IL-24 in COVID-19 Patients: Correlations with Disease Progression
Abstract
1. Introduction
2. Results
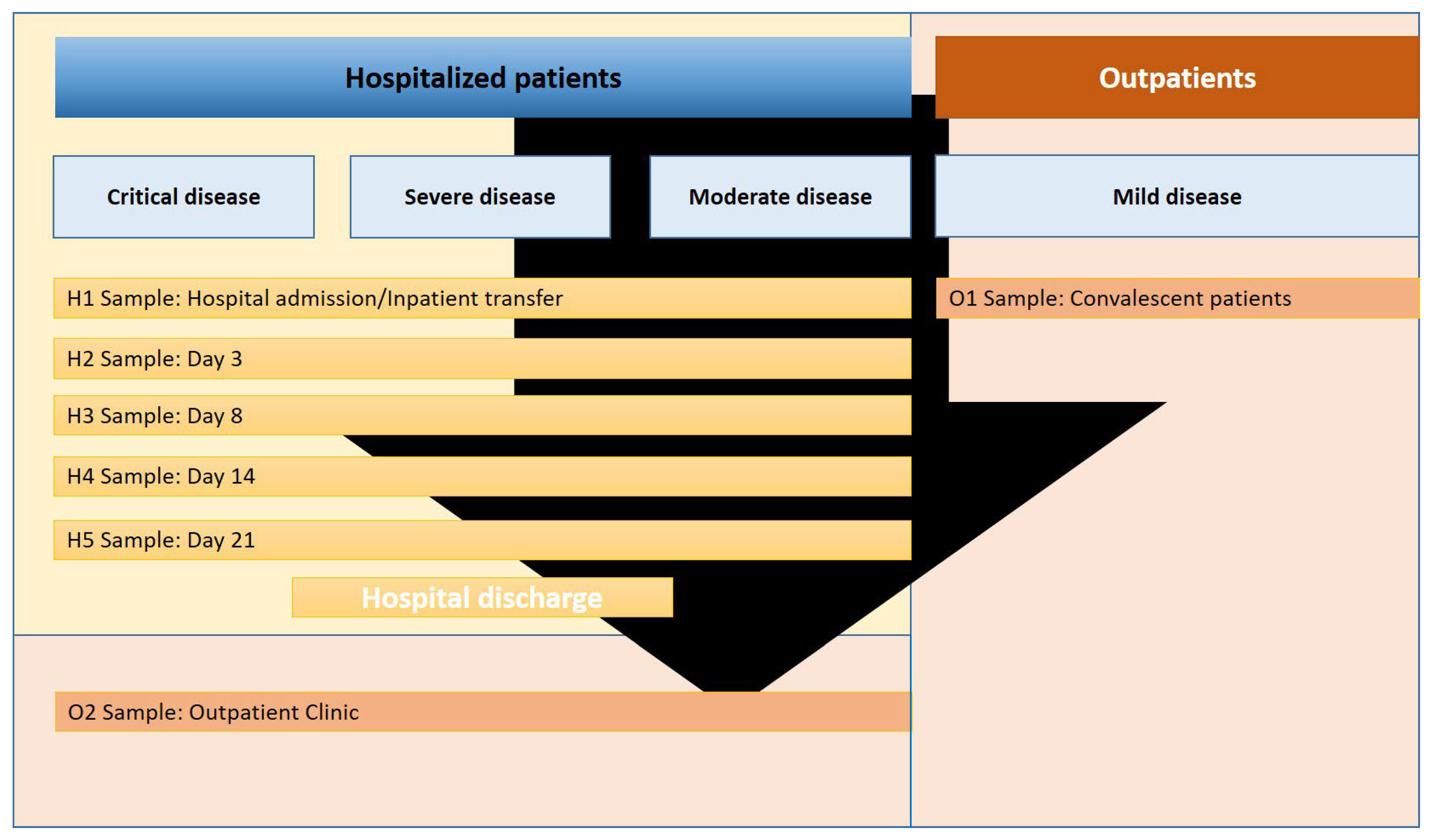
2.1. Patients Characteristics
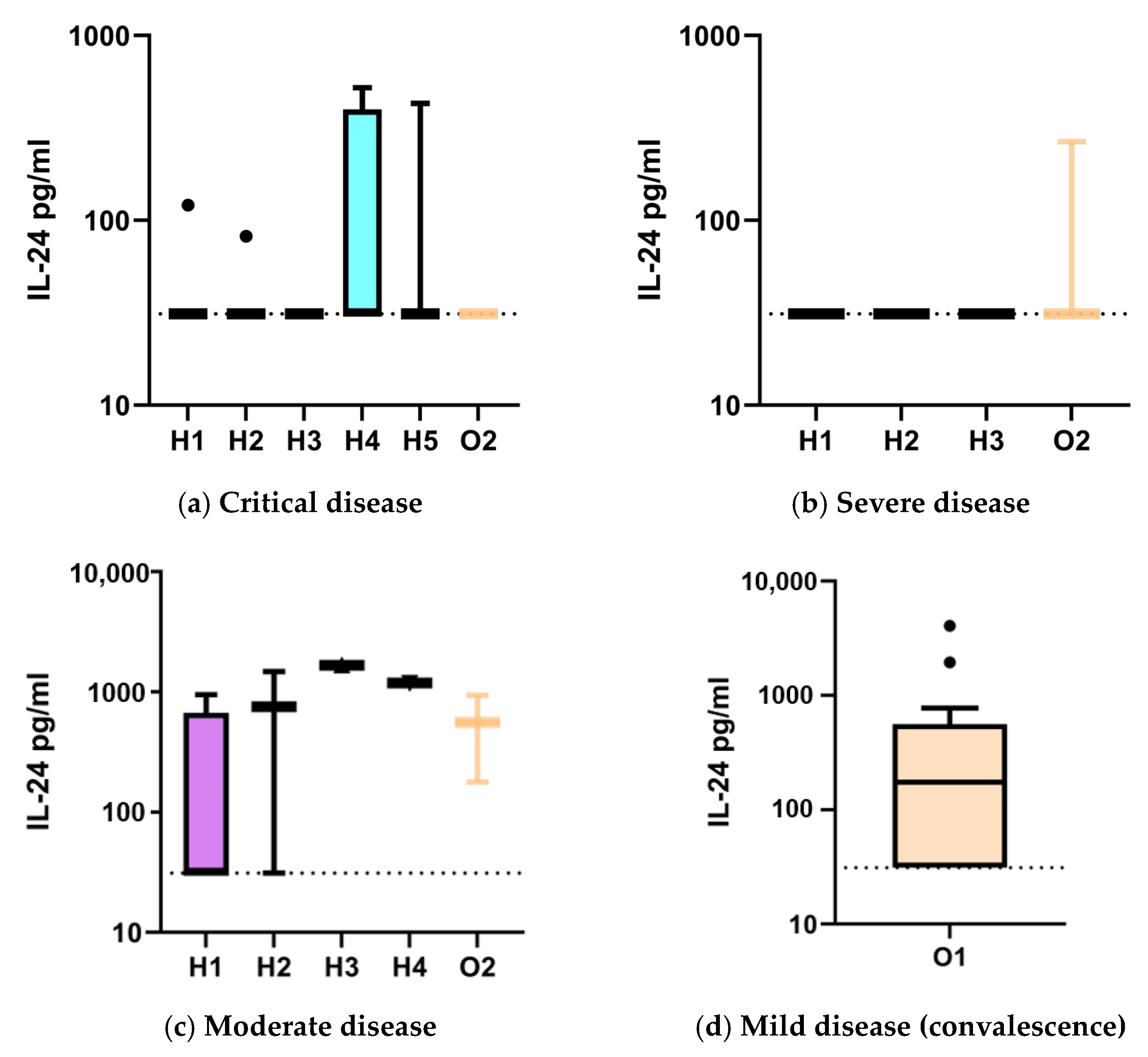
2.2. IL-24 Serum Levels in COVID-19 Patients by Disease Severity and Sampling Time: Higher Proportion Above Lower Limit of Quantification in Mild and Moderate Cases
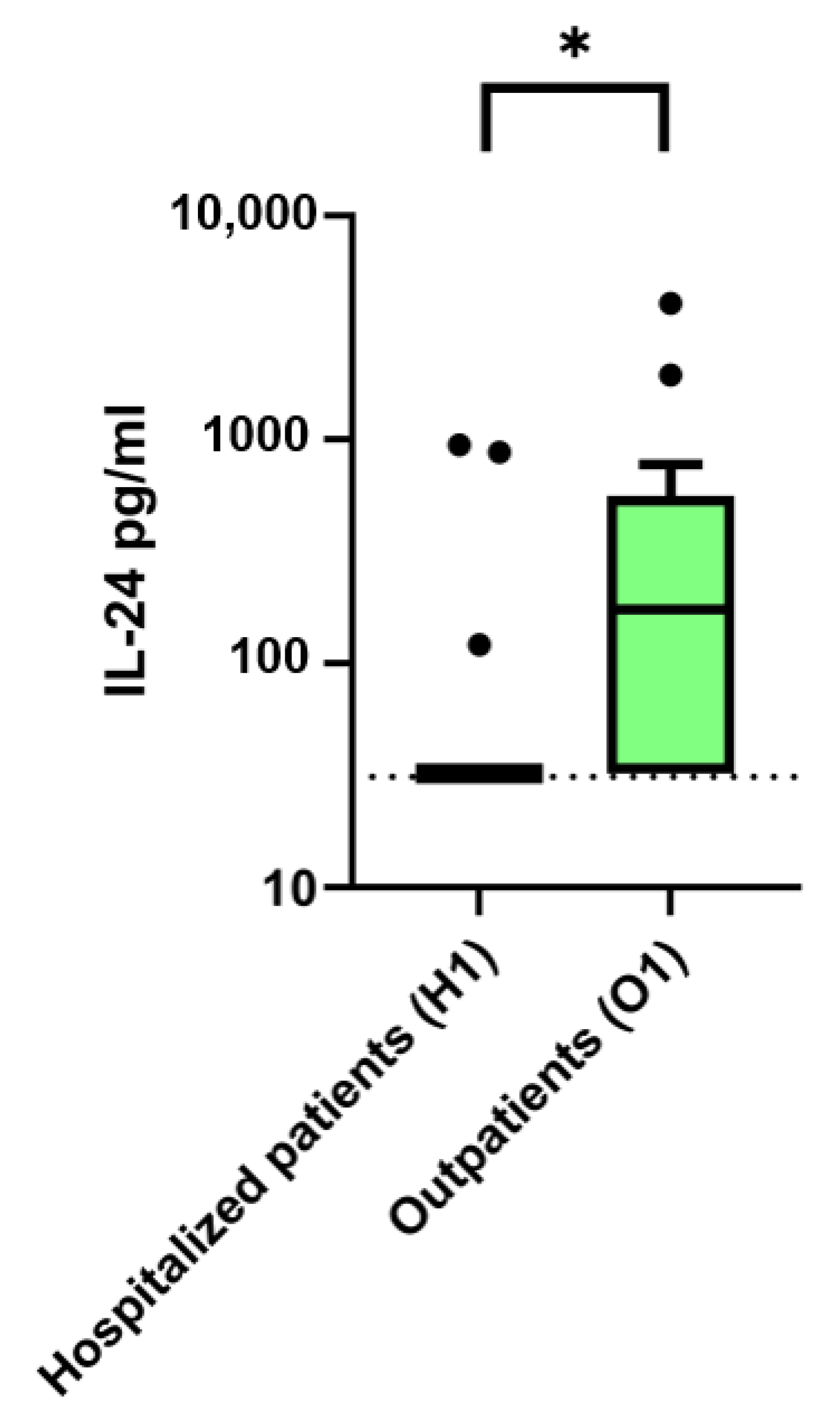
2.3. Comparison of IL-24 Serum Levels and Symptom-to-Sampling Times in Hospitalized vs. Convalescent Outpatient COVID-19 Patients
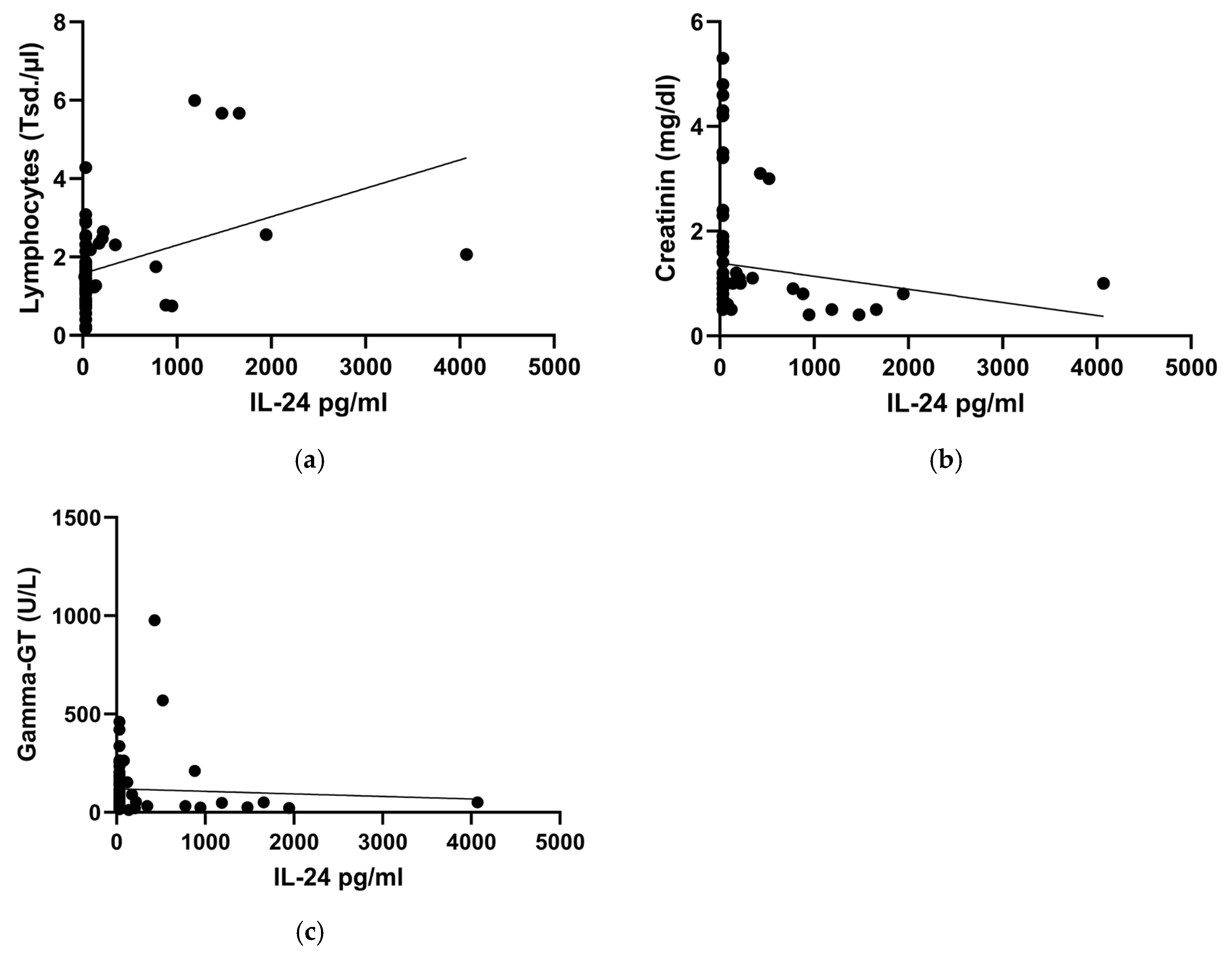
2.4. Reduced IL-24 Serum Levels in COVID-19 Patients Are Associated with Indicators of Liver-/Kidney Damage and Lymphocytopenia
3. Discussion
4. Materials and Methods
4.1. Study Subjects and Samples
4.2. Blood Collection and Sample Processing
4.3. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vollenberg, R.; Matern, P.; Nowacki, T.; Fuhrmann, V.; Padberg, J.-S.; Ochs, K.; Schütte-Nütgen, K.; Strauß, M.; Schmidt, H.; Tepasse, P.-R. Prone Position in Mechanically Ventilated COVID-19 Patients: A Multicenter Study. J. Clin. Med. 2021, 10, 1046. [Google Scholar] [CrossRef]
- Ranieri, V.M.; Rubenfeld, G.D.; Thompson, B.T.; Ferguson, N.D.; Caldwell, E.; Fan, E.; Camporota, L.; Slutsky, A.S. Acute respiratory distress syndrome: The Berlin Definition. JAMA 2012, 307, 2526–2533. [Google Scholar] [CrossRef]
- Batah, S.S.; Fabro, A.T. Pulmonary pathology of ARDS in COVID-19: A pathological review for clinicians. Respir. Med. 2021, 176, 106239. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.-J.; Dong, X.; Liu, G.-H.; Gao, Y.-D. Risk and Protective Factors for COVID-19 Morbidity, Severity, and Mortality. Clin. Rev. Allergy Immunol. 2023, 64, 90–107. [Google Scholar] [CrossRef] [PubMed]
- Kessel, C.; Vollenberg, R.; Masjosthusmann, K.; Hinze, C.; Wittkowski, H.; Debaugnies, F.; Nagant, C.; Corazza, F.; Vély, F.; Kaplanski, G.; et al. Discrimination of COVID-19 From Inflammation-Induced Cytokine Storm Syndromes Using Disease-Related Blood Biomarkers. Arthritis Rheumatol. 2021, 73, 1791–1799. [Google Scholar] [CrossRef] [PubMed]
