholar]
- Saggu, S.; Divekar, H.; Gupta, V.; Sawhney, R.; Banerjee, P.; Kumar, R. Adaptogenic and safety evaluation of seabuckthorn (Hippophae rhamnoides) leaf extract: A dose dependent study. Food Chem. Toxicol. 2007, 45, 609–617. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, N.K.; Kumar, R.; Siddiqui, M.; Gupta, A. Mechanism of wound-healing activity of Hippophae rhamnoides L. leaf extract in experimental burns. Evid. Based Complement. Altern. Med. 2009. [Google Scholar] [CrossRef]
- Yadav, V.; Sah, V.; Singh, A.; Sharma, S. Variations in morphological and biochemical characters of seabuckthorn (Hippophae salicifolia D. Don.) populations growing in Harsil area of Garhwal Himalaya in India. Trop. Agric. Res. Ext. 2006, 9, 1–7. [Google Scholar]
- Yang, M.; Xiong, Y.; Wang, H.; Wu, X.; Xue, Z.; Mu, S.; Liu, Z.; Ge, J. Genetic diversity of Hippophae rhamnoides L. sub-populations on loess plateau under effects of varying meteorologic conditions. J. Appl. Ecol. 2008, 19, 1–7. [Google Scholar]
- Yang, B.; Kallio, H.P. Fatty acid composition of lipids in sea buckthorn (Hippophaë rhamnoides L.) berries of different origins. J. Agric. Food Chem. 2001, 49, 1939–1947. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Wang, Y.; Zhao, C.; Korpelainen, H.; Li, C. Genetic diversity of Hippophae rhamnoides populations at varying altitudes in the Wolong natural reserve of China as revealed by ISSR markers. Silvae Genet. 2008, 57, 29–36. [Google Scholar] [CrossRef]
- Yang, Y.; Yao, Y.; Xu, G.; Li, C. Growth and physiological responses to drought and elevated ultraviolet B in two contrasting populations of Hippophae rhamnoides. Physiol. Plant. 2005, 124, 431–440. [Google Scholar] [CrossRef]
- Guo, W.; Li, B.; Zhang, X.; Wang, R. Architectural plasticity and growth responses of Hippophae rhamnoides and Caragana intermedia seedlings to simulated water stress. J. Arid Environ. 2007, 69, 385–399. [Google Scholar] [CrossRef]
- Sharma, V.; Dwivedi, S.; Awasthi, O.; Verma, M. Variation in nutrient composition of seabuckthorn (Hippophae rhamnoides L.) leaves collected from different locations of Ladakh. Indian J. Hortic. 2014, 71, 421–423. [Google Scholar]
- Jia, D.R.; Abbott, R.J.; Liu, T.L.; Mao, K.S.; Bartish, I.V.; Liu, J.Q. Out of the Qinghai–Tibet Plateau: Evidence for the origin and dispersal of Eurasian temperate plants from a phylogeographic study of Hippophaë rhamnoides (Elaeagnaceae). New Phytol. 2012, 194, 1123–1133. [Google Scholar] [CrossRef] [PubMed]
- Qiong, L.; Zhang, W.; Wang, H.; Zeng, L.; Birks, H.J.B.; Zhong, Y. Testing the effect of the Himalayan Mountains as a physical barrier to gene flow in Hippophae tibetana Schlect. (Elaeagnaceae). PLoS ONE 2017, 12, e0172948. [Google Scholar] [CrossRef] [PubMed]
- Bauert, M.; Kälin, M.; Baltisberger, M.; Edwards, P. No genetic variation detected within isolated relict populations of Saxifraga cernua in the Alps using RAPD markers. Mol. Ecol. 1998, 7, 1519–1527. [Google Scholar] [CrossRef]
- Su, H.; Qu, L.; He, K.; Zhang, Z. The Great Wall of China: A physical barrier to gene flow? Heredity 2003, 90, 212. [Google Scholar] [CrossRef] [PubMed]
