Abstract
A polymerase chain reaction (PCR) based diagnostic assay was used to develop markers for detection of Fusarium verticillioides (=F. moniliforme), a fumonisin producing fungus in maize tissues. Species-specific primers were designed based on sequence data from the polyketide synthase (PKS) gene (FUM1- previously FUM5) responsible for fumonisin production in fungi. Four sets of oligonucleotide primers were tested for their specificity using 24 strains of F. verticillioides, 10 F. proliferatum, and 12 of other Fusarium species. In addition, 13 species of other fungal genera, from four phyla, were tested as negative controls. Among the four sets, primer set B consistently amplified a 419-bp fragment from the DNA 96% of all F. verticillioides strains and 83% of F. proliferatum. All other fungi tested were negative using primer set B. A total of 38% of the F. verticillioides strains grown on a selective liquid medium produced fumonisin and 92% formed the toxin on standard rice medium. When fumonisin formed in culture, PCR assay using primer set B detected every strain of F. verticillioides, but only amplified 80% of F. proliferatum strains that produced the toxin. PCR detection was consistent at 100 pg/μl concentration of genomic DNA from 4 F. verticillioides strains, but varied at 10 pg/μl. Two duplicate greenhouse tests using artificially inoculated maize plants, had greater levels of F. verticillioides detected after re-evaluting using primer set B than from culturing of the tissues. The molecular protocols described in this study requires only 1 day for completion compared to approximately 10 days for cultural work and morphological determination. In conclusion, conventional PCR assay using primer set B provides a sensitive and accurate detection assay that can be used as a primary or secondary confirmation method for identification and occurrence of F. verticillioides within the maize tissues. However, studies using primer set B for fumonisin production determined by strains of F. verticillioides and F. proliferatum will require further verification.
1. Introduction
Fusarium spp. are commonly associated with maize (Zea mays L.) causing root, stalk, ear and kernel rots [1]. In Mississippi, the most common species infecting maize is Fusarium verticillioides (Sacc.) Nirenberg (= Fusarium moniliforme Sheld.), but Fusarium proliferatum (T. Matsushima) Nirenberg is also commonly isolated from infected maize [2]. The teleomorphic stage Giberella fujikuroi (Sawada) Ito in Ito & K. Kimura, consists of a complex containing up to nine (A-G) intersterile mating populations, which produce high levels of fumonisin [3–5]. Within this complex, F. verticillioides and F. proliferatum belong to mating populations A and D, respectively [6].
Fusarium verticillioides, which occurs worldwide, can colonize all portions of maize plants including the kernels, but infections are often symptomless [7]. The fungus was reported to occur systemically in all plant parts of maize [8]. However, a simplified method can be used to eliminate F. verticillioides from seed prior to planting for resistance studies [9]. In addition to damaging plant tissues, F. verticillioides forms fumonisins (FBs), a group of mycotoxins which cause a variety of health problems in animals and humans [10–12]. Fumonisins are water soluble toxins which consist of aminopolyols with a core structure containing either 19 or 20 carbon backbones with hydroxyl, methyl, and tricarballylic acid moieties at different positions along the carbon backbone [13,14]. Biochemical analyses indicated that fumonisins are products of a polyketide synthase (PKS) gene called FUM1 (previously FUM5) [15–17].
Four main groups of fumonisins occur in nature, but the fumonisin B series contains the most important ones consisting of FB1, FB2, FB3 and FB4, with the most common being FB1 and FB2 [11,18,19]. The types that form, however, are strain dependent [7]. In several studies, 23 species of Fusarium were tested for fumonisin production and only F. verticillioides, F. proliferatum, and F. nygamai Burgess & Trimboli formed the toxin [20–22]. Unfortunately, infections by fumoninsin forming fungi are symptomless in kernels and supportive plant tissues that are unknowingly provided to animals and humans as feed or food, respectively [7].
Symptomless infections of kernels by F. verticillioides are generally much greater than visible infections responsible for observable tissue necrosis or damage [23]. Methods to morphologically and physiologically characterize fumonisin forming Fusarium spp. are considered labor intensive and slow in determination. Improved methods for rapid identification and detection of fumonisin forming fungi become essential especially since the grain is consumed by animals and humans.
