2.1 ATP docking and scoring
ATP docking was performed for 50 ATP-protein complexes (see Methods) and the generated ligand positions were compared with the experimental X-ray structures. The correctness of a solution was judged based on a cut-off of root-mean-square deviation (RMSD) from the reference structure. Thus, for the adenine and ribose moieties the cut-off value was set to 2.0 Å, while for the phosphate tail – to 3.0 Å, and for the whole ATP molecule – to 3.5 Å, taking into account the greater flexibility of the two latter cases. Applying these criteria to the results of docking demonstrated that both correct and wrong orientations of adenine and the whole ATP could be found for all 50 complexes, while for both ribose and phosphate tail there were only 46 such complexes. The fact that correct ATP positions were found in each of 50 complexes should not be considered to be in conflict with, for example, the absence of correct phosphate tail orientations in four complexes. The explanation is that the RMSD criterion applied to the whole molecule is more tolerant and, additionally, RMSD was averaged over the whole structure.
The fact that analysis of docking results for all 50 complexes revealed that in most cases both correct and wrong solutions were obtained indicates high efficiency of the ligand conformational search. Now, when we have in our hands quite a large and representative set of different docking solutions for ATP one needs a proper scoring method to select the correct ones among the majority of misleading positions. Scoring by the standard function “goldscore” [
24] ranked correct ATP positions on top only for 22 out of 50 complexes (
Table 1). To some extent such a poor hit rate (< 50%) can be explained by unusual docking parameters. Namely, the “goldscore” was trained on protein-ligand complexes including also coordinated metal ions, while in our study these ions were compulsorily removed, thereby distorting the “goldscore” values. Nevertheless, it would still be desirable to develop a new and more efficient ranking criterion to rank the results of ATP docking.
2.2 Design of fragmental scores
Moreover, taking into account our previous advantages in developing individual scoring criterion for the adenine moiety, it would also be interesting to see whether it is possible to design a fragmental scoring function. While in case of ATP it is easy to divide the molecule into parts differing in physical properties guided by intuitive considerations, it would also be desirable to elaborate some rigorous criteria for such a procedure. We propose to divide a molecule into fragments, or groups of atoms, based on their distinct hydrophobic/hydrophilic properties. It is rather easy to do using the Molecular Hydrophobicity Potential (MHP) approach [
22], when each atom has its own value of hydrophobicity (or hydrophilicity, if negative). After that atoms are clusterized by these values. In this study we applied a cut-off value of 0.2 MHP units to judge whether two atoms bound by a covalent bond belong to the same cluster. However this procedure is not perfect yet, and needs further substantial improvement. Thus, for the sake of better description of a molecule in case of ATP we had to compulsory fuse the hydrophilic amino group with the hydrophobic adenine cluster, which would otherwise remain as independent fragment.
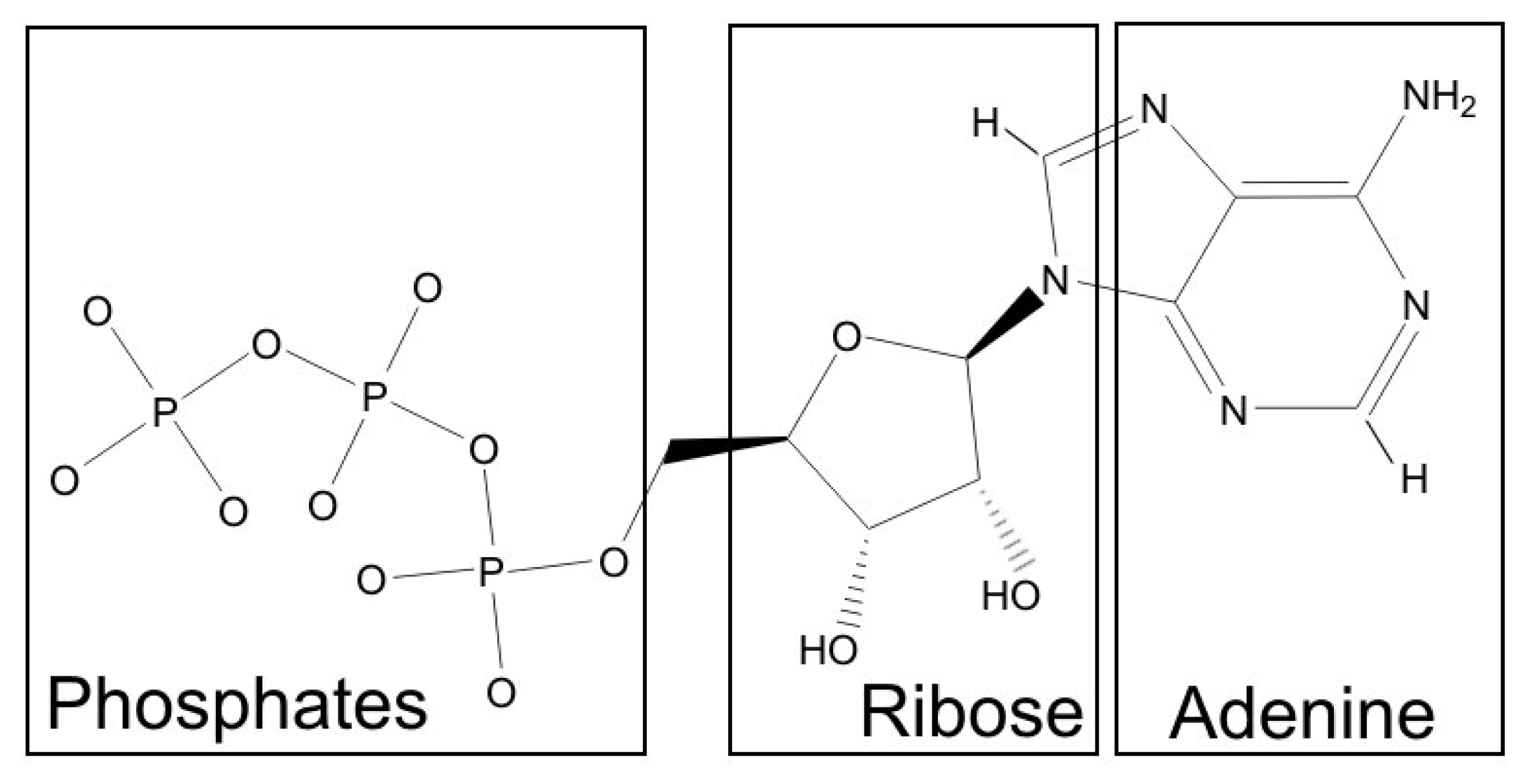
According to the procedure of dissecting a molecule into fragments, three fragments were identified in ATP. These corresponded well to the intuitively recognized parts with different physicochemical properties: the hydrophobic adenine ring, a slightly hydrophilic ribose moiety and a distinctly hydrophilic phosphate tail (