- Evers, G.; Schulze, A.B.; Osiaevi, I.; Harmening, K.; Vollenberg, R.; Wiewrodt, R.; Pistulli, R.; Boentert, M.; Tepasse, P.-R.; Sindermann, J.R.; et al. Sustained Impairment in Cardiopulmonary Exercise Capacity Testing in Patients after COVID-19: A Single Center Experience. Can. Respir. J. 2022, 2022, 2466789. [Google Scholar] [CrossRef]
- Laniece Delaunay, C.; Verdasca, N.; Monge, S.; Domegan, L.; Sève, N.; Buda, S.; Meijer, A.; Lucaccioni, H.; López Torrijos, M.; McKenna, A.; et al. COVID-19 Vaccine Effectiveness Against Medically Attended Symptomatic SARS-CoV-2 Infection Among Target Groups in Europe, October 2024-January 2025, VEBIS Primary Care Network. Influenza Other Respir. Viruses 2025, 19, e70120. [Google Scholar] [CrossRef]
- Liu, C.; Tsang, T.K.; Sullivan, S.G.; Cowling, B.J.; Yang, B. Comparative duration of neutralizing responses and protections of COVID-19 vaccination and correlates of protection. Nat. Commun. 2025, 16, 4748. [Google Scholar] [CrossRef]
- Volkman, H.R.; Nguyen, J.L.; Mustapha, M.M.; Yang, J.; Jodar, L.; McLaughlin, J.M. Effectiveness of a single COVID-19 mRNA vaccine dose in individuals with prior SARS-CoV-2 infection: A systematic review. Commun. Med. 2025, 5, 151. [Google Scholar] [CrossRef]
- Vollenberg, R.; Tepasse, P.-R.; Lorentzen, E.; Nowacki, T.M. Impaired Humoral Immunity with Concomitant Preserved T Cell Reactivity in IBD Patients on Treatment with Infliximab 6 Month after Vaccination with the SARS-CoV-2 mRNA Vaccine BNT162b2: A Pilot Study. J. Pers. Med. 2022, 12, 694. [Google Scholar] [CrossRef]
- Bausch-Jurken, M.; Alter, G. The immunological impact of revaccination in a hybrid-immune world. Front. Immunol. 2025, 16, 1588259. [Google Scholar] [CrossRef]
- Link-Gelles, R.; Chickery, S.; Webber, A.; Ong, T.C.; Rowley, E.A.K.; DeSilva, M.B.; Dascomb, K.; Irving, S.A.; Klein, N.P.; Grannis, S.J.; et al. Interim Estimates of 2024–2025 COVID-19 Vaccine Effectiveness Among Adults Aged≥ 18 Years—VISION and IVY Networks, September 2024–January 2025. MMWR Morb. Mortal. Wkly. Rep. 2025, 74, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Bertini, C.D.; Khawaja, F.; Sheshadri, A. Coronavirus Disease-2019 in the Immunocompromised Host. Clin. Chest Med. 2023, 44, 395–406. [Google Scholar] [CrossRef] [PubMed]