Traditional methods for identification of Fusarium spp. require considerable expertise and specialized growth media [20,24]. These procedures are labor intensive and require one or two weeks of growth before identifications can be established. Cost of laboratory supplies can be considerable for conducting research (e.g. variety testing) or for diagnostic determinations. Since F. verticillioides and other fumonisin producing fungi routinely form latent or asymptomatic infections in maize tissues, determining their occurrence and frequency on host tissue requires systematic sampling [25]. Therefore, improved and quicker methods for identifying fumonisin forming fungi from maize tissues has become important, especially since fumonisins are now being implicated in diseases and cancer of animals. Molecular methods such as polymerase chain reaction (PCR) are now widely used in fungal taxonomy [26–29] and more recently they have been used for detection of fungal pathogens in plant tissue [30,31]. Another important aspect of PCR techniques is that only small quantities of DNA are required to confirm the presence of a pathogen on host tissue that otherwise might not be detected [32]. In previous studies, PCR techniques have been used successfully for identifying specific pathotypes of the take-all pathogen Gaeumannomyces graminis (var.-tritici, avenae, and graminis) on wheat and turf [33]. Each of the three pathotypes differs in pathogenicity and host range and recommended control practices vary between these fungi. Knowledge of strain variation in the field is crucial for development of control strategies [28]. Methods for developing species specific primers have varied over the last 15 years, but one successful approach involves use of internal transcriber regions (ITS) [34]. The ribosomal DNA (rDNA) found in the nuclear genome of fungi consists of three repeating RNA genes transcribed as a single united code for the 18S, 5.8S and 28S RNA genes that are highly conserved and primers to these regions have been developed [35]. Between these coding regions, the ITS1 and ITS2 separate the three RNA genes. O’Donnell and Cigelnik [36] and O’Donnell et al. [37] recommended against the use of ITS regions for species-level identification of Fusarium due to non-orthologous copies of the ITS2 within the Gibberella clade. This limitation would prevent the development of primers for identification of fumonisin forming fungi. Another approach for PCR, random amplified polymorphic DNA (RAPD), or arbitrary sequences, has been shown to be an excellent method for identifying polymorphic differences within and between fungal species [34]. This method does not require previous knowledge of the genome and uses short oligodeoxynucleotides as arbitrary primers. In one study, Bayman and Cotty [38] used RAPD to correlate taxa relatedness based on differences in morphology, physiology, and toxigenic production potential. In addition, RAPDs were employed in other studies to distinguish fungal species and pathogenicity variability within species [29,39]. Preliminary studies showed that RAPD was not sensitive enough to distinguish fumonisin forming fungi (Baird, unpublished data). Therefore, specific primers developed from known sequences of genes for fumonisin forming fungi should be investigated [40,41]. González-Jaén et al. [40] compared DNA-based strategies for development of primers for identification of F. verticillioides fumonisin producing and non-producing strains.
Previous investigations have been successful in development of multiplex PCR assay for fumonisin forming fungi using the FUM1 gene consisting of a 7.8-kb coding region [42,43] or the intergenic spacer region (IGS), located between 16S–23S rRNA genes, for distinguishing fumonisin producing strains of F. verticillioides from non-producing strains [44]. During this period of multiple primer development, the following study was being conducted to identify additional primers to differentiate strains of fumonisin producing fungi, such as F. verticillioides from the FUM1 gene sequence involved in fumonisin biosynthesis. Additional verification molecular methods are essential for further verification of fumonisin forming Fusarium spp. associated with food crops such as maize. Therefore, the objective of this research was to develop a reliable and sensitive conventional diagnostic PCR assay for detection of fumonisin forming fungi. Furthermore, the study attempted to verify that the primers are specific to fumonisin producing strains regardless of their toxin activity on a selective medium.
2. Materials and Methods
Fungal Cultures
Strains of Fusarium spp. and other fungal genera were obtained from four locations: the USDA/ARS, National Center for Agricultural Utilization Research, Peoria, IL; Fusarium Research Center, The Pennsylvania State University, University Park, PA, USDA/ARS CHPRRU Unit (Mycotoxin research), Mississippi State University, MS and USDA/ARS Mycotoxin Research Unit, Stoneville, MS. All strains were grown at room temperature in Petri plates (10 × 100 mm) containing Difco potato dextrose agar. Cultures used for genomic DNA extraction were grown in a defined liquid medium that enhances fumonisin production of strains up to 18 hrs after incubation with F. verticillioides [45]. The medium included 2.5 mM KH2PO4, 2.5 mM MgSO4, 85 mM NaC1, 10 mM of ammonium phosphate (nitrogen source substitute) and 117 mM sucrose at pH 5.9. A total of 100 ml of liquid medium was added per flask with 5 replicate flasks per strain. All strains were grown for 14-d with shaking at 200 rpm and 26°C. Mycelia were harvested from the culture by filtration onto Whatman No. 1 filter paper, transferred to a mortar and frozen in liquid nitrogen. The hyphae were ground to a fine powder and stored at – 4°C placed in sterile vials (1.5ml). Total genomic DNA was obtained using the DNeasy®Plant Mini Kit by Qiagen (Valencia, CA) following a modified form of the manufactures protocol. A total of 0.15 g of mycelium was used per strain for extraction instead of the recommended 0.1 g. Genomic DNA was quantified by UV spectrophotometer (Amersham Biosciences, Newark, NJ) at 260nm.
Design of gene-specific oligonucleotide primers
The oligonucleotide primers used in the amplification reaction were derived from the FUM1 gene sequence (GenBank accession AF 155773-listed as FUM5) involved in fumonisin formation by F. verticillioides [15]. Lasergene expert sequence analysis software, version 5 (DNASTAR, Inc., Madison, WI) was used to select the optimal primers from the known gene sequence. Based on unique primer sites found within the FUM 1 sequence region, four sets of primers including A, B, C and D were compared. The selected primers varied from 21 to 28 nucleotides in length, minimum G + C content of 50%, and a minimum melting temperature of 66.1°C and were chosen to amplify fragments between 0.4 to 1.5 kb (Table 1).

Table 1.
