Examining the Roles of Genomic Context and Endogenous Regulatory Elements on IS1 Transposition Within the Escherichia coli Genome
Abstract
1. Introduction
2. Results
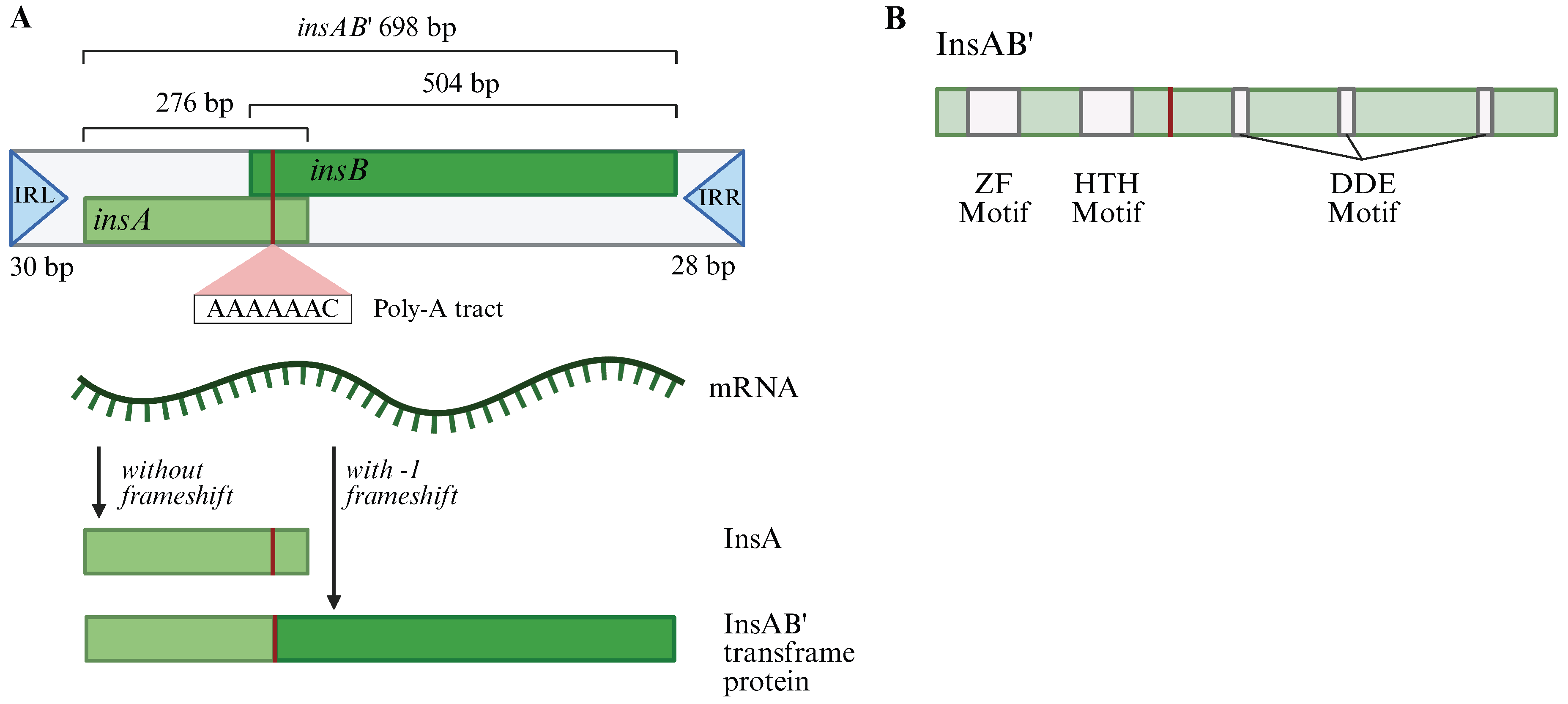
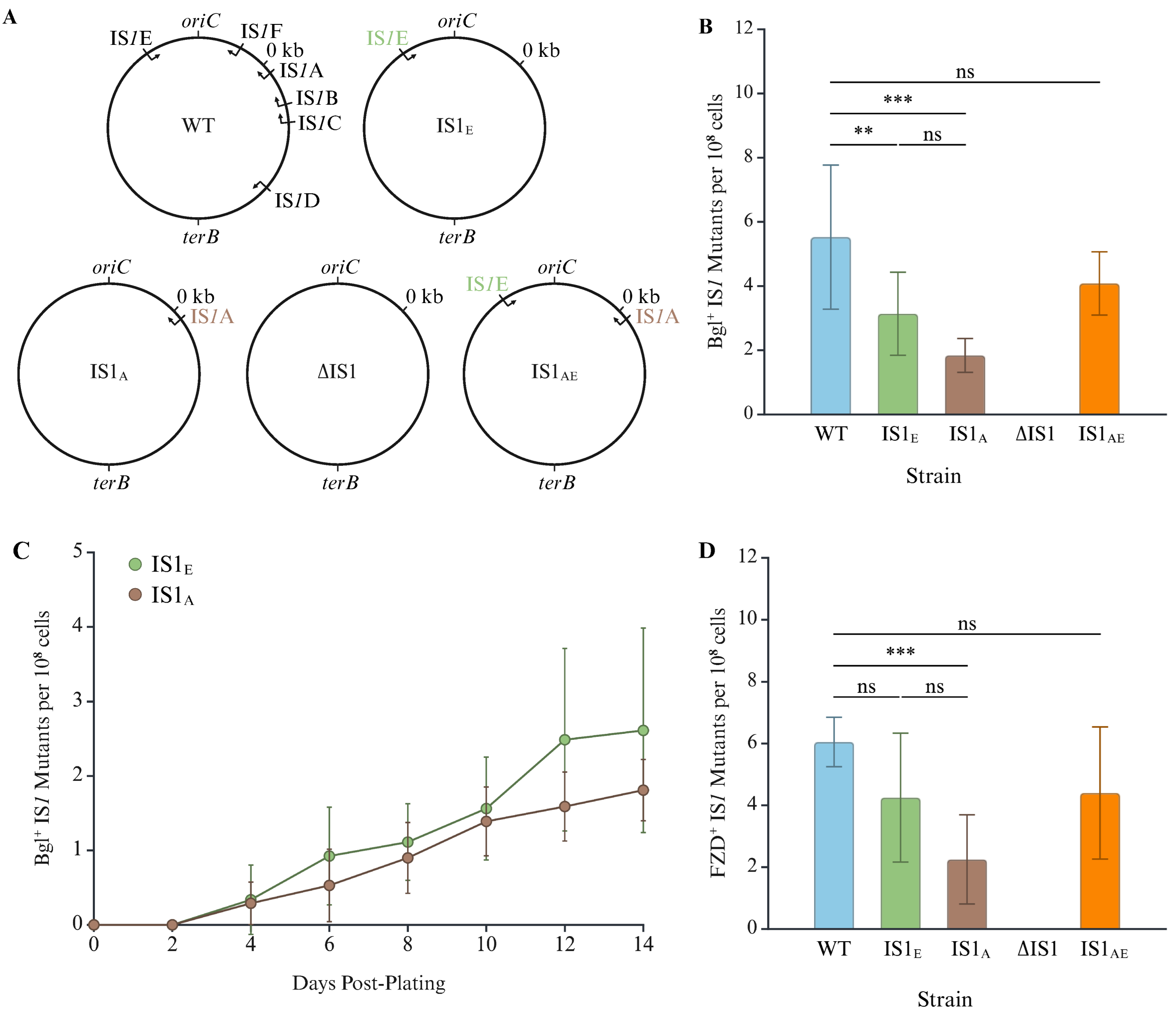
2.1. IS1A and IS1E Maintain the IS1 Consensus Sequence and Are the Primary Contributors to Transposition
2.2. IS1E Transposes at a Higher Frequency than IS1A Although Both Are Main Contributors for IS1 Transposition
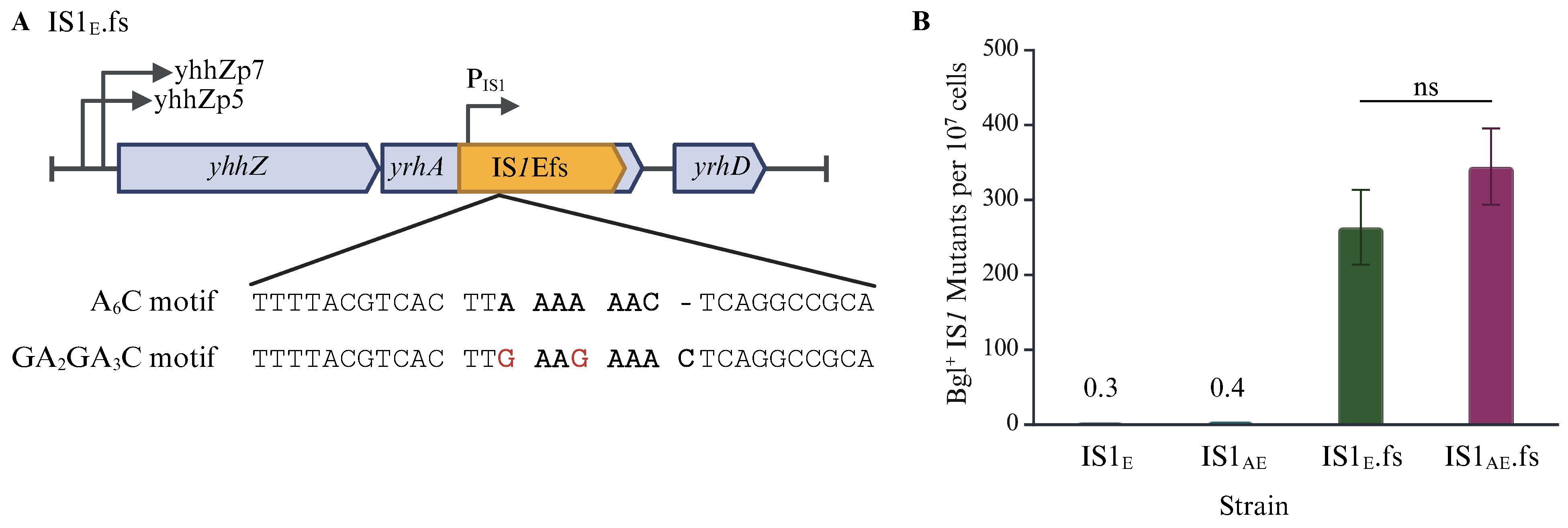
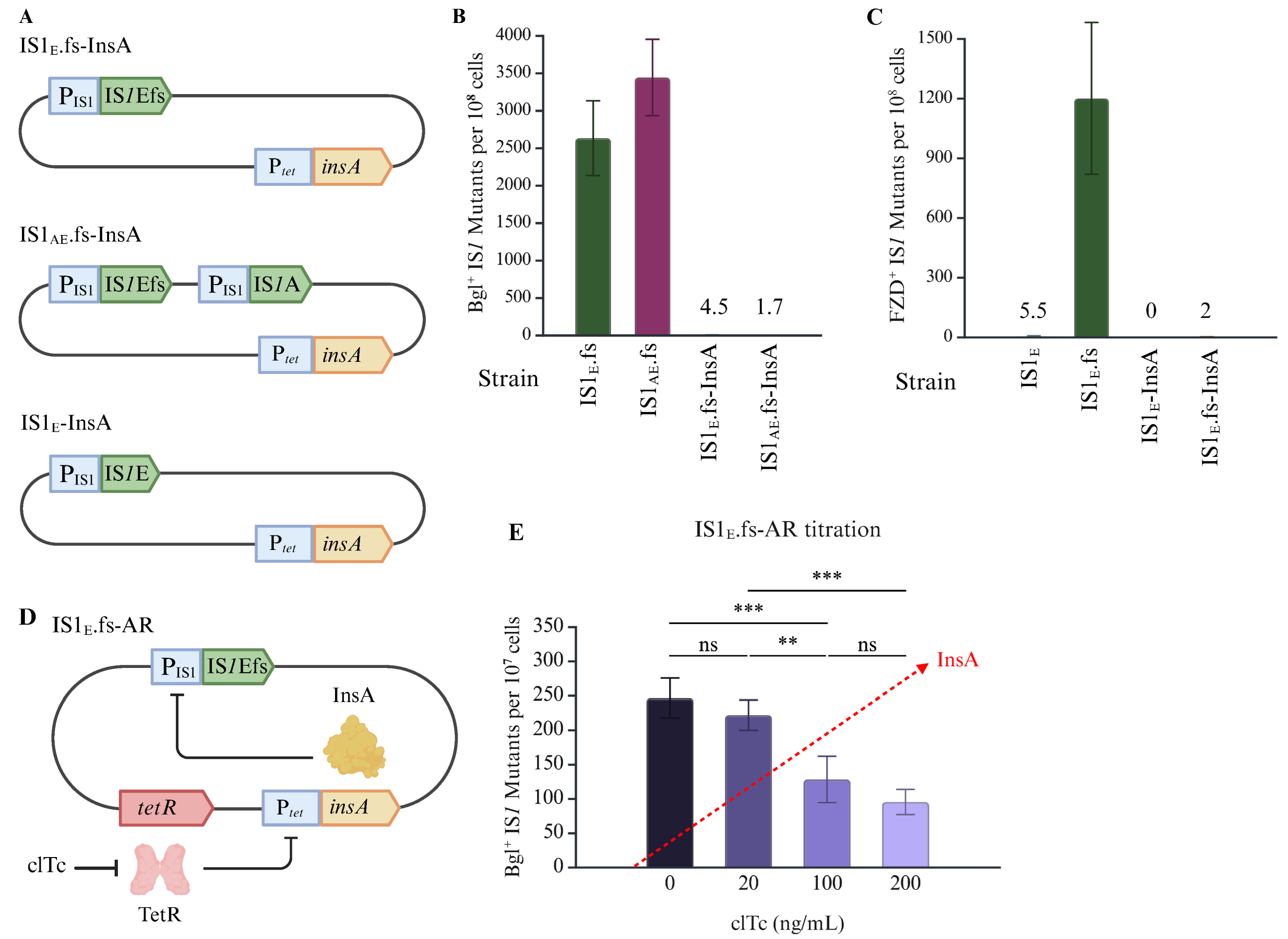
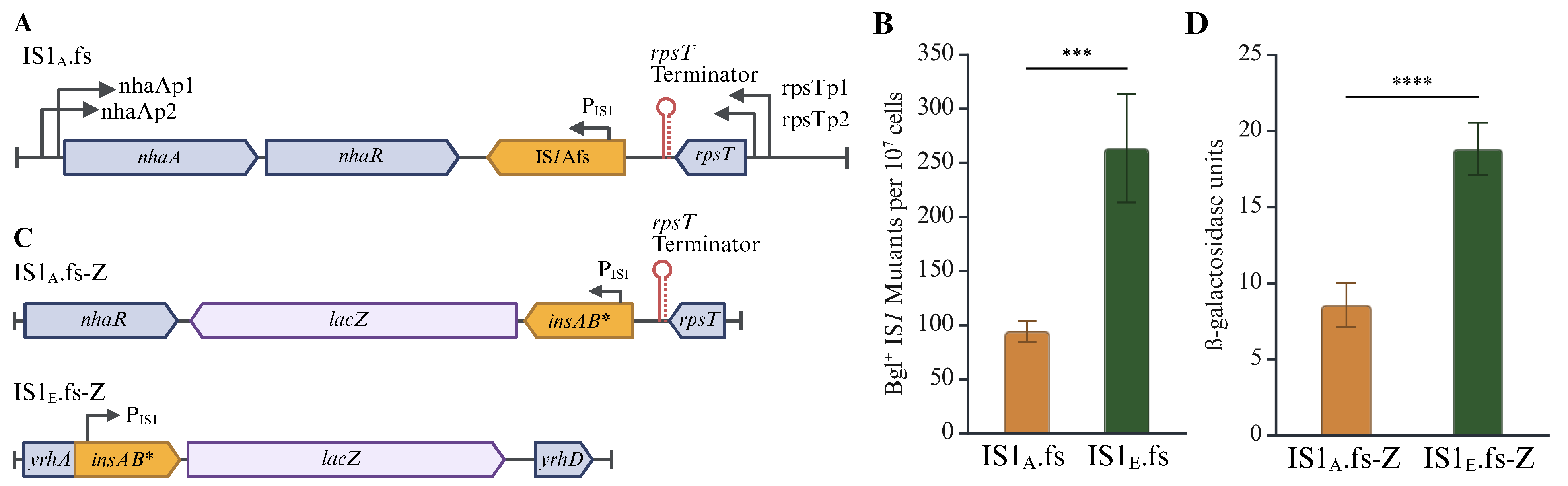
2.3. Translational Regulation Has a Large Effect on IS1 Transposition
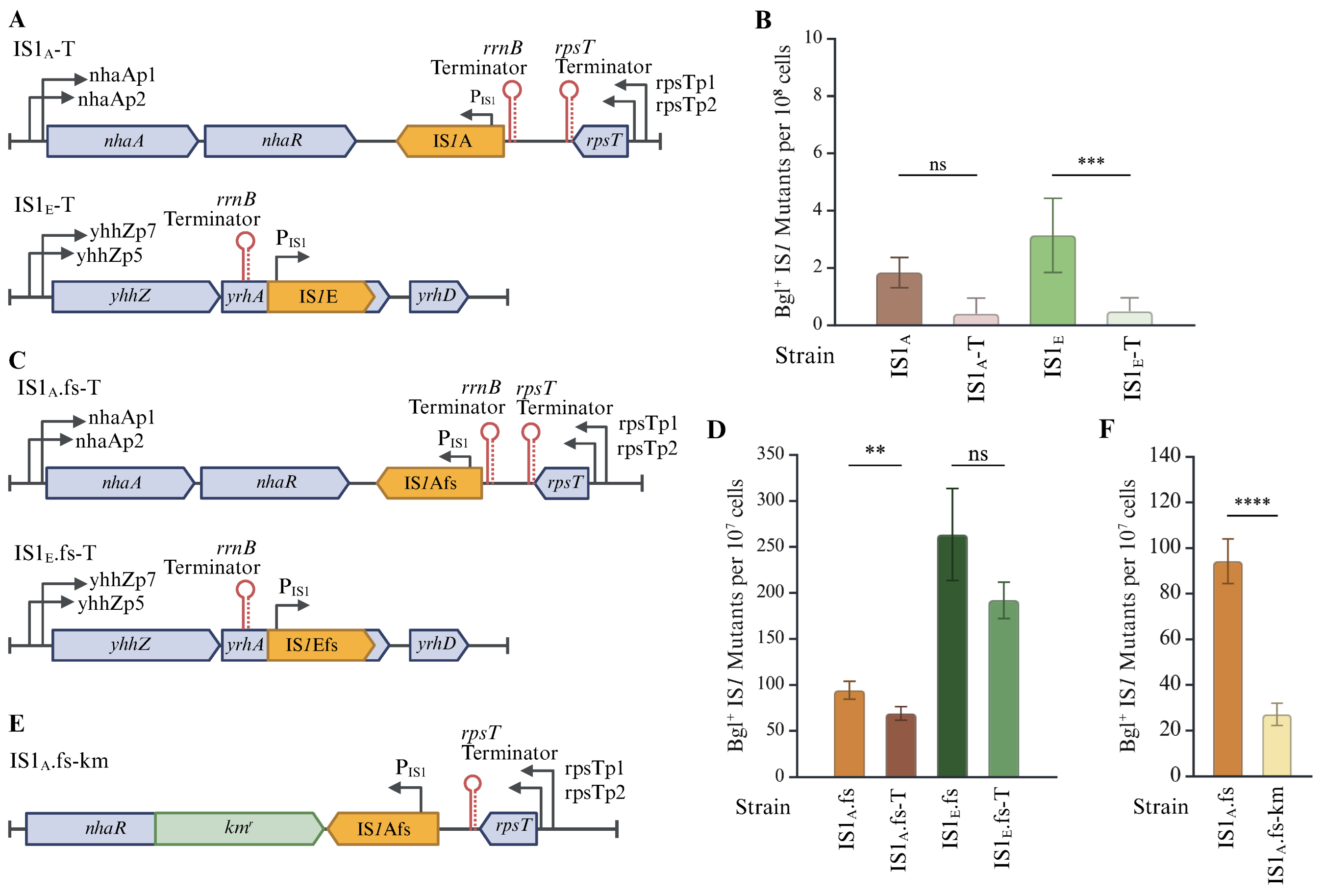
2.4. Transcriptional Regulation Has a Large Effect on IS1 Transposition
2.5. Effects of Genomic Contexts on IS1 Transposition
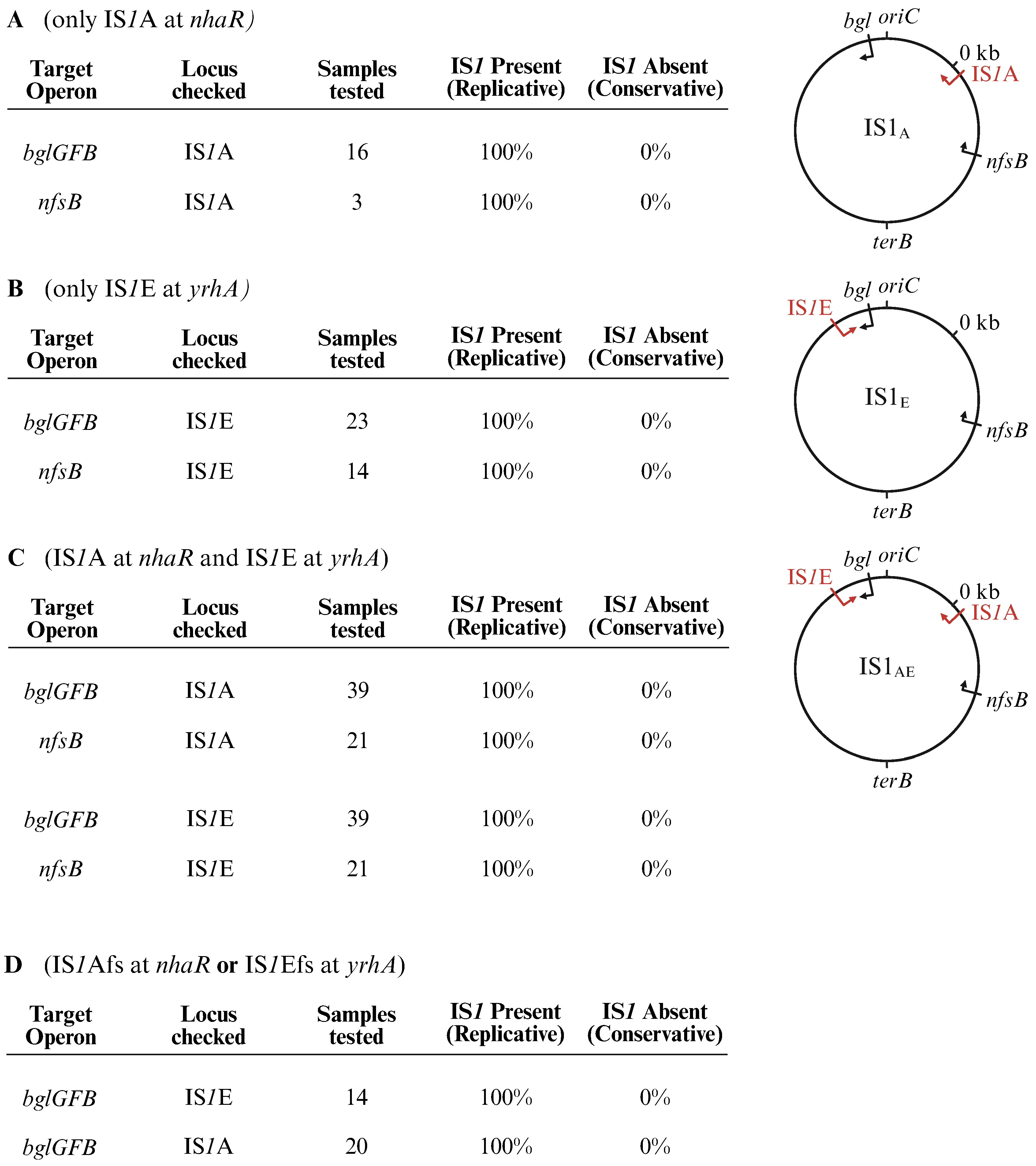
2.6. IS1 Mainly Uses a Replicative Mechanism for Transposition Within the Genome
3. Discussion
4. Materials and Methods
4.1. E. coli Strains and Growth Conditions
4.2. Deletions of IS1 Elements from Chromosome
4.3. Construction of IS1fs at yrhA Locus and IS1fs at the nhaR Locus
4.4. Construction of Ptet Driving insA at the intS Locus
4.5. Construction of Transcriptional Reporters Using lacZ
4.6. Addition of rrnB Terminator
4.7. β-Glucoside Growth Mutation (Bgl+) Assay
4.8. Swarming Mutation (SWM+) Assay
4.9. Furazolidone Resistance Mutation (FZD+) Assay
4.10. β-Galactosidase Assay
4.11. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Siguier, P.; Filée, J.; Chandler, M. Insertion Sequences in Prokaryotic Genomes. Curr. Opin. Microbiol. 2006, 9, 526–531. [Google Scholar] [CrossRef]
- Ooka, T.; Ogura, Y.; Asadulghani, M.; Ohnishi, M.; Nakayama, K.; Terajima, J.; Watanabe, H.; Hayashi, T. Inference of the Impact of Insertion Sequence (IS) Elements on Bacterial Genome Diversification through Analysis of Small-Size Structural Polymorphisms in Escherichia coli O157 Genomes. Genome Res. 2009, 19, 1809–1816. [Google Scholar] [CrossRef] [PubMed]
