Synthesis and Structural Characterization of (E)-4-[(2-Hydroxy-3-methoxybenzylidene)amino]butanoic Acid and Its Novel Cu(II) Complex
Abstract
1. Introduction
2. Results and Discussion
2.1. Synthesis
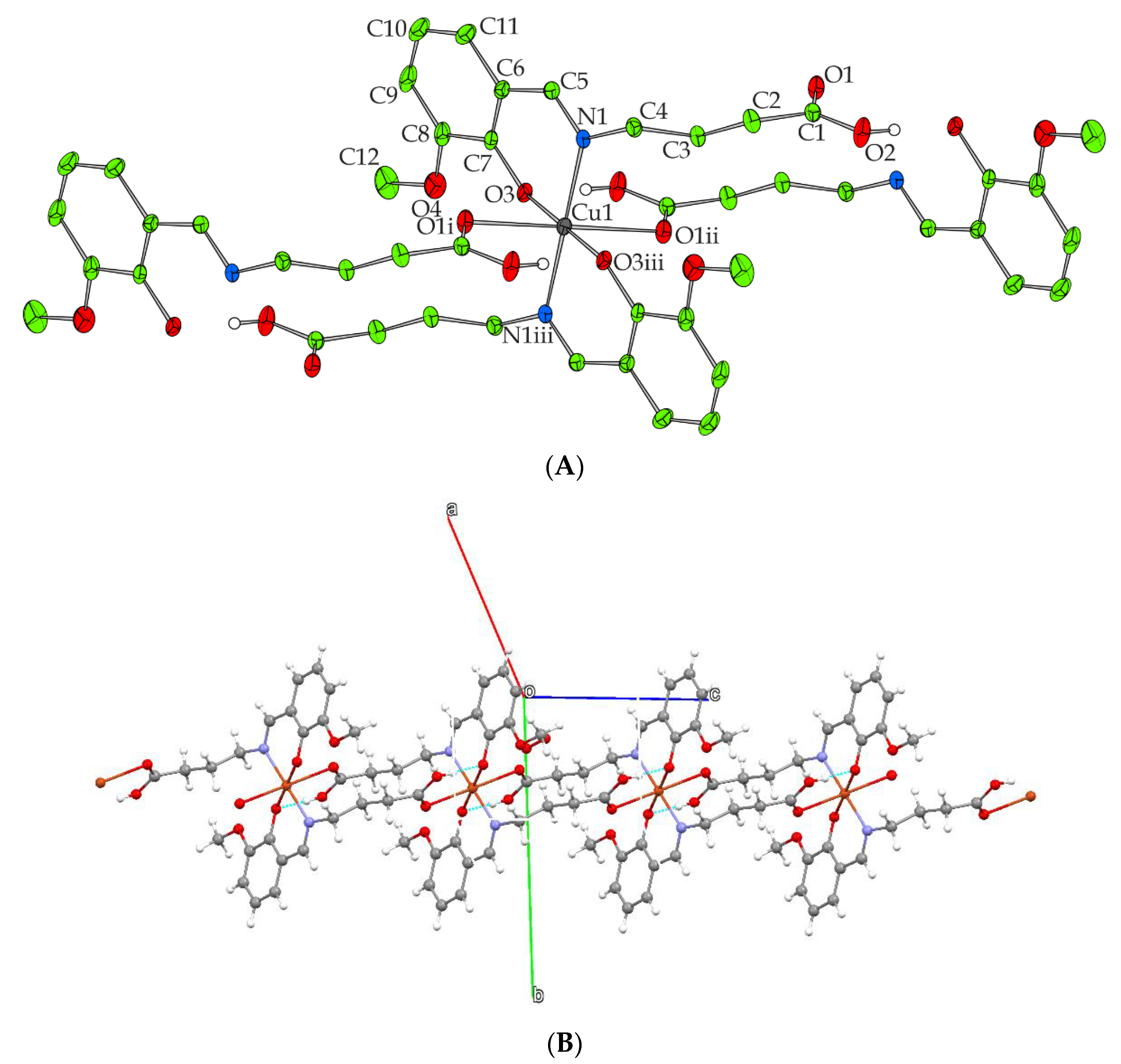
2.2. Description of X-ray Crystallographic Structure
2.3. FT-IR Spectroscopy
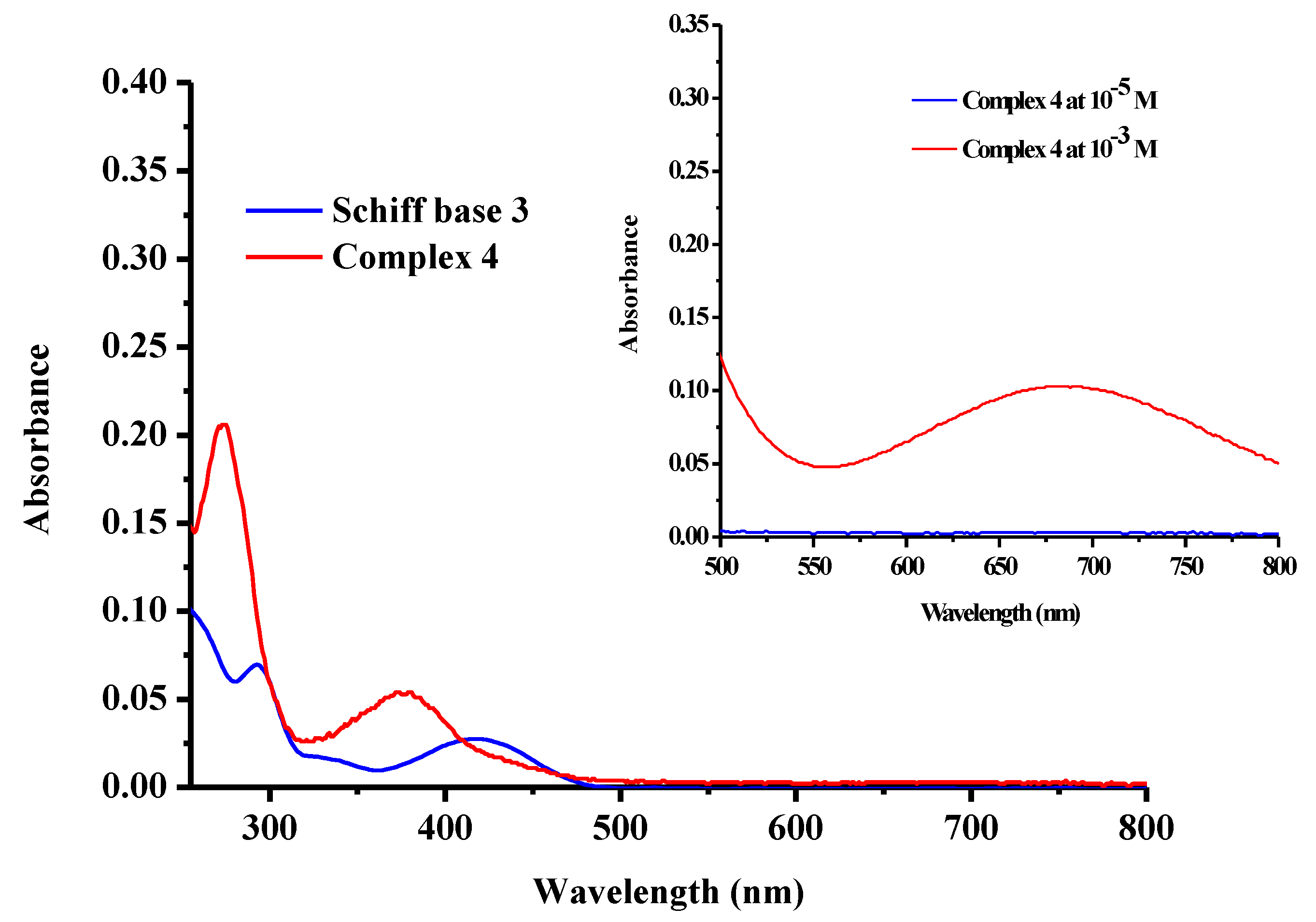
2.4. UV-Vis Spectroscopy
3. Materials and Methods
3.1. Synthesis
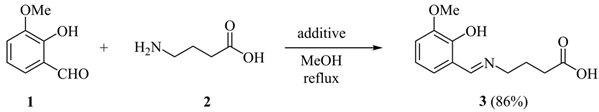
3.1.1. (E)-4-[(2-Hydroxy-3-methoxybenzylidene)amino]butanoic Acid (3)
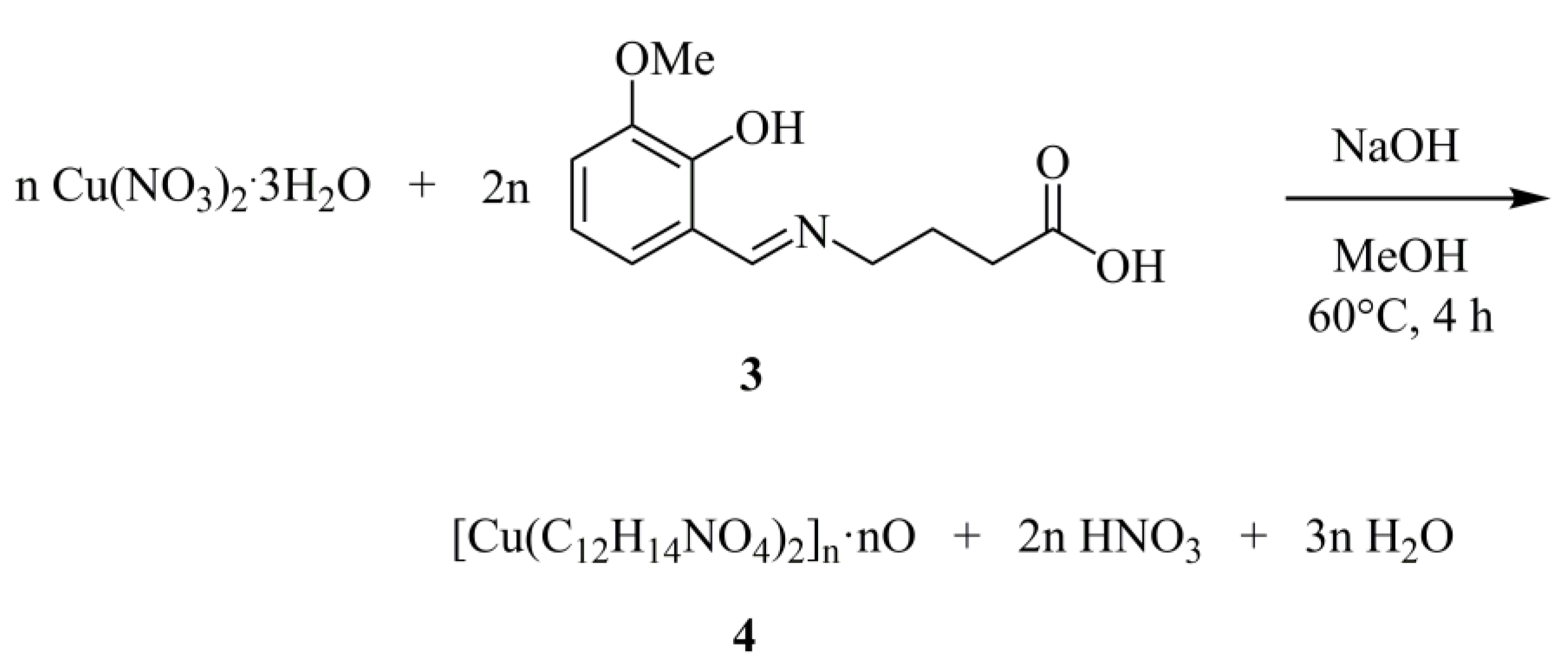
3.1.2. Synthesis of [Cu(C12H14NO4)2]n·nO (4)
3.2. X-ray Crystal Structure Determination
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Kostova, I.; Saso, L. Advances in research of Schiff-base metal complexes as potent antioxidants. Curr. Med. Chem. 2013, 20, 4609–4632. [Google Scholar] [CrossRef]
- Ejidike, I.P.; Ajibade, P.A. Transition metal complexes of symmetrical and asymmetrical Schiff bases as antibacterial, antifungal, antioxidant, and anticancer agents: Progress and prospects. Rev. Inorg. Chem. 2015, 35, 191–224. [Google Scholar] [CrossRef]
- Halevas, E.; Tsave, O.; Yavropoulou, M.P.; Hatzidimitriou, A.; Yovos, J.G.; Psycharis, V.; Gabriel, C.; Salifoglou, A. Design, synthesis and characterization of novel binary V(V)-Schiff base materials linked with insulin-mimetic vanadium-induced differentiation of 3T3-L1 fibroblasts to adipocytes. Structure–function correlations at the molecular level. J. Inorg. Biochem. 2015, 147, 99–115. [Google Scholar] [CrossRef] [PubMed]