- Commins, S.; Steinke, J.W.; Borish, L. The extended IL-10 superfamily: IL-10, IL-19, IL-20, IL-22, IL-24, IL-26, IL-28, and IL-29. J. Allergy Clin. Immunol. 2008, 121, 1108–1111. [Google Scholar] [CrossRef] [PubMed]
- Huang, E.Y.; Madireddi, M.T.; Gopalkrishnan, R.V.; Leszczyniecka, M.; Su, Z.; Lebedeva, I.V.; Kang, D.; Jiang, H.; Lin, J.J.; Alexandre, D.; et al. Genomic structure, chromosomal localization and expression profile of a novel melanoma differentiation associated (mda-7) gene with cancer specific growth suppressing and apoptosis inducing properties. Oncogene 2001, 20, 7051–7063. [Google Scholar] [CrossRef]
- Park, S.-J.; Kumar, M.; Kwon, H.; Seong, R.-K.; Han, K.; Song, J.; Kim, C.-J.; Choi, Y.-K.; Shin, O.S. Dynamic changes in host gene expression associated with H5N8 avian influenza virus infection in mice. Sci. Rep. 2015, 5, 16512. [Google Scholar] [CrossRef]
- Weiss, R.; Laengle, J.; Sachet, M.; Shurygina, A.-P.; Kiselev, O.; Egorov, A.; Bergmann, M. Interleukin-24 inhibits influenza A virus replication in vitro through induction of toll-like receptor 3 dependent apoptosis. Antivir. Res. 2015, 123, 93–104. [Google Scholar] [CrossRef]
- Seong, R.-K.; Choi, Y.-K.; Shin, O.S. MDA7/IL-24 is an anti-viral factor that inhibits influenza virus replication. J. Microbiol. 2016, 54, 695–700. [Google Scholar] [CrossRef]
- Iwasaki, A.; Yang, Y. The potential danger of suboptimal antibody responses in COVID-19. Nat. Rev. Immunol. 2020, 20, 339–341. [Google Scholar] [CrossRef]
- Menezes, M.E.; Bhoopathi, P.; Pradhan, A.K.; Emdad, L.; Das, S.K.; Guo, C.; Wang, X.-Y.; Sarkar, D.; Fisher, P.B. Role of MDA-7/IL-24 a Multifunction Protein in Human Diseases. Adv. Cancer Res. 2018, 138, 143–182. [Google Scholar] [CrossRef]
- Persaud, L.; De Jesus, D.; Brannigan, O.; Richiez-Paredes, M.; Huaman, J.; Alvarado, G.; Riker, L.; Mendez, G.; Dejoie, J.; Sauane, M. Mechanism of Action and Applications of Interleukin 24 in Immunotherapy. Int. J. Mol. Sci. 2016, 17, 869. [Google Scholar] [CrossRef]