- Miller, A.J.; Schaal, B.A. Domestication and the distribution of genetic variation in wild and cultivated populations of the Mesoamerican fruit tree Spondias purpurea L. (Anacardiaceae). Mol. Ecol. 2006, 15, 1467–1480. [Google Scholar] [CrossRef] [PubMed]
- Dawson, I.K.; Hollingsworth, P.M.; Doyle, J.J.; Kresovich, S.; Weber, J.C.; Montes, C.S.; Pennington, T.D.; Pennington, R.T. Origins and genetic conservation of tropical trees in agroforestry systems: A case study from the Peruvian Amazon. Conserv. Genet. 2008, 9, 361–372. [Google Scholar] [CrossRef]
- Ekué, M.R.; Gailing, O.; Vornam, B.; Finkeldey, R. Assessment of the domestication state of ackee (Blighia sapida KD Koenig) in Benin based on AFLP and microsatellite markers. Conserv. Genet. 2011, 12, 475–489. [Google Scholar] [CrossRef]
- Bartish, I.; Jeppsson, N.; Nybom, H. Population genetic structure in the dioecious pioneer plant species Hippophae rhamnoides investigated by random amplified polymorphic DNA (RAPD) markers. Mol. Ecol. 1999, 8, 791–802. [Google Scholar] [CrossRef]
- Bartish, I.; Jeppsson, N.; Bartish, G.; Lu, R.; Nybom, H. Inter and intraspecific genetic variation in Hippophae (Elaeagnaceae) investigated by RAPD markers. Plant Syst. Evol. 2000, 225, 85–101. [Google Scholar] [CrossRef]
- Ruan, C.; Qin, P.; Zheng, J.; He, Z. Genetic relationships among some cultivars of sea buckthorn from China, Russia and Mongolia based on RAPD analysis. Sci. Hortic. 2004, 101, 417–426. [Google Scholar] [CrossRef]
- Sheng, H.; An, L.; Chen, T.; Xu, S.; Liu, G.; Zheng, X.; Pu, L.; Liu, Y.; Lian, Y. Analysis of the genetic diversity and relationships among and within species of Hippophae (Elaeagnaceae) based on RAPD markers. Plant Syst. Evol. 2006, 260, 25–37. [Google Scholar] [CrossRef]
- Singh, R.; Mishra, S.; Dwivedi, S.; Ahmed, Z. Genetic variation in seabuckthorn (Hippophae rhamnoides L.) populations of cold arid Ladakh (India) using RAPD markers. Curr. Sci. 2006, 91, 1321–1322. [Google Scholar]
- Ercişli, S.; Orhan, E.; Yildirim, N.; Ağar, G. Comparison of sea buckthorn genotypes (Hippophae rhamnoides L.) based on RAPD and FAME data. Turk. J. Agric. For. 2008, 32, 363–368. [Google Scholar]
- Ruan, C. Genetic relationships among sea buckthorn varieties from China, Russia and Mongolia using AFLP markers. J. Hortic. Sci. Biotechnol. 2006, 81, 409–414. [Google Scholar] [CrossRef]
- Shah, A.H.; Ahmad, S.D.; Khaliq, I.; Batool, F.; Hassan, L.; Pearce, R. Evaluation of phylogenetic relationship among sea buckthorn (Hippophae rhamnoides L. spp. turkestanica) wild ecotypes from Pakistan using amplified fragment length polymorphism (AFLP). Pak. J. Bot. 2009, 41, 2419–2426. [Google Scholar]
- Wang, A.; Zhang, Q.; Wan, D.; Liu, J. Nine microsatellite DNA primers for Hippophae rhamnoides ssp. sinensis (Elaeagnaceae). Conserv. Genet. 2008, 9, 969–971. [Google Scholar] [CrossRef]
- Li, H.; Ruan, C.-J.; da Silva, J.A.T. Identification and genetic relationship based on ISSR analysis in a germplasm collection of sea buckthorn (Hippophae L.) from China and other countries. Sci. Hortic. 2009, 123, 263–271. [Google Scholar] [CrossRef]
