New Insights into TFEB SUMOylation and Its Role in Lipid Metabolism and Cardiovascular Disease
Abstract
1. Introduction
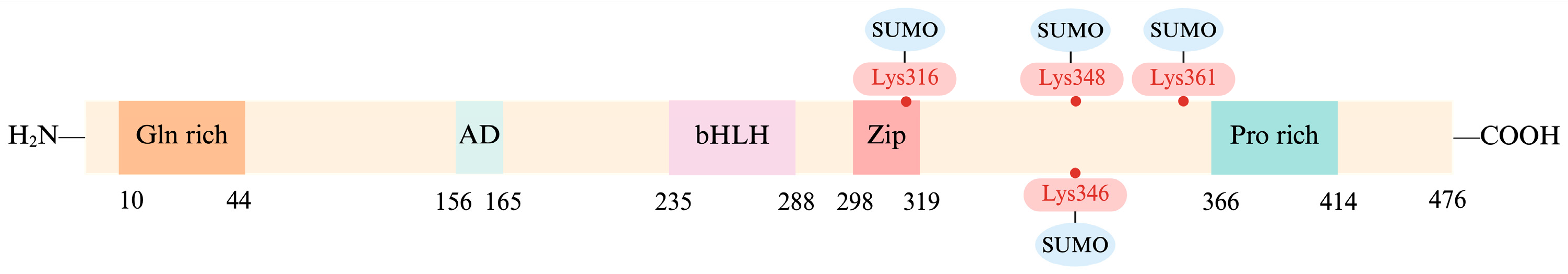
2. Structure and Function of TFEB
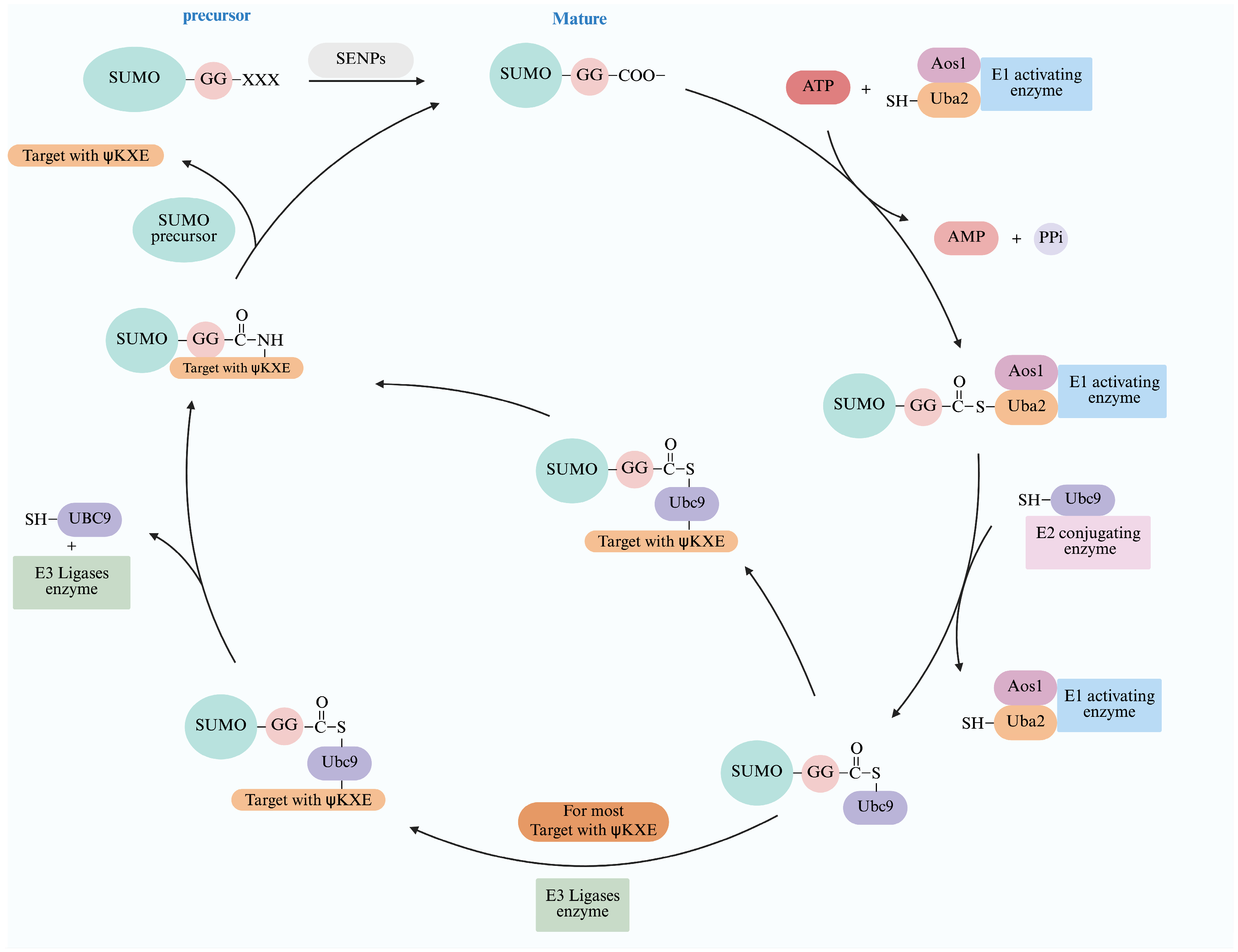
3. Process of SUMOylation
4. SUMOylation of TFEB
5. Biological Effects and Subcellular Localization of TFEB SUMOylation
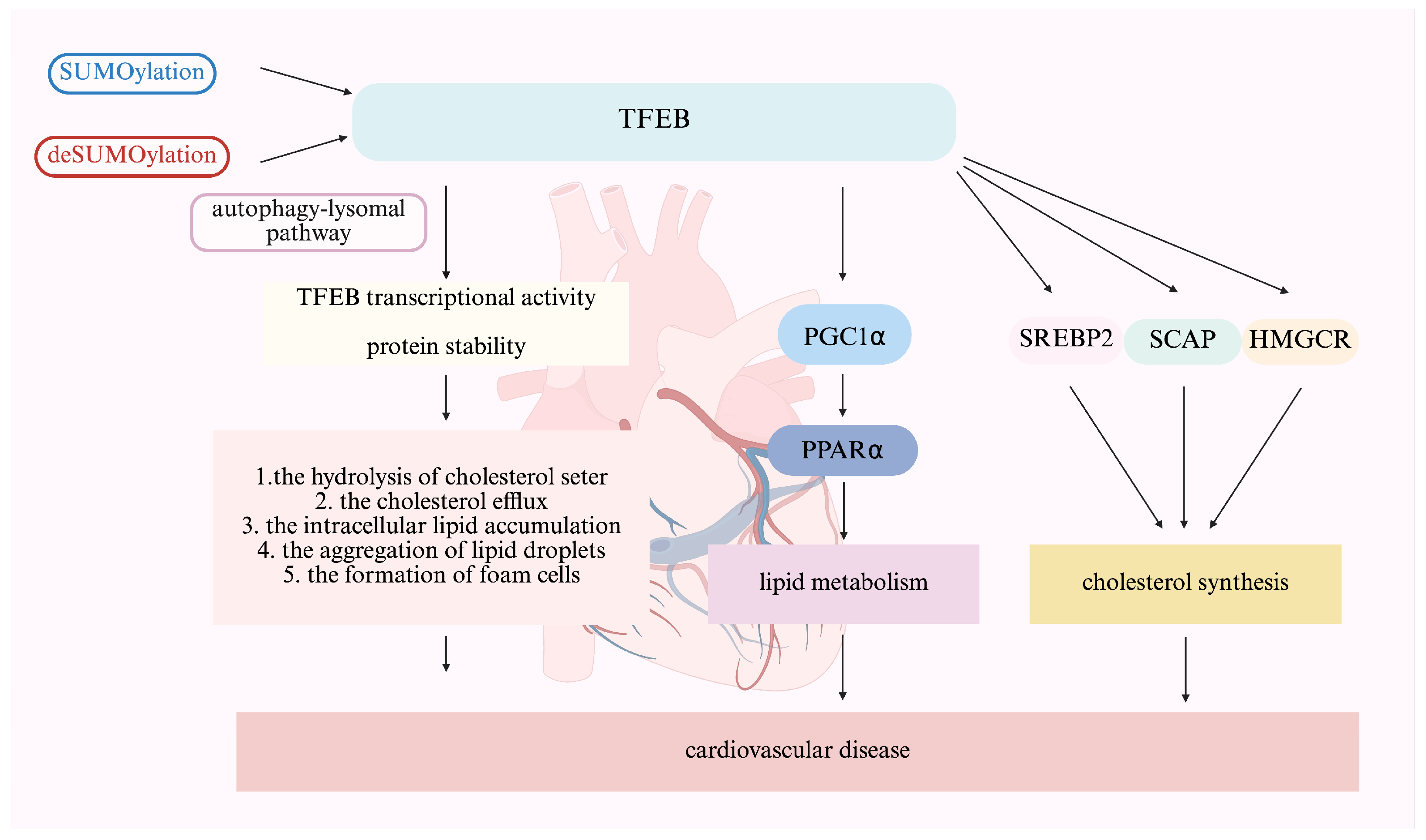
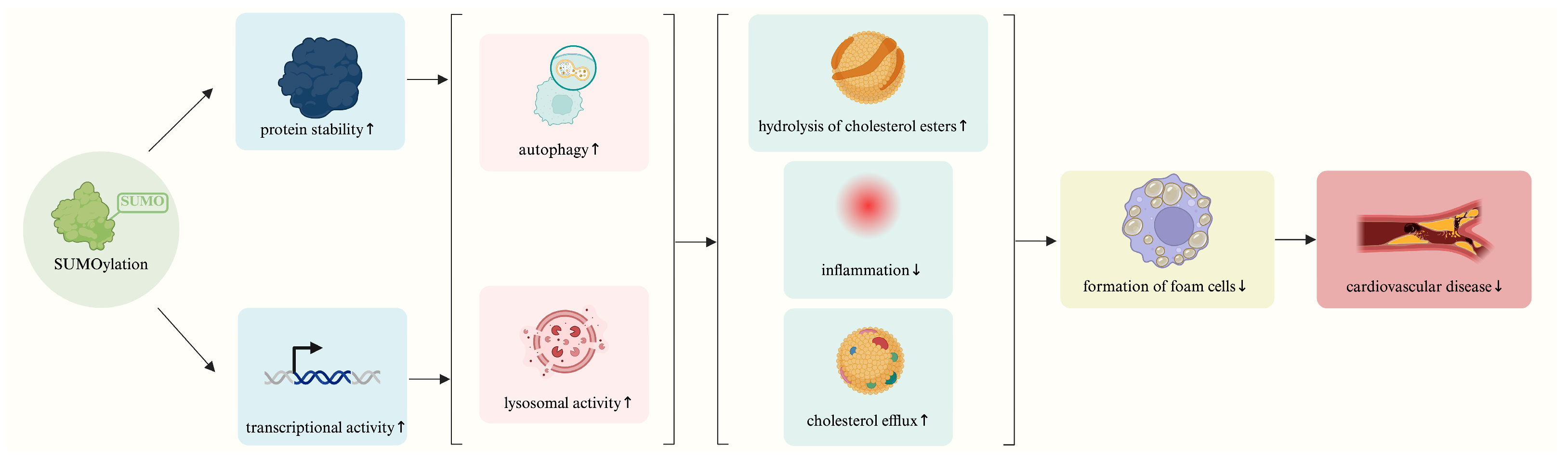
6. Roles of TFEB SUMOylation in Regulating Lipid Metabolism
7. Roles for TFEB SUMOylation in Cardiovascular Diseases
8. Conclusions and Future Perspective
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Powell-Wiley, T.M.; Poirier, P.; Burke, L.E.; Després, J.P.; Gordon-Larsen, P.; Lavie, C.J.; Lear, S.A.; Ndumele, C.E.; Neeland, I.J.; Sanders, P.; et al. Obesity and Cardiovascular Disease: A Scientific Statement From the American Heart Association. Circulation 2021, 143, e984–e1010. [Google Scholar] [CrossRef] [PubMed]
- Miao, G.; Zhao, X.; Chan, S.L.; Zhang, L.; Li, Y.; Zhang, Y.; Zhang, L.; Wang, B. Vascular smooth muscle cell c-Fos is critical for foam cell formation and atherosclerosis. Metabolism 2022, 132, 155213. [Google Scholar] [CrossRef]
- Kuznetsova, T.; Prange, K.H.M.; Glass, C.K.; de Winther, M.P.J. Transcriptional and epigenetic regulation of macrophages in atherosclerosis. Nat. Rev. Cardiol. 2020, 17, 216–228. [Google Scholar] [CrossRef]
- Jia, Y.; Zhang, L.; Liu, Z.; Mao, C.; Ma, Z.; Li, W.; Yu, F.; Wang, Y.; Huang, Y.; Zhang, W.; et al. Targeting macrophage TFEB-14-3-3 epsilon Interface by naringenin inhibits abdominal aortic aneurysm. Cell Discov. 2022, 8, 21. [Google Scholar] [CrossRef]