- Liang, J.; Huang, R.-L.; Huang, Q.; Peng, Z.; Zhang, P.-H.; Wu, Z.-X. Adenovirus-mediated human interleukin 24 (MDA-7/IL-24) selectively suppresses proliferation and induces apoptosis in keloid fibroblasts. Ann. Plast. Surg. 2011, 66, 660–666. [Google Scholar] [CrossRef]
- Nace, J.; Fortunato, S.J.; Maul, H.; Menon, R. The expression pattern of two novel cytokines (IL-24 and IL-29) in human fetal membranes. J. Perinat. Med. 2010, 38, 665–670. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Zuo, X.; Zhang, S.; Ouyang, Z.; Jiang, S.; Wang, F.; Wang, G. The Mechanism behind Influenza Virus Cytokine Storm. Viruses 2021, 13, 1362. [Google Scholar] [CrossRef] [PubMed]
- Suhre, K.; Sarwath, H.; Engelke, R.; Sohail, M.U.; Cho, S.J.; Whalen, W.; Alvarez-Mulett, S.; Krumsiek, J.; Choi, A.M.K.; Schmidt, F. Identification of Robust Protein Associations With COVID-19 Disease Based on Five Clinical Studies. Front. Immunol. 2021, 12, 781100. [Google Scholar] [CrossRef] [PubMed]
- André, S.; Picard, M.; Cezar, R.; Roux-Dalvai, F.; Alleaume-Butaux, A.; Soundaramourty, C.; Cruz, A.S.; Mendes-Frias, A.; Gotti, C.; Leclercq, M.; et al. T cell apoptosis characterizes severe COVID-19 disease. Cell Death Differ. 2022, 29, 1486–1499. [Google Scholar] [CrossRef]
- Cezar, R.; Kundura, L.; André, S.; Lozano, C.; Vincent, T.; Muller, L.; Lefrant, J.-Y.; Roger, C.; Claret, P.-G.; Duvnjak, S.; et al. T4 apoptosis in the acute phase of SARS-CoV-2 infection predicts long COVID. Front. Immunol. 2023, 14, 1335352. [Google Scholar] [CrossRef]
- Paolini, A.; Borella, R.; De Biasi, S.; Neroni, A.; Mattioli, M.; Lo Tartaro, D.; Simonini, C.; Franceschini, L.; Cicco, G.; Piparo, A.M.; et al. Cell Death in Coronavirus Infections: Uncovering Its Role during COVID-19. Cells 2021, 10, 1585. [Google Scholar] [CrossRef]
- Yuan, C.; Ma, Z.; Xie, J.; Li, W.; Su, L.; Zhang, G.; Xu, J.; Wu, Y.; Zhang, M.; Liu, W. The role of cell death in SARS-CoV-2 infection. Signal Transduct. Target. Ther. 2023, 8, 357. [Google Scholar] [CrossRef]