- Jain, A.; Ghangal, R.; Grover, A.; Raghuvanshi, S.; Sharma, P.C. Development of EST-based new SSR markers in seabuckthorn. Physiol. Mol. Biol. Plants 2010, 16, 375–378. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jain, A.; Chaudhary, S.; Sharma, P.C. Mining of microsatellites using next generation sequencing of seabuckthorn (Hippophae rhamnoides L.) transcriptome. Physiol. Mol. Biol. Plants 2014, 20, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Muchugi, A.; Kadu, C.; Kindt, R.; Kipruto, H.; Lemurt, S.; Olale, K.; Nyadoi, P.; Dawson, I.; Jamnadas, R. Molecular Markers for Tropical Trees: A Practical Guide to Principles and Procedures. ICRAF Technical Manual, 9th ed.; Dawson, I., Jamnadass, R., Eds.; World Agroforestry Centre: Nairobi, Kenya, 2008; ISBN 9290592257. [Google Scholar]
- Arzai, A.; Aliyu, B. The relationship between canopy width, height and trunk size in some tree species growing in the Savana zone of Nigeria. Bayero J. Pure Appl. Sci. 2010, 3, 260–263. [Google Scholar] [CrossRef]
- Hothorn, T.; Bretz, F.; Westfall, P. Simultaneous inference in general parametric models. Biom. J. 2008, 50, 346–363. [Google Scholar] [CrossRef] [PubMed]
- Herberich, E.; Sikorski, J.; Hothorn, T. A robust procedure for comparing multiple means under heteroscedasticity in unbalanced designs. PLoS ONE 2010, 5, e9788. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mangiafico, S. rcompanion: Functions to Support Extension Education Program Evaluation. In R Package Version 1.5. 0; The Comprehensive R Archive Network; 2017. [Google Scholar]
- Van Oosterhout, C.; Hutchinson, W.F.; Wills, D.P.; Shipley, P. Microchecker: Software for identifying and correcting genotyping errors in microsatellite data. Mol. Ecol. Notes 2004, 4, 535–538. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed]
- Kalinowski, S.T. HP-Rare 1.0: A computer program for performing rarefaction on measures of allelic richness. Mol. Ecol. Notes 2005, 5, 187–189. [Google Scholar] [CrossRef]
- Hedrick, P.W. A standardized genetic differentiation measure. Evol. Int. J. Org. Evol. 2005, 59, 1633–1638. [Google Scholar] [CrossRef]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [PubMed]
- Earl, D.A.; Holdt, V. Structure Harvester: A website and program for visualizing structure output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Du Toit, B.; Dovey, S.B. Effect of site management on leaf area, early biomass development, and stand growth efficiency of a Eucalyptus grandis plantation in South Africa. Can. J. For. Res. 2005, 35, 891–900. [Google Scholar] [CrossRef]
- Li, X.; Li, Y.; Zhang, Z.; Li, X. Influences of environmental factors on leaf morphology of Chinese jujubes. PLoS ONE 2015, 10, e0127825. [Google Scholar] [CrossRef] [PubMed]
- Pan, S.; Liu, C.; Zhang, W.; Xu, S.; Wang, N.; Li, Y.; Gao, J.; Wang, Y.; Wang, G. The scaling relationships between leaf mass and leaf area of vascular plant species change with altitude. PLoS ONE 2013, 8, e76872. [Google Scholar] [CrossRef] [PubMed]
- Shelley, C.E.; Gehring, N.R. Elevation’s Effect on Malosma Laurinais Leaf Size. Bachelor’s Thesis, Pepperdine University, Malibu, CA, USA, 2014. [Google Scholar]