- Jespersen, M.G.; Hayes, A.J.; Tong, S.Y.C.; Davies, M.R. Insertion Sequence Elements and Unique Symmetrical Genomic Regions Mediate Chromosomal Inversions in Streptococcus Pyogenes. Nucleic Acids Res. 2024, 52, 13128–13137. [Google Scholar] [CrossRef]
- Ngan, W.Y.; Parab, L.; Bertels, F.; Gallie, J. A More Significant Role for Insertion Sequences in Large-Scale Rearrangements in Bacterial Genomes. MBio 2024, 16, e0305224. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y.; Sabnis, A.; Mumin, Z.; Potterill, I.; Jauneikaite, E.; Brown, C.S.; Ellington, M.J.; Edwards, A.; Sriskandan, S. IS1-Related Large-Scale Deletion of Chromosomal Regions Harbouring the Oxygen-Insensitive Nitroreductase Gene NfsB Causes Nitrofurantoin Heteroresistance in Escherichia coli. Microb. Genom. 2023, 9, 001102. [Google Scholar] [CrossRef]
- Aubert, D.; Naas, T.; Héritier, C.; Poirel, L.; Nordmann, P. Functional Characterization of IS1999, an IS4 Family Element Involved in Mobilization and Expression of Beta-Lactam Resistance Genes. J. Bacteriol. 2006, 188, 6506–6514. [Google Scholar] [CrossRef]
- Whiteway, J.; Koziarz, P.; Veall, J.; Sandhu, N.; Kumar, P.; Hoecher, B.; Lambert, I.B. Oxygen-Insensitive Nitroreductases: Analysis of the Roles of NfsA and NfsB in Development of Resistance to 5-Nitrofuran Derivatives in Escherichia coli. J. Bacteriol. 1998, 180, 5529–5539. [Google Scholar] [CrossRef] [PubMed]
- Lipszyc, A.; Szuplewska, M.; Bartosik, D. How Do Transposable Elements Activate Expression of Transcriptionally Silent Antibiotic Resistance Genes? Int. J. Mol. Sci. 2022, 23, 8063. [Google Scholar] [CrossRef]
- Noel, H.R.; Petrey, J.R.; Palmer, L.D. Mobile Genetic Elements in Acinetobacter Antibiotic-Resistance Acquisition and Dissemination. Ann. N. Y. Acad. Sci. 2022, 1518, 166–182. [Google Scholar] [CrossRef]
