The Functional State of Thermoplasma acidophilum Pyruvate Kinase Relies on an Extra Carboxyl-Terminal Sequence
Abstract
1. Introduction
2. Results and Discussion
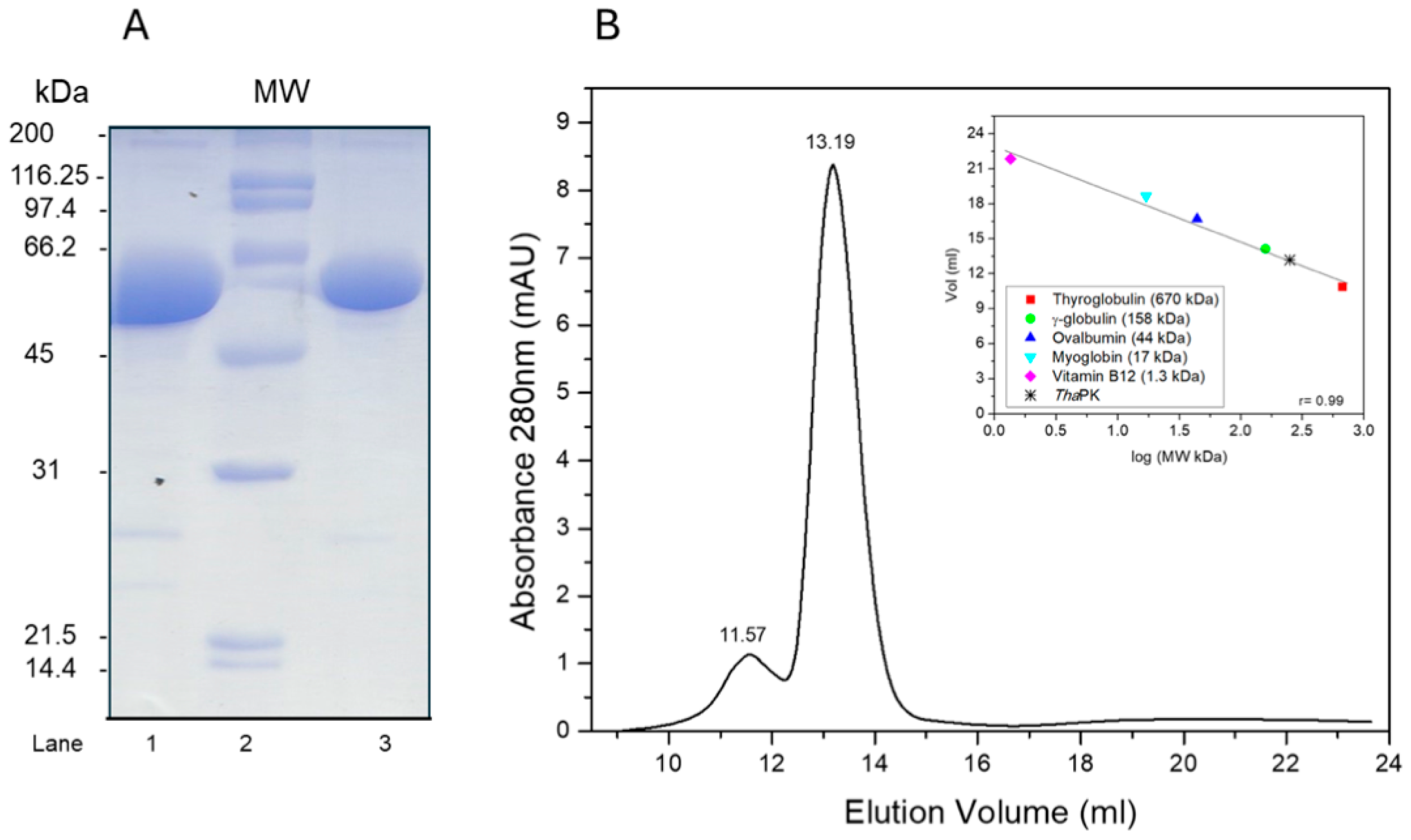
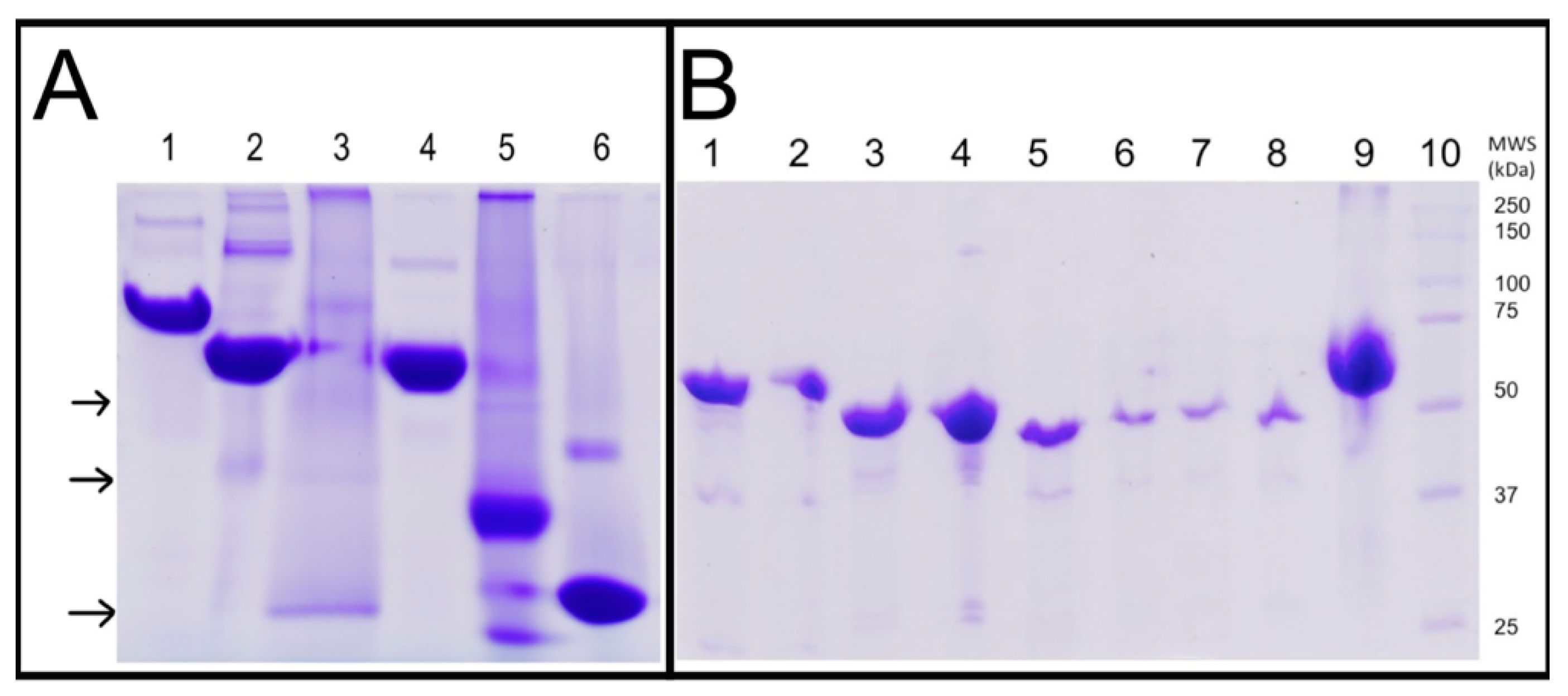
2.1. Purification and Oligomeric State of WT-ThaPK
2.2. pH Profile of WT-ThaPK
2.3. Effect of Monovalent Cations and Allosteric Effectors on WT-ThaPK
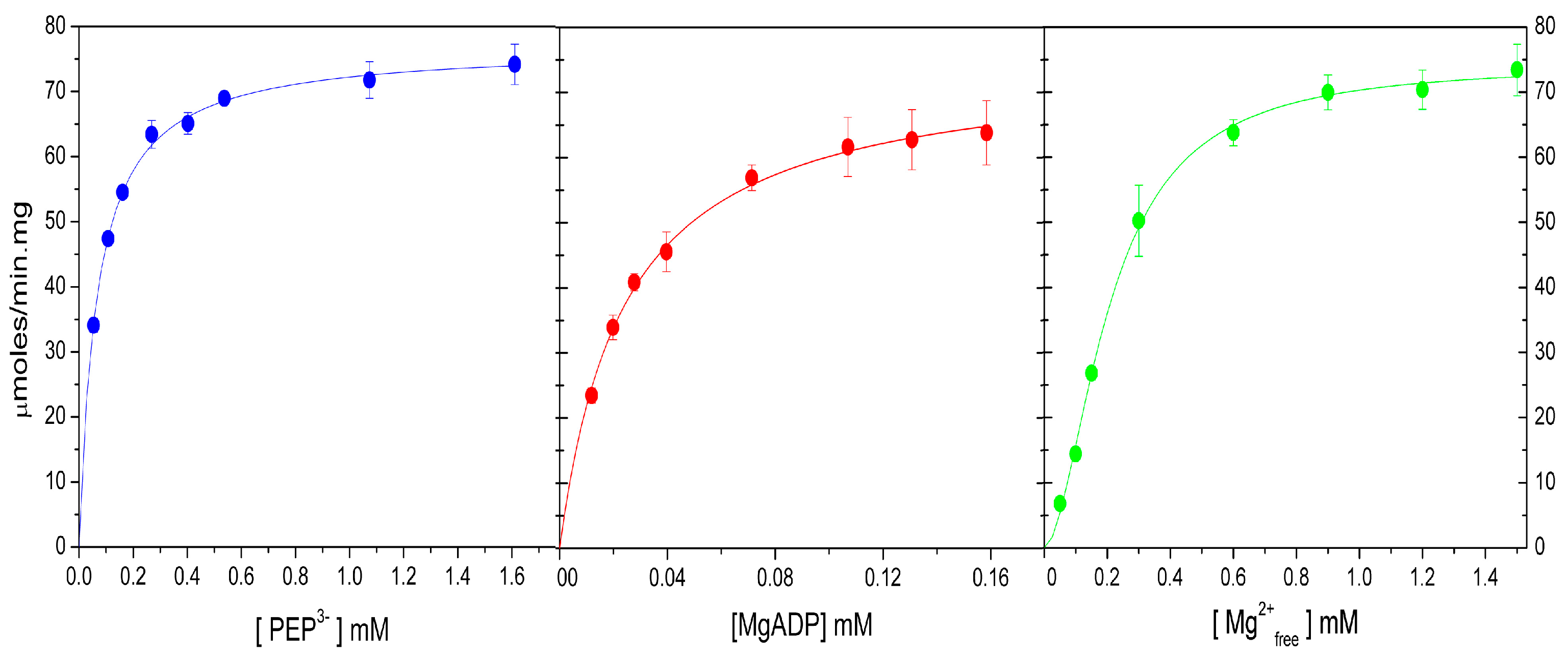
2.4. Kinetic Parameters of WT-ThaPK in the Presence of Mg2+ and Mn2+
2.5. Bi-Substrate Kinetics of WT-ThaPK
2.6. Dead-End Inhibition Studies of WT-ThaPK
2.7. The Structural Model of the WT-ThaPK
2.8. Kinetic Parameters for ECTS Truncated-ThaPK
2.9. Circular Dichroism Spectra and Differential Scanning Calorimetry of WT-ThaPK and ECTS Truncated-ThaPK
2.10. Molecular Dynamics Simulations of WT-ThaPK and ECTS Truncated-ThaPK
2.11. D Blue Native Gel Electrophoresis and 2D SDS-PAGE of WT-ThaPK and ECTS-Truncated ThaPK
2.12. Is the Extra Carboxyl-Terminal Sequence (ECTS) Widely Distributed Along the PKs?
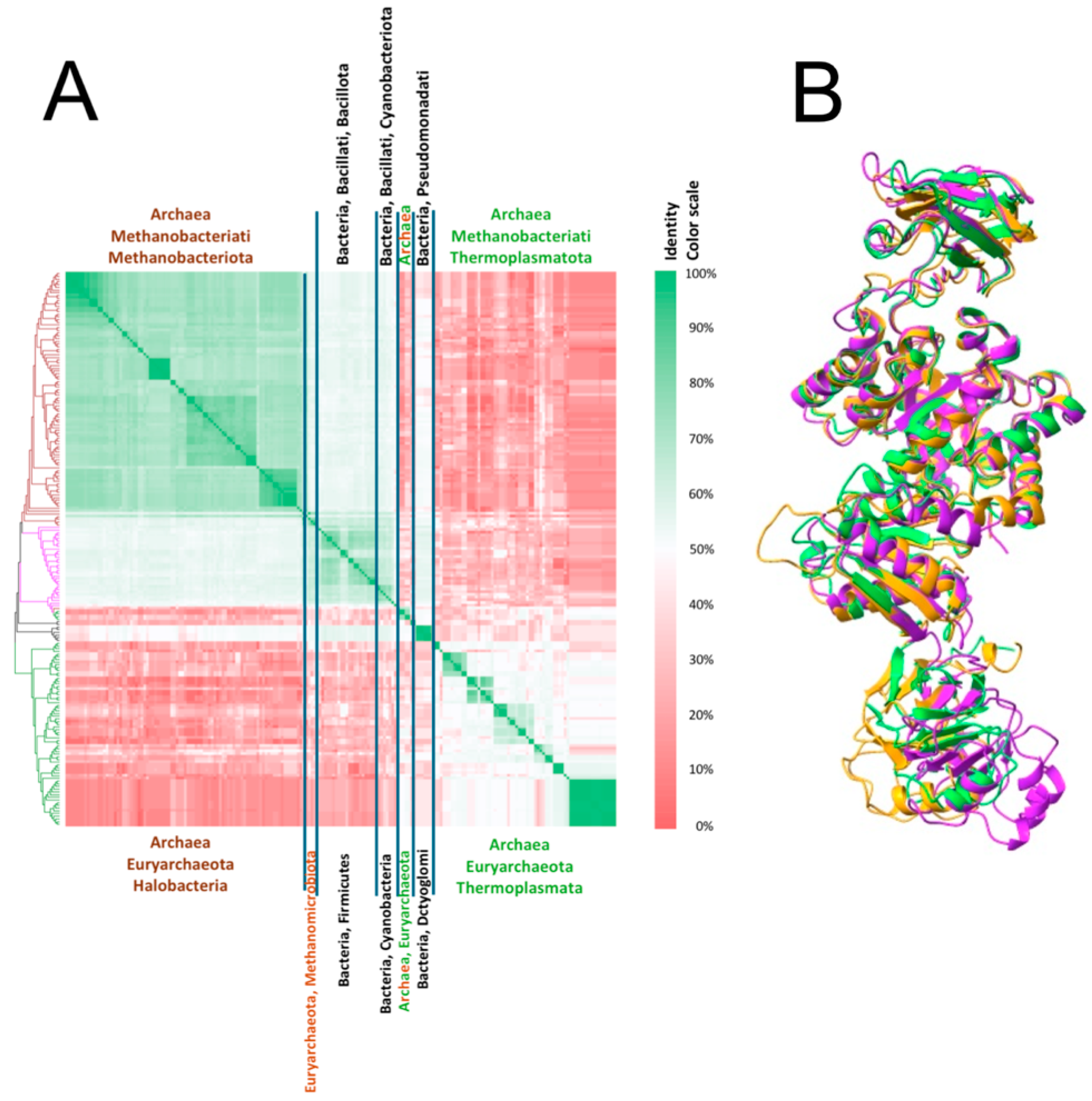
2.13. Structural Alignment in 3D of PKs That Possess ECTS and Belong to Different Taxonomic Groups
2.14. Are the Pyruvate Kinases That Possess an Extra C-Terminal Sequence (ECTS) Clustered Together in a Global Phylogenetic Tree?
3. Materials and Methods
3.1. Chemicals
3.2. Cloning and Expression of WT-ThaPK
3.3. Cloning and Expression of ECTS Truncated-ThaPK
3.4. Cell Growth and Purification of WT-ThaPK and ECTS Truncated-ThaPK
3.5. Assays of Pyruvate Kinase Activity
3.6. Kinetic Studies
3.7. AlphaFold Model of WT-ThaPK
3.8. Docking
3.9. Circular Dichroism Experiments
3.10. Differential Scanning Calorimetry (DSC)
3.11. Molecular Dynamics Simulations
3.12. 1D Blue Native-PAGE and 2D SDS-PAGE
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Oria-Hernández, J.; Riveros-Rosas, H.; Ramírez-Sílva, L. Dichotomic phylogenetic tree of the pyruvate kinase family: K+-dependent and -independent enzymes. J. Biol. Chem. 2006, 281, 30717–30724. [Google Scholar] [CrossRef]
- De la Vega-Ruíz, G.; Domínguez-Ramírez, L.; Riveros-Rosas, H.; Guerrero-Mendiola, C.; Torres-Larios, A.; Hernández-Alcántara, G.; García-Trejo, J.J.; Ramírez-Silva, L. New insights on the mechanism of the K+-independent activity of crenarchaeota pyruvate kinases. PLoS ONE 2015, 10, e0119233, Correction in PLoS ONE 2015, 10, e0129757. [Google Scholar] [CrossRef][Green Version]
- Göker, M.; Oren, A. Valid publication of names of two domains and seven kingdoms of prokaryotes. Int. J. Syst. Evol. Microbiol. 2024, 74, 006242. [Google Scholar] [CrossRef]
- Potter, S.; Fothergill-Gilmore, L.A. Purification and properties of pyruvate kinase from Thermoplasma acidophilum. FEMS Microbiol. Lett. 1992, 94, 235–240. [Google Scholar] [CrossRef]