- Halevas, E.; Nday, C.M.; Chatzigeorgiou, E.; Varsamis, V.; Eleftheriadou, D.; Jackson, G.E.; Litsardakis, G.; Lazari, D.; Ypsilantis, K.; Salifoglou, A. Chitosan encapsulation of essential oil “cocktails” with well-defined binary Zn(II)-Schiff base species targeting antibacterial medicinal nanotechnology. J. Inorg. Biochem. 2017, 176, 24–37. [Google Scholar] [CrossRef] [PubMed]
- Halevas, E.; Tsave, O.; Yavropoulou, M.; Yovos, J.G.; Hatzidimitriou, A.; Psycharis, V.; Salifoglou, A. In vitro structure-specific Zn(II)-induced adipogenesis and structure-function bioreactivity correlations. J. Inorg. Biochem. 2017, 177, 228–246. [Google Scholar] [CrossRef] [PubMed]
- Shamsi, M.; Yadav, S.; Arjmand, F. Synthesis and characterization of new transition metal Cu(II), Ni(II) and Co(II) L phenylalanine-DACH conjugate complexes: In vitro DNA binding, cleavage and molecular docking studies. J. Photochem. Photobiol. B 2014, 136, 1–11. [Google Scholar] [CrossRef]
- Fattuoni, C.; Vascellari, S.; Pivetta, T. Synthesis, protonation constants and biological activity determination of amino acid–salicylaldehyde-derived Schiff bases. Amino Acids 2020, 52, 397–407. [Google Scholar] [CrossRef]
- Gupta, K.C.; Kumar, A.S.; Lin, C.C. Polymer-supported Schiff base complexes in oxidation reactions. Coord. Chem. Rev. 2009, 253, 1923–1946. [Google Scholar] [CrossRef]
- Abdel-Rahman, L.H.; El-Khati, R.M.; Nassr, L.A.E.; Abu-Dief, A.M. DNA binding ability mode, spectroscopic studies, hydrophobicity, and in vitro antibacterial evaluation of some new Fe(II) complexes bearing ONO donors amino acid Schiff bases. Arab. J. Chem. 2017, 10, S1835–S1846. [Google Scholar] [CrossRef]
