Analysis of Mechanisms of T-2 Toxin Toxicity Using Yeast DNA Microarrays
Abstract
1. Introduction
2. Results and Discussion
2.1. Conditions for T-2 toxin treatment
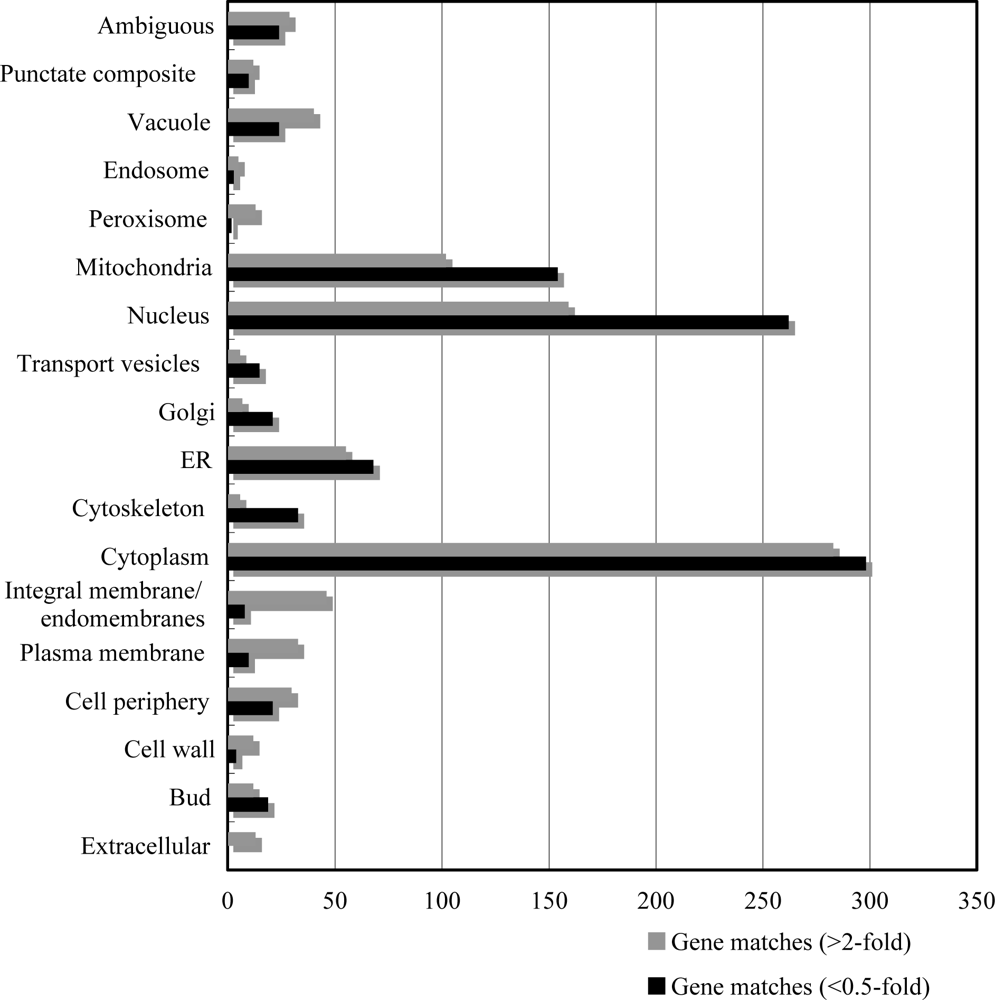
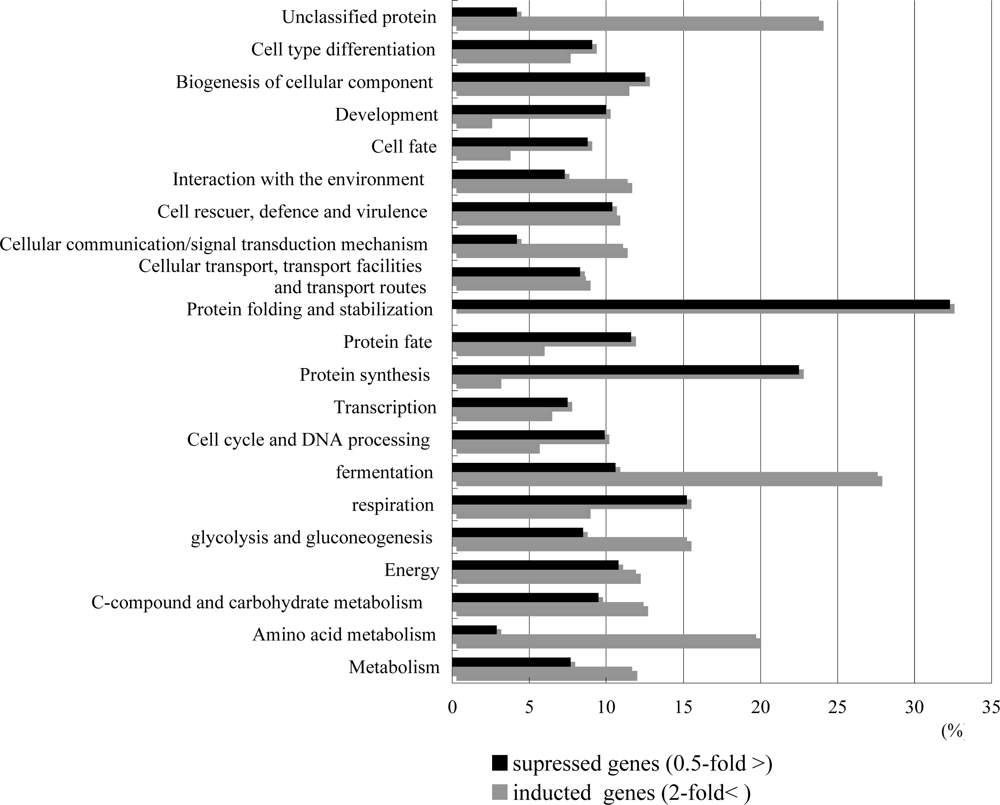
2.2. Overview of T-2 toxin induced genes, cellular location and functional distribution
2.3. T-2 toxin causes the oxidative stress, and causes the energy scarcity in yeast
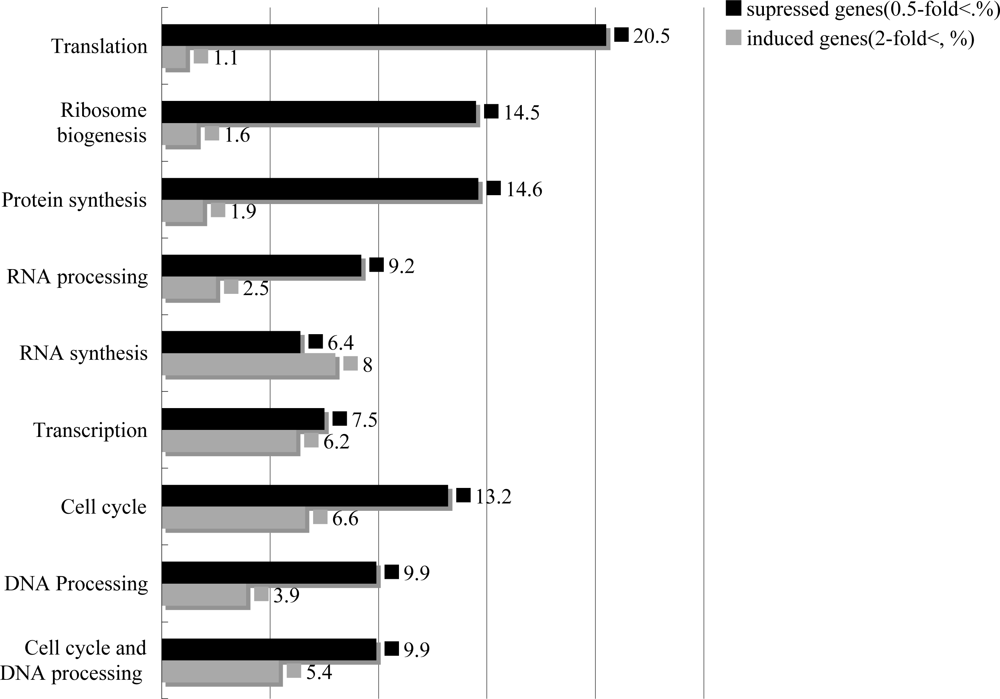
2.4. T-2 toxin induced cell arrest, decreased DNA repair system, and inhibited the synthesis and the processing of the ribosome
2.5. Mutagenicity in the T-2 toxin
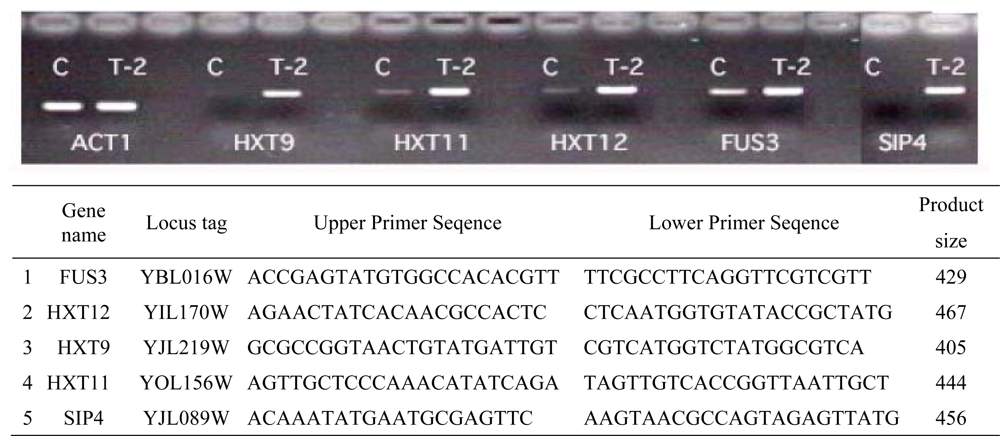
2.6. Biomarker Genes for T-2 toxin and cluster analysis
3. Experimental Section
3.1. Strains, growth conditions and T-2 toxin treatment
3.2. DNA microarray analysis
4. Conclusions
Acknowledgments
References and Notes
- Sugita-Konishi, Y. Toxicity and control trichothecene mycotoxins. Mycotoxins 2008, 58, 23–28. [Google Scholar]
- Sehata, S; Kiyosawa, N; Makino, T; Atsumi, F; Ito, K; Yamoto, T; Teranishi, M; Baba, Y; Uetsuka, K; Nakayama, H; Doi, K; Author, A. Morphological and microarray analysis of T-2 toxin-induced rat fetal brain lesion. Food Chem. Toxicol 2004, 42, 1727–1736. [Google Scholar]
- Ahmadi, K; Riazipour, M. Effects of T-2 toxin on cytokine production by mice peritoneal macrophages and lymph node T-cells. Iran J. Immunol 2008, 5, 177–180. [Google Scholar]
- Fu, YT; Lin, WG; BaoCheng, Z; Quan, G. The effect of T-2 toxin on IL-1beta and IL-6 secretion in human fetal chondrocytes. Int. Orthop 2001, 25, 199–201. [Google Scholar]
