Neurologist’s Black Swan: Molecular Basis of Prenatal Seizures
Abstract
1. Introduction
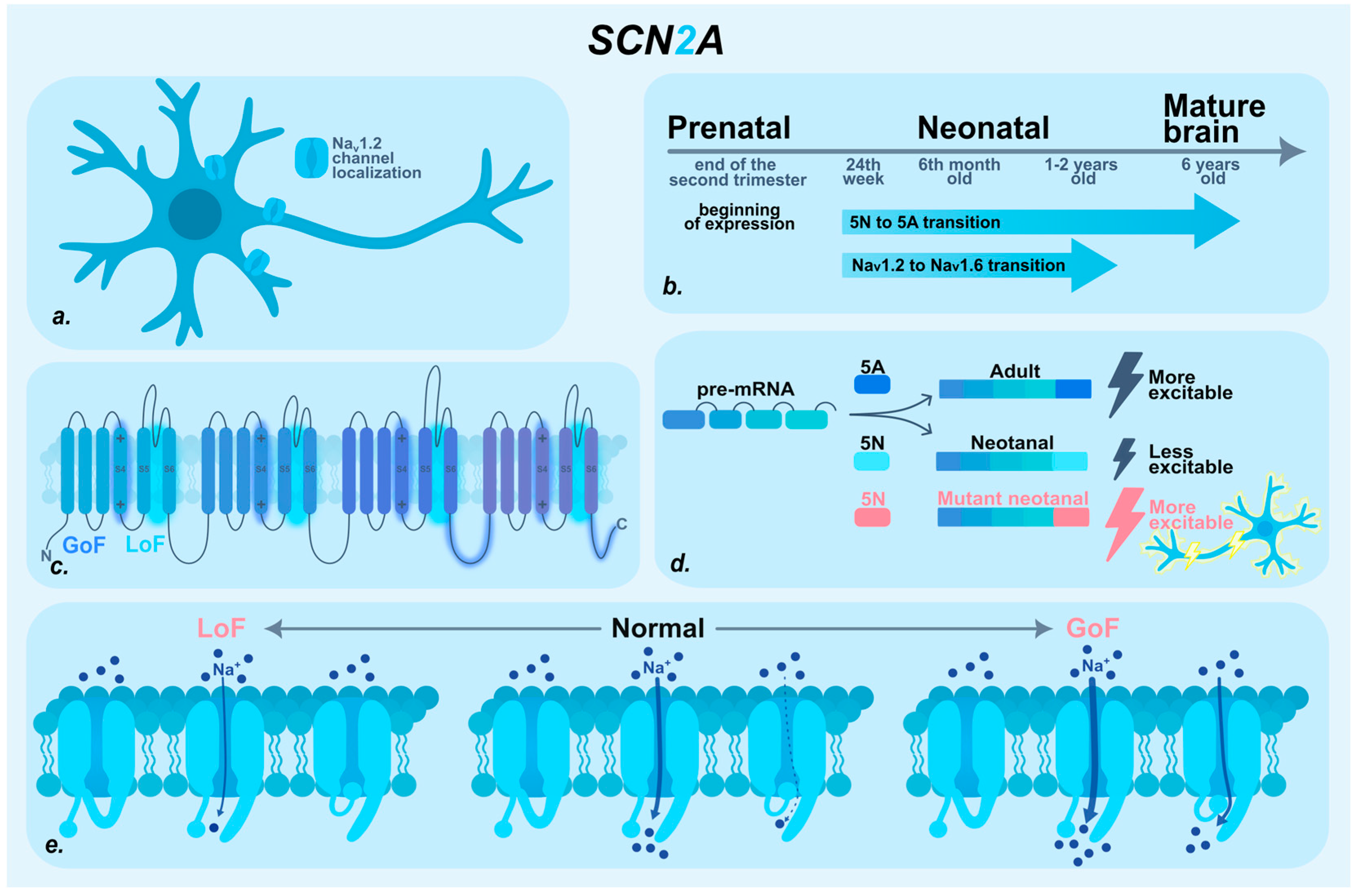
2. SCN2A
2.1. SCN2A Patients with Prenatal Seizures
2.2. Molecular Basis of SCN2A-Associated Epilepsy
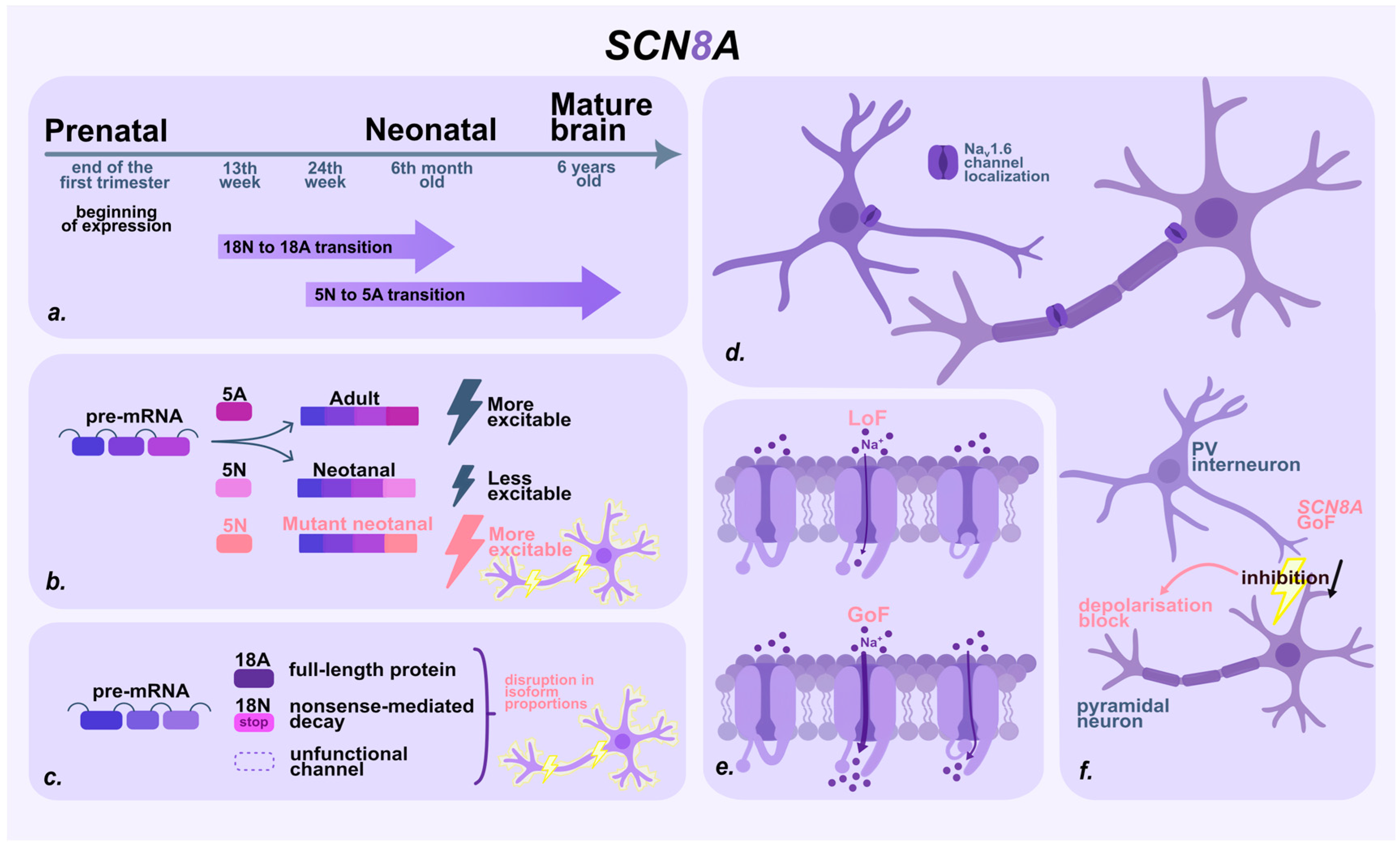
3. SCN8A
3.1. SCN8A Patients with Prenatal Seizures
3.2. Molecular Basis of SCN8A-Associated Epilepsy
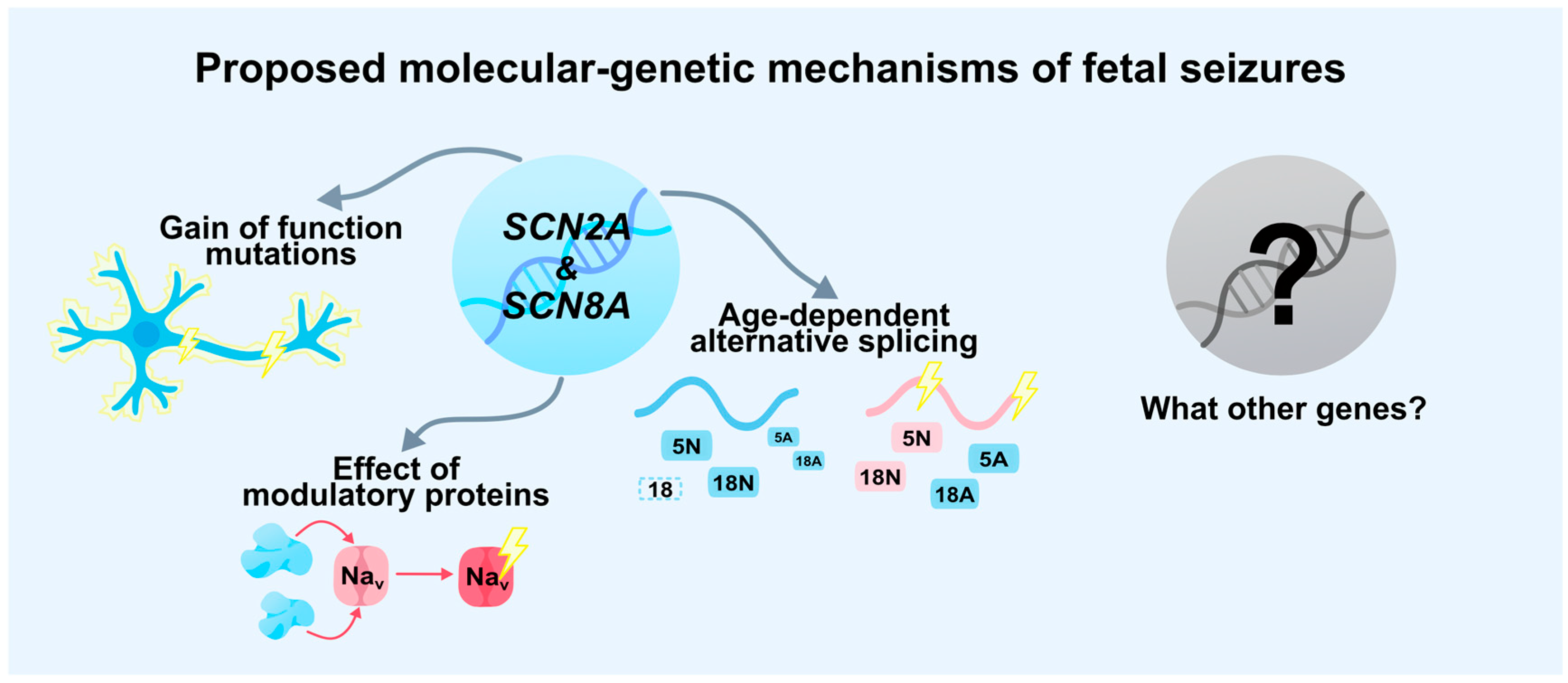
4. Discussion
- Conducting prospective genomic studies in cohorts of infants with seizures within the first 72 h of life and/or prenatally verified seizure patterns.
- Developing experimental models (based on iPSCs and brain organoids with identified mutations) to investigate pathogenesis and explore therapies that modify disease progression rather than merely suppress symptoms.
- Establishing clear criteria for collecting clinical history and identifying characteristic features of prenatal epilepsy.