- Franco-Juárez, B.; Coronel-Cruz, C.; Hernández-Ochoa, B.; Gómez-Manzo, S.; Cárdenas-Rodríguez, N.; Arreguin-Espinosa, R.; Bandala, C.; Canseco-Ávila, L.M.; Ortega-Cuellar, D. TFEB; Beyond Its Role as an Autophagy and Lysosomes Regulator. Cells 2022, 11, 3153. [Google Scholar] [CrossRef]
- Li, X.; Zhu, R.; Jiang, H.; Yin, Q.; Gu, J.; Chen, J.; Ji, X.; Wu, X.; Fu, H.; Wang, H.; et al. Autophagy enhanced by curcumin ameliorates inflammation in atherogenesis via the TFEB-P300-BRD4 axis. Acta Pharm. Sin. B 2022, 12, 2280–2299. [Google Scholar] [CrossRef]
- Zhao, J.; Hu, B.; Xiao, H.; Yang, Q.; Cao, Q.; Li, X.; Zhang, Q.; Ji, A.; Song, S. Fucoidan reduces lipid accumulation by promoting foam cell autophagy via TFEB. Carbohydr. Polym. 2021, 268, 118247. [Google Scholar] [CrossRef]
- Ariano, C.; Riganti, C.; Corà, D.; Valdembri, D.; Mana, G.; Astanina, E.; Serini, G.; Bussolino, F.; Doronzo, G. TFEB controls integrin-mediated endothelial cell adhesion by the regulation of cholesterol metabolism. Angiogenesis 2022, 25, 471–492. [Google Scholar] [CrossRef]
- Li, M.; Wang, Z.; Wang, P.; Li, H.; Yang, L. TFEB: A Emerging Regulator in Lipid Homeostasis for Atherosclerosis. Front. Physiol. 2021, 12, 639920. [Google Scholar] [CrossRef]
- Astanina, E.; Bussolino, F.; Doronzo, G. Multifaceted activities of transcription factor EB in cancer onset and progression. Mol. Oncol. 2021, 15, 327–346. [Google Scholar] [CrossRef]
- Wang, K. TFEB SUMOylation in Macrophage Affects Foam Cell Formation and Atherosclerosis. Doctoral Thesis, Shanghai Jiaotong University, Shanghai, China, 2020. [Google Scholar]
- Pang, Q. Role of TFEB SUMOylation in macrophage foam cell formation. Doctoral Thesis, Shanghai Jiaotong University, Shanghai, China, 2015. [Google Scholar]
- Napolitano, G.; Ballabio, A. TFEB at a glance. J. Cell Sci. 2016, 129, 2475–2481. [Google Scholar] [CrossRef]