- Iwahashi, Y; Hosoda, H; Park, JH; Lee, JH; Suzuki, Y; Kitagawa, E; Murata, SM; Jwa, NS; Gu, MB; Iwahashi, H. Mechanisms of patulin toxicity under conditions that inhibit yeast growth. J. Agric. Food Chem 2006, 54, 1936–1942. [Google Scholar]
- Iwahashi, H; Kitagawa, E; Suzuki, Y; Ueda, Y; Ishizawa, YH; Nobumasa, H; Kuboki, Y; Hosoda, H; Iwahashi, Y. Evaluation of toxicity of the mycotoxin citrinin using yeast ORF DNA microarray and Oligo DNA microarray. BMC Genomics 2007, 5, 95. [Google Scholar]
- Sehata, S; Kiyosawa, N; Sakuma, K; Ito, K; Yamoto, T; Teranishi, M; Uetsuka, K; Nakayama, H; Doi, K. Gene expression profiles in pregnant rats treated with T-2 toxin. Exp. Toxicol. Pathol 2004, 55, 357–366. [Google Scholar]
- Stephanie, R; Jacqueline, K; Hans-Joachim, S. Transcriptional activators Cat8 and Sip4 discriminate between sequence variants of the carbon source-responsive promoter element in the yeast Saccharomyces cerevisiae. Curr. Genet 2004, 45, 121–128. [Google Scholar]
- Kahn, BB; Alquier, T; Carling, D; Hardie, DG. AMP-activated protein kinase: Ancient energy gauge provides clues to modern understanding of metabolism. Cell Metab 2005, 1, 15–25. [Google Scholar]
- Celenza, JL; Carlson, M. Mutational analysis of the Saccharomyces cerevisiae SNF1 protein kinase and evidence for functional interaction with the SNF4 protein. Mol. Cell Biol 1989, 9, 5034–5044. [Google Scholar]
- Jiang, R; Carlson, M. Glucose regulates protein interactions within the yeast SNF1 protein kinase complex. Genes Dev 1996, 10, 3105–3115. [Google Scholar]