- Johnsen, U.; Reinhardt, A.; Landan, G.; Tria, F.D.K.; Turner, J.M.; Davies, C.; Schónheit, P. New views on an old enzyme: Allosteric regulation and evolution of archaeal pyruvate kinases. FEBS J. 2019, 286, 2471–2489. [Google Scholar] [CrossRef]
- Ramírez-Silva, L.; Hernández-Alcántara, G.; Guerrero-Mendiola, C.; González-Andrade, M.; Rodríguez-Romero, A.; Rodríguez-Hernández, A.; Lugo-Munguía, A.; Gómez-Coronado, P.A.; Rodríguez-Méndez, C.; Vega-Segura, A. The K+-Dependent and -Independent Pyruvate Kinases Acquire the Active Conformation by Different Mechanisms. Int. J. Mol. Sci. 2022, 23, 1347. [Google Scholar] [CrossRef]
- Sakai, H.; Ohta, T. Molecular cloning and nucleotide sequence of the gene for pyruvate kinase of Bacillus stearothermophilus and the production of the enzyme in Escherichia coli. Evidence that the genes for phosphofructokinase and pyruvate kinase constitute an operon. Eur. J. Biochem. 1993, 211, 851–859. [Google Scholar] [CrossRef]
- Tanaka, K.; Sakai, H.; Ohta, T.; Matsuzawa, H. Molecular cloning of the genes for pyruvate kinase of two bacilli, Bacillus psychrophilus and Bacillus licheniformis, and comparison of the properties of the enzymes produced in Escherichia coli. Biosci. Biotechnol. Biochem. 1995, 59, 1536–1542. [Google Scholar] [CrossRef]
- Muñoz, M.E.; Le Borgne, S.; Bolívar, F.; Valle, F. Molecular cloning of the gene that codes for the pyruvate kinase of Bacillus subtilis, primary characterization of a strain carrying this gene insertionally inactivated. Rev. Latinoam. Microbiol. 1997, 39, 129–140. [Google Scholar]
- Muñoz, M.E.; Ponce, E. Pyruvate kinase: Current status of regulatory and functional properties. Comp. Biochem. Physiol. B 2003, 135, 197–218. [Google Scholar] [CrossRef]
- Zoraghi, T.; Swayze, R.; Finlay, B.B.; Brunham, R.C.; McMaster, W.R.; Reiner, N.E. Functional analysis, overexpression and kinetic characterization of pyruvate kinase from Methicillin-Resistant Staphylococcus aureus. Biochemistry 2010, 49, 7733–7747. [Google Scholar] [CrossRef]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Guerrero-Mendiola, C.; García-Trejo, J.J.; Encalada, R.; Saavedra, E.; Ramírez-Silva, L. The contribution of two isozymes to the pyruvate kinase activity of Vibrio cholerae: One K+-dependent constitutively active and another K+-independent with essential allosteric activation. PLoS ONE 2017, 12, e0178673. [Google Scholar] [CrossRef]
- Ramírez-Silva, L.; Guerrero-Mendiola, C.; Cabrera, N. The Importance of Polarity in the Evolution of the K+ Binding Site of Pyruvate Kinase. Int. J. Mol. Sci. 2014, 15, 22214–22226. [Google Scholar] [CrossRef]
- Kayne, F.J. Thallium (I) activation of pyruvate kinase. Arch. Biochem. Biophys. 1971, 143, 232–239. [Google Scholar] [CrossRef]