- Verastegui, C.; Bertolotto, C.; Bille, K.; Abbe, P.; Ortonne, J.P.; Ballotti, R. TFE3, a transcription factor homologous to microphthalmia, is a potential transcriptional activator of tyrosinase and TyrpI genes. Mol. Endocrinol. 2000, 14, 449–456. [Google Scholar] [CrossRef]
- Abokyi, S.; Ghartey-Kwansah, G.; Tse, D.Y. TFEB is a central regulator of the aging process and age-related diseases. Ageing Res. Rev. 2023, 89, 101985. [Google Scholar] [CrossRef]
- Tan, A.; Prasad, R.; Lee, C.; Jho, E.H. Past, present, and future perspectives of transcription factor EB (TFEB): Mechanisms of regulation and association with disease. Cell Death Differ. 2022, 29, 1433–1449. [Google Scholar] [CrossRef]
- Scheschonka, A.; Tang, Z.; Betz, H. Sumoylation in neurons: Nuclear and synaptic roles? Trends Neurosci. 2007, 30, 85–91. [Google Scholar] [CrossRef]
- Zeng, M.; Liu, W.; Hu, Y.; Fu, N. Sumoylation in liver disease. Clin. Chim. Acta 2020, 510, 347–353. [Google Scholar] [CrossRef]
- Johnson, E.S. Protein modification by SUMO. Annu. Rev. Biochem. 2004, 73, 355–382. [Google Scholar] [CrossRef]
- Shi, Y.; Yasen, M.; Wang, Z.; Du, T.; Ding, Y.; Li, X.; Chai, Z.; Jie, C.; Ju, G.; Ji, M. The allosteric effect of the upper half of SENP1 contributes to its substrate selectivity for SUMO1 over SUMO2. J. Biomol. Struct. Dyn. 2023, 41, 12372–12386. [Google Scholar] [CrossRef] [PubMed]
- Hickey, C.M.; Wilson, N.R.; Hochstrasser, M. Function and regulation of SUMO proteases. Nat. Rev. Mol. Cell Biol. 2012, 13, 755–766. [Google Scholar] [CrossRef] [PubMed]
- Tokarz, P.; Woźniak, K. SENP Proteases as Potential Targets for Cancer Therapy. Cancers 2021, 13, 2059. [Google Scholar] [CrossRef] [PubMed]
- Kunz, K.; Piller, T.; Müller, S. SUMO-specific proteases and isopeptidases of the SENP family at a glance. J. Cell Sci. 2018, 131, jcs211904. [Google Scholar] [CrossRef]