- Da Silva, C.M.; da Silva, D.L.; Modolo, L.V.; Alves, R.B.; de Resende, M.A.; Martins, C.V.B.; de Fátima, Ȃ. Schiff bases: A short review of their antimicrobial activities. J. Adv. Res. 2011, 2, 1–8. [Google Scholar] [CrossRef]
- Li, Y.; Dong, J.; Zhao, P.; Hu, P.; Yang, D.; Gao, L.; Li, L. Synthesis of amino acid Schiff base nickel (II) complexes as potential anticancer drugs in vitro. Bioinorg. Chem. Appl. 2020, 2020, 8834859. [Google Scholar] [CrossRef] [PubMed]
- Abu-Dief, A.M.; Mohamed, I.M.A. A review on versatile applications of transition metal complexes incorporating Schiff bases. Beni-Suef Univ. J. Basic Appl. Sci. 2015, 4, 119–133. [Google Scholar] [CrossRef] [PubMed]
- Zou, Y.; Han, J.; Saghyan, A.S.; Mkrtchyan, A.F.; Konno, H.; Moriwaki, H.; Izawa, K.; Soloshonok, V.A. Asymmetric synthesis of tailor-made amino acids using chiral Ni(II) complexes of Schiff bases. An update of the recent literature. Molecules 2020, 25, 2739. [Google Scholar] [CrossRef]
- Puterová-Tokárová, Z.; Mrázová, V.; Boča, R. Magnetism of novel Schiff-base copper(II) complexes derived from aminoacids. Polyhedron 2013, 61, 87–93. [Google Scholar] [CrossRef]
- Wang, Y.-Y.; Xu, F.-Z.; Zhu, Y.-Y.; Song, B.; Luo, D.; Yu, G.; Chen, S.; Xue, W.; Wu, J. Pyrazolo[3,4-d]pyrimidine derivatives containing a Schiff base moiety as potential antiviral agents. Bioorg. Med. Chem. Lett. 2018, 28, 2979–2984. [Google Scholar] [CrossRef]