- Hardie, DG; Carling, D; Carlson, M. The AMP-activated/SNF1 protein kinase subfamily: metabolic sensors of the eukaryotic cell? Annu. Rev. Biochem 1998, 67, 821–855. [Google Scholar]
- Kahn, BB; Alquier, T; Carling, D; Hardie, DG. AMP-activated protein kinase: Ancient energy gauge provides clues to modern understanding of metabolism. Cell Metab 2005, 1, 15–25. [Google Scholar]
- Hoffman, M; Chiang, HL. Isolation of degradation-deficient mutants defective in the targeting of fructose-1,6-bisphosphatase into the vacuole for degradation in Saccharomyces cerevisiae. Genetics 1996, 143, 1555–1566. [Google Scholar]
- Hung, GC; Brown, CR; Wolfe, AB; Liu, J; Chiang, HL. Degradation of the gluconeogenic enzymes fructose-1,6-bisphosphatase and malate dehydrogenase is mediated by distinct proteolytic pathways and signaling events. J. Biol. Chem 2004, 279, 49138–49150. [Google Scholar]
- Santt, O; Pfirrmann, T; Braun, B; Juretschke, J; Kimmig, P; Scheel, H; Hofmann, K; Thumm, M; Wolf, DH. The yeast GID complex, a novel ubiquitin ligase (E3) involved in the regulation of carbohydrate metabolism. Mol. Biol. Cell 2008, 19, 3323–3333. [Google Scholar]
- Fujimura, HA. The DAC2/FUS3 protein kinase is not essential for transcriptional activation of the mating pheromone response pathway in Saccharomyces cerevisiae. Mol. Gen. Genet 1992, 235, 450–452. [Google Scholar]
- Elion, EA; Satterberg, B; Kranz, JE. FUS3 phosphorylates multiple components of the mating signal transduction cascade: evidence for STE12 and FAR1. Mol. Biol. Cell 1993, 4, 495–510. [Google Scholar]
