Double Pathogenic or Likely Pathogenic Variants in Cancer Predisposition Genes in Hungarian Cancer Patients
Abstract
1. Introduction
2. Results
2.1. Cohort Characteristics and Detection Rate
2.2. Double Heterozygosity
3. Discussion
4. Materials and Methods
4.1. Patients
4.2. Genetic Analysis
4.3. Statistical Analyses
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Garutti, M.; Foffano, L.; Mazzeo, R.; Michelotti, A.; Da Ros, L.; Viel, A.; Miolo, G.; Zambelli, A.; Puglisi, F. Hereditary Cancer Syndromes: A Comprehensive Review with a Visual Tool. Genes 2023, 14, 1025. [Google Scholar] [CrossRef] [PubMed]
- Nagy, R.; Sweet, K.; Eng, C. Highly Penetrant Hereditary Cancer Syndromes. Oncogene 2004, 23, 6445–6470. [Google Scholar] [CrossRef] [PubMed]
- Imyanitov, E.N.; Kuligina, E.S.; Sokolenko, A.P.; Suspitsin, E.N.; Yanus, G.A.; Iyevleva, A.G.; Ivantsov, A.O.; Aleksakhina, S.N. Hereditary Cancer Syndromes. World J. Clin. Oncol. 2023, 14, 40–68. [Google Scholar] [CrossRef]
- Ragupathi, A.; Singh, M.; Perez, A.M.; Zhang, D. Targeting the BRCA1/2 Deficient Cancer with PARP Inhibitors: Clinical Outcomes and Mechanistic Insights. Front. Cell Dev. Biol. 2023, 11, 1133472. [Google Scholar] [CrossRef]
- Infante, M.; Arranz-Ledo, M.; Lastra, E.; Abella, L.E.; Ferreira, R.; Orozco, M.; Hernández, L.; Martínez, N.; Durán, M. Increased Co-Occurrence of Pathogenic Variants in Hereditary Breast and Ovarian Cancer and Lynch Syndromes: A Consequence of Multigene Panel Genetic Testing? Int. J. Mol. Sci. 2022, 23, 11499. [Google Scholar] [CrossRef]
- McGuigan, A.; Whitworth, J.; Andreou, A.; Hearn, T.; Genomics England Research Consortium; Ambrose, J.C.; Arumugam, P.; Bevers, R.; Bleda, M.; Boardman-Pretty, F.; et al. Multilocus Inherited Neoplasia Allele Syndrome (MINAS): An Update. Eur. J. Hum. Genet. 2022, 30, 265–270. [Google Scholar] [CrossRef]
- Whitworth, J.; Skytte, A.-B.; Sunde, L.; Lim, D.H.; Arends, M.J.; Happerfield, L.; Frayling, I.M.; Van Minkelen, R.; Woodward, E.R.; Tischkowitz, M.D.; et al. Multilocus Inherited Neoplasia Alleles Syndrome: A Case Series and Review. JAMA Oncol. 2016, 2, 373–379. [Google Scholar] [CrossRef]
- Rebbeck, T.R.; Friebel, T.M.; Mitra, N.; Wan, F.; Chen, S.; Andrulis, I.L.; Apostolou, P.; Arnold, N.; Arun, B.K.; Barrowdale, D.; et al. Inheritance of Deleterious Mutations at Both BRCA1 and BRCA2 in an International Sample of 32,295 Women. Breast Cancer Res. 2016, 18, 112. [Google Scholar] [CrossRef]
