6-Bromo-N-(2-methyl-2H-benzo[d][1,2,3]triazol-5-yl)quinolin-4-amine
Abstract
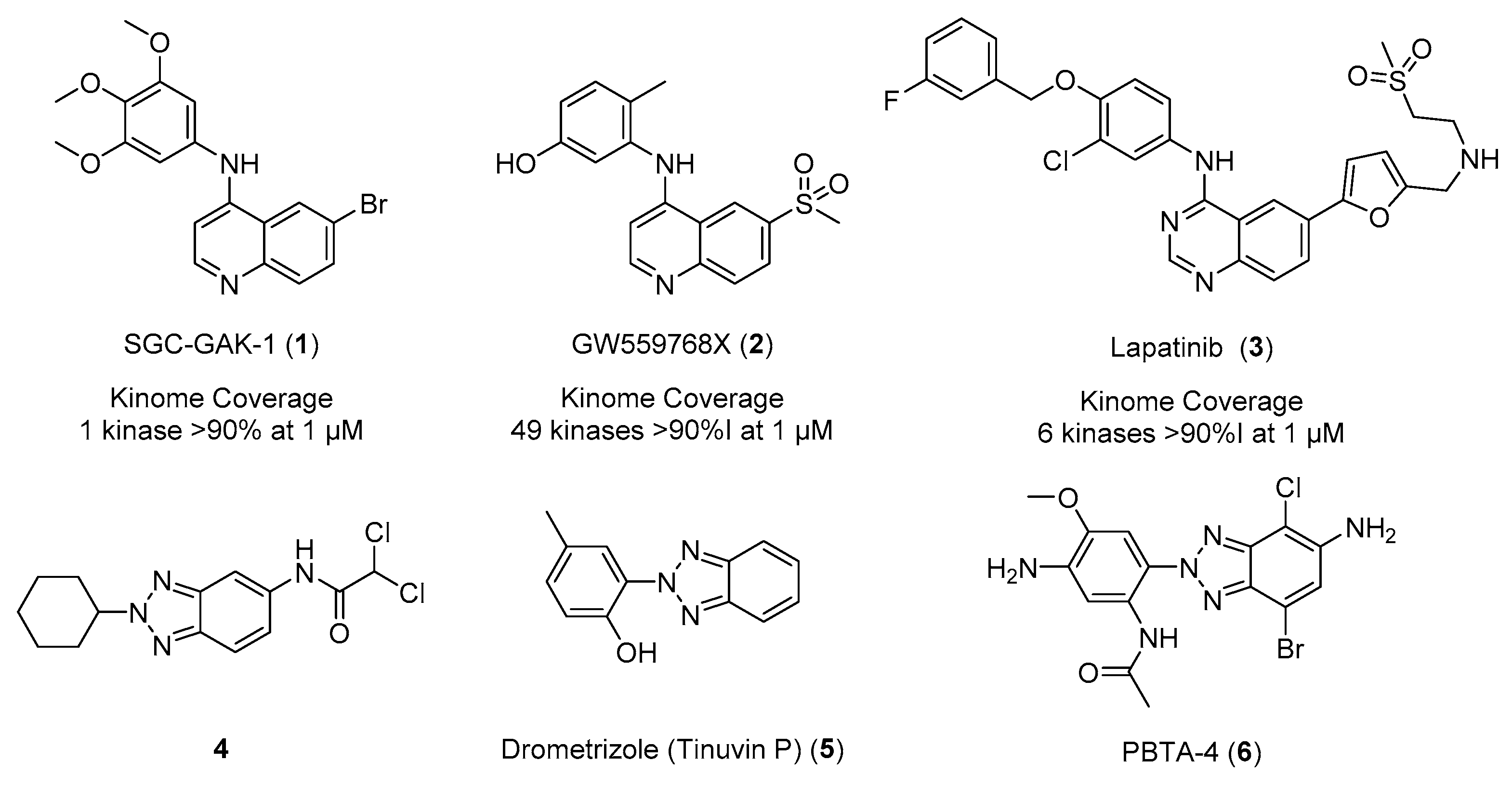
:1. Introduction
2. Results
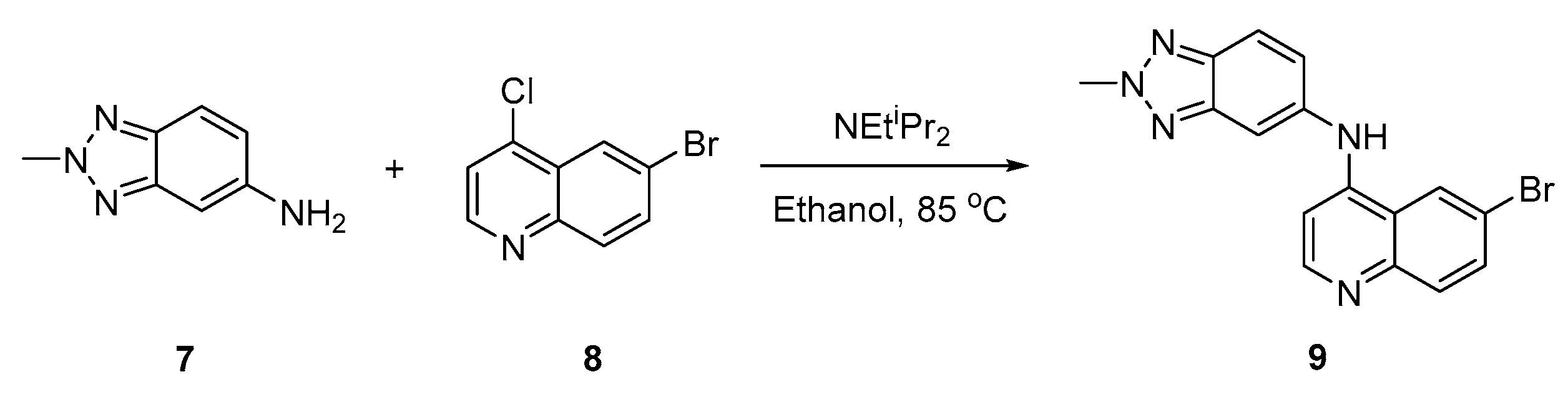
2.1. Synthesis of 9
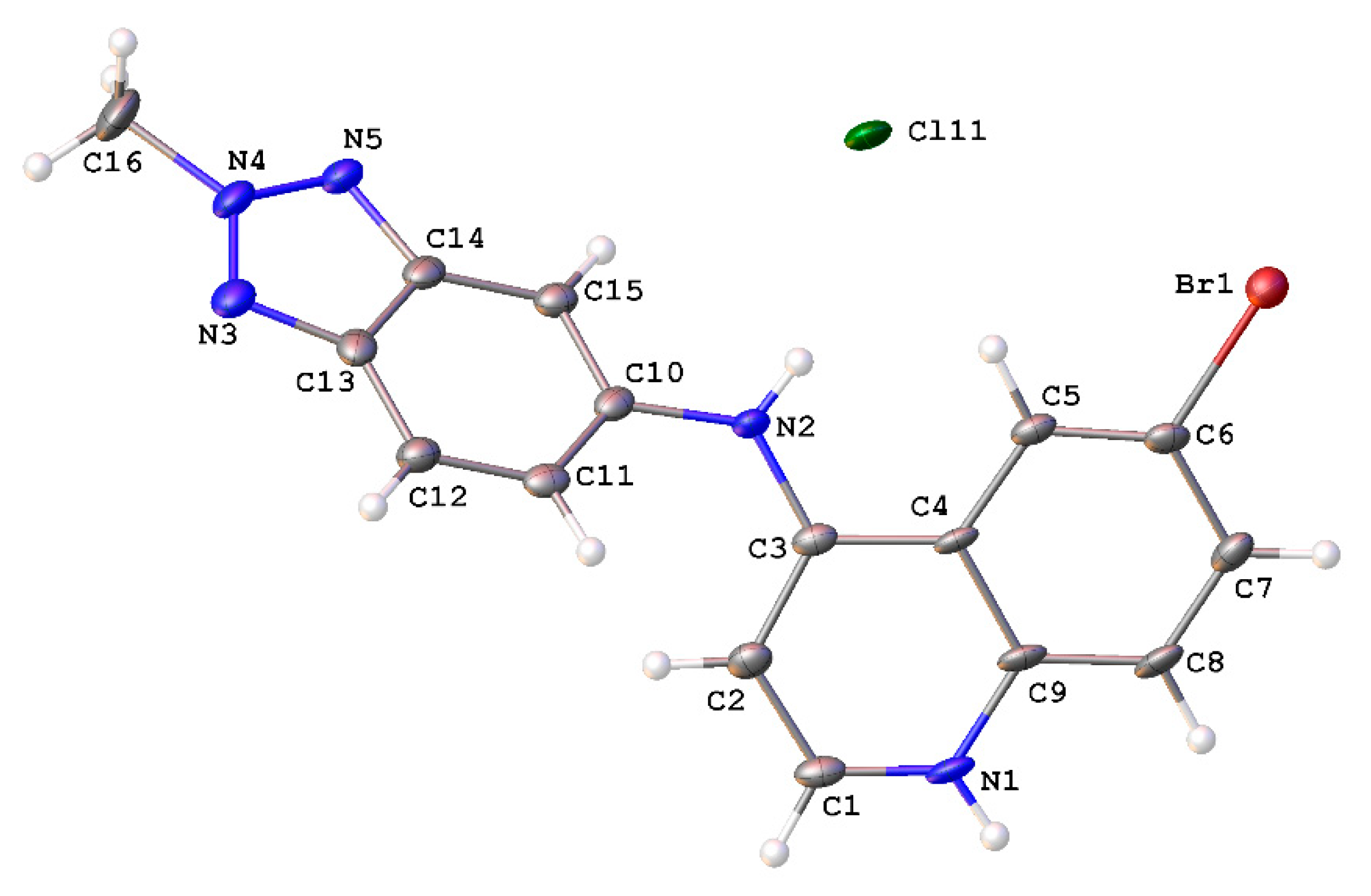
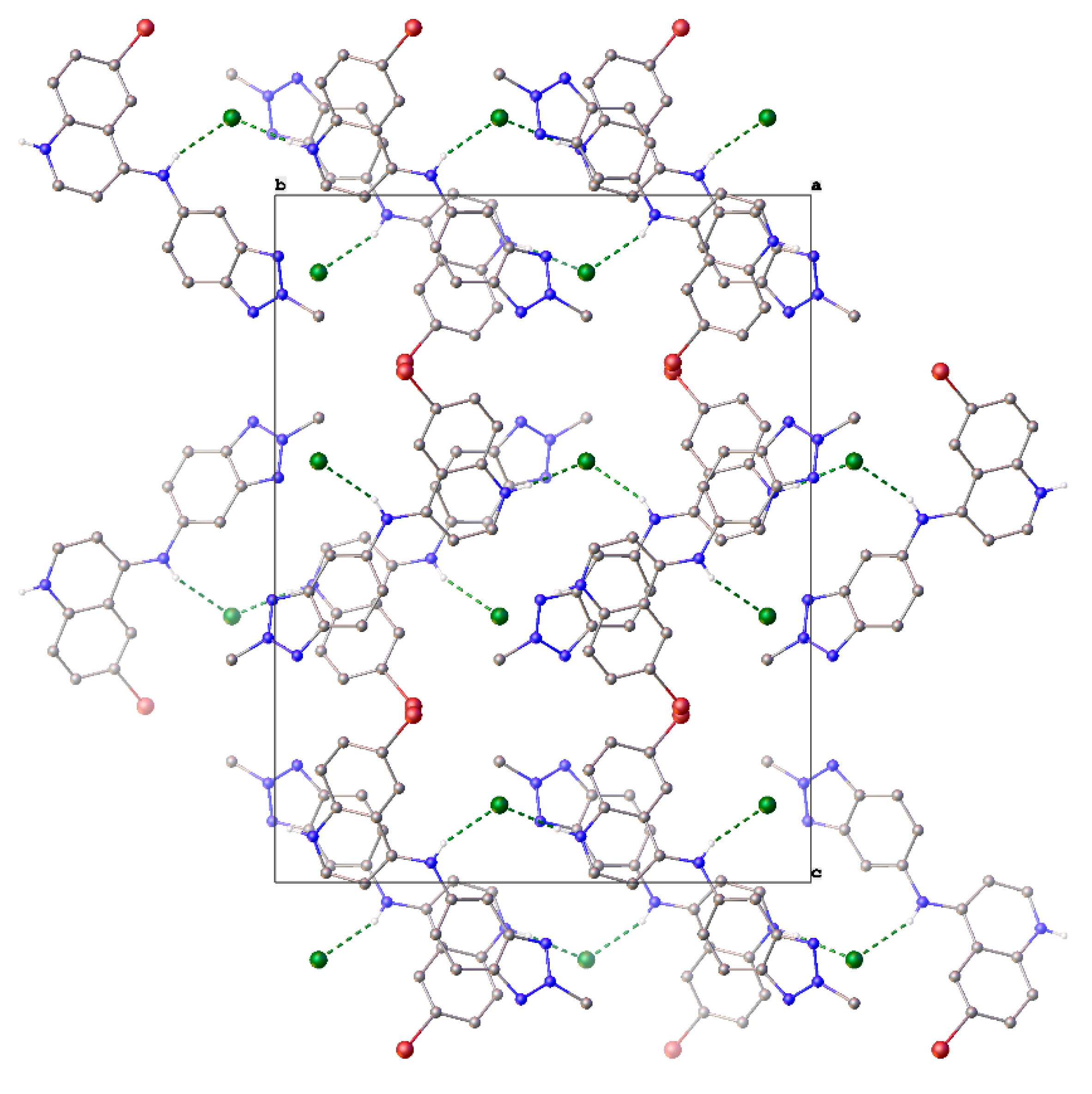
2.2. Crystal Structure of 9
3. Discussion
4. Experimental Section
4.1. Chemistry
4.2. Crystallography
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ferguson, F.M.; Gray, N.S. Kinase inhibitors: The road ahead. Nat Rev Drug Discov. 2018, 17, 353–377. [Google Scholar] [CrossRef] [PubMed]
- Cohen, P.; Alessi, D.R. Kinase drug discovery—What’s next in the field? ACS Chem. Biol. 2013, 8, 96–104. [Google Scholar] [CrossRef] [PubMed]
- Asquith, C.R.M.; Laitinen, T.; Bennett, J.M.; Godoi, P.H.; East, M.P.; Tizzard, G.J.; Graves, L.M.; Johnson, G.L.; Dornsife, R.E.; Wells, C.I.; et al. Identification and Optimization of 4-Anilinoquinolines as Inhibitors of Cyclin G Associated Kinase. ChemMedChem 2018, 13, 48–66. [Google Scholar] [CrossRef] [PubMed]
- Asquith, C.R.M.; Berger, B.T.; Wan, J.; Bennett, J.M.; Capuzzi, S.J.; Crona, D.J.; Drewry, D.H.; East, M.P.; Elkins, J.M.; Fedorov, O.; et al. SGC-GAK-1: A Chemical Probe for Cyclin G Associated Kinase (GAK). J. Med. Chem. 2019, 62, 2830–2836. [Google Scholar] [CrossRef] [PubMed]