- Ramírez-Silva, L.; Oria, J.; Gómez-Puyou, A.; Tuena de Gómez-Puyou, M. The contribution of water to the selectivity of pyruvate kinase for Na+ and K+. Eur. J. Biochem. 1997, 250, 583–589. [Google Scholar] [CrossRef]
- Mildvan, A.S.; Cohn, M. Kinetic and magnetic resonance studies of the pyruvate kinase reaction: I. Divalent Metal Complexes of Pyruvate Kinase. J. Biol. Chem. 1965, 240, 238–246. [Google Scholar] [CrossRef]
- Dupont, C.L.; Butcher, A.; Vaalas, R.E.; Bourne, P.E.; Caetano-Anallés, G. History of biological metal utilization inferred through phylogenomic analysis of protein structures. Proc. Natl. Acad. Sci. USA 2010, 107, 10567–10572. [Google Scholar] [CrossRef]
- Reynard, A.M.; Hass, L.F.; Jacobsen, D.D.; Boyer, P.D. The correlation of reaction kinetics and substrate binding with the mechanism of pyruvate kinase. J. Biol. Chem. 1961, 236, 2277–2283. [Google Scholar] [CrossRef]
- Ainsworth, S.; MacFarlane, N. A kinetic study of rabbit muscle pyruvate kinase. Biochem. J. 1973, 131, 223–236. [Google Scholar] [CrossRef]
- Oria-Hernández, J.; Cabrera, N.; Pérez-Montfort, R.; Ramírez-Silva, L. Pyruvate kinase revisited. The activating effect of K+. J. Biol. Chem. 2005, 280, 37294–37929. [Google Scholar] [CrossRef] [PubMed]
- Cleland, W.W. The Enzymes, 3rd ed.; Academic Press Inc.: New York, NY, USA, 1970; Volume 2, pp. 1–65. [Google Scholar]
- Reed, G.H.; Morgan, S.D. Kinetic and magnetic resonance studies of the interaction of oxalate with pyruvate kinase. Biochemistry 1974, 13, 3537–3541. [Google Scholar] [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef]
- Pocalyko, D.J.; Carroll, L.J.; Martin, B.M.; Babbitt, P.C.; Dunaway-Mariano, D. Analysis of sequence homologies in plant and bacterial pyruvate phosphate dikinase, enzyme I of the bacterial phosphoenolpyruvate: Sugar phosphotransferase system and other PEP-utilizing enzymes. Identification of potential catalytic and regulatory motifs. Biochemistry 1990, 29, 10757–10765. [Google Scholar] [CrossRef] [PubMed]
- Sakai, H. Possible structure and function of the extra C-terminal sequence of pyruvate kinase from Bacillus stearothermophilus. J. Biochem. 2004, 136, 471–476. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, K.; Ito, S.; Shimizu-Ibuka, A.; Sakai, H. Crystal Structure of Pyruvate Kinase from Geobacillus stearothermophilus. J. Biochem. 2008, 144, 305–312. [Google Scholar] [CrossRef]
- Zoraghi, R.; Worrall, L.; See, R.H.; Strangman, W.; Popplewell, W.L.; Gong, H.; Samaai, T.; Swayze, R.D.; Kaur, S.; Vuckovic, M.; et al. Methicillin-resistant Staphylococcus aureus (MRSA) Pyruvate Kinase as a Target for Bis-indole Alkaloids with Antibacterial Activities. J. Biol. Chem. 2011, 286, 44716–44725. [Google Scholar] [CrossRef]