- Jamil, D.M.; Al-Okbi, A.K.; Al-Baghdadi, S.B.; Al-Amiery, A.A.; Kadhim, A.; Gaaz, T.S.; Kadhum, A.A.H.; Mohamad, A.B. Experimental and theoretical studies of Schiff bases as corrosion inhibitors. Chem. Cent. J. 2018, 12, 7. [Google Scholar] [CrossRef]
- Mobinikhaledi, A.; Jabbarpour, M.; Hamta, A. Synthesis of some novel and biologically active Schiff bases bearing a 1,3,4-thiadiazole moiety under acidic and PTC conditions. J. Chil. Chem. Soc. 2011, 56, 812–814. [Google Scholar] [CrossRef]
- Galić, N.; Cimerman, Z.; Tomišić, V. Spectrometric study of tautomeric and protonation equilibria of o-vanillin Schiff base derivatives and their complexes with Cu(II). Spectrochim. Acta A Mol. Biomol. Spectrosc. 2008, 71, 1274–1280. [Google Scholar] [CrossRef]
- Pis-Diez, R.; Echeverria, G.A.; Piro, O.E.; Jios, J.L. Parajón-Costa, B.S. A structural, spectroscopic and theoretical study of an o-vanillin Schiff base derivative involved in enol-imine and keto-amine tautomerism. New J. Chem. 2016, 40, 2730–2740. [Google Scholar] [CrossRef]
- Muche, S.; Harms, K.; Biernasiuk, A.; Malm, A.; Popiołek, Ł.; Hordyjewska, A.; Olszewska, A.; Hołyńska, M. New Pd(II) Schiff base complexes derived from ortho-vanillin and L-tyrosine or L-glutamic acid: Synthesis, characterization, crystal structures and biological properties. Polyhedron 2018, 151, 465–477. [Google Scholar] [CrossRef]
- Halevas, E.; Pekou, A.; Papi, R.; Mavroidi, B.; Hatzidimitriou, A.G.; Zahariou, G.; Litsardakis, G.; Sagnou, M.; Pelecanou, M.; Pantazaki, A.A. Synthesis, physicochemical characterization and biological properties of two novel Cu(II) complexes based on natural products curcumin and quercetin. J. Inorg. Biochem. 2020, 208, 111083. [Google Scholar] [CrossRef] [PubMed]