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sun, Y.; Vestergaard, M.; Christensen, J.; Nahmias, A.J.; Olsen, J. Prenatal Exposure to Maternal Infections and Epilepsy in Childhood: A Population-Based Cohort Study. Pediatrics 2008, 121, e1100–e1107. [Google Scholar] [CrossRef]
- Sauvestre, F.; Moutton, S.; Badens, C.; Broussin, B.; Carles, D.; Houcinat, N.; Lacoste, C.; Marguet, F.; Pecheux, C.; Villard, L.; et al. In Utero Seizures Revealing Dentato-Olivary Dysplasia Caused by SCN2A Mutation. Neuropathol. Appl. Neurobiol. 2017, 43, 631–635. [Google Scholar] [CrossRef]
- McNally, M.A.; Johnson, J.; Huisman, T.A.; Poretti, A.; Baranano, K.W.; Baschat, A.A.; Stafstrom, C.E. SCN8A Epileptic Encephalopathy: Detection of Fetal Seizures Guides Multidisciplinary Approach to Diagnosis and Treatment. Pediatr. Neurol. 2016, 64, 87–91. [Google Scholar] [CrossRef] [PubMed]
- Hadjipanteli, A.; Theodosiou, A.; Papaevripidou, I.; Evangelidou, P.; Alexandrou, A.; Salameh, N.; Kallikas, I.; Kakoullis, K.; Frakala, S.; Oxinou, C.; et al. Sodium Channel Gene Variants in Fetuses with Abnormal Sonographic Findings: Expanding the Prenatal Phenotypic Spectrum of Sodium Channelopathies. Genes 2024, 15, 119. [Google Scholar] [CrossRef] [PubMed]
- Bernardo, S.; Marchionni, E.; Prudente, S.; Liso, P.; De Spalice, A.; Giancotti, A.; Manganaro, L.; Pizzuti, A. Unusual Association of SCN2A Epileptic Encephalopathy with Severe Cortical Dysplasia Detected by Prenatal MRI. Eur. J. Paediatr. Neurol. 2017, 21, 587–590. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Jayapal, S.; Goyal, S.; Jungbluth, H.; Lascelles, K. Early-Onset Movement Disorder and Epileptic Encephalopathy Due to de Novo Dominant SCN8A Mutation. Seizure 2015, 26, 69–71. [Google Scholar] [CrossRef]
- Denis, J.; Villeneuve, N.; Cacciagli, P.; Mignon-Ravix, C.; Lacoste, C.; Lefranc, J.; Napuri, S.; Damaj, L.; Villega, F.; Pedespan, J.; et al. Clinical Study of 19 Patients with SCN8A-related Epilepsy: Two Modes of Onset Regarding EEG and Seizures. Epilepsia 2019, 60, 845–856. [Google Scholar] [CrossRef]
- Jung, E.; Lee, B.Y.; Huh, C.Y. Prenatal Diagnosis of Fetal Seizure: A Case Report. J. Korean Med. Sci. 2008, 23, 906. [Google Scholar] [CrossRef]
- Falsaperla, R.; Ruggieri, M.; Polizzi, A.; Praticò, A.D. From Abnormal Fetal Movements to Neonatal Seizures: A Literature Review. Epilepsy Res. 2025, 214, 107557. [Google Scholar] [CrossRef]
- Abrams, L.; Balducci, J. Fetal Seizures: A Case Study. Obstet. Gynecol. 1996, 88, 661–663. [Google Scholar] [CrossRef]
- Sheizaf, B.; Mazor, M.; Landau, D.; Burstein, E.; Bashiri, A.; Hershkovitz, R. Early Sonographic Prenatal Diagnosis of Seizures. Ultrasound Obstet. Gynecol. 2007, 30, 1007–1009. [Google Scholar] [CrossRef] [PubMed]
- Whitehead, C.L.; Cohen, N.; Visser, G.H.A.; Farine, D. Are Increased Fetal Movements Always Reassuring? J. Matern. Fetal Neonatal Med. 2020, 33, 3713–3718. [Google Scholar] [CrossRef]
- Takahashi, H.; Matsubara, T.; Horie, K.; Kobayashi, A.; Furukawa, R.; Matsubara, S. Fetal Seizure: A Video Recording and Possible Etiology. J. Obstet. Gynaecol. Res. 2019, 45, 1619–1625. [Google Scholar] [CrossRef]
- Patane, L.; Ghidini, A. Fetal Seizures: Case Report and Literature Review. J. Matern.-Fetal Neonatal Med. 2001, 10, 287–289. [Google Scholar] [CrossRef]
- Librizzi, L.; Verde, D.V.; de Curtis, M. Causal Correlation between Seizure Activity and Brain Damage. Neurol. Sci. 2025, 46, 4259–4267. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Anttonen, A.K.; Liukkonen, E.; Gaily, E.; Maljevic, S.; Schubert, S.; Bellan-Koch, A.; Petrou, S.; Ahonen, V.E.; Lerche, H.; et al. SCN2A Mutation Associated with Neonatal Epilepsy, Late-Onset Episodic Ataxia, Myoclonus, and Pain. Neurology 2010, 75, 1454–1458. [Google Scholar] [CrossRef] [PubMed]
- Tian, C.; Wang, K.; Ke, W.; Guo, H.; Shu, Y. Molecular Identity of Axonal Sodium Channels in Human Cortical Pyramidal Cells. Front. Cell. Neurosci. 2014, 8, 297. [Google Scholar] [CrossRef]
- Spratt, P.W.E.; Ben-Shalom, R.; Keeshen, C.M.; Burke, K.J.; Clarkson, R.L.; Sanders, S.J.; Bender, K.J. The Autism-Associated Gene Scn2a Contributes to Dendritic Excitability and Synaptic Function in the Prefrontal Cortex. Neuron 2019, 103, 673–685.e5. [Google Scholar] [CrossRef]
- Brunklaus, A.; Du, J.; Steckler, F.; Ghanty, I.I.; Johannesen, K.M.; Fenger, C.D.; Schorge, S.; Baez-Nieto, D.; Wang, H.R.; Allen, A.; et al. Biological Concepts in Human Sodium Channel Epilepsies and Their Relevance in Clinical Practice. Epilepsia 2020, 61, 387–399. [Google Scholar] [CrossRef]
- Heighway, J.; Sedo, A.; Garg, A.; Eldershaw, L.; Perreau, V.; Berecki, G.; Reid, C.A.; Petrou, S.; Maljevic, S. Sodium Channel Expression and Transcript Variation in the Developing Brain of Human, Rhesus Monkey, and Mouse. Neurobiol. Dis. 2022, 164, 105622. [Google Scholar] [CrossRef]
- Lerche, H.; Shah, M.; Beck, H.; Noebels, J.; Johnston, D.; Vincent, A. Ion Channels in Genetic and Acquired Forms of Epilepsy. J. Physiol. 2013, 591, 753–764. [Google Scholar] [CrossRef]
- Meisler, M.H.; Hill, S.F.; Yu, W. Sodium Channelopathies in Neurodevelopmental Disorders. Nat. Rev. Neurosci. 2021, 22, 152–166, Erratum in Nat. Rev. Neurosci. 2021, 22, 256. https://doi.org/10.1038/s41583-021-00449-5. [Google Scholar] [CrossRef]