- Flotho, A.; Melchior, F. Sumoylation: A regulatory protein modification in health and disease. Annu. Rev. Biochem. 2013, 82, 357–385. [Google Scholar] [CrossRef]
- Chang, H.M.; Yeh, E.T.H. SUMO: From Bench to Bedside. Physiol. Rev. 2020, 100, 1599–1619. [Google Scholar] [CrossRef]
- Tang, J.; Baba, M. MiT/TFE Family Renal Cell Carcinoma. Genes 2023, 14, 151. [Google Scholar] [CrossRef]
- Zhang, L. Role of SUMOylation in Regulating TFEB Biological Function and Autophagy in Aged Heart. Master’s Thesis, Fourth Military Medical University, Xi’an, China, 2015. [Google Scholar]
- Jansen, N.S.; Vertegaal, A.C.O. A Chain of Events: Regulating Target Proteins by SUMO Polymers. Trends Biochem. Sci. 2021, 46, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Xie, M.; Yu, J.; Ge, S.; Huang, J.; Fan, X. SUMOylation homeostasis in tumorigenesis. Cancer Lett. 2020, 469, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Hilgarth, R.S.; Murphy, L.A.; Skaggs, H.S.; Wilkerson, D.C.; Xing, H.; Sarge, K.D. Regulation and function of SUMO modification. J. Biol. Chem. 2004, 279, 53899–53902. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Zhou, W.; Hu, G.; Wang, L.; Cai, R.; Tian, T. TFEB SUMOylation in macrophages accelerates atherosclerosis by promoting the formation of foam cells through inhibiting lysosomal activity. Cell. Mol. Life Sci. 2023, 80, 358. [Google Scholar] [CrossRef]
- Miller, A.J.; Levy, C.; Davis, I.J.; Razin, E.; Fisher, D.E. Sumoylation of MITF and its related family members TFE3 and TFEB. J. Biol. Chem. 2005, 280, 146–155. [Google Scholar] [CrossRef]