- Asquith, C.R.M.; Treiber, D.K.; Zuercher, W.J. Utilizing comprehensive and mini-kinome panels to optimize the selectivity of quinoline inhibitors for cyclin G associated kinase (GAK). Bioorg. Med. Chem. Lett. 2019, 29, 1727–1731. [Google Scholar] [CrossRef] [PubMed]
- Drewry, D.H.; Wells, C.I.; Andrews, D.M.; Angell, R.; Al-Ali, H.; Axtman, A.D.; Capuzzi, S.J.; Elkins, J.M.; Ettmayer, P.; Frederiksen, M.; et al. Progress towards a public chemogenomic set for protein kinases and a call for contributions. PLoS ONE 2017, 12, e0181585. [Google Scholar] [CrossRef]
- Fabian, M.A.; Biggs, W.H., 3rd; Treiber, D.K.; Atteridge, C.E.; Azimioara, M.D.; Benedetti, M.G.; Carter, T.A.; Ciceri, P.; Edeen, P.T.; Floyd, M.; et al. A small molecule–kinase interaction map for clinical kinase inhibitors. Nat. Biotechnol. 2005, 23, 329–336. [Google Scholar] [CrossRef]
- Agalave, S.G.; Maujan, S.R.; Pore, V.S. Click chemistry: 1,2,3-triazoles as pharmacophores. Chem. Asian J. 2011, 6, 2696–2718. [Google Scholar] [CrossRef]
- Dheer, D.; Singh, V.; Shankar, R. Medicinal attributes of 1,2,3-triazoles: Current developments. Bioorg. Chem. 2017, 71, 30–54. [Google Scholar] [CrossRef]