- Butty, AC; Pryciak, PM; Huang, LS; Herskowitz, I; Peter, M. The role of Far1p in linking the heterotrimeric G protein to polarity establishment proteins during yeast mating. Science 1998, 282, 1511–1516. [Google Scholar]
- Boulton, TG; Yancopoulos, GD; Gregory, JS; Slaughter, C; Moomaw, C; Hsu, J; Cobb, MH. An insulin-stimulated protein kinase similar to yeast kinases involved in cell cycle control. Science 1990, 249, 64–67. [Google Scholar]
- Mendenhall, MD. Cyclin-dependent kinase inhibitors of Saccharomyces cerevisiae and Schizosaccharomyces pombe. Curr. Top Microbiol. Immunol 1998, 227, 1–24. [Google Scholar]
- Peter, M; Herskowitz, I. Direct inhibition of the yeast cyclin-dependent kinase Cdc28-Cln by Far1. Science 1994, 265, 1228–1231. [Google Scholar]
- Chang, F; Herskowitz, I. Identification of a gene necessary for cell cycle arrest by a negative growth factor of yeast: FAR1 is an inhibitor of a G1 cyclin, CLN2. Cell 1990, 63, 999–1011. [Google Scholar]
- Henchoz, S; Chi, Y; Catarin, B; Herskowitz, I; Deshaies, RJ; Peter, M. Phosphorylation- and ubiquitin-dependent degradation of the cyclin-dependent kinase inhibitor Far1p in budding yeast. Genes Dev 1997, 11, 3046–3060. [Google Scholar]
- Peter, M; Gartner, A; Horecka, J; Ammerer, G; Herskowitz, I. FAR1 links the signal transduction pathway to the cell cycle machinery in yeast. Cell 1993, 73, 747–760. [Google Scholar]
- Butty, AC; Pryciak, PM; Huang, LS; Herskowitz, I; Peter, M. The role of Far1p in linking the heterotrimeric G protein to polarity establishment proteins during yeast mating. Science 1998, 282, 1511–1516. [Google Scholar]