- Łobocka, M.B.; Rose, D.J.; Plunkett, G., 3rd; Rusin, M.; Samojedny, A.; Lehnherr, H.; Yarmolinsky, M.B.; Blattner, F.R. Genome of Bacteriophage P1. J. Bacteriol. 2004, 186, 7032–7068. [Google Scholar] [CrossRef]
- Iida, S.; Meyer, J.; Arber, W. The Insertion Element IS1 Is a Natural Constituent of Coliphage P1 DNA. Plasmid 1978, 1, 357–365. [Google Scholar] [CrossRef] [PubMed]
- Brügger, K.; Redder, P.; She, Q.; Confalonieri, F.; Zivanovic, Y.; Garrett, R. Mobile Elements in Archaeal Genomes. FEMS Microbiol. Lett. 2002, 206, 131–141. [Google Scholar] [CrossRef] [PubMed]
- Prentki, P.; Teter, B.; Chandler, M.; Galas, D.J. Functional Promoters Created by the Insertion of Transposable Element ISI. J. Mol. BioE. 1986, 191, 383–393. [Google Scholar] [CrossRef]
- Olliver, A.; Vallé, M.; Chaslus-Dancla, E.; Cloeckaert, A. Overexpression of the Multidrug Efflux Operon AcrEF by Insertional Activation with IS1 or IS10 Elements in Salmonella Enterica Serovar Typhimurium DT204 AcrB Mutants Selected with Fluoroquinolones. Antimicrob. Agents Chemother. 2005, 49, 289–301. [Google Scholar] [CrossRef]
- Fan, C.; Wu, Y.-H.; Decker, C.M.; Rohani, R.; Gesell Salazar, M.; Ye, H.; Cui, Z.; Schmidt, F.; Huang, W.E. Defensive Function of Transposable Elements in Bacteria. ACS Synth. Biol. 2019, 8, 2141–2151. [Google Scholar] [CrossRef]
- Ton-Hoang, B.; Turlan, C.; Chandler, M. Functional Domains of the IS1 Transposase: Analysis in Vivo and in Vitro. Mol. Microbiol. 2004, 53, 1529–1543. [Google Scholar] [CrossRef] [PubMed]
- Kopkowski, P.W.; Zhang, Z.; Saier, M.H., Jr. The Effect of DNA—Binding Proteins on Insertion Sequence Element Transposition Upstream of the Bgl Operon in Escherichia coli. Front. Microbiol. 2024, 15, 1388522. [Google Scholar] [CrossRef]