- Munde, A.S.; Jagdale, A.N.; Jadhav, S.M.; Chondhekar, T.K. Synthesis, characterization and thermal study of some transition metal complexes of an asymmetrical tetradentate Schiff base ligand. J. Serb. Chem. Soc. 2010, 75, 349–359. [Google Scholar] [CrossRef]
- Bruker Analytical X-Ray Systems, Inc. Apex2, Version 2 User Manual: M86-E01078; Bruker Analytical X-Ray Systems, Inc.: Madison, WI, USA, 2006. [Google Scholar]
- Siemens Industrial Automation, Inc. SADABS: Area-Detector Absorption Correction; Siemens Industrial Automation, Inc.: Madison, WI, USA, 1996. [Google Scholar]
- Betteridge, P.W.; Carruthers, J.R.; Cooper, R.I.; Prout, K.; Watkin, D.J. CRYSTALS version 12: Software for guided crystal structure analysis. J. Appl. Cryst. 2003, 36, 1487. [Google Scholar] [CrossRef]
- Palatinus, L.; Chapuis, G. SUPERFLIP–a computer program for the solution of crystal structures by charge flipping in arbitrary dimensions. Appl. Cryst. 2007, 40, 786–790. [Google Scholar] [CrossRef]
- Watkin, D.J.; Prout, C.K.; Pearce, L.J. CAMERON; Chemical Crystallography Laboratory: Oxford, UK, 1996. [Google Scholar]

 | ||
| Entry | Additive | Yield (%) b |
| 1 | none | 76 |
| 2 | CH3COOH | 86 |
| 3 | H2SO4 c | trace amount |
| 4 | piperidine | 58 c |
| Bond Lengths (Å) | |||
| Cu(1)—O(1) i | 2.6167 (19) | O(4)—C(12) | 1.429 (3) |
| Cu(1)—O(1) ii | 2.6167 (19) | O(5)—O(5) iv | 1.449 (5) |
| Cu(1)—N(1) iii | 2.005 (2) | C(1)—C(2) | 1.505 (4) |