- Symington, LS. Role of RAD52 epistasis group genes in homologous recombination and double-strand break repair. Microbiol. Mol. Biol. Rev 2002, 66, 630–670. [Google Scholar]
- Kitagawa, E; Momose, Y; Iwahashi, H. Correlation of the structures of agricultural fungicides to gene expression in Saccharomyces cerevisiae upon exposure to toxic dose. Environ. Sci. Technol 2003, 37, 2788–2793. [Google Scholar]
- Murata, Y; Watanabe, T; Sato, M; Momose, Y; Nakahara, T; Oka, S; Iwahashi, H. DMSO exposure facilitates phospholipid biosynthesis and cellular membrane proliferation in yeast cells. J. Biol. Chem 2003, 278, 33185–33193. [Google Scholar]
- Momose, Y; Iwahashi, H. Bioassay of cadmium using a DNA microarray: genome wide expression patterns of Saccharomyces cerevisiae response to cadmium. Environ. Toxicol. Chem 2001, 20, 2533–2360. [Google Scholar]
- Kitagawa, E; Takahashi, J; Momose, Y; Iwahashi, H. Effects of the pesticide thiuram: Genome-wide screening of indicator genes by yeast DNA microarray. Environ. Sci. Technol 2002, 36, 3908–3915. [Google Scholar]
- Parveen, M; Hasan, K; Takahashi, J; Murata, Y; Kitagawa, E; Kodama, O; Iwahashi, H. Response of Saccharomyces cerevisiae to a monoterpene: Evaluation of antifungal potential by DNA microarray analysis. J. Antimicrob. Chemother 2004, 54, 46–55. [Google Scholar]
- http://www.silicongenetics.com/cgi/SiG.cgi/index.smf; accessed October 2007.
- Munich Information Center for Protein Sequences; http://mips.gsf.de/; accessed October 2007.
- Yeast Genome Database; http://www.yeastgenome.org/; accessed October 2007.