- Simmons, R.; Martinez, A.B.; Barkovich, J.; Numis, A.L.; Cilio, M.R.; Glenn, O.A.; Gano, D.; Rogers, E.E.; Glass, H.C. Disorders of Neuronal Migration/Organization Convey the Highest Risk of Neonatal Onset Epilepsy Compared with Other Congenital Brain Malformations. Pediatr. Neurol. 2022, 127, 20–27. [Google Scholar] [CrossRef] [PubMed]
- DiStefano, N.; Cooper, J.N.; Elisha, D.H.; Zalta, M.; Mittal, J.; Cohen, D.; Monterrubio, A.; Hossain, R.; Sangadi, A.; Mittal, R.; et al. Decoding SCN2A Variants: Bridging Genetics and Phenotypes in Autism Spectrum Disorder. J. Clin. Med. 2025, 14, 3790. [Google Scholar] [CrossRef] [PubMed]
- Hamosh, A.; Scott, A.F.; Amberger, J.; Valle, D.; McKusick, V.A. Online Mendelian Inheritance in Man (OMIM). Hum. Mutat. 2000, 15, 57–61. [Google Scholar] [CrossRef]
- Winquist, R.J.; Cohen, C.J. Integration of Biological/Pathophysiological Contexts to Help Clarify Genotype-Phenotype Mismatches in Monogenetic Diseases. Childhood Epilepsies Associated with SCN2A as a Case Study. Biochem. Pharmacol. 2018, 151, 252–262. [Google Scholar] [CrossRef]
- Liang, L.; Darbandi, S.F.; Pochareddy, S.; Gulden, F.O.; Gilson, M.C.; Sheppard, B.K.; Sahagun, A.; An, J.-Y.; Werling, D.M.; Rubenstein, J.L.R.; et al. Developmental Dynamics of Voltage-Gated Sodium Channel Isoform Expression in the Human and Mouse Brain. Genome Med. 2021, 13, 135. [Google Scholar] [CrossRef]
- Kearney, J.A.; Plummer, N.W.; Smith, M.R.; Kapur, J.; Cummins, T.R.; Waxman, S.G.; Goldin, A.L.; Meisler, M.H. A Gain-of-Function Mutation in the Sodium Channel Gene Scn2a Results in Seizures and Behavioral Abnormalities. Neuroscience 2001, 102, 307–317. [Google Scholar] [CrossRef]
- Hu, W.; Tian, C.; Li, T.; Yang, M.; Hou, H.; Shu, Y. Distinct Contributions of Nav1.6 and Nav1.2 in Action Potential Initiation and Backpropagation. Nat. Neurosci. 2009, 12, 996–1002. [Google Scholar] [CrossRef]
- Catterall, W.A. Voltage-Gated Sodium Channels at 60: Structure, Function and Pathophysiology. J. Physiol. 2012, 590, 2577–2589. [Google Scholar] [CrossRef]
- Bender, K.J.; Trussell, L.O. The Physiology of the Axon Initial Segment. Annu. Rev. Neurosci. 2012, 35, 249–265. [Google Scholar] [CrossRef]
- Kole, M.H.P.; Stuart, G.J. Signal Processing in the Axon Initial Segment. Neuron 2012, 73, 235–247. [Google Scholar] [CrossRef]
- Wei, F.; Yan, L.M.; Su, T.; He, N.; Lin, Z.J.; Wang, J.; Shi, Y.W.; Yi, Y.H.; Liao, W.P. Ion Channel Genes and Epilepsy: Functional Alteration, Pathogenic Potential, and Mechanism of Epilepsy. Neurosci. Bull. 2017, 33, 455–477. [Google Scholar] [CrossRef]
- Thompson, C.H.; Potet, F.; Abramova, T.V.; Dekeyser, J.M.; Ghabra, N.F.; Vanoye, C.G.; Millichap, J.J.; George, A.L. Epilepsy-Associated SCN2A (NaV1.2) Variants Exhibit Diverse and Complex Functional Properties. J. Gen. Physiol. 2023, 155, e202313375. [Google Scholar] [CrossRef]
- Clatot, J.; Goldberg, E.M. No Pain, No Gain of Function: Epilepsy-Associated Variants in SCN2A Defy Classification. Epilepsy Curr. 2024, 24, 126–128. [Google Scholar] [CrossRef] [PubMed]
- Pablo, J.L.; Wang, C.; Presby, M.M.; Pitt, G.S. Polarized Localization of Voltage-Gated Na+ Channels Is Regulated by Concerted FGF13 and FGF14 Action. Proc. Natl. Acad. Sci. USA 2016, 113, E2665–E2674. [Google Scholar] [CrossRef]
- Hongyue, L.; Ling, Q.; Rui, F.; Jaehoon, S.; Xuan, H.; Xiaoxue, X.; Dongyi, Z. Increased NaV1.2 Expression and Its Interaction with CaM Contribute to the Hyperexcitability Induced by Prolonged Inhibition of CaMKII. Epilepsia 2025, 66, 2521–2537. [Google Scholar]
- Sanders, S.J.; Campbell, A.J.; Cottrell, J.R.; Moller, R.S.; Wagner, F.F.; Auldridge, A.L.; Bernier, R.A.; Catterall, W.A.; Chung, W.K.; Empfield, J.R.; et al. Progress in Understanding and Treating SCN2A-Mediated Disorders. Trends Neurosci. 2018, 41, 442–456. [Google Scholar] [CrossRef] [PubMed]
- Gazina, E.V.; Leaw, B.T.W.; Richards, K.L.; Wimmer, V.C.; Kim, T.H.; Aumann, T.D.; Featherby, T.J.; Churilov, L.; Hammond, V.E.; Reid, C.A.; et al. ‘Neonatal’ Nav1.2 Reduces Neuronal Excitability and Affects Seizure Susceptibility and Behaviour. Hum. Mol. Genet. 2015, 24, 1457–1468. [Google Scholar] [CrossRef]
- Nelson, A.D.; Catalfio, A.M.; Gupta, J.P.; Min, L.; Caballero-Florán, R.N.; Dean, K.P.; Elvira, C.C.; Derderian, K.D.; Kyoung, H.; Sahagun, A.; et al. Physical and Functional Convergence of the Autism Risk Genes Scn2a and Ank2 in Neocortical Pyramidal Cell Dendrites. Neuron 2024, 112, 1133–1149.e6. [Google Scholar] [CrossRef]
- Vacher, H.; Mohapatra, D.P.; Trimmer, J.S. Localization and Targeting of Voltage-Dependent Ion Channels in Mammalian Central Neurons. Physiol. Rev. 2008, 88, 1407–1447. [Google Scholar] [CrossRef]
- Gazina, E.V.; Richards, K.L.; Mokhtar, M.B.C.; Thomas, E.A.; Reid, C.A.; Petrou, S. Differential Expression of Exon 5 Splice Variants of Sodium Channel α Subunit MRNAs in the Developing Mouse Brain. Neuroscience 2010, 166, 195–200. [Google Scholar] [CrossRef]
- Copley, R.R. Evolutionary Convergence of Alternative Splicing in Ion Channels. Trends Genet. 2004, 20, 171–176. [Google Scholar] [CrossRef] [PubMed]
- Thompson, C.H.; Ben-Shalom, R.; Bender, K.J.; George, A.L. Alternative Splicing Potentiates Dysfunction of Early-Onset Epileptic Encephalopathy SCN2A Variants. J. Gen. Physiol. 2020, 152, e201912442. [Google Scholar] [CrossRef]