- Subramanian, L.; Benson, M.D.; Iñiguez-Lluhí, J.A. A synergy control motif within the attenuator domain of CCAAT/enhancer-binding protein alpha inhibits transcriptional synergy through its PIASy-enhanced modification by SUMO-1 or SUMO-3. J. Biol. Chem. 2003, 278, 9134–9141. [Google Scholar] [CrossRef]
- Hay, R.T. SUMO: A history of modification. Mol. Cell 2005, 18, 1–12. [Google Scholar] [CrossRef]
- Zhang, P.; Torres, K.; Liu, X.; Liu, C.G.; Pollock, R.E. An Overview of Chromatin-Regulating Proteins in Cells. Curr. Protein Pept. Sci. 2016, 17, 401–410. [Google Scholar] [CrossRef]
- Katsuragi, Y.; Ichimura, Y.; Komatsu, M. p62/SQSTM1 functions as a signaling hub and an autophagy adaptor. FEBS J. 2015, 282, 4672–4678. [Google Scholar] [CrossRef] [PubMed]
- Carey, K.L.; Paulus, G.L.C.; Wang, L.; Balce, D.R.; Luo, J.W.; Bergman, P.; Ferder, I.C.; Kong, L.; Renaud, N.; Singh, S.; et al. TFEB Transcriptional Responses Reveal Negative Feedback by BHLHE40 and BHLHE41. Cell Rep. 2020, 33, 108371. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.Y.; Yao, R.Q.; Li, Y.X.; Zhao, P.Y.; Ren, C.; Du, X.H.; Yao, Y.M. The Role and Regulatory Mechanism of Transcription Factor EB in Health and Diseases. Front. Cell Dev. Biol. 2021, 9, 667750. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.T.; Chiang, M.F.; Chang, C.C.; Yang, S.Y.; Huang, S.W.; Liao, N.S.; Shih, H.M.; Hsu, W.; Lin, K.I. SENP2 restrains the generation of pathogenic Th17 cells in mouse models of colitis. Commun. Biol. 2023, 6, 629. [Google Scholar] [CrossRef]
- Ptak, C.; Wozniak, R.W. SUMO and Nucleocytoplasmic Transport. Adv. Exp. Med. Biol. 2017, 963, 111–126. [Google Scholar]
- Zuo, Y. The Mechanism of SENP1-TFEB Axis Regulating Autophagic Lysosome Production in Macrophages and Its Relationship with Atherosclerosis; Approval number:81870321; National Natural Science Foundation of China: Beijing, China, 2022.