Description of four primer sets used for PCR amplification of F. verticillioides DNA
PCR Amplification
Reactions were performed in volumes of 50 μl and contained 20 ng of fungal DNA for each reaction. The reaction mixture consisted of a 10X buffer (Taq Polymerase Kit – Fisher Scientific, NJ) solution consisting of 50 mM KCl and 10 mM Tris-HCl (pH 9), 0.2 mM of each dATP, dCTP, dGTP, and dTTP, 0.5 mM of each forward and reverse primer pairs (Table 1), and 1.5 mM of MgCl2.
Amplification was performed using the Mastercycler® personal (Eppendorf AG, Hamburg). The PCR program included the following temperature regime: 95°C for 3 min, hold at 72°C for the addition of 5 units of Taq DNA polymerase at 5 u/μl (Fisher Scientific, New Jersey) to each 0.5 ml microfuge tube. The remaining steps in the program were repeated 32 times and consisted of 95°C for 1 min, 60°C for 1 min, 72°C for 3 min, and the final extension was at 72ºC for 5 min. The PCR products (15 μl) were separated by gel electrophoresis in 2.0 % agarose at 1 X TAE gel and stained with ethidium bromide for photography.
Primer pairs were tested for their ability to amplify different levels of target DNA using strains M-2326, M-5496, F1 (FCRB13) and F7 (FCR 14). Each PCR reaction was replicated three times and compared using DNA levels of 30 ng/μl, 20 ng/μl, 10 ng/μl, 1 ng/μl, 100 pg/μl, 10 pg/μl, and 1 pg/μl.
Fumonisin Production in vitro
Fumonisin production was determined for each strain during the investigation using previously described methods [2]. All strains were grown on two selective media (discussed below) to verify their potential to produce fumonisin (Table 2). Following the 14-d incubation of the strains with shaking at 200 rpm and 26°C, culture filtrates (liquid medium) were collected for assay. A total of 100 ml were collected per strain from 5 flasks. All samples were stored at –20°C until processing to determine presence of fumonisin. The liquid within each tube for the strains was removed by a Savant Speedvac (RC 210B). The dried samples were stored at –20°C for further processing. The second method used to determine fumonisin production in strains was with rice previously described [46]. All isolates used four replicate flasks were pooled into a composite sample. Each pooled sample was assayed twice to confirm the results.

Table 2.
Isolate designation, original host, geographic origin, source, PCR reaction, and fumonisin production in vitro of fungi assayed in this study.
The analytical method used in determining the fumonisins (FB1, FB2, and FB3) grown on two media was by liquid chromatography/electrospray ionization/mass spectrometry (LC/ESI/MS). Aliquots (500 μL) from cultural filtrates were dried under nitrogen, redissolved in 500 μL water, vortexed for 30 sec., followed by the addition of 500 μL of methanol to extract fumonisins and their derivatives. Samples (10 g) from the mold rice samples were extracted with 50 ml of 70% methanol and filtered through No. 1 Whatman filter paper. An aliquot (10 ml) was applied to a SAX clean-up column (Varian, Harbor City, CA) and analyzed with some modifications as reported by Plattner [47]. Extracts were diluted 2-fold to prevent fumonisin carry-over. The LC/ESI/MS analysis was performed on a Thermo Finnigan LCQ Advantage, coupled to a Thermo Finnigan Surveyor MS, and a Thermo Finnigan Surveyor MS Pump. In full-scan mode, a 10 μl partial loop injection was used to run each sample, using the appropriate mass range for the fumonisins (722.4, B1; 706, B2 and B3). The column used for fumonisin analysis was a MetaChem Intersil 5u ODS-3. The mobile phase consisted of A:100% HPLC H2O, C:1% AcOH in MeOH, and D:100% MeOH. Gradient elution starting conditions were 65%A, 35%C followed by an increase of MeOH 60%D, 35%C, 5%A for 10 min. and holding this constant for 10 min., then returning to the initial starting conditions for 10 min. The standards FB1, FB2, and FB3 (Sigma, St. Louis, MO)were injected at 0.2 ng/μl for qualitative purposes.
Isolation of F. verticillioides from inoculated maize tissue
For initial inoculum preparation, a strain of F. verticillioides (M 2326) was grown on PDA in Petri plates (15 × 100 mm) for one week at room temperature. Following growth of the fungus, another set of plates with PDA was prepared containing 10 sterile toothpicks laying on the surface of each plate. The plates were then inoculated with a 2 cm agar plug of the F. verticillioides cultures from the actively growing margins. The plates were stored at room temperature for two weeks to allow the fungus to completely colonize the toothpicks. Control plates containing toothpicks uninoculated were also prepared and left at room temperature. A preliminary study showed that the F. verticillioides could be routinely reisolated from the toothpicks on PDA following colonization after two weeks.