- Xu, R.; Thomas, E.A.; Jenkins, M.; Gazina, E.V.; Chiu, C.; Heron, S.E.; Mulley, J.C.; Scheffer, I.E.; Berkovic, S.F.; Petrou, S. A Childhood Epilepsy Mutation Reveals a Role for Developmentally Regulated Splicing of a Sodium Channel. Mol. Cell. Neurosci. 2007, 35, 292–301. [Google Scholar] [CrossRef] [PubMed]
- Ganguly, S.; Thompson, C.H.; George, A.L. Enhanced Slow Inactivation Contributes to Dysfunction of a Recurrent SCN2A Mutation Associated with Developmental and Epileptic Encephalopathy. J. Physiol. 2021, 599, 4375–4388. [Google Scholar] [CrossRef] [PubMed]
- Jérôme, C.; Christopher, T.; Susan, S.; Jinan, J.; Marina, T.; Simona, B. Rare Dysfunctional SCN2A Variants Are Associated with Malformation of Cortical Development. Epilepsia 2024, 66, 914–928. [Google Scholar] [CrossRef]
- Howell, K.B.; McMahon, J.M.; Carvill, G.L.; Tambunan, D.; Mackay, M.T.; Rodriguez-Casero, V.; Webster, R.; Clark, D.; Freeman, J.L.; Calvert, S.; et al. SCN2A Encephalopathy: A Major Cause of Epilepsy of Infancy with Migrating Focal Seizures. Neurology 2015, 85, 958–966. [Google Scholar] [CrossRef]
- Ben-Shalom, R.; Keeshen, C.M.; Berrios, K.N.; An, J.Y.; Sanders, S.J.; Bender, K.J. Opposing Effects on Na V 1.2 Function Underlie Differences Between SCN2A Variants Observed in Individuals With Autism Spectrum Disorder or Infantile Seizures. Biol. Psychiatry 2017, 82, 224–232. [Google Scholar] [CrossRef]
- Asadollahi, R.; Delvendahl, I.; Muff, R.; Tan, G.; Rodríguez, D.G.; Turan, S.; Russo, M.; Oneda, B.; Joset, P.; Boonsawat, P.; et al. Pathogenic SCN2A Variants Cause Early-Stage Dysfunction in Patient-Derived Neurons. Hum. Mol. Genet. 2023, 32, 2192–2204. [Google Scholar] [CrossRef]
- Géza, B.; Tao, E.; Katherine, H.; Rohini, C.; Erik, A.; Kris, K.; Markus, W.; Ben, C.; Steven, P. Nav1.2 Channel Mutations Preventing Fast Inactivation Lead to SCN2A Encephalopathy. Brain 2025, 148, 212–226. [Google Scholar]
- Backwell, L.; Marsh, J.A. Diverse Molecular Mechanisms Underlying Pathogenic Protein Mutations: Beyond the Loss-of-Function Paradigm. Annu. Rev. Genom. Hum. Genet. 2022, 23, 475–498. [Google Scholar] [CrossRef]
- Gerasimavicius, L.; Livesey, B.J.; Marsh, J.A. Loss-of-Function, Gain-of-Function and Dominant-Negative Mutations Have Profoundly Different Effects on Protein Structure. Nat. Commun. 2022, 13, 3895. [Google Scholar] [CrossRef]
- Orhan, G.; Bock, M.; Schepers, D.; Ilina, E.I.; Reichel, S.N.; Löffler, H.; Jezutkovic, N.; Weckhuysen, S.; Mandelstam, S.; Suls, A.; et al. Dominant-Negative Effects of KCNQ2 Mutations Are Associated with Epileptic Encephalopathy. Ann. Neurol. 2014, 75, 382–394. [Google Scholar] [CrossRef]
- Zhu, B.; Mak, J.C.H.; Morris, A.P.; Marson, A.G.; Barclay, J.W.; Sills, G.J.; Morgan, A. Functional Analysis of Epilepsy-Associated Variants in STXBP1/Munc18-1 Using Humanized Caenorhabditis Elegans. Epilepsia 2020, 61, 810–821. [Google Scholar] [CrossRef]
- Liu, A.; Xu, X.; Yang, X.; Jiang, Y.; Yang, Z.; Liu, X.; Wu, Y.; Wu, X.; Wei, L.; Zhang, Y. The Clinical Spectrum of Female Epilepsy Patients with PCDH19 Mutations in a Chinese Population. Clin. Genet. 2017, 91, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Carvill, G.L.; Matheny, T.; Hesselberth, J.; Demarest, S. Haploinsufficiency, Dominant Negative, and Gain-of-Function Mechanisms in Epilepsy: Matching Therapeutic Approach to the Pathophysiology. Neurotherapeutics 2021, 18, 1500–1514. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, J.E.; Meisler, M.H. Sodium Channel SCN8A (Nav1.6): Properties and de Novo Mutations in Epileptic Encephalopathy and Intellectual Disability. Front. Genet. 2013, 4, 213. [Google Scholar] [CrossRef]
- Goldfarb, M.; Schoorlemmer, J.; Williams, A.; Diwakar, S.; Wang, Q.; Huang, X.; Giza, J.; Tchetchik, D.; Kelley, K.; Vega, A.; et al. Fibroblast Growth Factor Homologous Factors Control Neuronal Excitability through Modulation of Voltage-Gated Sodium Channels. Neuron 2007, 55, 449–463. [Google Scholar] [CrossRef] [PubMed]
- Laezza, F.; Gerber, B.R.; Lou, J.-Y.; Kozel, M.A.; Hartman, H.; Craig, A.M.; Ornitz, D.M.; Nerbonne, J.M. The FGF14 F145S Mutation Disrupts the Interaction of FGF14 with Voltage-Gated Na + Channels and Impairs Neuronal Excitability. J. Neurosci. 2007, 27, 12033–12044. [Google Scholar] [CrossRef]
- Liu, J.-P.; Laufer, E.; Jessell, T.M. Assigning the Positional Identity of Spinal Motor Neurons. Neuron 2001, 32, 997–1012. [Google Scholar] [CrossRef]
- Lou, J.Y.; Laezza, F.; Gerber, B.R.; Xiao, M.; Yamada, K.A.; Hartmann, H.; Craig, A.M.; Nerbonne, J.M.; Ornitz, D.M. Fibroblast Growth Factor 14 Is an Intracellular Modulator of Voltage-Gated Sodium Channels. J. Physiol. 2005, 569, 179–193. [Google Scholar] [CrossRef]
- Laezza, F.; Lampert, A.; Kozel, M.A.; Gerber, B.R.; Rush, A.M.; Nerbonne, J.M.; Waxman, S.G.; Dib-Hajj, S.D.; Ornitz, D.M. FGF14 N-Terminal Splice Variants Differentially Modulate Nav1.2 and Nav1.6-Encoded Sodium Channels. Mol. Cell. Neurosci. 2009, 42, 90–101. [Google Scholar] [CrossRef]
- Bergren, S.K.; Chen, S.; Galecki, A.; Kearney, J.A. Genetic Modifiers Affecting Severity of Epilepsy Caused by Mutation of Sodium Channel Scn2a. Mamm. Genome 2005, 16, 683–690. [Google Scholar] [CrossRef]
- Larsen, J.; Carvill, G.L.; Gardella, E.; Kluger, G.; Schmiedel, G.; Barisic, N.; Depienne, C.; Brilstra, E.; Mang, Y.; Nielsen, J.E.K.; et al. The Phenotypic Spectrum of SCN8A Encephalopathy. Neurology 2015, 84, 480–489. [Google Scholar] [CrossRef]