- Wang, T.; Cao, Y.; Zheng, Q.; Tu, J.; Zhou, W.; He, J.; Zhong, J.; Chen, Y.; Wang, J.; Cai, R.; et al. SENP1-Sirt3 Signaling Controls Mitochondrial Protein Acetylation and Metabolism. Mol. Cell 2019, 75, 823–834.e5. [Google Scholar] [CrossRef]
- Wei, B.; Huang, C.; Liu, B.; Wang, Y.; Xia, N.; Fan, Q.; Chen, G.Q.; Cheng, J. Mitotic Phosphorylation of SENP3 Regulates DeSUMOylation of Chromosome-Associated Proteins and Chromosome Stability. Cancer Res. 2018, 78, 2171–2178. [Google Scholar] [CrossRef]
- Ma, G.; Li, S.; Han, Y.; Li, S.; Yue, T.; Wang, B.; Jiang, J. Regulation of Smoothened Trafficking and Hedgehog Signaling by the SUMO Pathway. Dev. Cell 2016, 39, 438–451. [Google Scholar] [CrossRef]
- Conti, L.; Nelis, S.; Zhang, C.; Woodcock, A.; Swarup, R.; Galbiati, M.; Tonelli, C.; Napier, R.; Hedden, P.; Bennett, M.; et al. Small Ubiquitin-like Modifier protein SUMO enables plants to control growth independently of the phytohormone gibberellin. Dev. Cell 2014, 28, 102–110. [Google Scholar] [CrossRef] [PubMed]
- Pelisch, F.; Tammsalu, T.; Wang, B.; Jaffray, E.G.; Gartner, A.; Hay, R.T. A SUMO-Dependent Protein Network Regulates Chromosome Congression during Oocyte Meiosis. Mol. Cell 2017, 65, 66–77. [Google Scholar] [CrossRef]
- Vejpongsa, P.; Yeh, E.T. Wrestling with heart failure: SUMO-1 to the rescue. Circ. Res. 2014, 114, 1561–1563. [Google Scholar] [CrossRef]
- Lee, J.Y.; Won, H.Y.; Park, J.H.; Kim, H.Y.; Choi, H.J.; Shin, D.H.; Kang, J.H.; Woo, J.K.; Oh, S.H.; Son, T.; et al. MEL-18 loss mediates estrogen receptor-α downregulation and hormone independence. J. Clin. Investig. 2015, 125, 1801–1814. [Google Scholar] [CrossRef]
- Liu, B.; Wang, T.; Mei, W.; Li, D.; Cai, R.; Zuo, Y.; Cheng, J. Small ubiquitin-like modifier (SUMO) protein-specific protease 1 de-SUMOylates Sharp-1 protein and controls adipocyte differentiation. J. Biol. Chem. 2014, 289, 22358–22364. [Google Scholar] [CrossRef]
- Settembre, C.; De Cegli, R.; Mansueto, G.; Saha, P.K.; Vetrini, F.; Visvikis, O.; Huynh, T.; Carissimo, A.; Palmer, D.; Klisch, T.J.; et al. TFEB controls cellular lipid metabolism through a starvation-induced autoregulatory loop. Nat. Cell Biol. 2013, 15, 647–658, Erratum in Nat. Cell Biol. 2013, 15, 1016. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, B.; Schulze, R.J.; Weller, S.G.; Sletten, A.C.; Casey, C.A.; McNiven, M.A. The small GTPase Rab7 as a central regulator of hepatocellular lipophagy. Hepatology 2015, 61, 1896–1907. [Google Scholar] [CrossRef]
- Evans, T.D.; Zhang, X.; Jeong, S.J.; He, A.; Song, E.; Bhattacharya, S.; Holloway, K.B.; Lodhi, I.J.; Razani, B. TFEB drives PGC-1α expression in adipocytes to protect against diet-induced metabolic dysfunction. Sci. Signal 2019, 12, eaau2281. [Google Scholar] [CrossRef] [PubMed]