To establish the maize plants in the greenhouse, four seeds (Terral TV 2100) were sown 2 cm deep in each of 20 pots (20 × 100 cm) containing 1:1 presterilized Promix ™ (Premier Horticulture Ltd., Dorval, Canada) and vermiculite. After two weeks, plants were culled to one per pot. Pots were watered daily as required to maintain soil moisture above 50% of available moisture capacity. Growth temperatures ranged from 20°C to 28°C during the investigation in the greenhouse. Twenty-eight days after planting, the toothpicks containing strain M-2326 were used to inoculate 15 replicate plants and uninfested toothpicks were inserted into 5 replicate control plants. Each toothpick was inserted into the main stem 6 cm from the soil surface, equal lengths were left on both sides of the stem. The experimental design was a randomized complete block. After two weeks, the 20 plants were sampled and examined for survival of F. verticillioides within the tissues. From each plant a 5 cm long section containing the inoculation point as the midpoint was removed. Measuring from the point of inoculation two 1-cm tissue samples were then collected. One sample was used to attempt cultural isolation and identification of F. verticillioides and the second was used for PCR determination of the fungus using primer set B. A replicate experiment (test 2) using same methods was conducted using 15 plants for inoculation and 5 as controls. Environmental conditions in the second study were the same as the first.
For cultural isolation attempts, each 1 cm stem piece was placed into 0.524% sodium hypoclorite solution (w/v) for 3 min. The pieces were then plated onto PDA and left at room temperature for two weeks. All fungi growing from the tissues were subcultured onto PDA and identified by comparing with the original culture using macroscopic and microscopic characters.
For PCR comparison, 0.15 g of tissues from the inoculation point from each of the 20 plants was prepared and assayed using the methods described above for DNA isolation and PCR. A preliminary study showed that the additional tissue was necessary to ensure adequate amounts of ng/μL of fungal DNA were extracted.
3. Results
Four primer sets (A, B, C, and D) selected from the polyketide synthase (PKS) gene sequence (FUM 1) were tested for their specificity to F. verticillioides (Table 1) strains and other fungal taxa. The PCR primer set A and C amplification products were inconsistent or absent when compared between the different Fusarium spp. strains and other fungi tested (Data not shown). Only set B containing primers, Fum 5F + Fum 6R, consistently amplified a similar sized single fragment (419 bp: Figure 1) from the DNA strain extracts of F. verticillioides. Strain M-3441 did not react using primer set B and fumonisin could not be detected from the liquid or rice media (Table 2). Strain M-5496 was amplified using both primer sets B and D, but fumonisin was not detected using either medium. The strain was reexamined and confirmed as F verticillioides that was sent from Penn State and later confirmed using methods previously discussed [48]. When the 10 strains of F. proliferatum were tested, a similar PCR product was observed in 83% of the isolates analyzed with primer set B. Primer set D, Fum1F + Fum6R, amplified 83% of the F. verticillioides isolates evaluated in the study. However, no isolates of F. proliferatum (Table 2) were amplified using primer set D.
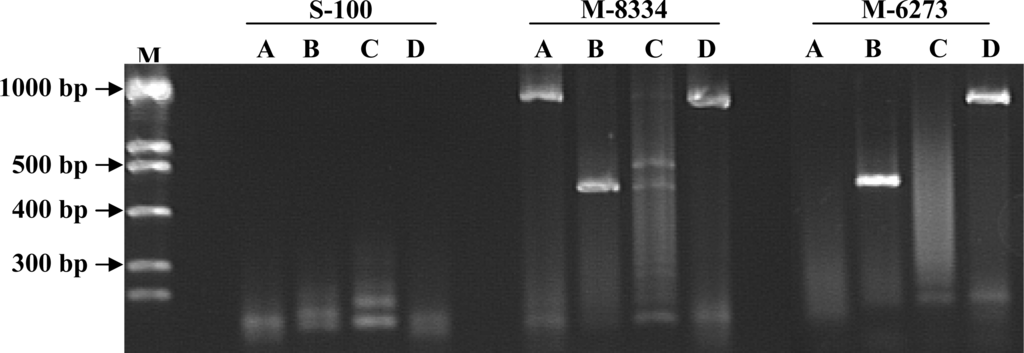
Figure 1.
Amplification of DNA from Fusarium solani (S-100) and Fusarium verticillioides (M-8334, M-6273) isolates using four primer sets. A, B, C, and D refer to primer sets Primer set B (Fum 5F and Fum6R) consistently produced 419 bp bands for F. verticillioides and the majority of F. proliferatum isolates tested.
Fumonisin production on rice medium occurred for 92% of F. verticillioides strains compared with 37.5% from liquid medium used for growing the strain tested in this study. For F. proliferatum strains, no fumonisin was produced in liquid medium [45], but the toxin was detected in 57% of the strains on rice [46]. When fumonisin was formed on rice, PCR amplification always occurred using primer set B for F. verticillioides, but were inconsistent for F. proliferatum. Nepal isolate M-5496 of F. verticillioides was amplified with primer sets B and D, but fumonisin was not detected on liquid or rice media. Also, fumonisin did not occur for the other Fusarium spp. or for the fungi used as negative controls.
Sensitivity of the PCR reactions was evaluated during the investigation. Decreasing amounts of the genomic DNA of F. verticillioides (M-2326) were used in order to determine the minimum amount of input DNA required to produce detectable product with primer set B. Using 10 replicate samples per DNA concentration, amplification products of the primer set B were visible on gels from 30 ng/μl, 20 ng/μl, 10 ng/μl, 1 ng/μl, and 100 pg/μl of F. verticillioides (M-2326) DNA. At 10 pg/μl and 1pg/μl, PCR detectable products were not consistently observed across the 10 replicate reactions per primer set (Data not shown).