- Gardella, E.; Marini, C.; Trivisano, M.; Fitzgerald, M.P.; Alber, M.; Howell, K.B.; Darra, F.; Siliquini, S.; Bölsterli, B.K.; Masnada, S.; et al. The Phenotype of SCN8A Developmental and Epileptic Encephalopathy. Neurology 2018, 91, e1112–e1124. [Google Scholar] [CrossRef] [PubMed]
- Barker, B.S.; Ottolini, M.; Wagnon, J.L.; Hollander, R.M.; Meisler, M.H.; Patel, M.K. The SCN8A Encephalopathy Mutation p.Ile1327Val Displays Elevated Sensitivity to the Anticonvulsant Phenytoin. Epilepsia 2016, 57, 1458–1466. [Google Scholar] [CrossRef] [PubMed]
- Wagnon, J.L.; Barker, B.S.; Ottolini, M.; Park, Y.; Volkheimer, A.; Valdez, P.; Swinkels, M.E.M.; Patel, M.K.; Meisler, M.H. Loss-of-Function Variants of SCN8A in Intellectual Disability without Seizures. Neurol. Genet. 2017, 3, e170. [Google Scholar] [CrossRef]
- Kurjak, A.; Pooh, R.K.; Merce, L.T.; Carrera, J.M.; Salihagic-Kadic, A.; Andonotopo, W. Structural and Functional Early Human Development Assessed by Three-Dimensional and Four-Dimensional Sonography. Fertil. Steril. 2005, 84, 1285–1299. [Google Scholar] [CrossRef] [PubMed]
- Royeck, M.; Horstmann, M.T.; Remy, S.; Reitze, M.; Yaari, Y.; Beck, H. Role of Axonal NaV1.6 Sodium Channels in Action Potential Initiation of CA1 Pyramidal Neurons. J. Neurophysiol. 2008, 100, 2361–2380. [Google Scholar] [CrossRef]
- Wagnon, J.L.; Meisler, M.H. Recurrent and Non-Recurrent Mutations of SCN8A in Epileptic Encephalopathy. Front. Neurol. 2015, 6, 104. [Google Scholar] [CrossRef]
- Meisler, M.H.; Helman, G.; Hammer, M.F.; Fureman, B.E.; Gaillard, W.D.; Goldin, A.L.; Hirose, S.; Ishii, A.; Kroner, B.L.; Lossin, C.; et al. SCN8A Encephalopathy: Research Progress and Prospects. Epilepsia 2016, 57, 1027–1035. [Google Scholar] [CrossRef]
- Katz, E.; Stoler, O.; Scheller, A.; Khrapunsky, Y.; Goebbels, S.; Kirchhoff, F.; Gutnick, M.J.; Wolf, F.; Fleidervish, I.A. Role of Sodium Channel Subtype in Action Potential Generation by Neocortical Pyramidal Neurons. Proc. Natl. Acad. Sci. USA 2018, 115, E7184–E7192. [Google Scholar] [CrossRef] [PubMed]
- Bunton-Stasyshyn, R.K.A.; Wagnon, J.L.; Wengert, E.R.; Barker, B.S.; Faulkner, A.; Wagley, P.K.; Bhatia, K.; Jones, J.M.; Maniaci, M.R.; Parent, J.M.; et al. Prominent Role of Forebrain Excitatory Neurons in SCN8A Encephalopathy. Brain 2019, 142, 362–375. [Google Scholar] [CrossRef] [PubMed]
- Menezes, L.F.S.; Sabiá Júnior, E.F.; Tibery, D.V.; Carneiro, L.d.A.; Schwartz, E.F. Epilepsy-Related Voltage-Gated Sodium Channelopathies: A Review. Front. Pharmacol. 2020, 11, 1276. [Google Scholar] [CrossRef]
- Liu, Y.; Koko, M.; Lerche, H. A SCN8A Variant Associated with Severe Early Onset Epilepsy and Developmental Delay: Loss- or Gain-of-Function? Epilepsy Res. 2021, 178, 106824. [Google Scholar] [CrossRef] [PubMed]
- Fry, A.E.; Marra, C.; Derrick, A.V.; Pickrell, W.O.; Higgins, A.T.; te Water Naude, J.; McClatchey, M.A.; Davies, S.J.; Metcalfe, K.A.; Tan, H.J.; et al. Missense Variants in the N-Terminal Domain of the A Isoform of FHF2/FGF13 Cause an X-Linked Developmental and Epileptic Encephalopathy. Am. J. Hum. Genet. 2021, 108, 176–185. [Google Scholar] [CrossRef]
- Miralles, R.M.; Boscia, A.R.; Kittur, S.; Jessica, C.H.; Payal, S.P. Parvalbumin Interneuron Impairment Causes Synaptic Transmission Deficits and Seizures in SCN8A Developmental and Epileptic Encephalopathy. JCI Insight 2024, 9, e181005. [Google Scholar] [CrossRef]
- Vanoye, C.G.; Abramova, T.V.; DeKeyser, J.M. Molecular and Cellular Context Influences SCN8A Variant Function. JCI Insight 2024, 9, e177530. [Google Scholar] [CrossRef]
- Plummer, N.W.; McBurney, M.W.; Meisler, M.H. Alternative Splicing of the Sodium Channel SCN8A Predicts a Truncated Two-Domain Protein in Fetal Brain and Non-Neuronal Cells. J. Biol. Chem. 1997, 272, 24008–24015. [Google Scholar] [CrossRef]
- Liu, Y.; Schubert, J.; Sonnenberg, L.; Helbig, K.L.; Hoei-Hansen, C.E.; Koko, M.; Rannap, M.; Lauxmann, S.; Huq, M.; Schneider, M.C.; et al. Neuronal Mechanisms of Mutations in SCN8A Causing Epilepsy or Intellectual Disability. Brain 2019, 142, 376–390. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Gao, H.; Bao, X.; Zhang, Q.; Li, J.; Wei, L.; Wu, X.; Chen, Y.; Yu, S. SCN8A Mutations in Chinese Patients with Early Onset Epileptic Encephalopathy and Benign Infantile Seizures. BMC Med. Genet. 2017, 18, 104. [Google Scholar] [CrossRef]
- Veeramah, K.R.; O’Brien, J.E.; Meisler, M.H.; Cheng, X.; Dib-Hajj, S.D.; Waxman, S.G.; Talwar, D.; Girirajan, S.; Eichler, E.E.; Restifo, L.L.; et al. De Novo Pathogenic SCN8A Mutation Identified by Whole-Genome Sequencing of a Family Quartet Affected by Infantile Epileptic Encephalopathy and SUDEP. Am. J. Hum. Genet. 2012, 90, 502–510. [Google Scholar] [CrossRef]
- Blanchard, M.G.; Willemsen, M.H.; Walker, J.B.; Dib-Hajj, S.D.; Waxman, S.G.; Jongmans, M.C.J.; Kleefstra, T.; van de Warrenburg, B.P.; Praamstra, P.; Nicolai, J.; et al. De Novo Gain-of-Function and Loss-of-Function Mutations of SCN8A in Patients with Intellectual Disabilities and Epilepsy. J. Med. Genet. 2015, 52, 330–337. [Google Scholar] [CrossRef]
- Meisler, M.H. SCN8A Encephalopathy: Mechanisms and Models. Epilepsia 2019, 60, S86–S91. [Google Scholar] [CrossRef]