- Palmieri, M.; Impey, S.; Kang, H.; di Ronza, A.; Pelz, C.; Sardiello, M.; Ballabio, A. Characterization of the CLEAR network reveals an integrated control of cellular clearance pathways. Hum. Mol. Genet. 2011, 20, 3852–3866. [Google Scholar] [CrossRef]
- Ouimet, M.; Franklin, V.; Mak, E.; Liao, X.; Tabas, I.; Marcel, Y.L. Autophagy regulates cholesterol efflux from macrophage foam cells via lysosomal acid lipase. Cell Metab. 2011, 13, 655–667. [Google Scholar] [CrossRef]
- Khawar, M.B.; Gao, H.; Li, W. Autophagy and Lipid Metabolism. Adv. Exp. Med. Biol. 2019, 1206, 359–374. [Google Scholar]
- Emanuel, R.; Sergin, I.; Bhattacharya, S.; Turner, J.; Epelman, S.; Settembre, C.; Diwan, A.; Ballabio, A.; Razani, B. Induction of lysosomal biogenesis in atherosclerotic macrophages can rescue lipid-induced lysosomal dysfunction and downstream sequelae. Arterioscler. Thromb. Vasc. Biol. 2014, 34, 1942–1952. [Google Scholar] [CrossRef]
- Wang, J.Q.; Lin, Z.C.; Li, L.L.; Zhang, S.F.; Li, W.H.; Liu, W.; Song, B.L.; Luo, J. SUMOylation of the ubiquitin ligase IDOL decreases LDL receptor levels and is reversed by SENP1. J. Biol. Chem. 2021, 296, 100032. [Google Scholar] [CrossRef]
- Franco-Juárez, B.; Mejía-Martínez, F.; Moreno-Arriola, E.; Hernández-Vázquez, A.; Gómez-Manzo, S.; Marcial-Quino, J.; Arreguín-Espinosa, R.; Velázquez-Arellano, A.; Ortega-Cuellar, D. A high glucose diet induces autophagy in a HLH-30/TFEB-dependent manner and impairs the normal lifespan of C. elegans. Aging 2018, 10, 2657–2667. [Google Scholar] [CrossRef]
- Nakamura, S.; Yoshimori, T. Autophagy and Longevity. Mol. Cells 2018, 41, 65–72. [Google Scholar]
- Ji, J.; Petropavlovskaia, M.; Khatchadourian, A.; Patapas, J.; Makhlin, J.; Rosenberg, L.; Maysinger, D. Type 2 diabetes is associated with suppression of autophagy and lipid accumulation in β-cells. J. Cell Mol. Med. 2019, 23, 2890–2900. [Google Scholar] [CrossRef]
- Wang, Y.; Gunewardena, S.; Li, F.; Matye, D.J.; Chen, C.; Chao, X.; Jung, T.; Zhang, Y.; Czerwiński, M.; Ni, H.M.; et al. An FGF15/19-TFEB regulatory loop controls hepatic cholesterol and bile acid homeostasis. Nat. Commun. 2020, 11, 3612. [Google Scholar] [CrossRef] [PubMed]
- Ouimet, M.; Barrett, T.J.; Fisher, E.A. HDL and Reverse Cholesterol Transport. Circ. Res. 2019, 124, 1505–1518. [Google Scholar] [CrossRef] [PubMed]
- Rosenson, R.S.; Brewer, H.B., Jr.; Davidson, W.S.; Fayad, Z.A.; Fuster, V.; Goldstein, J.; Hellerstein, M.; Jiang, X.C.; Phillips, M.C.; Rader, D.J.; et al. Cholesterol efflux and atheroprotection: Advancing the concept of reverse cholesterol transport. Circulation 2012, 125, 1905–1919. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Gong, K.; Xu, S.; Zhang, F.; Meng, X.; Han, J. Regulation of cholesterol homeostasis in health and diseases: From mechanisms to targeted therapeutics. Signal Transduct. Target. Ther. 2022, 7, 265. [Google Scholar] [CrossRef]
- Yuan, Y.; Li, P.; Ye, J. Lipid homeostasis and the formation of macrophage-derived foam cells in atherosclerosis. Protein Cell 2012, 3, 173–181. [Google Scholar] [CrossRef]