- Le Page, C.; Rahimi, K.; Rodrigues, M.; Heinzelmann-Schwarz, V.; Recio, N.; Tommasi, S.; Bataillon, G.; Portelance, L.; Golmard, L.; Meunier, L.; et al. Clinicopathological Features of Women with Epithelial Ovarian Cancer and Double Heterozygosity for BRCA1 and BRCA2: A Systematic Review and Case Report Analysis. Gynecol. Oncol. 2020, 156, 377–386. [Google Scholar] [CrossRef]
- Laish, I.; Friedman, E.; Levi-Reznick, G.; Kedar, I.; Katz, L.; Levi, Z.; Halpern, N.; Parnasa, S.; Abu-Shatya, A.; Half, E.; et al. Double Heterozygotes of BRCA1/BRCA2 and Mismatch Repair Gene Pathogenic Variants: Case Series and Clinical Implications. Breast Cancer Res. Treat. 2021, 188, 685–694. [Google Scholar] [CrossRef] [PubMed]
- Bang, Y.J.; Kwon, W.K.; Nam, S.J.; Kim, S.W.; Chae, B.-J.; Lee, S.K.; Ryu, J.M.; Kim, J.-W.; Yu, J.; Lee, J.E. Clinicopathological Characterization of Double Heterozygosity for BRCA1 and BRCA2 Variants in Korean Breast Cancer Patients. Cancer Res. Treat. 2022, 54, 827–833. [Google Scholar] [CrossRef] [PubMed]
- Van Der Merwe, N.C.; Buccimazza, I.; Rossouw, B.; Araujo, M.; Ntaita, K.S.; Schoeman, M.; Vorster, K.; Napo, K.; Kotze, M.J.; Oosthuizen, J. Clinical Relevance of Double Heterozygosity Revealed by Next-Generation Sequencing of Homologous Recombination Repair Pathway Genes in South African Breast Cancer Patients. Breast Cancer Res. Treat. 2024, 207, 331–342. [Google Scholar] [CrossRef]
- Laitman, Y.; Niskakoski, A.; Bernstein-Molho, R.; Koskinen, L.; Rabina, D.; Koskenvuo, J.; Friedman, E. Phenotypes of Carriers of Two Mutated Alleles in Major Cancer Susceptibility Genes. Breast Cancer Res. Treat. 2024, 208, 589–595. [Google Scholar] [CrossRef] [PubMed]
- Megid, T.B.C.; Barros-Filho, M.C.; Pisani, J.P.; Achatz, M.I. Double Heterozygous Pathogenic Variants Prevalence in a Cohort of Patients with Hereditary Breast Cancer. Front. Oncol. 2022, 12, 873395. [Google Scholar] [CrossRef]
- Agaoglu, N.B.; Doganay, L. Concurrent Pathogenic Variations in Patients with Hereditary Cancer Syndromes. Eur. J. Med. Genet. 2021, 64, 104366. [Google Scholar] [CrossRef]