| Cu(1)—O3 iii | 1.9272 (16) | C(2)—C(3) | 1.514 (3) |
| Cu(1)—N(1) | 2.005 (2) | C(3)—C(4) | 1.510 (3) |
| Cu(1)—O(3) | 1.9272 (16) | C(5)—C(6) | 1.443 (4) |
| N(1)—C(4) | 1.476 (3) | C(6)—C(7) | 1.392 (4) |
| N(1)—C(5) | 1.282 (3) | C(6)—C(11) | 1.406 (4) |
| O(1)—C(1) | 1.203 (3) | C(7)—C(8) | 1.421 (3) |
| O(2)—C(1) | 1.308 (3) | C(8)—C(9) | 1.389 (5) |
| O(3)—C(7) | 1.325 (3) | C(9)—C(10) | 1.373 (5) |
| O(4)—C(8) | 1.363 (4) | C(10)—C(11) | 1.354 (5) |
| Angles (°) | |||
| O(1) i—Cu(1)—O(1) ii | 180 | C(8)—O(4)—C(12) | 117.5 (3) |
| O(1) i—Cu(1)—N(1) iii | 93.65 (7) | O(2)—C(1)—O(1) | 122.9 (2) |
| O(1) ii—Cu(1)—N(1) iii | 86.35 (7) | O(2)—C(1)—C(2) | 114.0 (2) |
| O(1) i—Cu(1)—O(3) iii | 83.34 (7) | O(1)—C(1)—C(2) | 123.1 (2) |
| O(1) ii—Cu(1)—O(3) iii | 96.66 (7) | C(1)—C(2)—C(3) | 112.2 (2) |
| N(1) iii—Cu(1)—O(3) iii | 90.23 (8) | C(2)—C(3)—C(4) | 112.8 (2) |
| O(1) i—Cu(1)—N(1) | 86.35 (7) | C(3)—C(4)—N(1) | 110.5 (2) |
| O(1) ii—Cu(1)—N(1) | 93.65 (7) | N(1)—C(5)—C(6) | 126.7 (2) |
| N(1) iii—Cu(1)—N(1) | 180 | C(5)—C(6)—C(7) | 121.9 (2) |
| O(3) iii—Cu(1)—N(1) | 89.77 (8) | C(5)—C(6)—C(11) | 117.6 (3) |
| O(1) i—Cu(1)—O(3) | 96.66 (7) | C(7)—C(6)—C(11) | 120.5 (3) |
| O(1) ii—Cu(1)—O(3) | 83.34 (7) | C(6)—C(7)—O(3) | 123.9 (2) |
| N(1) iii—Cu(1)—O(3) | 89.77 (8) | C(6)—C(7)—C(8) | 118.2 (3) |
| O(3) iii—Cu(1)—O(3) | 180 | O(3)—C(7)—C(8) | 117.8 (3) |
| N(1)—Cu(1)—O(3) | 90.23 (8) | C(7)—C(8)—O(4) | 114.6 (3) |
| Cu(1)—N(1)—C(4) | 120.85 (16) | C(7)—C(8)—C(9) | 119.2 (3) |
| Cu(1)—N(1)—C(5) | 122.71 (18) | O(4)—C(8)—C(9) | 126.2 (3) |