- Liang, K.; Barnett, K.C.; Hsu, M.; Chou, W.-C.; Bais, S.S.; Riebe, K.; Xie, Y.; Nguyen, T.T.; Oguin, T.H.; Vannella, K.M.; et al. Initiator cell death event induced by SARS-CoV-2 in the human airway epithelium. Sci. Immunol. 2024, 9, eadn0178. [Google Scholar] [CrossRef]
- Giamarellos-Bourboulis, E.J.; Netea, M.G.; Rovina, N.; Akinosoglou, K.; Antoniadou, A.; Antonakos, N.; Damoraki, G.; Gkavogianni, T.; Adami, M.-E.; Katsaounou, P.; et al. Complex Immune Dysregulation in COVID-19 Patients with Severe Respiratory Failure. Cell Host Microbe 2020, 27, 992–1000.e3. [Google Scholar] [CrossRef]
- Zhong, Y.; Zhang, X.; Chong, W. Interleukin-24 Immunobiology and Its Roles in Inflammatory Diseases. Int. J. Mol. Sci. 2022, 23, 627. [Google Scholar] [CrossRef]
- Wu, B.; Huang, C.; Kato-Maeda, M.; Hopewell, P.C.; Daley, C.L.; Krensky, A.M.; Clayberger, C. IL-24 modulates IFN-gamma expression in patients with tuberculosis. Immunol. Lett. 2008, 117, 57–62. [Google Scholar] [CrossRef]
- Wang, C.-W.; Chen, C.-B.; Lu, C.-W.; Chen, W.-T.; Hui, R.C.-Y.; Chiu, T.-M.; Chi, M.-H.; Lin, J.-C.; Huang, Y.-H.; Chang, Y.-C.; et al. Characteristics of immune response profile in patients with immediate allergic and autoimmune urticarial reactions induced by SARS-CoV-2 vaccines. J. Autoimmun. 2023, 138, 103054. [Google Scholar] [CrossRef]
- Ouyang, W.; Rutz, S.; Crellin, N.K.; Valdez, P.A.; Hymowitz, S.G. Regulation and functions of the IL-10 family of cytokines in inflammation and disease. Annu. Rev. Immunol. 2011, 29, 71–109. [Google Scholar] [CrossRef] [PubMed]
- Bortolotti, D.; Gentili, V.; Rizzo, S.; Schiuma, G.; Beltrami, S.; Strazzabosco, G.; Fernandez, M.; Caccuri, F.; Caruso, A.; Rizzo, R. TLR3 and TLR7 RNA Sensor Activation during SARS-CoV-2 Infection. Microorganisms 2021, 9, 1820. [Google Scholar] [CrossRef] [PubMed]
- Takemura, H.; Ai, T.; Kimura, K.; Nagasaka, K.; Takahashi, T.; Tsuchiya, K.; Yang, H.; Konishi, A.; Uchihashi, K.; Horii, T.; et al. Evaluation of cell count and classification capabilities in body fluids using a fully automated Sysmex XN equipped with high-sensitive Analysis (hsA) mode and DI-60 hematology analyzer system. PLoS ONE 2018, 13, e0195923. [Google Scholar] [CrossRef] [PubMed]