- Escoubas, J.M.; Prère, M.F.; Fayet, O.; Salvignol, I.; Galas, D.; Zerbib, D.; Chandler, M. Translational Control of Transposition Activity of the Bacterial Insertion Sequence IS1. EMBO J. 1991, 10, 705–712. [Google Scholar] [CrossRef]
- Giedroc, D.P.; Theimer, C.A.; Nixon, P.L. Structure, Stability and Function of RNA Pseudoknots Involved in Stimulating Ribosomal Frameshifting. J. Mol. Biol. 2000, 298, 167–185. [Google Scholar] [CrossRef]
- Ohta, S.; Yoshimura, E.; Ohtsubo, E. Involvement of Two Domains with Helix-Turn-Helix and Zinc Finger Motifs in the Binding of IS1 Transposase to Terminal Inverted Repeats. Mol. Microbiol. 2004, 53, 193–202. [Google Scholar] [CrossRef]
- Zerbib, D.; Polard, P.; Escoubas, J.M.; Galas, D.; Chandler, M. The Regulatory Role of the IS1-Encoded InsA Protein in Transposition. Mol. Microbiol. 1990, 4, 471–477. [Google Scholar] [CrossRef]
- Siguier, P.; Gagnevin, L.; Chandler, M. The New IS1595 Family, Its Relation to IS1 and the Frontier between Insertion Sequences and Transposons. Res. Microbiol. 2009, 160, 232–241. [Google Scholar] [CrossRef] [PubMed]
- Grindley, N.D. IS1 Insertion Generates Duplication of a Nine Base Pair Sequence at Its Target Site. Cell 1978, 13, 419–426. [Google Scholar] [CrossRef]
- Hu, M.; Deonier, R.C. Comparison of IS1, IS2 and IS3 Copy Number in Escherichia coli Strains K-12, B and C. Gene 1981, 16, 161–170. [Google Scholar] [CrossRef]
- Schneider, D.; Duperchy, E.; Depeyrot, J.; Coursange, E.; Lenski, R.; Blot, M. Genomic Comparisons among Escherichia coli Strains B, K-12, and O157:H7 Using IS Elements as Molecular Markers. BMC Microbiol. 2002, 2, 18. [Google Scholar] [CrossRef]
- Karp, P.D.; Billington, R.; Caspi, R.; Fulcher, C.A.; Latendresse, M.; Kothari, A.; Keseler, I.M.; Krummenacker, M.; Midford, P.E.; Ong, Q.; et al. The BioCyc Collection of Microbial Genomes and Metabolic Pathways. Brief. Bioinform. 2019, 20, 1085–1093. [Google Scholar] [CrossRef] [PubMed]