- Stradella, A.; Del Valle, J.; Rofes, P.; Feliubadaló, L.; Grau Garces, È.; Velasco, À.; González, S.; Vargas, G.; Izquierdo, Á.; Campos, O.; et al. Does Multilocus Inherited Neoplasia Alleles Syndrome Have Severe Clinical Expression? J. Med. Genet. 2019, 56, 521–525. [Google Scholar] [CrossRef] [PubMed]
- Kwong, A.; Ho, C.Y.S.; Au, C.-H.; Ma, E.S.K. Double Heterozygosity for Germline Mutations in Chinese Breast Cancer Patients. Cancers 2024, 16, 2547. [Google Scholar] [CrossRef]
- Abe, A.; Imoto, I.; Ueki, A.; Nomura, H.; Kanao, H. Moderate-Risk Genes for Hereditary Ovarian Cancers Involved in the Homologous Recombination Repair Pathway. IJMS 2022, 23, 11790. [Google Scholar] [CrossRef]
- Daly, M.B.; Pal, T.; Berry, M.P.; Buys, S.S.; Dickson, P.; Domchek, S.M.; Elkhanany, A.; Friedman, S.; Goggins, M.; Hutton, M.L.; et al. Genetic/Familial High-Risk Assessment: Breast, Ovarian, and Pancreatic, Version 2.2021, NCCN Clinical Practice Guidelines in Oncology. J. Natl. Compr. Cancer Netw. 2021, 19, 77–102. [Google Scholar] [CrossRef]
- LaDuca, H.; Polley, E.C.; Yussuf, A.; Hoang, L.; Gutierrez, S.; Hart, S.N.; Yadav, S.; Hu, C.; Na, J.; Goldgar, D.E.; et al. A Clinical Guide to Hereditary Cancer Panel Testing: Evaluation of Gene-Specific Cancer Associations and Sensitivity of Genetic Testing Criteria in a Cohort of 165,000 High-Risk Patients. Genet. Med. 2020, 22, 407–415. [Google Scholar] [CrossRef]
- Rijken, J.A.; Niemeijer, N.D.; Jonker, M.A.; Eijkelenkamp, K.; Jansen, J.C.; Van Berkel, A.; Timmers, H.J.L.M.; Kunst, H.P.M.; Bisschop, P.H.L.T.; Kerstens, M.N.; et al. The Penetrance of Paraganglioma and Pheochromocytoma in SDHB Germline Mutation Carriers. Clin. Genet. 2018, 93, 60–66. [Google Scholar] [CrossRef]
- Maniam, P.; Zhou, K.; Lonergan, M.; Berg, J.N.; Goudie, D.R.; Newey, P.J. Pathogenicity and Penetrance of Germline SDHA Variants in Pheochromocytoma and Paraganglioma (PPGL). J. Endocr. Soc. 2018, 2, 806–816. [Google Scholar] [CrossRef] [PubMed]
- Lavie, O.; Narod, S.; Lejbkowicz, F.; Dishon, S.; Goldberg, Y.; Gemer, O.; Rennert, G. Double Heterozygosity in the BRCA1 and BRCA2 Genes in the Jewish Population. Ann. Oncol. 2011, 22, 964–966. [Google Scholar] [CrossRef] [PubMed]
- Leegte, B. Phenotypic Expression of Double Heterozygosity for BRCA1 and BRCA2 Germline Mutations. J. Med. Genet. 2005, 42, e20. [Google Scholar] [CrossRef]