- von Eckardstein, A.; Roth, H.J.; Jones, G.; Preston, S.; Szekeres, T.; Imdahl, R.; Conti, M.; Blanckaert, N.; Jose, D.; Thiery, J.; et al. Cobas 8000 Modular analyzer series evaluated under routine-like conditions at 14 sites in Australia, Europe, and the United States. J. Lab. Autom. 2013, 18, 306–327. [Google Scholar] [CrossRef]
- Rao, L.-Z.; Wang, Y.; Zhang, L.; Wu, G.; Zhang, L.; Wang, F.-X.; Chen, L.-M.; Sun, F.; Jia, S.; Zhang, S.; et al. IL-24 deficiency protects mice against bleomycin-induced pulmonary fibrosis by repressing IL-4-induced M2 program in macrophages. Cell Death Differ. 2021, 28, 1270–1283, Correction in Cell Death Differ. 2021, 28, 2989. [Google Scholar] [CrossRef]
| Patients n = 41 (Samples n = 88) | Hospitalized | Outpatients | p-Value | |||
|---|---|---|---|---|---|---|
| Critical Disease | Severe Disease | Moderate Disease | Mild Disease | |||
| Oxygen therapy | No oxygen therapy | |||||
| Patient characteristics | Patients, No. | 11 | 9 | 8 | 13 | |
| Samples, No. | 37 | 24 | 14 | 13 | ||
| Sex, male, No. (%) | 10 (91) | 8 (89) | 6 (75) | 13 (100) | 0.19 | |
| Age, median (min–max) | 59 (41–72) | 53 (44–76) | 50 (30–66) | 47 (31–70) | 0.314 | |
| BMI, median (IQR) | 26 (23–29) | 24 (23–26) | 25 (22–27) | 29 (25–31) | 0.057 | |
| Time symptom onset to sample (1. Sample) | 12 (11–17) | 11 (3–17) | 8 (2–8) | 38 (36–46) | <0.01 | |
| Pre-existing conditions | Diabetes (%) | 1 (9) | 0 (0) | 1 (13) | 0 (0) | |
| Chronic inflammatory bowel disease (%) | 1 (9) | 1 (11) | 0 (0) | 0 (0) | ||
| Coronary heart disease (%) | 2 (18) | 1 (11) | 1 (13) | 0 (0) | ||
| Respiratory disease (%) | 1 (9) | 0 (0) | 1 (13) | 0 (0) | ||
| Neoplasm (%) | 1 (9) | 0 (0) | 0 (0) | 0 (0) | ||
| Arterial hypertension (%) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | ||
| Patients n = 41 (1. Sample) | Critical Disease | Severe Disease | Moderate Disease | Mild Disease | p-Value |
|---|---|---|---|---|---|
| Leukocytes (109/L), median (IQR) | 8.7 (5.3–12.7) | 5.4 (3.4–7.6) | 4.1 (3.5–5.2) | 5.6 (5.1–7.4) | 0.014 |