- Nesmelova, I.V.; Hackett, P.B. DDE Transposases: Structural Similarity and Diversity. Adv. Drug Deliv. Rev. 2010, 62, 1187–1195. [Google Scholar] [CrossRef] [PubMed]
- Hickman, A.B.; Dyda, F. Mechanisms of DNA Transposition. Microbiol. Spectr. 2015, 3, MDNA3-0034–2014. [Google Scholar] [CrossRef] [PubMed]
- Machida, Y.; Machida, C.; Ohtsubo, H.; Ohtsubo, E. Factors Determining Frequency of Plasmid Cointegration Mediated by Insertion Sequence IS1. Proc. Natl. Acad. Sci. USA 1982, 79, 277–281. [Google Scholar] [CrossRef]
- Biel, S.W.; Berg, D.E. Mechanism of IS1 Transposition in E. Coli: Choice between Simple Insertion and Cointegration. Genetics 1984, 108, 319–330. [Google Scholar] [CrossRef]
- Shiga, Y.; Sekine, Y.; Ohtsubo, E. Transposition of IS1 Circles. Genes Cells 1999, 4, 551–561. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.H.; Hsu, W.B.; Hwang, J.L. Two Amino Acid Residues of Transposase Contributing to Differential Transposability of IS1 Elements in Escherichia coli. J. Bacteriol. 1998, 180, 5279–5283. [Google Scholar] [CrossRef]
- Onstead, J.; Zhang, Z.; Huo, J.; Ord, J.W.; Smith, S.; Saier, M.H., Jr. Investigating How Genomic Contexts Impact IS5 Transposition within the Escherichia coli Genome. Microorganisms 2024, 12, 2600. [Google Scholar] [CrossRef]
- Curcio, M.J.; Derbyshire, K.M. The Outs and Ins of Transposition: From Mu to Kangaroo. Nat. Rev. Mol. Cell Biol. 2003, 4, 865–877. [Google Scholar] [CrossRef] [PubMed]
- Derbyshire, K.M.; Grindley, N.D. Replicative and Conservative Transposition in Bacteria. Cell 1986, 47, 325–327. [Google Scholar] [CrossRef]
- Vandecraen, J.; Chandler, M.; Aertsen, A.; Van Houdt, R. The Impact of Insertion Sequences on Bacterial Genome Plasticity and Adaptability. Crit. Rev. Microbiol. 2017, 43, 709–730. [Google Scholar] [CrossRef]