- Agaoglu, N.B.; Bychkovsky, B.L.; Horton, C.; Lo, M.-T.; Polfus, L.; Carraway, C.; Hemyari, P.; Young, C.; Richardson, M.E.; Scheib, R.; et al. Cancer Burden in Individuals with Single versus Double Pathogenic Variants in Cancer Susceptibility Genes. Genet. Med. Open 2024, 2, 101829. [Google Scholar] [CrossRef]
- Yuen, J.; Zhou, S.; Caeser, R.; Venkatramani, M.; Bte Ishak, D.N.; Li, S.-T.; Zhang, Z.; Chiang, J.; Chan, S.H.; Ngeow, J. Multi-Locus Inherited Neoplasia Alleles Syndromes in Cancer: Implications for Clinical Practice. Eur. J. Hum. Genet. 2025, 33, 289–296. [Google Scholar] [CrossRef]
- Nagy, P.; Papp, J.; Grolmusz, V.K.; Bozsik, A.; Pócza, T.; Oláh, E.; Patócs, A.; Butz, H. Comprehensive Clinical Genetics, Molecular and Pathological Evaluation Efficiently Assist Diagnostics and Therapy Selection in Breast Cancer Patients with Hereditary Genetic Background. Int. J. Mol. Sci. 2024, 25, 12546. [Google Scholar] [CrossRef]
- Horti-Oravecz, K.; Bozsik, A.; Pócza, T.; Vereczkey, I.; Strausz, T.; Tóth, E.; Sedlackova, T.; Rusnakova, D.; Szemes, T.; Likó, I.; et al. Whole Genome Sequencing Completes the Molecular Genetic Testing Workflow of Patients with Lynch Syndrome. npj Genom. Med. 2025, 10, 5. [Google Scholar] [CrossRef] [PubMed]
- Bilyalov, A.; Nikolaev, S.; Shigapova, L.; Khatkov, I.; Danishevich, A.; Zhukova, L.; Smolin, S.; Titova, M.; Lisica, T.; Bodunova, N.; et al. Application of Multigene Panels Testing for Hereditary Cancer Syndromes. Biology 2022, 11, 1461. [Google Scholar] [CrossRef]
- Syed, H.; Sommovilla, J.; Burke, C.A.; McGee, S.; Macaron, C.; Heald, B.; Lyu, R.; Schmit, S.L.; Nair, K.; Kamath, S.; et al. Referral, Uptake, and Outcome of Genetic Counseling and Testing in Patients With Early-Onset Colorectal Cancer. J. Natl. Compr. Cancer Netw. 2023, 21, 1156–1163.e5. [Google Scholar] [CrossRef]
- Aguilar, D.; Garza-Rodríguez, M.L.; Pérez-Ibave, D.C.; Muñiz-Garza, C.E.; Treviño, V.; Villarreal-Garza, C.M.; Vidal-Gutiérrez, O.; Burciaga-Flores, C.H. Landscape of Multilocus Inherited Neoplasia Allele Syndrome in Mexican Population. JCO Glob. Oncol. 2025, 11, e2400065. [Google Scholar] [CrossRef]