In the two replicate greenhouse studies isolation frequencies of F. verticillioides from 1-cm maize tissues collected two weeks after inoculation ranged from 60% in Test 1 and 53% in Test 2. No isolations occurred from any of the control samples following both tests. Using primer set B on the duplicate set of tissues, amplification was observed from 67% in Test 1 and 72% in Test 2. No PCR product was observed from any of the control tissue samples. In an attempt to improve PCR efficiency the DNA samples from both tests were diluted 10 x and 100 x in nanopure water and PCR amplification products occurred in 73% of pieces from Test 1 and 80% in Test 2 at both concentrations. No amplification occurred from the control plant tissues. These findings suggest that the total genomic DNA samples contain some PCR inhibitors.the total genomic DNA samples contain some PCR inhibitors.
4. Discussion
This study further demonstrates that PCR analysis is an effective and fast way to detect fumonisin producing F. verticillioides strains. PCR analysis had better sensitivity compared to conventional cultural approaches with and without maize tissue. Even though the emphasis of the research was on the development of a rapid detection method for F. verticillioides to use in maize resistance studies, isolates of F. proliferatum could be amplified. In addition, all isolates that produced the toxin in vitro were positive in PCR analysis for F. verticillioides. Isolate M-5496 from Nepal was amplified using primer sets B and D, but fumonisin was not detected on either selective media. Previous studies reported that fumonisin was not formed by M-5496 in culture [49,50]. Since PCR amplification occurred for M-5496, the FUM 1 gene must be present enabling the primers to amplify the DNA.
The results of this study indicate the presence of genetic variations among strains of Fusarium used in this study. Of the 24 isolates of F. verticillioides, only M-5496 and M-3441 yielded a PCR product with primer sets B or D, did not synthesize toxin in vitro on liquid or rice media. These results were expected for M-5496 but uncertain why M-3441 did not produce fumonisin. Primer set D was less effective for detection of F. verticillioides and did not amplify DNA of F. proliferatum strains. In this study, the polyketide synthase gene FUM1 region was selected since it was thought to have unique primer binding sites for distinguishing fumonisin forming fungi and development of primers could target toxin production. Previously, sequences from internal transcribed spacer (ITS) regions ITS1 and ITS2 were successfully used for design of specific primers to identify select Fusarium spp. [25,42,43,51,52]. However, none of the PCR assays were capable of differentiating fumonisin producing from non-producing strains. More recently, Patiňo et al. [44] successfully designed primers from the IGS region that identifies fumonisin producing isolates from non-producing ones of F. verticillioides. Also, Bezuidonhout et al. [53] developed a multiplex PCR detection method for identification of fumonisin forming fungi. However, further study is needed to confirm these findings from the previous research.
Differing from those studies, the current investigation evaluated additional representatives from all fungal phyla, including the fungal-like Oomycota, to insure specificity to F. verticillioides. Multiplex PCR is a newly developed approach for evaluating fumonisin forming Fusarium spp. Unfortunately, optimization of PCR equipment may vary per laboratories and specialized real-time PCR equipment for multiplex may not be available at each location. Therefore use of species specific primers provides the most simplified method to enable many laboratories to utilize these diagnostic capabilities.
Contrary to the results using primer set B for F. verticillioides, PCR positive frequency of F. proliferatum strains was 80%. These results indicate that genetic differences in this species may be the reason for the reduced amplification rate for F. proliferatum. Diversity and genetic variation are well known among Fusarium species. These species are heterothallic and have dimictic mating systems [54]. In some Fusarium species, even entire chromosomes can be deleted [55]. Consequently, genetic variation in the genes dedicated to toxin production has been frequently observed [55,56]. For example, multiple mutants blocked at various steps of fumonisin biosynthesis were identified in natural populations or generated by UV mutagenesis [57]. Isolates of F. proliferatum were reported to produce fumonisin [55,58]. However, none of the strains included in this study produced fumonisin in vitro based on LC/ECI/MS analysis. The cluster involved in fumonisin biosynthesis is composed of 15 genes [17]. It is predicted that some deletions, transposon insertions, or mutations exist in some of the genes in isolates used in this study, consequently resulting in deficiency in fumonisin production. Interestingly, PCR product was obtained from 80% of the F. proliferatum isolates in this study, suggesting that intact or portion of the FUM 1 gene is present in the PCR-positive isolates. More intensive genetic analysis of these isolates is needed to verify this belief.
The results of the in vitro work determined that fumonisin production on defined liquid medium [45] was not consistent compared to strains of F. verticillioides grown on rice [46]. The results using the liquid medium from this study were not consistent with previous research. Only 33% of F. verticillioides isolates produced fumonisin on the liquid medium and all of them were also PCR-positive using primer set B. In the previous study, toxin production for one F. verticillioides (mating type A) strain was detected within 75 hrs after inoculation. Since a single strain was evaluated in that study it could not be determined if isolate variability in fumonisin formation may result from different nutrient or growth condition requirements for the isolates. Several factors can affect in vitro production of fumonisin including moderate water activity, limited nitrogen and pH levels [59]. The defined medium developed by Shim and Woloshuk [45] and used in this study has a pH of 5.9, but Miller [59] reported that a pH of 2 is optimizal for fumonisin synthesis in vitro. The higher pH of the liquid medium might have been an important or limiting factor in fumonisin production during this investigation. Therefore, the fumonisin data from the liquid medium is not considered reliable and rice data should be used.