- Iwahashi, H; Odani, M; Ishidou, E; Kitagawa, E. Adaptation of Saccharomyces cerevisiae to high hydrostatic pressure causing growth inhibition. FEBS Lett 2005, 579, 2847–2852. [Google Scholar]
- Sehata, S; Kiyosawa, N; Atsumi, F; Ito, K; Yamoto, T; Teranishi, M; Uetsuka, K; Nakayama, H; Doi, K. Microarray analysis of T-2 toxin-induced liver, placenta and fetal liver lesions in pregnant rats. Exp. Toxicol. Pathol 2005, 57, 15–28. [Google Scholar]

| Systematic | Normalized | Common | Description |
|---|---|---|---|
| YJL089W | 15.7 | SIP4 | interacts with SNF1 protein kinase |
| YOL156W | 15.7 | HXT11 | Hxt family protein with intrinsic hexose transport activity |
| YJL219W | 12.8 | HXT9 | Hxt family protein with intrinsic hexose transport activity |
| YNL279W | 12.4 | PRM1 | similarity to S. pombe coiled-coil protein of unknown function |
| YOL052C-A | 12.0 | DDR2 | heat shock protein DDRA2 |
| YHR137W | 11.4 | ARO9 | aromatic amino acid aminotransferase II |
| YIL170W | 10.7 | HXT12 | strong similarity to sugar transport proteins |
| YFL058W | 9.4 | THI5 | pyrimidine biosynthesis protein |
| YBL016W | 8.4 | FUS3 | mitogen-activated protein kinase (MAP kinase) |
| YBR296C | 8.2 | PHO89 | Na+/phosphate co-transporter |
| YOR303W | 7.8 | CPA1 | arginine-specific carbamoylphosphate synthase, small chain |
| YOR100C | 7.5 | CRC1 | mitochondrial carnitine carrier |
| YMR017W | 7.5 | SPO20 | Dbf2p interacting protein |
| YGL255W | 7.4 | ZRT1 | zinc transporter I |
| YFL026W | 7.3 | STE2 | pheromone alpha-factor receptor |
| YHR018C | 7.3 | ARG4 | arginosuccinate lyase |
| YGR213C | 7.2 | RTA1 | involved in 7-aminocholesterol resistance |
| YHL021C | 6.9 | FMP12 | weak similarity to Pseudomonas gamma-butyrobetaine hydroxylase |
| YJR078W | 6.8 | BNA2 | tryptophan 2,3-dioxygenase |
| YAR020C | 6.7 | PAU7 | strong similarity to members of the Srp1p/Tip1p family |
| YJR156C | 6.6 | THI11 | thiamine regulated gene, homologous to nmt1a in S. pombe |
| YOR388C | 6.6 | FDH1 | strong similarity to H.polymorpha formate dehydrogenase |
| YDR380W | 6.5 | ARO10 | similarity to Pdc6p, Thi3p and to pyruvate decarboxylases |
| YPL280W | 6.5 | HSP32 | strong similarity to YMR322c and YDR533c |
| YOL119C | 6.3 | MCH4 | similarity to monocarboxylate transporter proteins |
| YOR222W | 6.3 | ODC2 | mitochondrial 2-oxodicarboxylate carrier |
| YJR109C | 6.2 | CPA2 | arginine-specific carbamoylphosphate synthase, large chain |
| YPL250C | 6.2 | ICY2 | interacting with the cytoskeleton |
| YJL088W | 6.2 | ARG3 | ornithine carbamoyltransferase |
| YIL117C | 6.2 | PRM5 | similarity to hypothetical protein YNL058c |
| YGR055W | 6.0 | MUP1 | high affinity methionine permease |
| YHR021W-A | 6.0 | ECM12 | probably involved in cell wall structure or biogenesis |
| YFL053W | 6.0 | DAK2 | dihydroxyacetone kinase |
| YCL055W | 5.9 | KAR4 | regulatory protein required for pheromone induction of karyogamy genes |