- Heidemann, S.; Fischer, C.; Engel, C.; Fischer, B.; Harder, L.; Schlegelberger, B.; Niederacher, D.; Goecke, T.O.; Doelken, S.C.; Dikow, N.; et al. Double Heterozygosity for Mutations in BRCA1 and BRCA2 in German Breast Cancer Patients: Implications on Test Strategies and Clinical Management. Breast Cancer Res. Treat. 2012, 134, 1229–1239. [Google Scholar] [CrossRef]
- Rainville, I.; Hatcher, S.; Rosenthal, E.; Larson, K.; Bernhisel, R.; Meek, S.; Gorringe, H.; Mundt, E.; Manley, S. High Risk of Breast Cancer in Women with Biallelic Pathogenic Variants in CHEK2. Breast Cancer Res. Treat. 2020, 180, 503–509. [Google Scholar] [CrossRef] [PubMed]
- Ozanne, E.M.; O’Connell, A.; Bouzan, C.; Bosinoff, P.; Rourke, T.; Dowd, D.; Drohan, B.; Millham, F.; Griffin, P.; Halpern, E.F.; et al. Bias in the Reporting of Family History: Implications for Clinical Care. J. Genet. Couns. 2012, 21, 547–556. [Google Scholar] [CrossRef] [PubMed]
- Brock, P.; Geurts, J.L.; Van Galen, P.; Blouch, E.; Welch, J.; Kunz, A.; Desrosiers, L.; Gauerke, J.; Hyde, S. Hereditary Endocrine Tumours: Current State-of-the-art and Research Opportunities: Challenges and Opportunities in Genetic Counseling for Hereditary Endocrine Neoplasia Syndromes. Endocr. Relat. Cancer 2020, 27, T65–T75. [Google Scholar] [CrossRef]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and Guidelines for the Interpretation of Sequence Variants: A Joint Consensus Recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef] [PubMed]
- Oleinikov, K.; Grozinsky-Glasberg, S. Hereditary Syndromes Associated with Neuroendocrine Tumors. Curr. Opin. Endocr. Metab. Res. 2021, 18, 230–235. [Google Scholar] [CrossRef]
- Patócs, A.; Nagy, P.; Papp, J.; Bozsik, A.; Antal, B.; Grolmusz, V.K.; Pócza, T.; Butz, H. Cost-Effectiveness of Genetic Testing of Endocrine Tumor Patients Using a Comprehensive Hereditary Cancer Gene Panel. J. Clin. Endocrinol. Metab. 2024, 109, 3220–3233. [Google Scholar] [CrossRef]
- Bychkovsky, B.L.; Agaoglu, N.B.; Horton, C.; Zhou, J.; Yussuf, A.; Hemyari, P.; Richardson, M.E.; Young, C.; LaDuca, H.; McGuinness, D.L.; et al. Differences in Cancer Phenotypes Among Frequent CHEK2 Variants and Implications for Clinical Care—Checking CHEK2. JAMA Oncol. 2022, 8, 1598–1606. [Google Scholar] [CrossRef]
- Rozen, P.; Naiman, T.; Strul, H.; Taussky, P.; Karminsky, N.; Shomrat, R.; Samuel, Z.; Yaron, Y.; Orr-Urtreger, A. Clinical and Screening Implications of the I1307K Adenomatous Polyposis Coli Gene Variant in Israeli Ashkenazi Jews with Familial Colorectal Neoplasia. Evidence for a Founder Effect. Cancer 2002, 94, 2561–2568. [Google Scholar] [CrossRef]