Figure 1). It should be noted, however, that the “ribose fragment” includes also a part of the adenine ring, and the hydrophobic CH
2 group separating ribose from phosphates was not included into any fragment. For each of these fragments, an individual scoring function was then constructed. With this aim, according to our previous results, we decided to seek for the new criteria combining hydrophobic/hydrophilic complementarity, hydrogen bonds, and aromatic interactions.
New scoring functions were designed as linear combinations of the corresponding interaction terms. The weighting coefficients of the terms were adjusted by the least-squares fitting procedure based on docking results. In the regression procedure, each scoring criterion was fitted to a switch function so as to yield the value of 1 in case of correct prediction and the value of 0 otherwise. For that purpose all fragment poses were divided into three groups corresponding to different values of the switch function. Correct structures, characterized by RMSD from the reference within 2.0 Å for adenine and ribose and 3.0 Å for the phosphate tail, were assigned the value of 1. Since it was hard to judge whether the poses with RMSD greater than the correctness threshold but less then 5.0 Å are correct or not, they were assigned an intermediate value of 0.5. Finally, all positions with the greater RMSD were assumed incorrect and therefore corresponding to the value of 0.
It is also important to note that the final formulas of the fragmental scores comprise different interaction terms and not all terms were included into each equation. This needs a more detailed explanation. Starting with the term that had the best hit rate alone, other terms were added subsequently until the increase of correlation between the score and the switch fitness function was greater than 0.01. According to the data presented in
Table 1, hydrophobic complementarity was chosen as the starting point for adenine and ribose, and for phosphates – different acceptor hydrogen bonds and hydrophilic complementarity (the last being the most successful run). Such approach left only those terms that are really relevant for discriminating the correct solutions. This is necessary because otherwise including minor terms into the equation could lead to an overfitting effect, i.e. the performance of the proposed score will increase for the training set in exchange for decreasing in other cases. The resulting criterion for adenine includes the terms of hydrophobic complementarity, hydrogen bonds with the amino group and stacking and pi-cation interactions with Arg. This agrees well with our previous results [
10]. The equation describing this criterion is (definitions of the terms are given in Methods):
The corresponding equations for the ribose and phosphate fragments are:
In all these equations the names of interaction terms correspond exactly to the terms in
Table 1. Surprisingly, the scores for ribose and phosphates do not include any hydrogen bond term. This may seem confusing at first glance, but to all appearance in case of phosphates these bonds are implicitly taken into account by the hydrophilic interactions terms.
The performance of the proposed scores is represented in
Table 1. Since the values of “goldscore” are related not to parts but to the whole ATP molecule, it is not quite correct to compare them with the fragmental scores, however it still can serve as a plausible reference point. It is clearly seen that the ADE_score and PHOS_score perform better than the “goldscore” while the RIB_score demonstrates very poor results.