- Adasme, M.F.; Linnemann, K.L.; Bolz, S.N.; Kaiser, F.; Salentin, S.; Haupt, V.J.; Schroeder, M. PLIP 2021: Expanding the scope of the protein–ligand interaction profiler to DNA and RNA. Nucleic Acids Res. 2021, 49, W530–W534. [Google Scholar] [CrossRef]
- Geenfield, N. Using circular dichroism spectra to estimate protein secondary structure. Nat. Protoc. 2006, 1, 2876–2890. [Google Scholar] [CrossRef]
- Goyal, M.; Chaudhuri, T.K.; Kuwajima, K. Irreversible denaturation of methanodextrin glucosidase studied by differential scanning calorimetry, circular dichroism, and turbidity measurements. PLoS ONE 2014, 9, e115877. [Google Scholar] [CrossRef]
- Roe, D.R.; Cheatham, T.E., III. PTRAJ and CPPTRAJ: Software for Processing and Analysis of Molecular Dynamics Trajectory Data. J. Chem. Theory Comput. 2013, 9, 3084–3095. [Google Scholar] [CrossRef]
- Bremer, N.; Martin, W.F.; Steel, M. Surprising effects of different loss in genome evolution: The last-one-out. FEMS Microbiol. Lett. 2025, 372, fnaf051. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.M.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Kasahara, M.; Penefsky, H.S. High affinity binding of monovalent Pi by beef heart mitochondrial adenosine triphosphatase. J. Biol. Chem. 1978, 253, 4180–4187. [Google Scholar] [CrossRef]
- Ramírez-Silva, L.; de Gómez-Puyou, M.T.; Gómez-Puyou, A. Water-induced transitions in the K+ requirements for the activity of pyruvate kinase entrapped in reverse micelles. Biochemistry 1993, 32, 5332–5338. [Google Scholar] [CrossRef]
- Büchner, T.; Pleiderer, G. Methods in Enzymol; Academic Press: New York, NY, USA, 1955; Volume 1, pp. 435–440. [Google Scholar]
- Schoemakers, J.M.; Visser, G.J.; Flik, G.; Theuvenet, P.R. CHELATOR: An improved method for computing metal ion concentrations in physiological solutions. Biotechniques 1992, 12, 870–879. [Google Scholar]
- Susan-Resiga, D.; Nowak, T. Proton donor in yeast pyruvate kinase: Chemical and kinetic properties of the active site Thr298 to Cys mutant. Biochemistry 2004, 43, 15230–15245. [Google Scholar] [CrossRef] [PubMed]
- Dougherty, T.M.; Cleland, W.W. pH Studies of the chemical mechanism of rabbit muscle pyruvate kinase. 2. Physiological substrates and phosphoenol-αketobutyrate. Biochemistry 1985, 24, 5875–5880. [Google Scholar] [CrossRef] [PubMed]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]
- Hanwell, M.D.; Curtis, D.E.; Lonie, D.C.; Vandermeersch, T.; Zurek, E.; Hutchison, G.R. Avogadro: An advanced semantic chemical editor, visualization, and analysis platform. J. Cheminform. 2012, 4, 17–49. [Google Scholar] [CrossRef]
- Masters, L.; Eagon, S.; Heying, M. Evaluation of consensus scoring methods for AutoDock Vina, smina and idock. J. Mol. Graph. Model. 2020, 96, 107532–107541. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general amber force field. J. Comput. Chem. 2004, 25, 1157–1174. [Google Scholar] [CrossRef]