| YPL135W | 5.8 | ISU1 | strong similarity to nitrogen fixation protein (nifU) |
| YLR142W | 5.8 | PUT1 | proline oxidase |
| YMR096W | 5.7 | SNZ1 | stationary phase protein |
| YML042W | 5.7 | CAT2 | carnitine O-acetyltransferase |
| YPL223C | 5.7 | GRE1 | induced by osmotic stress |
| YMR159C | 5.7 | ATG16 | coiled-coil protein required for autophagy |
| YDL244W | 5.6 | THI13 | strong similarity to Thi5p, YJR156c, YNL332w and A. parasiticus, S. pombe nmt1 protein |
| YGR161C | 5.3 | RTS3 | hypothetical protein |
| YNL125C | 5.3 | ESBP6 | similarity to YKL221w and human X-linked PEST-containing transporter |
| YKL217W | 5.0 | JEN1 | Lactate and pyruvate permease |
| Systematic No. | Fold | Common name | Description | Function |
|---|---|---|---|---|
| C-compound and carbohydrate metabolism | ||||
| YOR388C | 6.6 | FDH1 | NADH regeneration | detoxification |
| YFL053W | 6.0 | DAK2 | dihydroxyacetone kinase | stress |
| YDL243C | 4.3 | AAD4 | strong similarity to aryl-alcohol dehydrogenase | oxidative stress |
| YFL056C | 4.0 | AAD6 | strong similarity to aryl-alcohol dehydrogenases | oxidative stress |
| YCR105W | 3.8 | ADH7 | NADP(H)-dependent alcohol dehydrogenase | oxidative stress |
| YFL014W | 3.7 | HSP12 | heat shock protein | heat shock |
| YFL057C | 3.1 | AAD16 | aryl-alcohol dehydrogenase | oxidative stress |
| YAL021C | 3.1 | CCR4 | transcriptional regulator | response to drug |
| YOR178C | 2.8 | GAC1 | ser/thr phosphoprotein phosphatase 1, regulatory chain | heat shock protein binding |
| YNL331C | 2.6 | AAD14 | strong similarity aryl-alcohol reductase | oxidative stress |
| YKL062W | 2.5 | MSN4 | transcriptional activator | oxidative stress |
| YDL066W | 2.5 | IDP1 | isocitrate dehydrogenase (NADP+), mitochondrial | NADPH production |
| YER073W | 2.4 | ALD5 | aldehyde dehydrogenase (NAD+), mitochondrial | oxidative stress |
| YML070W | 2.3 | DAK1 | dihydroxyacetone kinase, induced in high salt | response to stress |
| YGR019W | 2.3 | UGA1 | 4-aminobutyrate aminotransferase | oxidative stress |
| Glycolysis and glyconeogenesis | ||||
| YJL089W | 15.7 | SIP4 | interacts with SNF1 protein kinase | the positive regulation of gluconeogenesis |
| YCR105W | 3.8 | ADH7 | NADP(H)-dependent alcohol dehydrogenase | oxidative stress |
| YBR105C | 3.4 | VID24 | required for vacuolar import and degradation of Fbp1p | negative regulation of gluconeogenesis |
| YCL040W | 2.9 | GLK1 | aldohexose specific glucokinase | glucose import |
| YLR377C | 2.9 | FBP1 | fructose-1,6-bisphosphatase | gluconeogenesis |
| YOR347C | 2.9 | PYK2 | pyruvate kinase, glucose-repressed isoform | glycolysis |
| YJL155C | 2.1 | FBP26 | fructose-2,6-bisphosphatase | gluconeogenesis |
| Systematic No. | Normalized | Common | Description |
|---|---|---|---|
| YDR076W | 2.5 | RAD55 | DNA repair protein |
| YER095W | 1.5 | RAD51 | DNA repair protein |
| YDR004W | 1.5 | RAD57 | DNA repair protein |