- Telias, A.; Hoover, E.; Rother, D. Plant and environmental factors influencing the pattern of pigment accumulation in ‘Honeycrisp’ apple peels using a novel color analyzer software tool. HortScience 2008, 43, 1441–1445. [Google Scholar]
- McCallum, S.; Woodhead, M.; Hackett, C.A.; Kassim, A.; Paterson, A.; Graham, J. Genetic and environmental effects influencing fruit colour and QTL analysis in raspberry. Theor. Appl. Genet. 2010, 121, 611–627. [Google Scholar] [CrossRef] [PubMed]
- Paul, V.; Pandey, R. Role of internal atmosphere on fruit ripening and storability—A review. J. Food Sci. Technol. 2014, 51, 1223–1250. [Google Scholar] [CrossRef] [PubMed]
- Tanksley, S.D. The genetic, developmental, and molecular bases of fruit size and shape variation in tomato. Plant Cell 2004, 16, S181–S189. [Google Scholar] [CrossRef] [PubMed]
- Naegele, R.P.; Mitchell, J.; Hausbeck, M.K. Genetic diversity, population structure, and heritability of fruit traits in Capsicum annuum. PLoS ONE 2016, 11, e0156969. [Google Scholar] [CrossRef] [PubMed]
- Sabir, S.; Ahmed, S.; Lodhi, N.; Jäger, A. Morphological and biochemical variation in sea buckthorn Hippophae rhamnoides ssp. turkestanica, a multipurpose plant for fragile mountains of Pakistan. S. Afr. J. Bot. 2003, 69, 587–592. [Google Scholar] [CrossRef]
- Yao, Y.; Tigerstedt, P.M. Geographical variation of growth rhythm, height, and hardiness, and their relations in Hippophae rhamnoides. J. Am. Soc. Hortic. Sci. 1995, 120, 691–698. [Google Scholar]
- Ahmad, S.; Kamal, M. Morpo-molecular characterization of local genotypes Hippophae rhamnoides spp. turkestanica a multipurpose plant from northern areas of Pakistan. J. Biol. Sci. 2002, 2, 351–354. [Google Scholar]
- Ahmad, S.; Jasra, A.; Imtiaz, A. Genetic diversity in Pakistani genotypes of Hippophae rhamnoides L. spp. turkestanica. Int. J. Agric. Biol. 2003, 5, 10–13. [Google Scholar]
- Turchetto, C.; Segatto, A.L.A.; Beduschi, J.; Bonatto, S.L.; Freitas, L.B. Genetic differentiation and hybrid identification using microsatellite markers in closely related wild species. AoB Plants 2015. [Google Scholar] [CrossRef] [PubMed]
- Dillon, N.L.; Innes, D.J.; Bally, I.S.; Wright, C.L.; Devitt, L.C.; Dietzgen, R.G. Expressed sequence tag-simple sequence repeat (EST-SSR) marker resources for diversity analysis of mango (Mangifera indica L.). Diversity 2014, 6, 72–87. [Google Scholar] [CrossRef]
- Hamrick, J.L.; Godt, M.J.W.; Sherman-Broyles, S.L. Factors influencing levels of genetic diversity in woody plant species. New For. 1992, 6, 95–124. [Google Scholar] [CrossRef]
- Breton, C.; Pinatel, C.; Medail, F.; Bonhomme, F.; Berville, A. Comparison between classical and Bayesian methods to investigate the history of olive cultivars using SSR-polymorphisms. Plant Sci. 2008, 175, 524–532. [Google Scholar] [CrossRef]
- Wang, Y.; Jiang, H.; Peng, S.; Korpelainen, H. Genetic structure in fragmented populations of Hippophae rhamnoides ssp. sinensis in China investigated by ISSR and cpSSR markers. Plant Syst. Evol. 2011, 295, 97–107. [Google Scholar] [CrossRef]