The greenhouse studies confirm that primer set B could be successfully used for detection of F. verticillioides from maize tissue. The advantage of PCR over culturing has been discussed previously including speed of detection and amplification of the fungus when it is at low levels in tissues. The 100 picogram detection level verified during this investigation indicates that low levels of F. verticillioides DNA from the tissues can be consistently amplified.
In conclusion, primer set B consistently produced a product of bp 419 that was effective for identification of F. verticillioides associated with maize tissues. No PCR products were produced within the closely and distantly related taxa. Primer set D did not consistent identify all fumonisin forming strains when compared, but only amplified ones that produced the toxin. Since primer set B was shown to be effective for identification of F. verticillioides isolate M-2326 from maize tissue in the greenhouse studies, future research can now be conducted to determine cultivar resistance by evaluating colonization or movement of F. verticillioides and potentially other fumonisin forming fungi within maize plants.
References and Notes
- White, DG. Compendium of corn diseases, 3rd ed; American Phytopathological Society: St. Paul, MN, 1999. [Google Scholar]
- Abbas, HK; Williams, WP; Windham, GL; Pringle, HC; Xie, W; Shier, WT. Aflatoxin and fumonisin contamination of commercial corn (Zea mays) hybrids in Mississippi. J Agric Food Chem 2002, 50, 5246–5254. [Google Scholar]
- Leslie, JF. Mating populations in Gibberella fujikuroi (Fusarium section Liseola). Phytopathology 1991, 81, 1058–1060. [Google Scholar]
- Britz, HT; Coutinho, A; Wingfield, MJ; Marasas, WFO; Gordon, TR; Leslie, JF. Fusarium subglutinans f. sp. pini represents a distinct mating population in the Gibberella fujikuroi species complex. Appl Environ Microbiol 1999, 65, 198–1201. [Google Scholar]
- Leslie, JF. Interfertility of two mating populations in the Gibberella fujikuroi species complex. Euro J Plant Pathol 2004, 110, 611–618. [Google Scholar]
- Leslie, JF. Gibberella fujikuroi: available populations and variable traits. Can J Botany 1995, 73(Suppl. 1), S282–S291. [Google Scholar]
- Munkvold, GP; Hellmich, RL; Showers, WB. Reduced Fusarium ear and symptomless infection in kernels of maize genetically engineered for European cornborer resistance. Phytopathology 1997, 87, 1071–1077. [Google Scholar]
- Foley, DC. Systematic infection of corn by Fusarium moniliforme. Phytopathology 1999, 68, 1331–1335. [Google Scholar]
- Bacon, CW; Hinton, DM; Richardson, MD. A corn seedling assay for resistance to Fusarium moniliforme. Plant Dis 1994, 78, 302–305. [Google Scholar]
- Bezuidonhout, SC; Gelderblom, WCA; Gorst-Allman, CP; Horak, RM; Marasas, WFO. Structure elucidation of fumonisins, mycotoxins from Fusarium moniliforme. J Chem Soc Chem Commun 1988, 52, 743–745. [Google Scholar]
- Gelderblom, WCA; Jaskiewicz, K; Marasas, WFO; Thiel, PG; Horak, MJ; Vleggar, R; Kriek, NPK. Fumonisins - novel mycotoxins with cancer promoting activity produced by Fusarium moniliforme. Appl Environ Microbiol 1988, 54, 1806–1811. [Google Scholar]
- Riley, TR; Norred, WP; Bacon, CW. Fungal toxins in food: recent concerns. Annu Rev Nutr 1993, 13, 167–189. [Google Scholar]
- Abbas, HKT; Tanaka, T; Shier, WT. Biological activities of synthetic analogs of Alternaria alternata AAL-toxin and fumonisin in plant and mammalian cell cultures. Phytochemistry 1995, 40, 1681–1689. [Google Scholar]
- Shier, WT; Abbas, HK; Badria, FA. Complete structure of the sphingosine analog mycotoxins fumonisin B1 and AAL-toxin TA:absolute configuration of the side chains. Tetrahedron Lett 1995, 36, 1571–1574. [Google Scholar]
- Proctor, RH; Desjardins, AE; Plattner, RD; Hohn, TM. A polyketide synthase gene required for biosynthesis of fumonisin mycotoxins in Gibberella fujikuroi mating population A. Fungal Genet Biol 1999, 27, 100–112. [Google Scholar]
- Desjardins, AE; Munkvold, GP; Plattner, RD; Proctor, RH. Fum1: a gene required for fumonisin biosynthesis but not for maize ear rot and ear infection by Gibberella moniliforme in field tests. Mol Plant Microbe In 2002, 15, 1157–1164. [Google Scholar]
- Proctor, RH; Brown, DW; Plattner, RD; Desjardins, AE. Co-expression of 15 contiguous genes delineates a fumonisin biosynthetic gene cluster in Gibberella moniliformis. Fungal Genet Biol 2003, 38, 237–249. [Google Scholar]
- CAST (Council for Agriculture Science and Technology), Mycotoxins risks in plant, animal, and human systems; CAST, Task Force Report 139: Ames, IA, 2003.