- Case, D.A.; Aktulga, H.M.; Belfon, K.; Cerutti, D.S.; Cisneros, G.A.; Cruzeiro, V.W.D.; Forouzesh, N.; Giese, T.J.; Gotz, A.W.; Gohlke, H.; et al. AmberTools. J. Chem. Inf. Model. 2023, 63, 6183–6191. [Google Scholar] [CrossRef]
- Case, D.A.; Cheatham, T.E., 3rd; Darden, T.; Gohlke, H.; Luo, R.; Merz, K.M., Jr.; Onufriev, A.; Simmerling, C.; Wang, B.; Woods, R.J. The Amber biomolecular simulation programs. J. Comput. Chem. 2005, 26, 1668–1688. [Google Scholar] [CrossRef] [PubMed]
- Walker, R.C.; Crowley, M.F.; Case, D.A. The implementation of a fast and accurate QM/MM potential method in Amber. J. Comput. Chem. 2008, 29, 1019–1031. [Google Scholar] [CrossRef] [PubMed]
- Krieger, E.; Vriend, G. New ways to boost molecular dynamics simulations. J. Comput. Chem. 2015, 36, 996–1007. [Google Scholar] [CrossRef]
- Salomon-Ferrer, R.; Gotz, A.W.; Poole, D.; Le Grand, S.; Walker, R.C. Routine Microsecond Molecular Dynamics Simulations with AMBER on GPUs. 2. Explicit Solvent Particle Mesh Ewald. J. Chem. Theory Comput. 2013, 9, 3878–3888. [Google Scholar] [CrossRef]
- Maruyama, Y.; Igarashi, R.; Ushiku, Y.; Mitsutake, A. Analysis of Protein Folding Simulation with Moving Root Mean Square Deviation. J. Chem. Inf. Model. 2023, 63, 1529–1541. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera- a visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Schägger, H.; von Jagow, G. Blue native electrophoresis for isolation of membrane protein complexes in enzymatically active form. Anal. Biochem. 1991, 199, 223–231. [Google Scholar] [CrossRef]
- Edgar, R.C. Muscle5: High-accuracy alignment ensembles enable unbiased assessments of sequence homology and phylogeny. Nat. Commun. 2022, 13, 6968. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Suleski, M.; Sanderford, M.; Sharma, S.; Tamura, K. Molecular Evolutionary Genetics Analysis Version 12 for adaptive and green computing. Mol. Biol. Evol. 2024, 41, msae263. [Google Scholar] [CrossRef]
- Madej, T.; Marchler-Bauer, A.; Lanczycki, C.; Zhang, D.; Bryant, S.H. Biological Assembly Comparison With VAST. Methods Mol. Biol. 2020, 2112, 175–186. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]

 ), 0.0166 (
), 0.0166 ( ), 0.024 (
), 0.024 ( ), 0.40 (
), 0.40 ( ), and 0.071 mM (
), and 0.071 mM ( ). The variable fixed concentrations of PEP3- in plot (B) were 0.024 (
). The variable fixed concentrations of PEP3- in plot (B) were 0.024 ( ), 0.048 (
), 0.048 ( ), 0.097 (
), 0.097 ( ), and 0.48 mM (
), and 0.48 mM ( ). In both plots, the Mg2+free concentration was kept constant at 1.41 mM. The other experimental conditions were as shown in Figure 2. The addition of ThaPK initiated the reaction; the enzyme amounts ranged from 0.15 to 0.3 μg/mL. The fitted data are shown in Table 3.