- Berghuis, B.; de Kovel, C.G.F.; van Iterson, L.; Lamberts, R.J.; Sander, J.W.; Lindhout, D.; Koeleman, B.P.C. Complex SCN8A DNA-Abnormalities in an Individual with Therapy Resistant Absence Epilepsy. Epilepsy Res. 2015, 115, 141–144. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rudy, B.; Fishell, G.; Lee, S.H.; Hjerling-Leffler, J. Three Groups of Interneurons Account for Nearly 100% of Neocortical GABAergic Neurons. Dev. Neurobiol. 2011, 71, 45–61. [Google Scholar] [CrossRef] [PubMed]
- Tremblay, R.; Lee, S.; Rudy, B. GABAergic Interneurons in the Neocortex: From Cellular Properties to Circuits. Neuron 2016, 91. [Google Scholar] [CrossRef]
- Gouwens, N.W.; Sorensen, S.A.; Baftizadeh, F.; Budzillo, A.; Lee, B.R.; Jarsky, T.; Alfiler, L.; Baker, K.; Barkan, E.; Berry, K.; et al. Integrated Morphoelectric and Transcriptomic Classification of Cortical GABAergic Cells. Cell 2020, 183, 935–953.e19. [Google Scholar] [CrossRef]
- Scheffer, I.E.; Berkovic, S.; Capovilla, G.; Connolly, M.B.; French, J.; Guilhoto, L.; Hirsch, E.; Jain, S.; Mathern, G.W.; Moshé, S.L.; et al. ILAE Classification of the Epilepsies: Position Paper of the ILAE Commission for Classification and Terminology. Epilepsia 2017, 58, 512–521. [Google Scholar] [CrossRef]
- Scher, M.S. Neonatal Seizure Classification: A Fetal Perspective Concerning Childhood Epilepsy. Epilepsy Res. 2006, 70, 41–57. [Google Scholar] [CrossRef]
- Ohira, R.; Zhang, Y.H.; Guo, W.; Dipple, K.; Shih, S.L.; Doerr, J.; Huang, B.L.; Fu, L.J.; Abu-Khalil, A.; Geschwind, D.; et al. Human ARX Gene: Genomic Characterization and Expression. Mol. Genet. Metab. 2002, 77, 179–188. [Google Scholar] [CrossRef]
- Molinari, F.; Raas-Rothschild, A.; Rio, M.; Fiermonte, G.; Encha-Razavi, F.; Palmieri, L.; Palmieri, F.; Ben-Neriah, Z.; Kadhom, N.; Vekemans, M.; et al. Impaired Mitochondrial Glutamate Transport in Autosomal Recessive Neonatal Myoclonic Epilepsy. Am. J. Hum. Genet. 2005, 76, 334–339. [Google Scholar] [CrossRef]
- Kanaumi, T.; Takashima, S.; Iwasaki, H.; Itoh, M.; Mitsudome, A.; Hirose, S. Developmental Changes in KCNQ2 and KCNQ3 Expression in Human Brain: Possible Contribution to the Age-Dependent Etiology of Benign Familial Neonatal Convulsions. Brain Dev. 2008, 30, 362–369. [Google Scholar] [CrossRef]
- Jansen, L.A.; Hevner, R.F.; Roden, W.H.; Hahn, S.H.; Jung, S.; Gospe, S.M. Glial Localization of Antiquitin: Implications for Pyridoxine-Dependent Epilepsy. Ann. Neurol. 2014, 75, 22–32. [Google Scholar] [CrossRef]
- Mcleod, F.; Dimtsi, A.; Marshall, A.C.; Lewis-Smith, D.; Thomas, R.; Clowry, G.J.; Trevelyan, A.J. Altered Synaptic Connectivity in an in Vitro Human Model of STXBP1 Encephalopathy. Brain 2023, 146, 850–857. [Google Scholar] [CrossRef] [PubMed]
- Brunklaus, A.; Brünger, T.; Feng, T.; Fons, C.; Lehikoinen, A.; Panagiotakaki, E.; Vintan, M.A.; Symonds, J.; Andrew, J.; Arzimanoglou, A.; et al. The Gain of Function SCN1A Disorder Spectrum: Novel Epilepsy Phenotypes and Therapeutic Implications. Brain 2022, 145, 3816–3831. [Google Scholar] [CrossRef] [PubMed]
- Geng, Y.; Hou, X. KCNQ2-Neonatal Epileptic Encephalopathy Complicated by Ventricular Tachycardia: A Case Report. Front Neurol 2020, 11, 263. [Google Scholar] [CrossRef]
- Zeng, Q.; Yang, X.; Zhang, J.; Liu, A.; Yang, Z.; Liu, X.; Wu, Y.; Wu, X.; Wei, L.; Zhang, Y. Genetic Analysis of Benign Familial Epilepsies in the First Year of Life in a Chinese Cohort. J. Hum. Genet. 2018, 63, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Miceli, F.; Striano, P.; Soldovieri, M.V.; Fontana, A.; Nardello, R.; Robbiano, A.; Bellini, G.; Elia, M.; Zara, F.; Taglialatela, M.; et al. A Novel KCNQ3 Mutation in Familial Epilepsy with Focal Seizures and Intellectual Disability. Epilepsia 2015, 56, e15–e20. [Google Scholar] [CrossRef]
- Maljevic, S.; Vejzovic, S.; Bernhard, M.K.; Bertsche, A.; Weise, S.; Döcker, M.; Lerche, H.; Lemke, J.R.; Merkenschlager, A.; Syrbe, S. Novel KCNQ3 Mutation in a Large Family with Benign Familial Neonatal Epilepsy: A Rare Cause of Neonatal Seizures. Mol. Syndromol. 2016, 7, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Alsaleem, M.; Carrion, V.; Weinstock, A.; Chandrasekharan, P. Infantile Refractory Seizures Due to de Novo KCNT 1 Mutation. BMJ Case Rep. 2019, 12, e231178. [Google Scholar] [CrossRef]
- Barcia, G.; Chemaly, N.; Kuchenbuch, M.; Eisermann, M.; Gobin-Limballe, S.; Ciorna, V.; Macaya, A.; Lambert, L.; Dubois, F.; Doummar, D.; et al. Epilepsy with Migrating Focal SeizuresKCNT1 Mutation Hotspots and Phenotype Variability. Neurol. Genet. 2019, 5, e363. [Google Scholar] [CrossRef]
- Di Meglio, C.; Lesca, G.; Villeneuve, N.; Lacoste, C.; Abidi, A.; Cacciagli, P.; Altuzarra, C.; Roubertie, A.; Afenjar, A.; Renaldo-Robin, F.; et al. Epileptic Patients with de Novo STXBP1 Mutations: Key Clinical Features Based on 24 Cases. Epilepsia 2015, 56, 1931–1940. [Google Scholar] [CrossRef]
- Kato, M.; Saitoh, S.; Kamei, A.; Shiraishi, H.; Ueda, Y.; Akasaka, M.; Tohyama, J.; Akasaka, N.; Hayasaka, K. A Longer Polyalanine Expansion Mutation in the ARX Gene Causes Early Infantile Epileptic Encephalopathy with Suppression-Burst Pattern (Ohtahara Syndrome). Am. J. Hum. Genet. 2007, 81, 361–366. [Google Scholar] [CrossRef]
- André, M.V.; Cacciagli, P.; Cano, A.; Vaugier, L.; Roussel, M.; Girard, N.; Chabrol, B.; Villard, L.; Milh, M. The Phenotype Caused by Recessive Variations in SLC25A22: Report of a New Case and Literature Review. Arch. Pediatr. 2021, 28, 87–92. [Google Scholar] [CrossRef]