- NTP (National Toxicology Program), Toxicology and carcinogenesis studies on fumonisin B1 in F344/N rats and B6CF1 mice (feed studies); Technical Report Series, n.496. NIH Publication No. 99-3955; U.S. Department of Health and Human Services, National Institutes of Health: Research Triangle Park, NC, 1999.
- Nelson, PE; Desjardins, AE; Plattner, RD. Fumonisins, mycotoxins produced by Fusarium species: Biology, chemistry and significance. Annu Rev Phytopathol 1993, 31, 233–252. [Google Scholar]
- Thiel, PG; Marasas, WFO; Sydenham, EW; Shepard, GS; Gelderblom, WCA; Nieuwenhuis, JJ. Survey of fumonisin production by Fusarium species. Appl Environ Microbiol 1991, 57, 1089–1093. [Google Scholar]
- Rheeder, JP; Marasas, WFO; Vismer, HF. Production of fumonisin analogs by Fusarium species. Appl Environ Microbiol 2002, 68, 2101–2105. [Google Scholar]
- Headrick, JM; Pataky, JK. Maternal influence on the resistance of sweet corn lines to kernel infection by Fusarium moniliforme. Phytopathology 1991, 81, 268–274. [Google Scholar]
- Booth, C. The genus Fusarium; Commonwealth Mycological Institute: Kew, United Kingdom, 1971. [Google Scholar]
- Grimm, C; Geisen, R. A PCR-ELISA for the detection of potential fumonisin producing Fusarium species. Lett Appl Microbiol 1988, 26, 456–462. [Google Scholar]
- Innis, MA; Gelfand, DH; Sninsky, JJ; White, TJ. PCR Protocols: A guide to method and applications; Innis, D, Gelfand, H, Sninsky, JJ, Eds.; Academic Press, Inc: N.Y., 1990. [Google Scholar]
- Bruns, TD; White, TJ; Taylor, JW. Fungal molecular systematics. Annu Rev Ecol and Syst 1991, 22, 525–564. [Google Scholar]
- Bateman, GL; Ward, E; Antoniwi, JF. Identification of Gaeumannomycos graminis var. tritici and G. graminis var. avenae using a DNA probe and non-molecular methods. Mycol Res 1992, 96, 737–742. [Google Scholar]
- Zhang, AW; Riccioni, GL; Chen, L; Ma, WD; Petersen, WL. Using PCR to distinguish Diaporthe phaseolorum and Phompsis longicolla from other soybean fungal pathogens and to detect them in soybean tissues. Plant Dis 1997, 81, 1143–1149. [Google Scholar]
- Beck, JJ; Ligon, JM. Polymerase chain reaction assays for the detection of Stagonospora nodorum and Septoria tritici in wheat. Phytopathology 1995, 85, 319–324. [Google Scholar]
- Schilling, AG; Moller, EM; Geiger, HH. Polymerase chain reaction-based assays for the detection for species-specific detection of Fusarium culmorum, F. graminearum, and F. avenaceum. Phytopathology 1996, 86, 515–522. [Google Scholar]
- Guthrie, PAI; Magill, GW; Fredericksen, RA; Oduody, GN. Random amplified polyorphic DNA markers: a system for identifying and differentiating isolates of Colletotrichum graminicola. Phytopathology 1992, 82, 832–835. [Google Scholar]
- Hornby, D; Bateman, G. Take-all of Cereals; Home-Grown Cereals Authority Research Review no. 2. Home-Grown Cereals Authority: London, 1991. [Google Scholar]
- Williams, JGK; Kubelik, AR; Rafalski, JA; Tingey, SV. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 1990, 18, 615–643. [Google Scholar]
- White, TJ; Bruns, T; Lee, S; Taylor, J. Amplification of direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols: A guide to method and applications. Innis, D, Gelfand, H, Sninsky, JJ, Eds.; Academic Press, Inc: New York, 1990. [Google Scholar]
- O’Donnell, K; Cigelnik, E. Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol and Phylogenet Evol 1997, 7, 103–116. [Google Scholar]
- O’Donnell, K; Cigelnik, E; Nirenberg, H. Molecular systematics and phylogeography of the Gibberella fujikuroi species complex. Mycologia 1998, 90, 465–493. [Google Scholar]
- Bayman, P; Cotty, PJ. Genetic diversity in Aspergillus flavus: association with aflatoxin production and morphology. Can J Botany 1993, 71, 23–34. [Google Scholar]
- Jones, MJ; Dunkle, LD. Analysis of Cochliobulus carbonum races by PCR amplification with arbitrary and gene-specific primers. Phytopathology 1993, 83, 366–370. [Google Scholar]
- González-Jaén, MT; Mirete, S; Patiňo, B; López-Errasquin, E; Vázquez, C. Genetic markers for the analysis of variability and for production of specific diagnotic sequences in fumonisin-producing strains of Fusarium verticillioides. Euro J of Plant Path 2004, 110, 525–532. [Google Scholar]