). In both plots, the Mg2+free concentration was kept constant at 1.41 mM. The other experimental conditions were as shown in Figure 2. The addition of ThaPK initiated the reaction; the enzyme amounts ranged from 0.15 to 0.3 μg/mL. The fitted data are shown in Table 3.
 ), 0.0166 (
), 0.0166 ( ), 0.024 (
), 0.024 ( ), 0.40 (
), 0.40 ( ), and 0.071 mM (
), and 0.071 mM ( ). The variable fixed concentrations of PEP3- in plot (B) were 0.024 (
). The variable fixed concentrations of PEP3- in plot (B) were 0.024 ( ), 0.048 (
), 0.048 ( ), 0.097 (
), 0.097 ( ), and 0.48 mM (
), and 0.48 mM ( ). In both plots, the Mg2+free concentration was kept constant at 1.41 mM. The other experimental conditions were as shown in Figure 2. The addition of ThaPK initiated the reaction; the enzyme amounts ranged from 0.15 to 0.3 μg/mL. The fitted data are shown in Table 3.
). In both plots, the Mg2+free concentration was kept constant at 1.41 mM. The other experimental conditions were as shown in Figure 2. The addition of ThaPK initiated the reaction; the enzyme amounts ranged from 0.15 to 0.3 μg/mL. The fitted data are shown in Table 3.
 ), 0.05 (
), 0.05 ( ), 0.1 (
), 0.1 ( ), 0.25 (
), 0.25 ( ), and 0.4 mM (
), and 0.4 mM ( ). The fixed concentrations of free Mg2+ and MgADP were 1.88 and 0.257 mM, respectively. In plot B, the variable concentrations of MgADP were 0.0125, 0.0166, 0.024, 0.0396, and 0.0713 mM. The fixed variable concentrations of oxalate were 0 (
). The fixed concentrations of free Mg2+ and MgADP were 1.88 and 0.257 mM, respectively. In plot B, the variable concentrations of MgADP were 0.0125, 0.0166, 0.024, 0.0396, and 0.0713 mM. The fixed variable concentrations of oxalate were 0 ( ), 20 (
), 20 ( ), 30 (
), 30 ( ), 40 (
), 40 ( ), and 60 (
), and 60 ( ). The fixed concentrations of free Mg2+ and total PEP were 1.88 and 1.127 mM, respectively. The reaction was initiated by the addition of ThaPK; the enzyme amounts of enzyme ranged from 0.15 to 0.3 μg/mL.
). The fixed concentrations of free Mg2+ and total PEP were 1.88 and 1.127 mM, respectively. The reaction was initiated by the addition of ThaPK; the enzyme amounts of enzyme ranged from 0.15 to 0.3 μg/mL.
 ), 0.05 (
), 0.05 ( ), 0.1 (
), 0.1 ( ), 0.25 (
), 0.25 ( ), and 0.4 mM (
), and 0.4 mM ( ). The fixed concentrations of free Mg2+ and MgADP were 1.88 and 0.257 mM, respectively. In plot B, the variable concentrations of MgADP were 0.0125, 0.0166, 0.024, 0.0396, and 0.0713 mM. The fixed variable concentrations of oxalate were 0 (
). The fixed concentrations of free Mg2+ and MgADP were 1.88 and 0.257 mM, respectively. In plot B, the variable concentrations of MgADP were 0.0125, 0.0166, 0.024, 0.0396, and 0.0713 mM. The fixed variable concentrations of oxalate were 0 ( ), 20 (
), 20 ( ), 30 (
), 30 ( ), 40 (
), 40 ( ), and 60 (
), and 60 ( ). The fixed concentrations of free Mg2+ and total PEP were 1.88 and 1.127 mM, respectively. The reaction was initiated by the addition of ThaPK; the enzyme amounts of enzyme ranged from 0.15 to 0.3 μg/mL.