- Tian, C.; Nan, P.; Shi, S.; Chen, J.; Zhong, Y. Molecular genetic variation in Chinese populations of three subspecies of Hippophae rhamnoides. Biochem. Genet. 2004, 42, 259–267. [Google Scholar] [CrossRef] [PubMed]
- Ohsawa, T.; Ide, Y. Global patterns of genetic variation in plant species along vertical and horizontal gradients on mountains. Glob. Ecol. Biogeogr. 2008, 17, 152–163. [Google Scholar] [CrossRef] [Green Version]
- Ryman, N.; Palm, S.; André, C.; Carvalho, G.R.; Dahlgren, T.G.; Jorde, P.E.; Laikre, L.; Larsson, L.C.; PalmÉ, A.; Ruzzante, D.E. Power for detecting genetic divergence: Differences between statistical methods and marker loci. Mol. Ecol. 2006, 15, 2031–2045. [Google Scholar] [CrossRef] [PubMed]
- Morin, P.A.; Leduc, R.G.; Archer, F.I.; Martien, K.K.; Huebinger, R.; Bickham, J.W.; Taylor, B.L. Significant deviations from Hardy-Weinberg equilibrium caused by low levels of microsatellite genotyping errors. Mol. Ecol. Res. 2009, 9, 498–504. [Google Scholar] [CrossRef] [PubMed]
- Boccacci, P.; Aramini, M.; Valentini, N.; Bacchetta, L.; Rovira, M.; Drogoudi, P.; Silva, A.; Solar, A.; Calizzano, F.; Erdoğan, V. Molecular and morphological diversity of on-farm hazelnut (Corylus avellana L.) landraces from southern Europe and their role in the origin and diffusion of cultivated germplasm. Tree Genet. Genomes 2013, 9, 1465–1480. [Google Scholar] [CrossRef]
- Gao, L.; Möller, M.; Zhang, X.M.; Hollingsworth, M.; Liu, J.; Mill, R.; Gibby, M.; Li, D.Z. High variation and strong phylogeographic pattern among cpDNA haplotypes in Taxus wallichiana (Taxaceae) in China and North Vietnam. Mol. Ecol. 2007, 16, 4684–4698. [Google Scholar] [CrossRef] [PubMed]
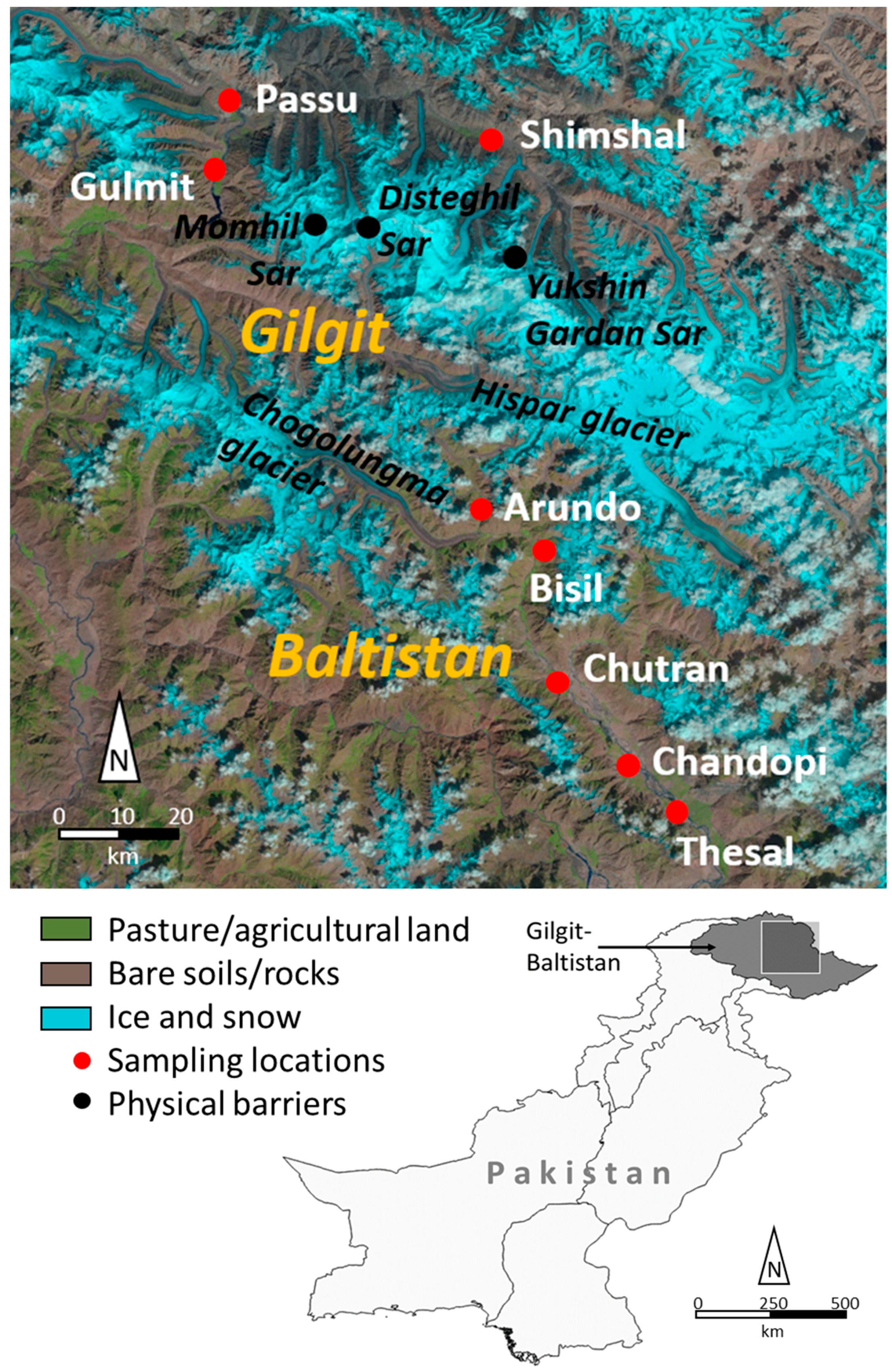

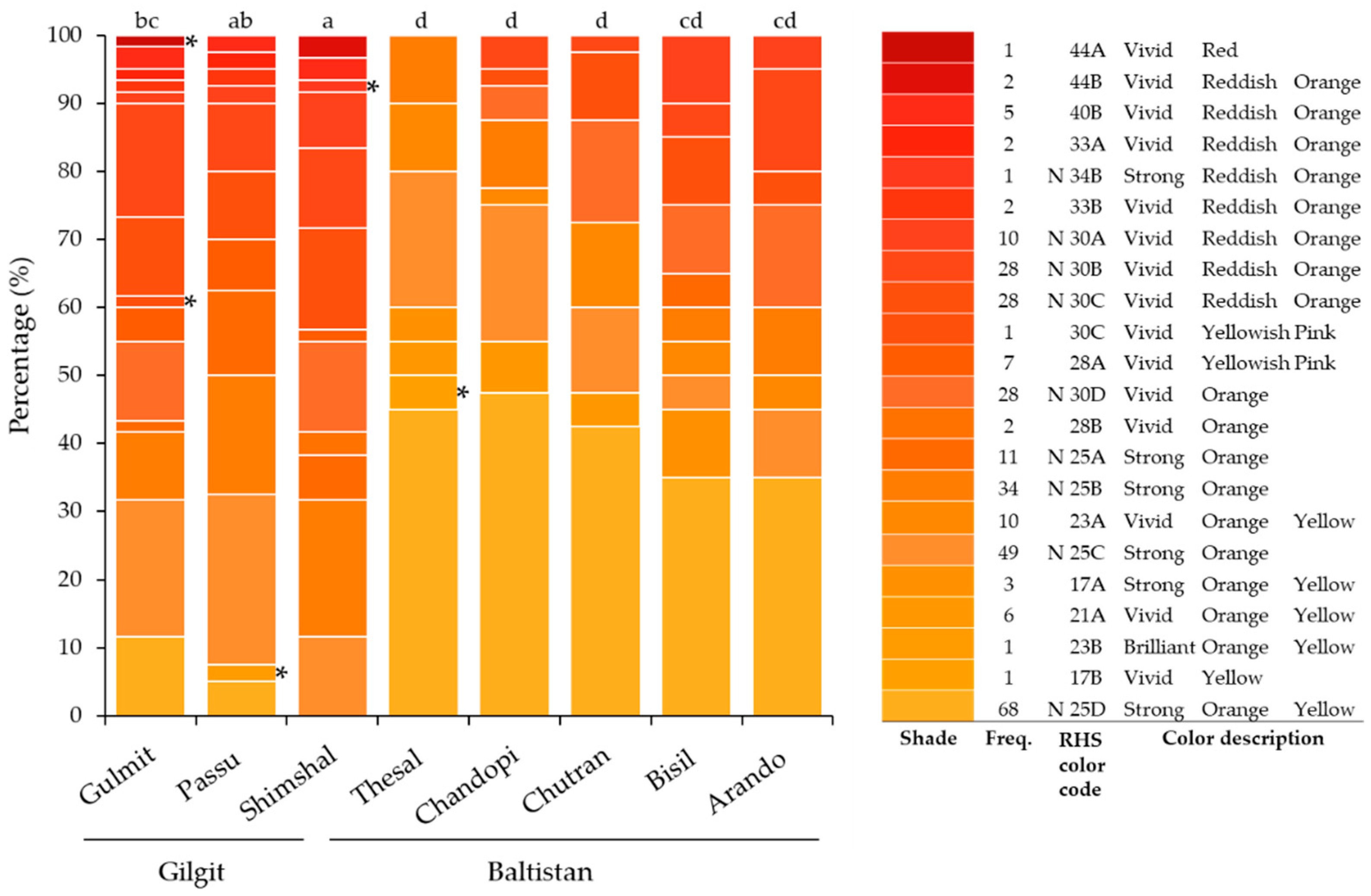
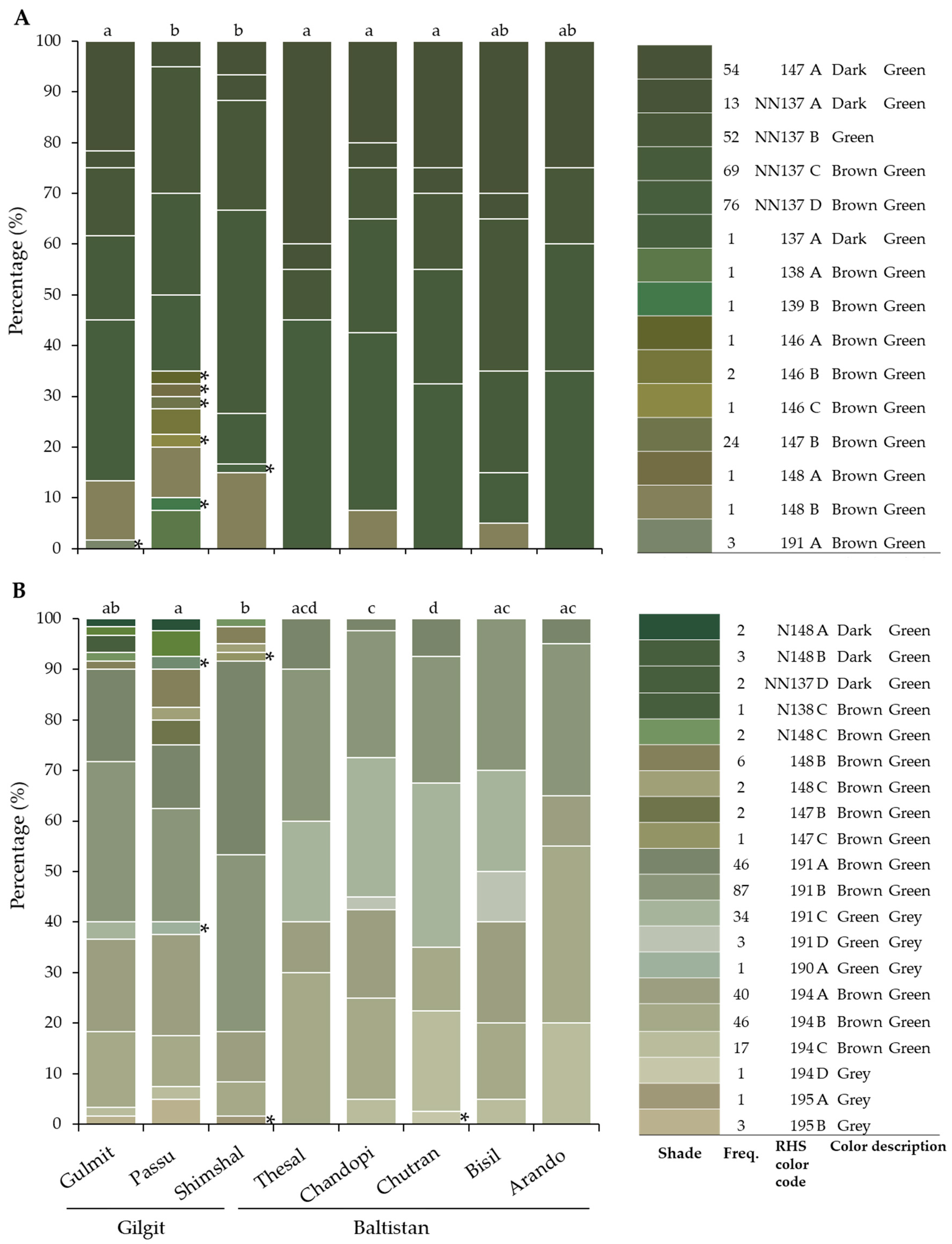
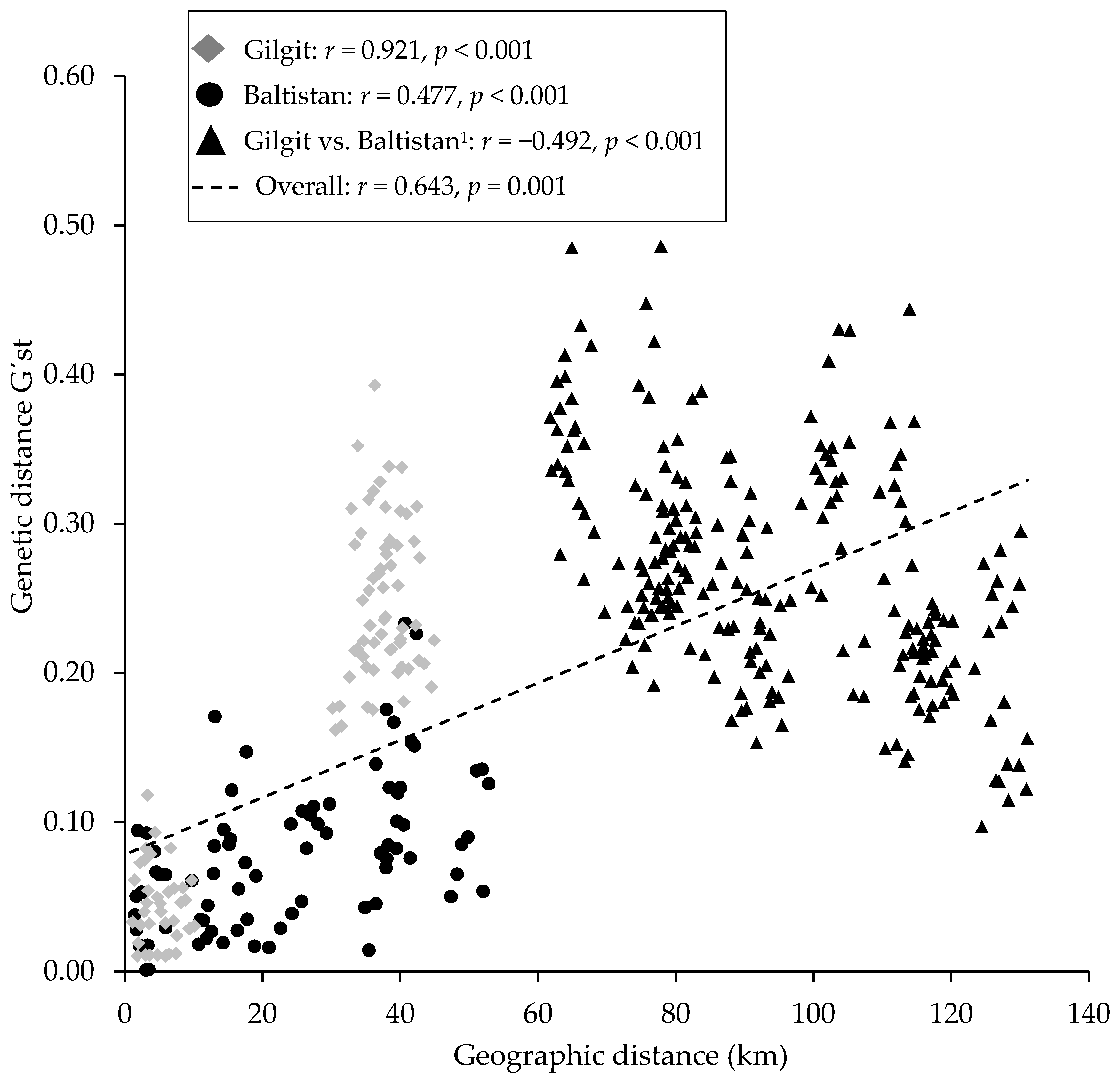
| Region | Population | No. of Individuals | Longitude E | Latitude N | Elevation, m.a.s.l. |
|---|---|---|---|---|---|
| Gilgit | Gulmit | 60 | 36.4049805 | 74.8734 | 2525 |
| Passu | 40 | 36.4558832 | 74.8997 | 2516 | |
| Shimshal | 60 | 36.4412537 | 75.3008 | 3112 | |
| Baltistan | Thesal | 20 | 35.4697578 | 75.6654 | 2275 |
| Chandopi | 40 | 35.5428859 | 75.5711 | 2330 | |
| Chutran | 40 | 35.7268528 | 75.4046 | 2435 | |
| Bisil | 20 | 35.8568179 | 75.4156 | 2655 | |
| Arando | 20 | 35.8695884 | 75.3677 | 2700 | |
| Total | 300 |
| Grouping | Population/Stand | Dendrometry | Leaf | Fruit | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Height, cm | Stem Diameter, cm | Canopy, m2 | Area, mm2 | Length to Width Ratio | Volume, mm3 | Length to Width Ratio | 20-berry wt., g | Moisture, % | ||||||||||||
| Populations | Gulmit | 146 | b | 5.1 | ab | 2.4 | ab | 85 | a | 7.5 | b | 75 | a | 1.1 | b | 1.6 | a | 66 | a | |
| Passu | 138 | ab | 4.4 | a | 1.5 | a | 93 | a | 7.2 | ab | 85 | a | 1.1 | b | 1.8 | a | 66 | ab | ||
| Shimshal | 174 | c | 4.9 | b | 2.1 | ab | 192 | e | 6.5 | a | 150 | c | 1.1 | a | 3.1 | b | 77 | c | ||
| Thesal | 193 | ac | 4.5 | ab | 3.1 | bc | 98 | ab | 7.9 | bc | 123 | bc | 1.2 | b | 2.5 | b | 67 | ab | ||