| Lymphocytes (rel., %), median (IQR) | 11.3 (7.2–14.5) | 21.6 (18.5–31.5) | 25.8 (19.3–32.4) | 32.5 (28.1–38) | 0.001 |
| Lymphocytes (abs. 109/L), median (IQR) | 1.5 (0.7–1.7) | 1.2 (0.9–1.6) | 0.96 (0.76–1.3) | 2.3 (1.4–2.5) | 0.013 |
| NLR | 6.7 (5.3–11.4) | 2.8 (1.5–2.9) | 2.2 (1.8–3.8) | 1.7 (1.4–2.2) | <0.001 |
| Bilirubin (mg/dL), median (IQR) | 0.5 (0.3–1.3) | 0.5 (0.5–0.6) | 0.5 (0.3–0.8) | 0.4 (0.4–0.5) | 0.471 |
| AST (U/L), median (IQR) | 81 (59–134) | 41 (32–62) | 29 (25–45) | 32 (25–42) | 0.001 |
| ALT (U/L), median (IQR) | 66 (27–114) | 34 (29–60) | 22 (18–47) | 29 (26–50) | 0.204 |
| Gamma-GT (U/L), median (IQR) | 97 (55–146) | 54 (30–60) | 41 (33–124) | 32 (21–52) | 0.014 |
| CRP (mg/dL), median (IQR) | 14 (6–30) | 5.2 (2.4–9.4) | 2 (0.8–3.3) | 0.5 (0.5–0.5) | <0.001 |
| Albumin (g/dL), median (IQR) | 2 (1–2.6) | 3.5 (3.1–4.3) | 4.0 (3.7–4.2) | 4.6 (4.5–4.8) | <0.001 |
| Ferritin (µg/L), median (IQR) | 999 (646–1937) | 779 (362–938) | 292 (191–557) | 248 (96–310) | <0.001 |
| Patients n = 41 (88 Samples) | Critical Disease | Severe Disease | Moderate Disease | Mild Disease |
|---|---|---|---|---|
| Hospitalized patients | ||||
| H1 Sample | 11 | 9 | 8 | |
| IL-24 * (pg/mL), median (IQR) | 0 (0–0) | 0 (0–0) | 0 (0–661) | |
| IL-24 # (pg/mL), median (IQR) | 31.3 (31.3–31.3) | 31.3 (31.3–31.3) | 31.3 (31.3–661) | |
| IL-24 values above LLOQ, n (%) | 1 (9) | 0 (0) | 2 (25) | |
| Time symptom onset to sample (days) | 12 (11–17) | 11 (3–17) | 8 (2–8) | |
| H2 Sample | 9 | 7 | 2 | |
| IL-24 * (pg/mL), median (IQR) | 0 (0–0) | 0 (0–0) | 754.1 (0) | |
| IL-24 # (pg/mL), median (IQR) | 31.3 (31.3–31.3) | 31.3 (31.3–31.3) | 754.1 (31.3) | |
| IL-24 values above LLOQ, n (%) | 1 (11) | 0 (0) | 1 (50) | |
| Time symptom onset to sample (days) | 17 (14–21) | 14 (4–22) | 4 (3) | |
| H3 Sample | 7 | 5 | 1 | |
| IL-24 * (pg/mL), median (IQR) | 0 (0–0) | 0 (0–0) | 1660 | |
| IL-24 # (pg/mL), median (IQR) | 31.3 (31.3–31.3) | 31.3 (31.3–31.3) | 1660 | |
| IL-24 values above LLOQ, n (%) | 0 (0) | 0 (0) | 1 (100) | |
| Time symptom onset to sample (days) | 22 (17–26) | 13 (11–26) | 8 | |
| H4 Sample | 4 | 1 | ||