Still, additional testing is required to make sure that overfitting was avoided. A common way to do that is to divide all the cases into two sets: the training and the test ones. However, still there is no generally accepted approach to perform such a division. So, in this study we propose to do that in the following way. Since overfitting is due to minor peculiarities of the training set, both sets should differ in minor features. At the same time, they should be similar in general properties like the strength of hydrophobic and/or hydrophilic interactions. To achieve this, all three sets were divided into two groups of an equal size by ranking them with the value of typical interactions observed in the X-ray structures (MHPphob-phob for adenine and MHPphil-phil for both ribose and phosphate tail). After that they were counted off by twos to create two sets. This procedure yielded similar distribution of hydrophobic/hydrophilic properties in both sets while being composed of different proteins they would likely differ in minor details such as the number of hydrogen bonds, etc.
These sets were then used in a cross test, where the weighting coefficients were trained with one set and then tested with another, and vice versa. The results are summarized in
Table 2. It is clearly seen that the relative errors of the weighting coefficients of the ADE score are acceptable, while those of the PHOS_score are a bit worse. Nevertheless, this is quite a good result, since exclusion of one half of the training cases is rather a rigorous test. Furthermore, both these scores demonstrate almost the same hit rate, which is definitely higher than that of the “goldscore”. These results indicate that the values of the weighting coefficients in the both equations lie in admissible ranges, and the values obtained with the full sets will be further used (
equations 1 and
3). The results for the RIB_score are completely unacceptable with very high relative error (> 0.70) of some coefficients. The last result is not surprising considering its relatively low hit rate (
Table 1).
In summary, efficient scoring criteria were designed to select correct poses of the adenine ring and the phosphate tail. Unlike that, our approach did not yield any improvement in ranking for the ribose moiety. The reason for that is not quite clear and would require additional investigation. However, one may speculate that ribose often does not play significant role in ATP recognition, but serves as a linker between adenine and phosphates. This effect has been identified in some complexes of ATP and its analogues with proteins [
3].
2.3 Performance of fragmental scores
Considering the quite encouraging improvement of ranking of ATP fragments it is interesting now to determine whether the same approach can be applied to the whole molecule. The same procedure was used in construction of a scoring criterion for the whole ligand molecule (results not shown). However, it exhibited only modest improvement as compared to “goldscore” increasing hit rate from 22 to 24, i.e. just by 4%.
Realizing that this methodology cannot yield sufficient improvement in scoring we decided to use another approach to construct a ranking criterion. Since the fragmental scores efficiently select correctly placed parts of ATP, the most obvious way was to combine them into an integral ranking function ATP_score. The best results were obtained for a mere sum of scores for adenine and phosphate – ADE_score and PHOS_score (see
equation 4) – increasing the hit rate for ATP up to 35 out of 50, which is 26% better than the “goldscore”. Interestingly, addition of the RIB_score greatly spoils the ranking.
The presented results allow a conclusion that a scoring function composed of individual scores of molecular fragments performs considerably better than the one based on intermolecular interactions of the whole molecule. Thus, the fragmental scores highlight those interactions that really determine their recognition by the receptor. At the same time, other interactions that could distort the resulting picture, are attenuated or excluded at all. For example, comparison of the ADE_score and PHOS_score demonstrates that MHPphil-phil term is important for recognition of phosphates. Meanwhile, in case the ATP molecule is not dissected into fragments, the value of MHPphil-phil is calculated over the whole ligand including the ribose ring and adenine amino group. In the last case it becomes unclear whether this term remains useful for scoring any longer.
To summarize, it would be useful to present the results on the overall performance of the ATP_score. Besides the improved hit-rate, it provides better separation between correct and misleading poses. To illustrate this, two types of ATP poses of all 50 complexes were considered - correct (RMSD < 3.5 Å, 692 cases) and definitely incorrect (RMSD > 5.0 Å, 1851 cases) solutions. Other solutions (between these RMSD cut-offs) lie in the twilight zone where it is hard to understand whether such RMSD is caused by slight displacement of the whole molecule or by completely incorrect orientation of its part.