| Chandopi | 198 | bc | 6.0 | ab | 3.5 | c | 131 | bc | 7.8 | bc | 135 | bc | 1.2 | b | 2.9 | b | 67 | ab | ||
| Chutran | 156 | b | 4.6 | ab | 2.6 | bc | 129 | bc | 7.9 | bc | 120 | b | 1.2 | b | 2.6 | b | 69 | ab | ||
| Bisil | 189 | bc | 4.1 | ab | 2.5 | ac | 186 | de | 8.8 | c | 129 | bc | 1.1 | ab | 2.9 | b | 70 | ab | ||
| Arando | 151 | bc | 3.7 | ab | 2.6 | ac | 155 | cd | 8.6 | c | 141 | bc | 1.2 | b | 2.9 | b | 71 | b | ||
| p-value | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | |||||||||||
| Stands | Wild | 160 | 4.8 | 2.5 | 126 | a | 7.7 | 117 | 1.1 | 2.5 | 70 | |||||||||
| Village | 177 | 5.7 | 2.5 | 141 | b | 7.8 | 122 | 1.1 | 2.6 | 69 | ||||||||||
| p-value | 0.050 | 0.140 | 0.950 | 0.009 | 0.527 | 0.257 | 0.817 | 0.780 | 0.130 | |||||||||||
| Populations*stands | Gulmit | Wild | 162 | ab | 6.2 | ab | 2.5 | ac | 84 | a | 7.3 | ace | 75 | a | 1.1 | ac | 1.7 | a | 68 | ac |
| Village | 131 | a | 3.9 | a | 2.3 | ac | 85 | a | 7.6 | bcd | 74 | a | 1.1 | ac | 1.5 | a | 64 | a | ||
| Passu | Wild | 125 | a | 3.6 | a | 1.3 | a | 80 | a | 6.8 | ac | 91 | ab | 1.1 | ac | 1.9 | ac | 67 | ac | |
| Village | 151 | ad | 5.2 | ab | 1.6 | ab | 106 | ab | 7.6 | ad | 80 | a | 1.2 | ac | 1.7 | ab | 66 | ab | ||
| Shimshal | Wild | 128 | a | 3.6 | a | 1.6 | ab | 171 | ef | 6.4 | a | 153 | e | 1.2 | ab | 3.2 | e | 82 | d | |
| Village | 221 | b | 6.2 | b | 2.5 | ac | 213 | g | 6.6 | ab | 148 | ce | 1.1 | a | 2.9 | de | 72 | c | ||
| Thesal | Wild | 155 | ab | 4.3 | a | 2.5 | ac | 92 | ab | 7.7 | ad | 104 | acd | 1.2 | ac | 2.3 | ade | 66 | ac | |
| Village | 231 | bd | 4.7 | a | 3.5 | bc | 105 | ac | 8.1 | ad | 141 | de | 1.2 | ac | 2.7 | bce | 68 | ac | ||
| Chandopi | Wild | 210 | bcd | 7.5 | ab | 4.2 | c | 143 | bcdf | 8.3 | cd | 147 | de | 1.1 | ac | 3.2 | e | 70 | bc | |
| Village | 185 | ab | 4.5 | a | 2.9 | ac | 120 | abd | 7.3 | ad | 124 | bde | 1.2 | c | 2.6 | ce | 64 | ab | ||
| Chutran | Wild | 174 | ab | 6.1 | ab | 2.9 | ac | 124 | abd | 7.6 | ad | 109 | ad | 1.1 | ac | 2.3 | acd | 65 | ab | |
| Village | 138 | ac | 3.1 | a | 2.3 | ab | 134 | bce | 8.1 | cd | 131 | de | 1.1 | ac | 2.8 | de | 73 | c | ||
| Bisil | Wild | 187 | ab | 4.1 | a | 2.5 | ac | 173 | deg | 8.6 | cd | 117 | ade | 1.1 | ac | 2.7 | bce | 69 | ac | |
| Village | 192 | ab | 4.0 | a | 2.5 | ac | 199 | fg | 9.1 | d | 141 | de | 1.1 | ac | 3.1 | de | 71 | ac | ||
| Arando | Wild | 137 | ab | 3.4 | a | 2.5 | ac | 145 | bcdf | 8.9 | de | 142 | de | 1.2 | bc | 3.1 | ce | 71 | ac | |
| Village | 165 | ab | 3.9 | a | 2.6 | ac | 166 | cdeg | 8.2 | bcd | 139 | de | 1.1 | ac | 2.8 | bce | 71 | ac | ||
| p-value | <0.001 | <0.001 | 0.030 | 0.028 | 0.095 | 0.021 | 0.173 | 0.042 | <0.001 | |||||||||||
| Grand mean | 165 | 5.3 | 2.5 | 132 | 7.5 | 117 | 1.1 | 2.5 | 69 | |||||||||||
| CV, % | 45 | 62 | 70 | 46 | 20 | 39 | 12 | 41 | 11 | |||||||||||
| Parameter | Dorsal Leaf Color | Ventral Leaf Color | Fruit Color |
|---|---|---|---|
| No. of shades | 20 | 15 | 22 |
| Df | 7 | 7 | 7 |
| χ2-value | 236 | 142 | 241 |
| p-value | <0.001 | 0.002 | <0.001 |
| EST-SSR | Primer Sequence (5′-3′) | Repeat Motif and Its Number in a Reference Sequence | Size Range, bp | Na | He | Ho | FIs |