- Matricardi, S.; De Liso, P.; Freri, E.; Costa, P.; Castellotti, B.; Magri, S.; Gellera, C.; Granata, T.; Musante, L.; Lesca, G.; et al. Neonatal Developmental and Epileptic Encephalopathy Due to Autosomal Recessive Variants in SLC13A5 Gene. Epilepsia 2020, 61, 2474–2485. [Google Scholar] [CrossRef] [PubMed]
- Toldo, I.; Bonardi, C.M.; Bettella, E.; Polli, R.; Talenti, G.; Burlina, A.; Sartori, S.; Murgia, A. Brain Malformations Associated to Aldh7a1 Gene Mutations: Report of a Novel Homozygous Mutation and Literature Review. Eur. J. Paediatr. Neurol. 2018, 22, 1042–1053. [Google Scholar] [CrossRef]
- Saeedi, M.; Mirnia, K.; Sedighzadeh, S.; Sangsari, R. Neonatal Refractory Seizures and Hyperammonemia in a Neonate With ALDH7A1 Deficiency. Clin. Case Rep. 2025, 13, e70852. [Google Scholar] [CrossRef]
- Medyanik, A.D.; Anisimova, P.E.; Kustova, A.O.; Tarabykin, V.S.; Kondakova, E.V. Developmental and Epileptic Encephalopathy: Pathogenesis of Intellectual Disability Beyond Channelopathies. Biomolecules 2025, 15, 133. [Google Scholar] [CrossRef] [PubMed]
- Bahi-Buisson, N.; Poirier, K.; Fourniol, F.; Saillour, Y.; Valence, S.; Lebrun, N.; Hully, M.; Fallet Bianco, C.; Boddaert, N.; Elie, C.; et al. The Wide Spectrum of Tubulinopathies: What Are the Key Features for the Diagnosis? Brain 2014, 137, 1676–1700. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Jancovski, N.; Jafar-Nejad, P.; Burbano, L.E.; Rollo, B.; Richards, K.; Drew, L.; Sedo, A.; Heighway, J.; Pachernegg, S.; et al. Antisense Oligonucleotide Therapy Reduces Seizures and Extends Life Span in an SCN2A Gain-of-Function Epilepsy Model. J. Clin. Investig. 2021, 131, e152079. [Google Scholar] [CrossRef] [PubMed]
- Lenk, G.M.; Jafar-Nejad, P.; Hill, S.F.; Huffman, L.D.; Smolen, C.E.; Wagnon, J.L.; Petit, H.; Yu, W.; Ziobro, J.; Bhatia, K.; et al. SCN8A Antisense Oligonucleotide Is Protective in Mouse Models of SCN8A Encephalopathy and Dravet Syndrome. Ann. Neurol. 2020, 87, 339–346. [Google Scholar] [CrossRef] [PubMed]

| Gene Names | Function | Earliest Reported Age | Phenotype/Epilepsy Type |
|---|---|---|---|
| SCN1A | Encodes the voltage-gated sodium channel α-subunit Nav1.1 essential for action potential initiation and neuronal excitability | Within the first 3 days of life [97] | Neonatal developmental and epileptic encephalopathy with MD and arthrogryposis (NDEEMA) [97] |
| KCNQ2 | Encodes the Kv7.2 subunit of M-type voltage-gated potassium channels that regulate neuronal excitability | 12 h postnatal [98] Postnatal day 1 [99] | Neonatal epileptic encephalopathy [98] Benign familial neonatal epilepsy (BFNE) [99] |
| KCNQ3 | Encodes the Kv7.3 subunit of voltage-gated potassium channels that co-assemble with Kv7.2 to form functional M-channels | Postnatal day 2 [100] Within the first week of life [101] | Familial epilepsy with focal seizures and intellectual disability [100] Benign familial neonatal epilepsy (BFNE) [101] |
| KCNT1 | Encodes a sodium-activated potassium channel (Slack/KNa1.1) involved in regulating neuronal firing patterns | Postnatal day 4 [102] From 1 h postnatal [103] | Nocturnal frontal lobe epilepsy and epilepsy of infancy with migrating focal seizures [102] Epilepsy of infancy with migrating focal seizures (EIMFS) [103] |
| STXBP1 | Encodes Munc18-1, an essential SNARE complex for synaptic vesicle docking and neurotransmitter release | From postnatal day 1 [104] | Early-onset epileptic encephalopathy (EOEE), Ohtahara syndrome, West syndrome [104] |
| ARX | Encodes a paired-like homeobox transcription factor involved in regulating neuronal proliferation, migration and differentiation during brain development | 40 min postnatal [105] | Ohtahara syndrome [105] |
| SLC25A22 | Encodes a mitochondrial glutamate carrier that imports glutamate into mitochondria | From postnatal day 1 [106] | Ohtahara syndrome [106] |
| SLC13A5 | Encodes the Na+-coupled citrate transporter (NaCT) that mediates cellular citrate uptake and links citrate availability to neuronal metabolism | Postnatal days 1–2 [107] | Neonatal onset DEE with nonmigrating clonic seizures [107] |
| ALDH7A1 | Encodes antiquitin (α-aminoadipic semialdehyde dehydrogenase), an enzyme in lysine catabolism | 3 h postnatal [108] Postnatal day 3 [109] | Pyridoxine-dependent epilepsy [108,109] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Kustova, A.O.; Medyanik, A.D.; Anisimova, P.E.; Tarabykin, V.S.; Kondakova, E.V. Neurologist’s Black Swan: Molecular Basis of Prenatal Seizures. Int. J. Mol. Sci. 2026, 27, 283. https://doi.org/10.3390/ijms27010283
Kustova AO, Medyanik AD, Anisimova PE, Tarabykin VS, Kondakova EV. Neurologist’s Black Swan: Molecular Basis of Prenatal Seizures. International Journal of Molecular Sciences. 2026; 27(1):283. https://doi.org/10.3390/ijms27010283
Chicago/Turabian StyleKustova, Angelina O., Alexandra D. Medyanik, Polina E. Anisimova, Victor S. Tarabykin, and Elena V. Kondakova. 2026. "Neurologist’s Black Swan: Molecular Basis of Prenatal Seizures" International Journal of Molecular Sciences 27, no. 1: 283. https://doi.org/10.3390/ijms27010283
APA StyleKustova, A. O., Medyanik, A. D., Anisimova, P. E., Tarabykin, V. S., & Kondakova, E. V. (2026). Neurologist’s Black Swan: Molecular Basis of Prenatal Seizures. International Journal of Molecular Sciences, 27(1), 283. https://doi.org/10.3390/ijms27010283