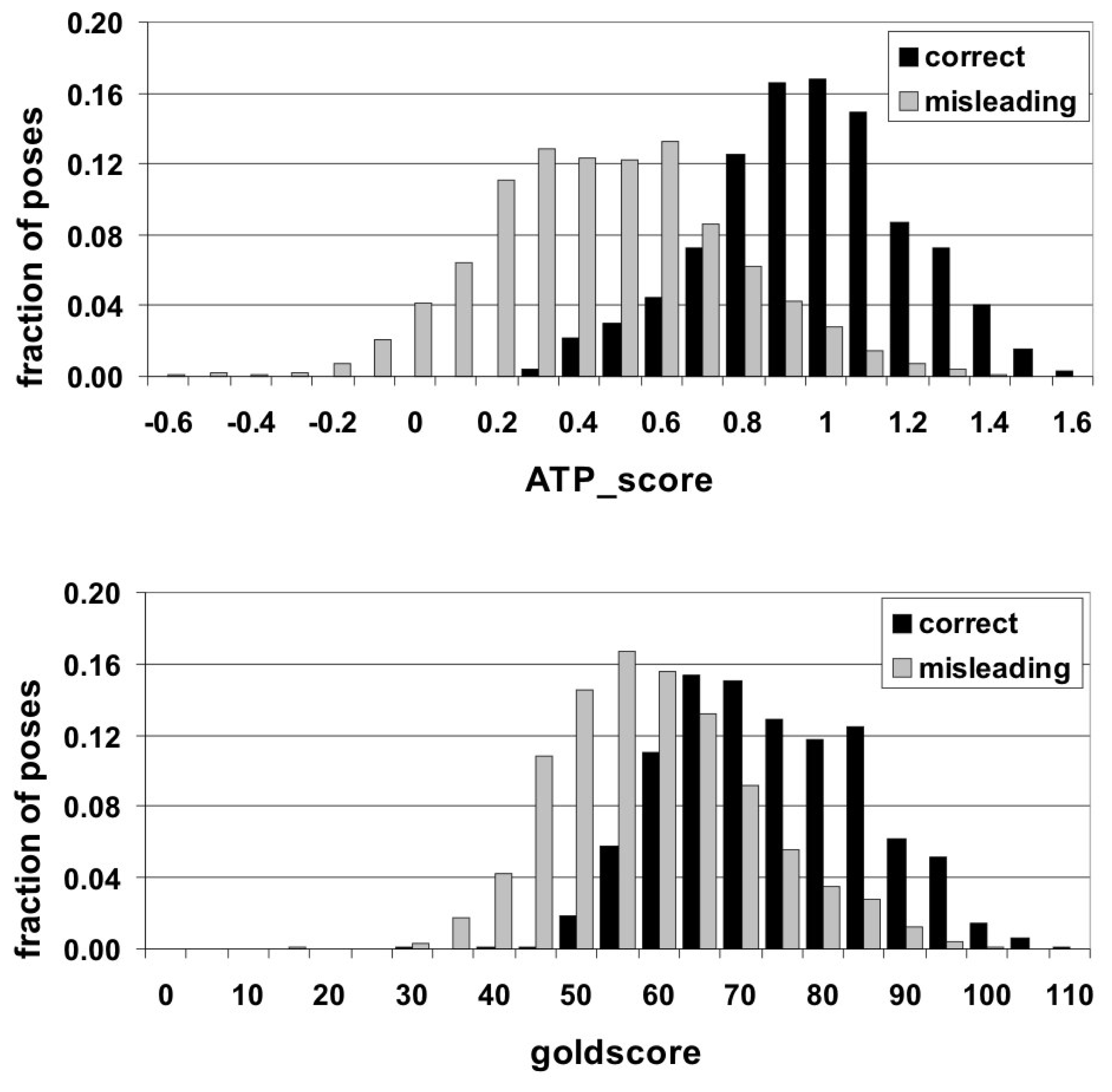
For all docking solutions the “goldscore” values fell into the range between 0 and 110 (in goldscore units). Theoretically, the ATP_score values should cover the interval between 0 and 2 (since in an ideal perfect case both ADE_score and PHOS_score are equal to 1). In practice, due to non-ideal interactions and inaccuracies of the fitting procedure they fell between −0.6 and 1.6. Comparing the distributions of both correct and misleading solutions over the values of the scoring functions (
Figure 2), it is clearly seen that they are quite similar for “goldscore”, whereas ATP_score efficiently separates them. Numerically, this conclusion can be supported by the χ
2 test yielding the values of 0.544 and 1.031 for “goldscore” and ATP_score, respectively. These results indicate that there can be proposed an ATP_score threshold for delimiting correct and misleading poses. The value ATP_score = 0.7 can be chosen as such a threshold since 90% of correct solutions have higher scores and 75% of misleading ones reveal lower values.
Similar distributions of correct and misleading positions were obtained for fragmental scoring functions for adenine and phosphates. It was found that the optimal ADE_score and PHOS_score cutoff values that separate correct and wrong solutions, are similar for both criteria and equal to 0.4 score units. The presented new ATP-oriented scoring function can be used to improve the results of docking ATP to different protein targets. It efficiently identifies those docking poses that are correctly oriented in the binding site. Further investigation will be aimed at improvement of the proposed score, e.g. considering interactions with metal ions, and development of similar approaches for a broader class of ligands.