|---|---|---|---|---|---|---|---|
| USMM1 | GGCGAAACTTGACTTGTTGC | (TAC)16 | 184–237 | 14 | 0.843 | 0.853 | 0.001 |
| ACCGATCAATACCGTTCTGC | |||||||
| USMM3 | AAGGATGTGGTCGATCCAAG | (TTC)10 | 155–169 | 7 | 0.704 | 0.729 | −0.036 |
| GTTTGCAGGCATTCCTTTGT | |||||||
| USMM7 | TCGCCGTCTGTTTCAGATAA | (AG)18 | 170–191 | 9 | 0.786 | 0.867 | −0.068 |
| GCTGATCCAACGGTCTCATT | |||||||
| USMM16 | CCAACCAACCTCATCGTTTC | (TC)14 | 157–169 | 7 | 0.744 | 0.769 | −0.004 |
| ATCGGAGGGATCGTTGAAAG | |||||||
| USMM24 | TAGCATTGCAGGCTCAGAGA | (AG)11 | 229–240 | 7 | 0.740 | 0.809 | −0.071 |
| ATCCGTGGTTAAGGTTGCAC | |||||||
| USMM25 | CGAGGTCCGAGTAGGAAGA | (AAG)10 | 214–237 | 9 | 0.676 | 0.653 | 0.036 * |
| CATTGGCCTTCAATCTCCTC | |||||||
| HrMS003 | GCTCTCATCCGATTTGATCC | (TCA)6 | 335–470 | 16 | 0.828 | 0.784 | 0.056 * |
| GTCGCAGTCTTCTTGGGTTC | |||||||
| HrMS004 | GTTTGAGGTCGGTGCTGAGT | (TC)8 | 260–304 | 9 | 0.739 | 0.492 | 0.331 *** |
| GGGTAACCCAACTCCTCCTT | |||||||
| HrMS010 | GGAAGCGTGAGGAAATGTC | (TGG)5 | 213–216 | 2 | 0.334 | 0.340 | −0.015 |
| GAACAGACAGACCATTAGAGAAC | |||||||
| HrMS012 | CTCCATCTCAATCATCACTGC | (CTT)11 | 133–169 | 7 | 0.722 | 0.721 | 0.034 * |
| TTAGGGATCCGGATGAAG | |||||||
| HrMS014 | ATACCTAGCTCGGCAACAAG | (TG)6(TA)8 | 227–239 | 4 | 0.444 | 0.454 | −0.011 |
| ACGACCCATGGCATAATAGTAC | |||||||
| HrMS018 | CAACATTGTTTCGTGCAG | (ATG)5 | 189–225 | 13 | 0.823 | 0.678 | 0.207 *** |
| ACTCACATAATCGATCTCAGC | |||||||
| Mean | 8 | 0.699 | 0.681 | 0.038 |
| Population | USMM1 | USMM3 | USMM7 | USMM16 | USMM24 | USMM25 | HrMS003 | HrMS004 | HrMS010 | HrMS012 | HrMS014 | HrMS018 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Df | χ2 | Df | χ2 | Df | χ2 | Df | χ2 | Df | χ2 | Df | χ2 | Df | χ2 | Df | χ2 | Df | χ2 | Df | χ2 | Df | χ2 | Df | χ2 | |
| Gulmit | 78 | 88 | 15 | 13 | 55 | 63 | 36 | 56 * | 15 | 14 | 45 | 48 | 136 | 280 *** | 45 | 174 *** | 1 | 1 | 36 | 30 | 15 | 12 | 55 | 203 *** |
| Passu | 55 | 68 | 15 | 17 | 55 | 43 | 28 | 19 | 15 | 7 | 36 | 25 | 66 | 46 | 45 | 142 *** | 1 | 0 | 15 | 14 | 6 | 1 | 45 | 60 |
| Shimshal | 91 | 82 | 21 | 26 | 45 | 51 | 15 | 14 | 21 | 24 | 36 | 37 | 15 | 10 | 36 | 96 *** | 1 | 0 | 28 | 34 | 10 | 16 | 45 | 44 |
| Thesal | 78 | 82 | 28 | 10 | 28 | 15 | 10 | 6 | 21 | 23 | 21 | 26 | 105 | 121 | 15 | 14 | 1 | 1 | 28 | 18 | 3 | 1 | 66 | 85 |
| Chandopi | 91 | 66 | 28 | 27 | 36 | 19 | 15 | 8 | 21 | 14 | 55 | 178 *** | 253 | 286 | 36 | 127 *** | 1 | 0 | 21 | 14 | 6 | 8 | 120 | 143 |
| Chutran | 171 | 156 | 21 | 18 | 66 | 47 | 21 | 11 | 21 | 29 | 55 | 59 | 325 | 295 | 36 | 77 *** | 1 | 1 | 36 | 54 * | 6 | 22 ** | 136 | 173 * |
| Bisil | 78 | 94 | 21 | 27 | 21 | 20 | 15 | 13 | 21 | 13 | 15 | 10 | 78 | 82 | 36 | 86 *** | 1 | 0 | 21 | 23 | 3 | 6 | 66 | 73 |
| Arando | 55 | 52 | 10 | 11 | 6 | 1 | 15 | 19 | 21 | 13 | 15 | 8 | 66 | 74 | 45 | 148 *** | 1 | 1 | 10 | 10 | 3 | 2 | 66 | 98 ** |
| Region | Population | n | Na | PNa | A | He | Ho | FIS | FIS without HrMS004 Marker |
|---|---|---|---|---|---|---|---|---|---|
| Gilgit | Gulmit | 60 | 9.3 | 8 | 7.5 | 0.708 | 0.668 | 0.063 *** | 0.038 |