- Liu, Y.; Yan, W.; Chen, Y.; Petersen, J.L.; Shi, X. Efficient synthesis of N-2-aryl-1,2,3-triazole fluorophores via post-triazole arylation. Org. Lett. 2008, 10, 5389–5392. [Google Scholar] [CrossRef]
- Wang, X.J.; Zhang, L.; Krishnamurthy, D.; Senanayake, C.H.; Wipf, P. General solution to the synthesis of N-2-substituted 1,2,3-triazoles. Org. Lett. 2010, 12, 4632–4635. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.J.; Sidhu, K.; Zhang, L.; Campbell, S.; Haddad, N.; Reeves, D.C.; Krishnamurthy, D.; Senanayake, C.H. Bromo-directed N-2 alkylation of NH-1,2,3-triazoles: Efficient synthesis of poly-substituted 1,2,3-triazoles. Org. Lett. 2009, 11, 5490–5493. [Google Scholar] [CrossRef] [PubMed]
- Cai, R.; Yan, W.; Bologna, M.G.; de Silva, K.; Ma, Z.; Finklea, H.O.; Petersen, J.L.; Li, M.; Shi, X. Synthesis and characterization of N-2-aryl-1,2,3-triazole based iridium complexes as photocatalysts with tunable photoredox potential. Org. Chem. Front. 2015, 2, 141–144. [Google Scholar] [CrossRef]
- Caliendo, G.; Novellino, E.; Sagliocco, G.; Santagada, V.; Silipo, C.; Vittoria, A. Synthesis, antimicrobial data and correlation analysis in a set of 2-alkyl-5-amidobenzotriazoles. Eur. J. Med. Chem. 1992, 27, 161–166. [Google Scholar] [CrossRef]