). The fixed concentrations of free Mg2+ and total PEP were 1.88 and 1.127 mM, respectively. The reaction was initiated by the addition of ThaPK; the enzyme amounts of enzyme ranged from 0.15 to 0.3 μg/mL.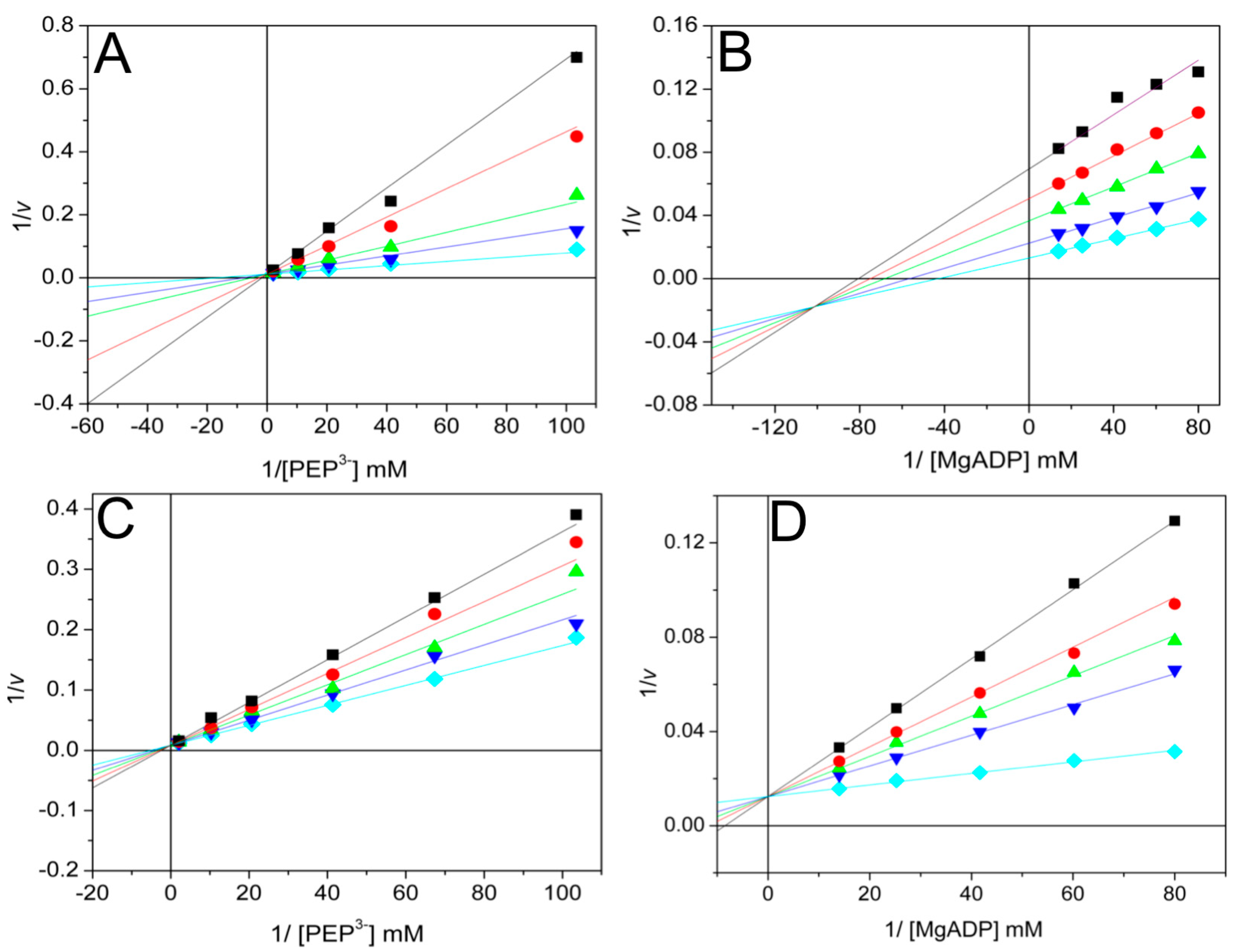

 ) and (C) of ECTS truncated-ThaPK (
) and (C) of ECTS truncated-ThaPK ( ). In (A), the spectra were obtained in mixtures containing 100 μg/mL of the WT-ThaPK or truncated-ThaPK in 20 mM NaH2PO4-NaOH, pH 7.0, at 25 °C in a 0.1 cm cell pathway. CD is expressed as θMRE (molar residue ellipticity). In (B,C), the protein concentrations were 1.0 mg/mL, 16.3 μM monomer of the wild type, and 19.41 μM monomer of the ECTS-truncated enzyme. The enzymes were diluted in 50 mM MOPS-(CH3)4NOH, pH 6.5, containing 0.3 mM oxalate, 0.1 mM ATP, and 1.05 mM Mg2+. The scan rate was 1.5 °C/min. The Tms were 79.2 and 78.4 °C for WT-ThaPK and truncated-ThaPK, respectively. ∆H (cal/mol °C) were 230,946.6 and 46,845.8 for the WT-ThaPK and truncated-ThaPK, respectively.
). In (A), the spectra were obtained in mixtures containing 100 μg/mL of the WT-ThaPK or truncated-ThaPK in 20 mM NaH2PO4-NaOH, pH 7.0, at 25 °C in a 0.1 cm cell pathway. CD is expressed as θMRE (molar residue ellipticity). In (B,C), the protein concentrations were 1.0 mg/mL, 16.3 μM monomer of the wild type, and 19.41 μM monomer of the ECTS-truncated enzyme. The enzymes were diluted in 50 mM MOPS-(CH3)4NOH, pH 6.5, containing 0.3 mM oxalate, 0.1 mM ATP, and 1.05 mM Mg2+. The scan rate was 1.5 °C/min. The Tms were 79.2 and 78.4 °C for WT-ThaPK and truncated-ThaPK, respectively. ∆H (cal/mol °C) were 230,946.6 and 46,845.8 for the WT-ThaPK and truncated-ThaPK, respectively.
 ) and (C) of ECTS truncated-ThaPK (
) and (C) of ECTS truncated-ThaPK ( ). In (A), the spectra were obtained in mixtures containing 100 μg/mL of the WT-ThaPK or truncated-ThaPK in 20 mM NaH2PO4-NaOH, pH 7.0, at 25 °C in a 0.1 cm cell pathway. CD is expressed as θMRE (molar residue ellipticity). In (B,C), the protein concentrations were 1.0 mg/mL, 16.3 μM monomer of the wild type, and 19.41 μM monomer of the ECTS-truncated enzyme. The enzymes were diluted in 50 mM MOPS-(CH3)4NOH, pH 6.5, containing 0.3 mM oxalate, 0.1 mM ATP, and 1.05 mM Mg2+. The scan rate was 1.5 °C/min. The Tms were 79.2 and 78.4 °C for WT-ThaPK and truncated-ThaPK, respectively. ∆H (cal/mol °C) were 230,946.6 and 46,845.8 for the WT-ThaPK and truncated-ThaPK, respectively.