| YER171W | 1.3 | RAD3 | DNA helicase/ATPase |
| YER162C | 1.3 | RAD4 | excision repair protein |
| YER173W | 1.2 | RAD24 | cell cycle checkpoint protein |
| YER095W | 1.5 | RAD51 | DNA repair protein |
| YDR004W | 1.5 | RAD57 | DNA repair protein |
| YER143W | 1.3 | DDI1 | induced in response to DNA alkylation damage |
| YPL164C | 1.2 | MLH3 | insertion and deletion mismatch repair protein |
| YBR272C | 1.1 | HSM3 | mismatch repair protein |
| YJR052W | 1.1 | RAD7 | nucleotide excision repair protein |
| YCR092C | 1.1 | MSH3 | DNA mismatch repair protein |
| YJR035W | 1.1 | RAD26 | DNA repair and recombination protein |
| YML032C | 1.0 | RAD52 | recombination and DNA repair protein |
| YGL163C | 1.0 | RAD54 | DNA-dependent ATPase of the Snf2p family |
| YML095C | 1.0 | RAD10 | DNA repair protein |
| YDL059C | 0.9 | RAD59 | recombination and DNA repair protein |
| YPL022W | 0.9 | RAD1 | component of the nucleotide excision repairosome |
| YDR030C | 0.8 | RAD28 | protein involved in the same pathway as Rad26p |
| YNL250W | 0.8 | RAD50 | DNA repair protein |
| YMR201C | 0.8 | RAD14 | nucleotide excision repair protein |
| YGR258C | 0.8 | RAD2 | structure-specific nuclease of the nucleotide excision repairosome |
| YBR114W | 0.8 | RAD16 | nucleotide excision repair protein |
| YLR032W | 0.7 | RAD5 | DNA helicase |
| YHR120W | 0.7 | MSH1 | DNA mismatch repair protein, mitochondrial |
| YGL058W | 0.6 | RAD6 | E2 ubiquitin-conjugating enzyme |
| YPL153C | 0.5 | RAD53 | ser/thr/tyr protein kinase |
| YEL037C | 0.5 | RAD23 | nucleotide excision repair protein (ubiquitin-like protein) |
| YER142C | 0.5 | MAG1 | 3-methyladenine DNA glycosylase |
| YNL082W | 0.4 | PMS1 | DNA mismatch repair protein |
| YDR217C | 0.4 | RAD9 | DNA repair checkpoint protein |
| YDR097C | 0.4 | MSH6 | DNA mismatch repair protein |
| YOL090W | 0.4 | MSH2 | DNA mismatch repair protein |
© 2008 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/). This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Iwahashi, Y.; Kitagawa, E.; Iwahashi, H. Analysis of Mechanisms of T-2 Toxin Toxicity Using Yeast DNA Microarrays. Int. J. Mol. Sci. 2008, 9, 2585-2600. https://doi.org/10.3390/ijms9122585
Iwahashi Y, Kitagawa E, Iwahashi H. Analysis of Mechanisms of T-2 Toxin Toxicity Using Yeast DNA Microarrays. International Journal of Molecular Sciences. 2008; 9(12):2585-2600. https://doi.org/10.3390/ijms9122585
Chicago/Turabian StyleIwahashi, Yumiko, Emiko Kitagawa, and Hitoshi Iwahashi. 2008. "Analysis of Mechanisms of T-2 Toxin Toxicity Using Yeast DNA Microarrays" International Journal of Molecular Sciences 9, no. 12: 2585-2600. https://doi.org/10.3390/ijms9122585
APA StyleIwahashi, Y., Kitagawa, E., & Iwahashi, H. (2008). Analysis of Mechanisms of T-2 Toxin Toxicity Using Yeast DNA Microarrays. International Journal of Molecular Sciences, 9(12), 2585-2600. https://doi.org/10.3390/ijms9122585