- Nuvole, A.; Sanna, P.; Paglietti, G.; Juliano, C.; Zanetti, S.; Cappuccinelli, P. 1,2,3-Triazolo [4,5-f]quinolines. II. Preparation and antimicrobial evaluation of 6-ethyl-6,9-dihydro-1(2)(3)-R-1(2) (3)H-triazolo[4,5-f]quinolin-9-one-8-carboxylic acids as anti-infectives of the urinary tract. Il Farmaco 1989, 44, 619–632. [Google Scholar]
- María, D.S.; Claramunt, R.M.; Bobosik, V.; Torralba, C.M.; Torres, M.R.; Alkorta, I.; Elguero, J. Synthesis and structural study of 2-arylbenzotriazoles related to Tinuvins. Tetrahedron 2013, 69, 3027–3038. [Google Scholar] [CrossRef]
- Nukaya, H.; Shiozawa, T.; Tada, A.; Terao, Y.; Ohe, T.; Watanabe, T.; Asanoma, M.; Sawanishi, H.; Katsuhara, T.; Sugimura, T.; et al. Identification of 2-[2-(acetylamino)-4-amino-5-methoxyphenyl]-5-amino-7-bromo-4-chloro-2H-benzotriazole (PBTA-4) as a potent mutagen in river water in Kyoto and Aichi prefectures, Japan. Mutat. Res. 2001, 492, 73–80. [Google Scholar] [CrossRef]
- Ohe, T.; Takeuchi, N.; Watanabe, T.; Tada, A.; Nukaya, H.; Terao, Y.; Sawanishi, H.; Hirayama, T.; Sugimura, T.; Wakabayashi, K. Quantification of two aromatic amine mutagens, PBTA-1 and PBTA-2, in the yodo river system. Environ. Health Perspect. 1999, 107, 701–704. [Google Scholar] [CrossRef]