- Lee, E.G.; Kim, K.H. Transposition of Transposable Element IS1 in Edwardsiella Piscicida Mutant Generated by CRISPR/Cas9 along with λ-Red Recombineering System. Res. Microbiol. 2025, 176, 104297. [Google Scholar] [CrossRef]
- Hallet, B.; Sherratt, D.J. Transposition and Site-Specific Recombination: Adapting DNA Cut-and-Paste Mechanisms to a Variety of Genetic Rearrangements. FEMS Microbiol. Rev. 2006, 21, 157–178. [Google Scholar] [CrossRef]
- Turlan, C.; Chandler, M. IS1-Mediated Intramolecular Rearrangements: Formation of Excised Transposon Circles and Replicative Deletions. EMBO J. 1995, 14, 5410–5421. [Google Scholar] [CrossRef] [PubMed]
- Prentki, P.; Pham, M.H.; Gamas, P.; Chandler, M.; Galas, D.J. Artificial Transposable Elements in the Study of the Ends of IS1. Gene 1987, 61, 91–101. [Google Scholar] [CrossRef]
- Machida, Y.; Machida, C.; Ohtsubo, E. Insertion Element IS1 Encodes Two Structural Genes Required for Its Transposition. J. Mol. Biol. 1984, 177, 229–245. [Google Scholar] [CrossRef] [PubMed]
- Ramirez-Santos, J.; Alvarez, G.; Cisneros, E.; Gomez-Eichelmann, M.C. Distribution of Insertion Sequence IS1 in Multiple-Antibiotic Resistant Clinical Enterobacteriaceae Strains. FEMS Microbiol. Lett. 1992, 72, 189–193. [Google Scholar] [CrossRef][Green Version]
- Ohta, S.; Tsuchida, K.; Choi, S.; Sekine, Y.; Shiga, Y.; Ohtsubo, E. Presence of a Characteristic D-D-E Motif in IS1 Transposase. J. Bacteriol. 2002, 184, 6146–6154. [Google Scholar] [CrossRef]
- Atkins, J.F.; Loughran, G.; Bhatt, P.R.; Firth, A.E.; Baranov, P.V. Ribosomal Frameshifting and Transcriptional Slippage: From Genetic Steganography and Cryptography to Adventitious Use. Nucleic 2016, 44, 7007–7078. [Google Scholar] [CrossRef]
- Vögele, K.; Schwartz, E.; Welz, C.; Schiltz, E.; Rak, B. High-Level Ribosomal Frameshifting Directs the Synthesis of IS150 Gene Products. Nucleic Acids Res. 1991, 19, 4377–4385. [Google Scholar] [CrossRef]
- Nagy, Z.; Chandler, M. Regulation of Transposition in Bacteria. Res. Microbiol. 2004, 155, 387–398. [Google Scholar] [CrossRef] [PubMed]