- Leitao, B.B.; Jones, M.C.; Brosens, J.J. The SUMO E3-ligase PIAS1 couples reactive oxygen species-dependent JNK activation to oxidative cell death. FASEB J. 2011, 25, 3416–3425. [Google Scholar] [CrossRef]
- Ouyang, J.; Gill, G. SUMO engages multiple corepressors to regulate chromatin structure and transcription. Epigenetics 2009, 4, 440–444. [Google Scholar] [CrossRef]
- Rosonina, E.; Akhter, A.; Dou, Y.; Babu, J.; Sri Theivakadadcham, V.S. Regulation of transcription factors by sumoylation. Transcription 2017, 8, 220–231. [Google Scholar] [CrossRef]
- Fu, X.; Wang, J.; Jiang, S.; Wu, J.; Mu, Z.; Tang, Y.; Wang, S.; Fu, H.; Ma, H.; Zhao, Y. Mortality trend analysis of ischemic heart disease in China between 2010 and 2019: A joinpoint analysis. BMC Public Health 2023, 23, 644. [Google Scholar] [CrossRef] [PubMed]
- Rawlings, N.; Lee, L.; Nakamura, Y.; Wilkinson, K.A.; Henley, J.M. Protective role of the deSUMOylating enzyme SENP3 in myocardial ischemia-reperfusion injury. PLoS ONE 2019, 14, e0213331. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Xue, X.; Zhang, H.; Che, X.; Luo, J.; Wang, P.; Xu, J.; Xing, Z.; Yuan, L.; Liu, Y.; et al. Neuronal-targeted TFEB rescues dysfunction of the autophagy-lysosomal pathway and alleviates ischemic injury in permanent cerebral ischemia. Autophagy 2019, 15, 493–509. [Google Scholar] [CrossRef] [PubMed]
- Bossis, G.; Melchior, F. Regulation of SUMOylation by reversible oxidation of SUMO conjugating enzymes. Mol. Cell 2006, 21, 349–357. [Google Scholar] [CrossRef]
- Xiao, H.; Zhou, H.; Zeng, G.; Mao, Z.; Zeng, J.; Gao, A. SUMOylation targeting mitophagy in cardiovascular diseases. J. Mol. Med. 2022, 100, 1511–1538. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Meng, Q.; Li, C.G.; Chen, X.; Cao, R.; Zhang, H.; Wang, P.; Jin, J. New Insights into TFEB SUMOylation and Its Role in Lipid Metabolism and Cardiovascular Disease. Int. J. Mol. Sci. 2026, 27, 347. https://doi.org/10.3390/ijms27010347
Meng Q, Li CG, Chen X, Cao R, Zhang H, Wang P, Jin J. New Insights into TFEB SUMOylation and Its Role in Lipid Metabolism and Cardiovascular Disease. International Journal of Molecular Sciences. 2026; 27(1):347. https://doi.org/10.3390/ijms27010347
Chicago/Turabian StyleMeng, Qingxiu, Chun Guang Li, Xiaolong Chen, Rui Cao, Haihong Zhang, Ping Wang, and Jing Jin. 2026. "New Insights into TFEB SUMOylation and Its Role in Lipid Metabolism and Cardiovascular Disease" International Journal of Molecular Sciences 27, no. 1: 347. https://doi.org/10.3390/ijms27010347
APA StyleMeng, Q., Li, C. G., Chen, X., Cao, R., Zhang, H., Wang, P., & Jin, J. (2026). New Insights into TFEB SUMOylation and Its Role in Lipid Metabolism and Cardiovascular Disease. International Journal of Molecular Sciences, 27(1), 347. https://doi.org/10.3390/ijms27010347