- Shiozawa, T.; Tada, A.; Nukaya, H.; Watanabe, T.; Takahashi, Y.; Asanoma, M.; Ohe, T.; Sawanishi, H.; Katsuhara, T.; Sugimura, T.; Wakabayashi, K.; Terao, Y. Isolation and identification of a new 2-phenylbenzotriazole-type mutagen (PBTA-3) in the Nikko river in Aichi, Japan. Chem. Res. Toxicol. 2000, 13, 535–540. [Google Scholar] [CrossRef]
- Asquith, C.R.M.; Naegeli, N.; East, M.P.; Laitinen, T.; Havener, T.M.; Wells, C.I.; Johnson, G.L.; Drewry, D.H.; Zuercher, W.J.; Morris, D.C. Design of a cyclin G associated kinase (GAK)/epidermal growth factor receptor (EGFR) inhibitor set to interrogate the relationship of EGFR and GAK in chordoma. J. Med. Chem. 2019, 62, 4772–4778. [Google Scholar] [CrossRef]
- Asquith, C.R.M.; Fleck, N.; Torrice, C.D.; Crona, D.J.; Grundner, C.; Zuercher, W.J. Anti-tubercular activity of novel 4-anilinoquinolines and 4-anilinoquinazolines. Bioorg. Med. Chem. Lett. 2019, 18, 2695–2699. [Google Scholar] [CrossRef] [PubMed]
- Asquith, C.R.M.; Maffuid, K.A.; Laitinen, T.; Torrice, C.D.; Tizzard, G.J.; Crona, D.J.; Zuercher, W.J. Targeting an EGFR water network using novel 4-anilinoquin(az)olines inhibitors for chordoma. ChemMedChem 2019, 14, 1693–1700. [Google Scholar] [CrossRef] [PubMed]
- Asquith, C.R.M.; Bennett, J.M.; Su, L.; Laitinen, T.; Elkins, J.M.; Pickett, J.E.; Wells, C.I.; Li, Z.; Willson, T.M.; Zuercher, W.J. Development of SGC-GAK-1 as an orally active in vivo probe for cyclin G associated kinase through cytochrome P450 inhibition. bioRxiv 2019, 629220. [Google Scholar] [CrossRef] [Green Version]