| Passu | 40 | 7.9 | 3 | 7.0 | 0.688 | 0.670 | 0.038 *** | 0.006 | |
| Shimshal | 60 | 7.8 | 10 | 6.3 | 0.713 | 0.731 | −0.016 *** | −0.044 ** | |
| Baltistan | Thesal | 20 | 7.8 | 10 | 7.8 | 0.690 | 0.717 | −0.013 | −0.032 |
| Chandopi | 40 | 9.7 | 8 | 7.7 | 0.684 | 0.663 | 0.062 *** | 0.034 | |
| Chutran | 40 | 10.8 | 11 | 8.6 | 0.711 | 0.653 | 0.097 *** | 0.083 *** | |
| Bisil | 20 | 7.7 | 4 | 7.7 | 0.707 | 0.675 | 0.071 *** | 0.028 | |
| Arando | 20 | 6.9 | 2 | 6.9 | 0.689 | 0.688 | 0.058 *** | 0.009 | |
| Mean | 8.5 | 7 | 7.4 | 0.699 | 0.683 | 0.045 | 0.016 | ||
| Stands | Wild | 150 | 11.3 | 27 | 14.7 | 0.739 | 0.695 | 0.063 ** | 0.035 *** |
| Village | 150 | 14.8 | 69 | 11.2 | 0.724 | 0.665 | 0.085 *** | 0.060 |
| Grouping | Source | Df | Sum of Squares | Estimated Variation | % | FST | FIS | FIT |
|---|---|---|---|---|---|---|---|---|
| Population | Among populations | 7 | 187 | 0.30 | 0.07 | |||
| Among individuals within populations | 292 | 1301 | 0.19 | 0.04 | ||||
| Within individuals | 300 | 1223 | 4.07 | 0.89 | ||||
| Total | 599 | 2711 | 4.57 | 1.00 | 0.067 *** | 0.045 *** | 0.108 *** | |
| Stands | Among stands | 1 | 76 | 0.24 | 0.05 | |||
| Among individuals within stands | 298 | 1413 | 0.33 | 0.07 | ||||
| Within individuals | 300 | 1223 | 4.07 | 0.88 | ||||
| Total | 599 | 2712 | 4.64 | 1.00 | 0.052 *** | 0.075 *** | 0.123 *** | |
| Populations*stands | Among populations*stands | 29 | 294 | 0.28 | 0.06 | |||
| Among individuals within populations*stands | 270 | 1195 | 0.17 | 0.04 | ||||
| Within individuals | 300 | 1223 | 4.07 | 0.90 | ||||
| Total | 599 | 2712 | 4.53 | 1.00 | 0.063 *** | 0.041 *** | 0.102 *** | |
| Cluster # | Among clusters | 2 | 141 | 0.34 | 0.08 | |||
| Among individuals within clusters | 297 | 1347 | 0.23 | 0.05 | ||||
| Within individuals | 300 | 1222 | 4.07 | 0.88 | ||||
| Total | 599 | 2711 | 4.65 | 1.00 | 0.075 *** | 0.054 *** | 0.125 *** |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nawaz, M.A.; Krutovsky, K.V.; Mueller, M.; Gailing, O.; Khan, A.A.; Buerkert, A.; Wiehle, M. Morphological and Genetic Diversity of Sea Buckthorn (Hippophae rhamnoides L.) in the Karakoram Mountains of Northern Pakistan. Diversity 2018, 10, 76. https://doi.org/10.3390/d10030076
Nawaz MA, Krutovsky KV, Mueller M, Gailing O, Khan AA, Buerkert A, Wiehle M. Morphological and Genetic Diversity of Sea Buckthorn (Hippophae rhamnoides L.) in the Karakoram Mountains of Northern Pakistan. Diversity. 2018; 10(3):76. https://doi.org/10.3390/d10030076
Chicago/Turabian StyleNawaz, Muhammad Arslan, Konstantin V. Krutovsky, Markus Mueller, Oliver Gailing, Asif Ali Khan, Andreas Buerkert, and Martin Wiehle. 2018. "Morphological and Genetic Diversity of Sea Buckthorn (Hippophae rhamnoides L.) in the Karakoram Mountains of Northern Pakistan" Diversity 10, no. 3: 76. https://doi.org/10.3390/d10030076
APA StyleNawaz, M. A., Krutovsky, K. V., Mueller, M., Gailing, O., Khan, A. A., Buerkert, A., & Wiehle, M. (2018). Morphological and Genetic Diversity of Sea Buckthorn (Hippophae rhamnoides L.) in the Karakoram Mountains of Northern Pakistan. Diversity, 10(3), 76. https://doi.org/10.3390/d10030076