- Ukaegbu, C.; Levi, Z.; Fehlmann, T.D.; Uno, H.; Chittenden, A.; Inra, J.A.; Grover, S.; Kastrinos, F.; Syngal, S.; Yurgelun, M.B. Characterizing Germline APC and MUTYH Variants in Ashkenazi Jews Compared to Other Individuals. Fam. Cancer 2021, 20, 111–116. [Google Scholar] [CrossRef] [PubMed]
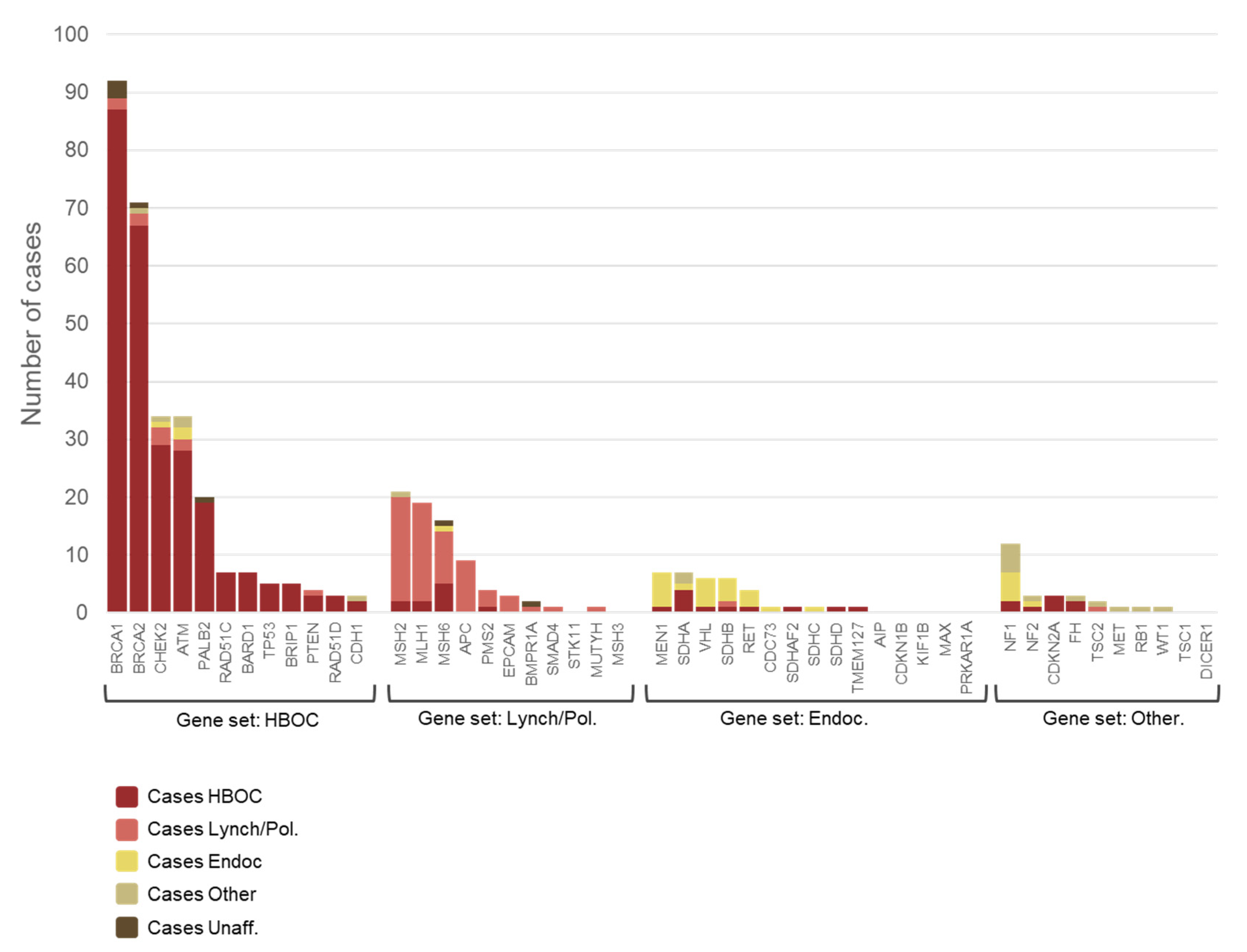

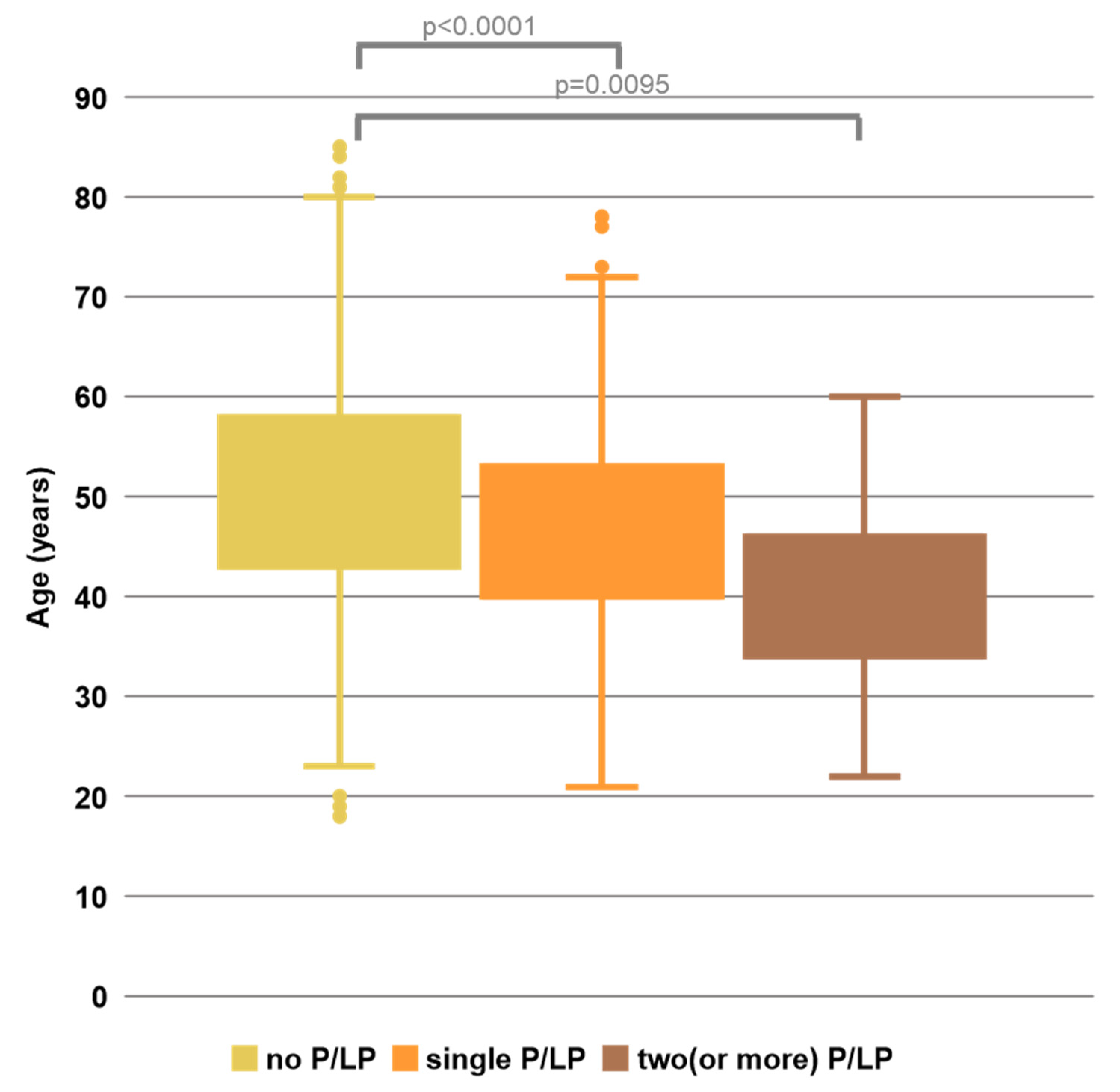
| HBOC | Lynch + Pol. | Endocrine | Other | Unaffected | Sum | |
|---|---|---|---|---|---|---|
| Case number | 1496 | 268 | 146 | 84 | 56 | 2050 |
| % | 73.0 | 13.1 | 7.1 | 4.1 | 2.7 | 100 |
| Gender | ||||||
| Female (number) | 1428 | 187 | 86 | 45 | 56 | 1802 |
| % | 95.5 | 69.8 | 58.9 | 53.6 | 100.0 | 87.9 |
| Male (number) | 68 | 81 | 60 | 39 | 0 | 248 |
| % | 4.5 | 30.2 | 41.1 | 46.4 | 0.0 | 12.1 |
| Mean age (years) | 49.5 | 51.9 | 41.2 | 41.9 | n.a. | |
| SD | 11.5 | 15.4 | 18.3 | 19.4 | n.a. | |
| Median age (years) | 48 | 54 | 42 | 44 | n.a. | 48 |
| min | 18 | 5 | 0 a | 0 b | n.a. | 0 b |
| max | 85 | 78 | 83 | 80 | n.a. | 85 |
| Muliple primaries | 203 | 49 | 5 | 22 | n.a. | 279 |
| % | 13.6 | 18.3 | 3.4 | 26.2 | n.a. | 13.6 |
| Number of P/LP cases | 277 | 73 | 31 | 18 | 6 | 405 |
| % | 18.5 | 27.2 | 21.2 | 21.4 | 10.7 | 19.8 |
| Number of DH cases | 13 | 1 | 0 | 1 | 1 | 16 |
| % | 0.9 | 0.4 | 0.0 | 1.2 | 1.8 | 0.8 |
| (A) | ||||||||||||
| # | Sex | Suspected Sy | Tumour_1 | Age | Tumour_2 | Age | MPs | Tumour Histology | Family History | |||
| 1 | FEM | HBOC | br | 44 | IDC; ER +; PR +; Her2 -; Ki67 80% | br (52 + 56); br (58) | ||||||
| 2 | FEM | Lynch/Polyposis | end | 36 | endometrioid cc with squamous differentiation | Lung (?); cr (43); st (37); st (53); br (60) + ov (60) | ||||||
| 3 | FEM | HBOC | br | 22 | IDC; ER -; PR -; Her2 -; Ki67 70% | Skin (45); br (77); cr (70) | ||||||
| 4 | FEM | HBOC | br | 37 | IDC; ER +; PR -; Her2 +; Ki67 80% | br (58); br (59 + 65); st (49); st (86) | ||||||
| 5 | FEM | unaff | unaff | unaffected | Br (42); br (?) | |||||||
| 6 | FEM | HBOC | br | 45 | NHL | 50 | IDC; ER -; PR -; Her2 -; Ki67 ?% | cr (60); br (57); gb (?); cr (?); ut (51) + ov (51) | ||||