- Sheldrick, G.M. ShelXT-Integrated space-group and crystal-structure determination. Acta Crystallogr. 2015, A71, 3–8. [Google Scholar] [CrossRef]
- Dolomanov, O.V.; Bourhis, L.J.; Gildea, R.J.; Howard, J.A.K.; Puschmann, H. Olex2: A complete structure solution, refinement and analysis program. J. Appl. Cryst. 2009, 42, 339–341. [Google Scholar] [CrossRef]
- Sheldrick, G.M. Crystal structure refinement with ShelXL. Acta Crystallogr. 2015, C27, 3–8. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Asquith, C.R.M.; Tizzard, G.J. 6-Bromo-N-(2-methyl-2H-benzo[d][1,2,3]triazol-5-yl)quinolin-4-amine. Molbank 2019, 2019, M1087. https://doi.org/10.3390/M1087
Asquith CRM, Tizzard GJ. 6-Bromo-N-(2-methyl-2H-benzo[d][1,2,3]triazol-5-yl)quinolin-4-amine. Molbank. 2019; 2019(4):M1087. https://doi.org/10.3390/M1087
Chicago/Turabian StyleAsquith, Christopher R. M., and Graham J. Tizzard. 2019. "6-Bromo-N-(2-methyl-2H-benzo[d][1,2,3]triazol-5-yl)quinolin-4-amine" Molbank 2019, no. 4: M1087. https://doi.org/10.3390/M1087
APA StyleAsquith, C. R. M., & Tizzard, G. J. (2019). 6-Bromo-N-(2-methyl-2H-benzo[d][1,2,3]triazol-5-yl)quinolin-4-amine. Molbank, 2019(4), M1087. https://doi.org/10.3390/M1087