- Surette, M.G.; Lavoie, B.D.; Chaconas, G. Action at a Distance in Mu DNA Transposition: An Enhancer-like Element Is the Site of Action of Supercoiling Relief Activity by Integration Host Factor (IHF). EMBO J. 1989, 8, 3483–3489. [Google Scholar] [CrossRef] [PubMed]
- Allison, R.G.; Chaconas, G. Role of the A Protein-Binding Sites in the in Vitro Transposition of Mu DNA. A Complex Circuit of Interactions Involving the Mu Ends and the Transpositional Enhancer. J. Biol. Chem. 1992, 267, 19963–19970. [Google Scholar] [CrossRef] [PubMed]
- Shiga, Y.; Sekine, Y.; Kano, Y.; Ohtsubo, E. Involvement of H-NS in Transpositional Recombination Mediated by IS1. J. Bacteriol. 2001, 183, 2476–2484. [Google Scholar] [CrossRef]
- Sekine, Y.; Ohtsubo, E. Frameshifting Is Required for Production of the Transposase Encoded by Insertion Sequence 1. Proc. Natl. Acad. Sci. USA 1989, 86, 4609–4613. [Google Scholar] [CrossRef]
- Plasterk, R.H. Frameshift Control of IS1 Transposition. Trends Genet. 1991, 7, 203–204. [Google Scholar] [CrossRef]
- Caliskan, N.; Peske, F.; Rodnina, M.V. Changed in Translation: MRNA Recoding by -1 Programmed Ribosomal Frameshifting. Trends Biochem. Sci. 2015, 40, 265–274. [Google Scholar] [CrossRef]
- Harger, J.W.; Meskauskas, A.; Dinman, J.D. An “Integrated Model” of Programmed Ribosomal Frameshifting. Trends Biochem. Sci. 2002, 27, 448–454. [Google Scholar] [CrossRef]
- Zhou, H.; Zhang, Z.; Velo, J.; Huo, J.; Smith, S.; Ho, A.; Saier, M.H., Jr. Transcriptional Mechanism by Which IS5 Activates the FucAO Operon in Escherichia coli. Nucleic Acids Res. 2025, 53, gkaf172. [Google Scholar] [CrossRef]