| 7 | FEM | HBOC | br | 47 | IDC; ER +; PR +; Her2 -; Ki67 40–50% | br (44) | ||||||
| 8 | FEM | HBOC | br | 30 | IDC; ER +; PR +; Her2 ?; Ki67 20% | br (53); br (66); cr (58); cr (50) | ||||||
| 9 | MAL | HBOC | prostate | 60 | adenocc. | br (49); nb (2) | ||||||
| 10 | FEM | HBOC | end | 54 | breast | 66 | y | adenocc.; LCIS + ILC; ER +; PR +; Her2 -; Ki67 1–2% | br (42); brain (43); ren (?); brain (?); st (?) | |||
| 11 | FEM | HBOC | br | 35 | IDC; ER -; PR -; Her2 -; Ki67 50% | br (68); lung (54); bone (?) | ||||||
| 12 | FEM | HBOC | br | 42 | IDC + DCIS; ER -; PR -; Her2 +; Ki67 10% | br (?); lung (?) | ||||||
| 13 | FEM | HBOC | br | 33 | IDC; ER -; PR -; Her2 -; Ki67 80% | br (55) | ||||||
| 14 | FEM | HBOC | br (bilat) | 37 | y | ILC; ER +; PR +; Her2 +; Ki67 10–15% | pros (46); panc (46); liver (46) | |||||
| 15 | FEM | HBOC | br | 43 | IDC; ER +; PR +; Her2 -; Ki67 10% | Cervix (44) | ||||||
| 16 | FEM | other | ut (leiomyoma) | 32 | FH deficient ut leiomyoma | br (48); br (66); larynx (52) | ||||||
| (B) | ||||||||||||
| # | Gene_1 | HGVS Transcript_1 | HGVS Protein_1 | Gene_2 | HGVS Transcript_2 | HGVS Protein_2 | Gene_3 | HGVS Transcript_3 | HGVS Protein_3 | |||
| 1 | ATM | NM_000051.4:c.6679C>T | NP_000042.3:p.(Arg2227Cys) | BARD1 | NM_000465.4:c.1932_1933del | NP_000456.2:p.(Cys645Ter) | ||||||
| 2 | MSH2 | NM_000251.3:c.2068C>T | NP_000242.1:p.(Gln690Ter) | SDHB | NM_003000.3:c.148G>T | NP_002991.2:p.(Asp50Tyr) | ||||||
| 3 | BRCA1 | NM_007294.4:c.5266dup | NP_009225.1:p.(Gln1756ProfsTer74) | MSH6 | NM_000179.3:c.3261del | NP_000170.1:p.(Phe1088SerfsTer2) | ||||||
| 4 | CHEK2 | NM_007194.4: del(exon 9–10) | NP_009125.1:p.? | CHEK2 | NM_007194.4:c.499G>A | NP_009125.1:p.(Gly167Arg) | ||||||
| 5 | BRCA1 | NM_007294.4:c.181T>G | NP_009225.1:p.(Cys61Gly) | MSH6 | NM_000179.3:c.3379_3438+5del | NP_000170.1:p.? | ||||||
| 6 | PALB2 | NM_024675.4: del(exon 11) | NP_078951.2:p.? | SDHA | NM_004168.4:c.91C>T | NP_004159.2:p.(Arg31Ter) | ||||||
| 7 | BRCA2 | NM_000059.4:c.5682C>G | NP_000050.3:p.(Tyr1894Ter) | BRIP1 | NM_032043.3:c.3525dup | NP_114432.2:p.(Ile1176TyrfsTer13) | ||||||
| 8 | BRCA2 | NM_000059.4:c.8249_8251del | NP_000050.3:p.(Lys2750del) | ATM | NM_000051.4:c.1564_1565del | NP_000042.3:p.(Glu522IlefsTer43) | ||||||