). In (A), the spectra were obtained in mixtures containing 100 μg/mL of the WT-ThaPK or truncated-ThaPK in 20 mM NaH2PO4-NaOH, pH 7.0, at 25 °C in a 0.1 cm cell pathway. CD is expressed as θMRE (molar residue ellipticity). In (B,C), the protein concentrations were 1.0 mg/mL, 16.3 μM monomer of the wild type, and 19.41 μM monomer of the ECTS-truncated enzyme. The enzymes were diluted in 50 mM MOPS-(CH3)4NOH, pH 6.5, containing 0.3 mM oxalate, 0.1 mM ATP, and 1.05 mM Mg2+. The scan rate was 1.5 °C/min. The Tms were 79.2 and 78.4 °C for WT-ThaPK and truncated-ThaPK, respectively. ∆H (cal/mol °C) were 230,946.6 and 46,845.8 for the WT-ThaPK and truncated-ThaPK, respectively.


| Mg2+ Complexes | |||||
|---|---|---|---|---|---|
| Substrate | Kapp (mM) | n | Vmax app (µmol/min▪mg) | kcat (s−1) | Log kcat/K (s−1/M) |
| PEP3 | 0.066 ± 0.003 | - | 77 ± 1 | 315 ± 3 | 6.68 |
| Mg-ADP | 0.024 ± 0.001 | - | 75 ± 1 | 307 ± 5 | 7.10 |
| Mg2+free | 0.188 ± 0.018 | 1.8 ± 0.1 | 73 ± 1 | 299 ± 6 | 6.20 |
| Mn2+ Complexes | |||||
|---|---|---|---|---|---|
| Substrate | Kapp (mM) | n | Vmax app (µmol/min▪mg) | kcat (s−1) | Log kcat/K (s−1/M) |
| PEP3− | 0.018 ± 0.001 | 1.7 ± 0.1 | 33 ± 1 | 134 ± 1 | 6.87 |
| Mn-ADP | 0.011 ± 0.001 | -- | 33 ± 1 | 137 ± 3 | 7.14 |
| Mn2+free | 0.082 ± 0.002 | 1.3 ± 0.1 | 25 ± 1 | 96 ± 1 | 6.08 |
| Substrate | Initial Velocity Intersecting Patterns 1/v Versus 1/[S] | Kinetic Mechanism | Vmax (µmol/min▪mg) | Km (mM) | kcat s−1 | Log kcat /Km s−1M−1 |
|---|---|---|---|---|---|---|
| PEP3− | Intersects to the left of the 1/v axis and on the 1/S axis | Random rapid equilibrium | 106 ± 3 | 0.031 ± 0.002 | 418 | 7.14 |
| ADP-Mg | 0.023 ± 0.002 | 7.25 |
| Dead End Analog of PEP: Oxalate | Dead End Analog of Mg-ADP: AMP | Ki (Oxalate) µM | Ki (AMP) mM | ||
|---|---|---|---|---|---|
| 1/v vs. 1/PEP, fixed Mg-ADP | 1/v vs. 1/MgADP, fixed PEP | 1/v vs. 1/PEP, fixed Mg-ADP | 1/v vs. 1/Mg-ADP, fixed PEP | ||
| C | MT | C | C | 44 ± 1 | 3.3 ± 0.3 |
| ECTS truncated-ThaPK | |||||
|---|---|---|---|---|---|
| Substrate | K0.5 app (mM) | n | Vmax app (µmol/min▪mg) | kcat (s−1) | Log kcat/K (s−1/M) |
| PEP3− | 3.15 ± 1 | 1.7 ± 0.4 | 8.8 ± 1.9 | 29± 6 | 3.96 |
| Mg-ADP | 0.17 ± 0.01 | 2.1 ± 0.2 | 6.5 ± 0.2 | 21 ± 1 | 5.09 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramírez-Silva, L.; Riveros-Rosas, H.; Hernández-Alcántara, G.; García-Trejo, J.J.; Vega-Segura, A.; González-Andrade, M.; Díaz-Salazar, A.J.; Salcedo-Barrientos, G. The Functional State of Thermoplasma acidophilum Pyruvate Kinase Relies on an Extra Carboxyl-Terminal Sequence. Int. J. Mol. Sci. 2025, 26, 8410. https://doi.org/10.3390/ijms26178410
Ramírez-Silva L, Riveros-Rosas H, Hernández-Alcántara G, García-Trejo JJ, Vega-Segura A, González-Andrade M, Díaz-Salazar AJ, Salcedo-Barrientos G. The Functional State of Thermoplasma acidophilum Pyruvate Kinase Relies on an Extra Carboxyl-Terminal Sequence. International Journal of Molecular Sciences. 2025; 26(17):8410. https://doi.org/10.3390/ijms26178410
Chicago/Turabian StyleRamírez-Silva, Leticia, Héctor Riveros-Rosas, Gloria Hernández-Alcántara, José J. García-Trejo, Alicia Vega-Segura, Martin González-Andrade, A. Jessica Díaz-Salazar, and Guillermo Salcedo-Barrientos. 2025. "The Functional State of Thermoplasma acidophilum Pyruvate Kinase Relies on an Extra Carboxyl-Terminal Sequence" International Journal of Molecular Sciences 26, no. 17: 8410. https://doi.org/10.3390/ijms26178410
APA StyleRamírez-Silva, L., Riveros-Rosas, H., Hernández-Alcántara, G., García-Trejo, J. J., Vega-Segura, A., González-Andrade, M., Díaz-Salazar, A. J., & Salcedo-Barrientos, G. (2025). The Functional State of Thermoplasma acidophilum Pyruvate Kinase Relies on an Extra Carboxyl-Terminal Sequence. International Journal of Molecular Sciences, 26(17), 8410. https://doi.org/10.3390/ijms26178410