- Waalwijk, C; van der Heide, R; deVries, I; van der Lee, T; Schoen, C; Costrel-de Corainville, G; Häuser-Hahn, I; Kastelein, P; Köhl, J; Lonnet, P; Demarquet, T; Kema, GHJ. Quantitative detection of Fusarium species in wheat using TaqMan. Eur J Plant Pathol 2004, 110, 481–494. [Google Scholar]
- Bluhm, BM; Flaherty, JE; Cousin, MA; Woloshuk, CP. Multiplex polymerase chain reaction assay for the differential detection of trichothecene- and fumonisin-producing species of Fusarium in cornmeal. J Food Protect 2002, 65, 1955–1961. [Google Scholar]
- Bluhm, BM; Cousin, MA; Woloshuk, CP. Multiplex real-time PCR detection of fumonisin-producing and trichothecene-producing groups of Fusarium species. J Food Protect 2004, 67, 536–543. [Google Scholar]
- Patiňo, B; Mirete, S; González-Jaén, MT; Mulé, EG; Vázquez, C. PCR detection assay of fumonisin-producing Fusarium verticillioides stains. J. Food Protect 2004, 67, 1278–1283. [Google Scholar]
- Shim, WB; Woloshuk, CP. Nitrogen repression of fumonisin B1 biosynthesis in Gibberella fujikuroi. FEMS Microbiol Lett 1999, 177, 109–116. [Google Scholar]
- Abbas, HK; Cartwright, RD; Xie, W; Mirocha, CJ; Richard, JL; Dvorak, TJ; Sciumbato, GL; Shier, WT. Mycotoxin production by Fusarium proliferatum isolates from rice with Fusarium sheath rot disease. Mycopathologia 1999, 147, 97–104. [Google Scholar]
- Plattner, RD. HPLC/MS Analysis of Fusarium mycotoxins, fumonisins and deoxynivalenol. Nat Toxins 1999, 7, 365–370. [Google Scholar]
- Geiser, DM; Jiménez-Gasco, MD; Kang, S; Makalowska, I; Veeraraghavan, N; Ward, TJ; Zhang, N; Kuldau, GA; O’Donnell, K. Fusarium-ID v. 1.0: A DNA sequence database for identifying Fusarium. Europ J Plant Pathol 2004, 110, 473–479. [Google Scholar]
- Nelson, PE; Plattner, RD; Shackelford, DD; Desjardins, AE. Production of fumonisin by Fusarium moniliforme strains from various substrates and geographic areas. Appl Environ Micobiol 1991, 57, 2410–2412. [Google Scholar]
- Falasconi, M; Gobbi, E; Pardo, M; Dell Torre, M; Bresciani, A; Sberveglieri, G. Sensor Actuat. B-Chem. Detection of toxigenic strains of Fusarium verticillioides in corn by electronic olfactory system. 2005, 108, 250–257. [Google Scholar]
- O’Donnell, K. Ribosomal DNA internal transcribed spacers are highly divergent in the phytopathogenic ascomycete Fusarium sambusinum (Gibberella pulicaris). Curr Genet 1992, 22, 213–220. [Google Scholar]
- O’Donnell, K; Gray, LE. Phylogenetic relationships of the soybean sudden death syndrome pathogen Fusarium solani f. sp. solani inferred from rDNA sequence data and PCR primers for its identification. Mol Plant Microbe In 1995, 5, 709–716. [Google Scholar]
- Bezuidonhout, SC; Prinsloo, M; Vanderwalt, AM. Multiplex PCR-based detection of potential fusarium producing Fusarium in traditional African vegetables. Environ Toxicol 2006, 21, 360–366. [Google Scholar]
- Kerenyi, Z; Keller, K; Hornok, L; Leslie, JF. Molecular standardization of mating type terminology in the Gibberella fujkuroi species complex. Appl Environ Micobiol 1999, 65, 4071–4076. [Google Scholar]
- Desjardins, AE; Proctor, RH. In Fusarium: Biochemistry and Genetics of Fusarium Toxins; Paul E. Nelson Memorial Symposium; Summerall, BA, Leslie, JF, Backhouse, D, Bryden, WL, Burgess, LW, Eds.; The American Phytopathological Society: St. Paul, MN, 2001. [Google Scholar]
- Chulze, SN; Ramirez, ML; Torres, A; Leslie, JF. Genetic variation in Fusarium section Liseola from no-till maize in Argentina. Appl Environ Microbiol 2000, 66, 5312–5315. [Google Scholar]
- Desjardins, AE; Plattner, RD; Proctor, RH. Fumonisins in food: Genetic and biochemical aspects of fumonisin production; Jackson, LS, Ed.; Plenum Press: NY, 1996. [Google Scholar]
- Mule, G; Susca, A; Stea, G; Moretti, A. Specific detection of the toxigenic species Fusarium proliferatum and F. oxysporum from asparagus plants using primers based on calmodulin gene sequences. FEMS Microbiol Lett 2004, 230, 235–240. [Google Scholar]
- Miller, SD. Factors that affect the occurrence of fumonisin. Environ Health Persp 2001, 109(Suppl.-2), 321–324. [Google Scholar]