| 9 | BRCA1 | NM_007294.4:c.181T>G | NP_009225.1:p.(Cys61Gly) | BRCA2 | NM_000059.4:c.6295A>T | NP_000050.3:p.(Arg2099Ter) | ||||||
| 10 | CHEK2 | NM_007194.4:c.1100del | NP_009125.1:p.(Thr367MetfsTer15) | SDHAF2 | NM_017841.4:c.446_447del | NP_060311.1:p.(Lys149ArgfsTer10) | ||||||
| 11 | BRCA2 | NM_000059.4:c.5073dup | NP_000050.3:p.(Trp1692MetfsTer3) | MSH6 | NM_000179.3:c.3379_3438+5del | NP_000170.1:p.? | ||||||
| 12 | BRCA1 | NM_007294.4:c.181T>G | NP_009225.1:p.(Cys61Gly) | ATM | NM_000051.4:c.7630-2A>C | NP_000042.3:p.? | SDHA | del (whole gene) | NP_004159.2:p.? | |||
| 13 | BRCA1 | NM_007294.4:c.181T>G | NP_009225.1:p.(Cys61Gly) | BRIP1 | NM_032043.3:c.1741C>T | NP_114432.2:p.(Arg581Ter) | ||||||
| 14 | BRCA2 | NM_000059.4:c.5645C>A | NP_000050.3:p.(Ser1882Ter) | CDH1 | NM_004360.5:c.1711+5G>A | NP_004351.1:p.? | ||||||
| 15 | CHEK2 | NM_007194.4:c.1337del | NP_009125.1:p.(Asn446ThrfsTer23) | CHEK2 | NM_007194.4:c.277del | NP_009125.1:p.(Trp93GlyfsTer17) | ||||||
| 16 | ATM | NM_000051.4:c.381del | NP_000042.3:p.(Val128Ter) | CDH1 | NM_004360.5:c.1063_1066dup | NP_004351.1:p.(Ser356IlefsTer13) | ||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pócza, T.; Papp, J.; Bozsik, A.; Grolmusz, V.K.; Nagy, P.; Patócs, A.; Butz, H. Double Pathogenic or Likely Pathogenic Variants in Cancer Predisposition Genes in Hungarian Cancer Patients. Int. J. Mol. Sci. 2025, 26, 8390. https://doi.org/10.3390/ijms26178390
Pócza T, Papp J, Bozsik A, Grolmusz VK, Nagy P, Patócs A, Butz H. Double Pathogenic or Likely Pathogenic Variants in Cancer Predisposition Genes in Hungarian Cancer Patients. International Journal of Molecular Sciences. 2025; 26(17):8390. https://doi.org/10.3390/ijms26178390
Chicago/Turabian StylePócza, Tímea, János Papp, Anikó Bozsik, Vince Kornél Grolmusz, Petra Nagy, Attila Patócs, and Henriett Butz. 2025. "Double Pathogenic or Likely Pathogenic Variants in Cancer Predisposition Genes in Hungarian Cancer Patients" International Journal of Molecular Sciences 26, no. 17: 8390. https://doi.org/10.3390/ijms26178390
APA StylePócza, T., Papp, J., Bozsik, A., Grolmusz, V. K., Nagy, P., Patócs, A., & Butz, H. (2025). Double Pathogenic or Likely Pathogenic Variants in Cancer Predisposition Genes in Hungarian Cancer Patients. International Journal of Molecular Sciences, 26(17), 8390. https://doi.org/10.3390/ijms26178390