| IL-24 * (pg/mL), median (IQR) | 0 (0–398.6) | 1188 | ||
| IL-24 # (pg/mL), median (IQR) | 31.3 (31.3–398.6)) | 1188 | ||
| IL-24 values above LLOQ, n (%) | 1 (25) | 1 (100) | ||
| Time symptom onset to sample (days) | 31 (27–37) | 10 | ||
| H5 Sample | 3 | |||
| IL-24 * (pg/mL), median (IQR) | 0 (0–0) | |||
| IL-24 # (pg/mL), median (IQR) | 31.3 (31.3–31.3) | |||
| IL-24 values above LLOQ, n (%) | 1 (33) | |||
| Time symptom onset to sample (days) | 46 (27) | |||
| Outpatients | ||||
| O1 Sample | 13 | |||
| IL-24 * (pg/mL), median (IQR) | 174 (0–561) | |||
| IL-24 # (pg/mL), median (IQR) | 174 (31.3–561) | |||
| IL-24 values above LLOQ, n (%) | 8 (62) | |||
| Time symptom onset to sample (days) | 38 (36–46) | |||
| O2 Sample | 3 | 3 | 2 | |
| IL-24 * (pg/mL), median (IQR) | 0 (0–0) | 0 (0) | 558 (178) | |
| IL-24 # (pg/mL), median (IQR) | 31.3 (31.3–31.) | 31.3 (31.3) | 558 (178) | |
| IL-24 values above LLOQ, n (%) | 0 (0) | 1 | 2 | |
| Time symptom onset to sample (days) | 140 (126) | 126 (122) | 137 (129) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vollenberg, R.; Schütte-Nütgen, K.; Strauss, M.; Trebicka, J.; Fischer, J.; Tepasse, P.-R. IL-24 in COVID-19 Patients: Correlations with Disease Progression. Int. J. Mol. Sci. 2025, 26, 8403. https://doi.org/10.3390/ijms26178403
Vollenberg R, Schütte-Nütgen K, Strauss M, Trebicka J, Fischer J, Tepasse P-R. IL-24 in COVID-19 Patients: Correlations with Disease Progression. International Journal of Molecular Sciences. 2025; 26(17):8403. https://doi.org/10.3390/ijms26178403
Chicago/Turabian StyleVollenberg, Richard, Katharina Schütte-Nütgen, Markus Strauss, Jonel Trebicka, Julia Fischer, and Phil-Robin Tepasse. 2025. "IL-24 in COVID-19 Patients: Correlations with Disease Progression" International Journal of Molecular Sciences 26, no. 17: 8403. https://doi.org/10.3390/ijms26178403
APA StyleVollenberg, R., Schütte-Nütgen, K., Strauss, M., Trebicka, J., Fischer, J., & Tepasse, P.-R. (2025). IL-24 in COVID-19 Patients: Correlations with Disease Progression. International Journal of Molecular Sciences, 26(17), 8403. https://doi.org/10.3390/ijms26178403


_Kim.png)