| C(4)—N(1)—C(5) | 116.4 (2) | C(8)—C(9)—C(10) | 121.5 (3) |
| Cu(1) v—O(1)—C(1) | 124.19 (17) | C(9)—C(10)—C(11) | 120.1 (3) |
| Cu(1)—O(3)—C(7) | 123.43 (16) | C(6)—C(11)—C(10) | 120.5 (3) |
| Cu(1)—O(1) i | 2.6167 (19) | O(4)—C(12) | 1.429 (3) |
| Cu(1)—O(1) ii | 2.6167 (19) | O(5)—O(5) iv | 1.449 (5) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Halevas, E.; Hatzidimitriou, A.; Mavroidi, B.; Sagnou, M.; Pelecanou, M.; Matiadis, D. Synthesis and Structural Characterization of (E)-4-[(2-Hydroxy-3-methoxybenzylidene)amino]butanoic Acid and Its Novel Cu(II) Complex. Molbank 2021, 2021, M1179. https://doi.org/10.3390/M1179
Halevas E, Hatzidimitriou A, Mavroidi B, Sagnou M, Pelecanou M, Matiadis D. Synthesis and Structural Characterization of (E)-4-[(2-Hydroxy-3-methoxybenzylidene)amino]butanoic Acid and Its Novel Cu(II) Complex. Molbank. 2021; 2021(1):M1179. https://doi.org/10.3390/M1179
Chicago/Turabian StyleHalevas, Eleftherios, Antonios Hatzidimitriou, Barbara Mavroidi, Marina Sagnou, Maria Pelecanou, and Dimitris Matiadis. 2021. "Synthesis and Structural Characterization of (E)-4-[(2-Hydroxy-3-methoxybenzylidene)amino]butanoic Acid and Its Novel Cu(II) Complex" Molbank 2021, no. 1: M1179. https://doi.org/10.3390/M1179
APA StyleHalevas, E., Hatzidimitriou, A., Mavroidi, B., Sagnou, M., Pelecanou, M., & Matiadis, D. (2021). Synthesis and Structural Characterization of (E)-4-[(2-Hydroxy-3-methoxybenzylidene)amino]butanoic Acid and Its Novel Cu(II) Complex. Molbank, 2021(1), M1179. https://doi.org/10.3390/M1179