- Sawers, R.G. Transcript Analysis of Escherichia coli K-12 Insertion Element IS5. FEMS Microbiol. Lett. 2005, 244, 397–401. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Machida, C.; Machida, Y.; Wang, H.C.; Ishizaki, K.; Ohtsubo, E. Repression of Cointegration Ability of Insertion Element IS1 by Transcriptional Readthrough from Flanking Regions. Cell 1983, 34, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.J.; Park, J.H.; Kim, K.S.; Chang, J.E.; Ko, J.H.; Kim, M.H.; Chung, D.H.; Chung, T.W.; Choe, I.S.; Lee, Y.C.; et al. Sequence Analysis of the 5′-Flanking Region of the Gene Encoding Human N-Acetylglucosaminyltransferase III. Gene 1996, 170, 281–283. [Google Scholar] [CrossRef]
- Chaconas, G. Studies on a “Jumping Gene Machine”: Higher-Order Nucleoprotein Complexes in Mu DNA Transposition. Biochem. Cell Biol. 1999, 77, 487–491. [Google Scholar] [CrossRef]
- Gueguen, E.; Rousseau, P.; Duval-Valentin, G.; Chandler, M. The Transpososome: Control of Transposition at the Level of Catalysis. Trends Microbiol. 2005, 13, 543–549. [Google Scholar] [CrossRef]
- Humayun, M.Z.; Zhang, Z.; Butcher, A.M.; Moshayedi, A.; Saier, M.H., Jr. Hopping into a Hot Seat: Role of DNA Structural Features on IS5-Mediated Gene Activation and Inactivation under Stress. PLoS ONE 2017, 12, e0180156. [Google Scholar] [CrossRef] [PubMed]
- Black, J.W.; Leff, P.; Shankley, N.P. Further Analysis of Anomalous PKB Values for Histamine H2-Receptor Antagonists on the Mouse Isolated Stomach Assay. Br. J. Pharmacol. 1985, 86, 581–587. [Google Scholar] [CrossRef] [PubMed]
- Saier, M.H., Jr.; Kukita, C.; Zhang, Z. Transposon-Mediated Directed Mutation in Bacteria and Eukaryotes. Front. Biosci. 2017, 22, 1458–1468. [Google Scholar] [CrossRef]
- Madan, R.; Kolter, R.; Mahadevan, S. Mutations That Activate the Silent Bgl Operon of Escherichia coli Confer a Growth Advantage in Stationary Phase. J. Bacteriol. 2005, 187, 7912–7917. [Google Scholar] [CrossRef]
- Wang, X.; Wood, T.K. IS5 Inserts Upstream of the Master Motility Operon FlhDC in a Quasi-Lamarckian Way. ISME J. 2011, 5, 1517–1525. [Google Scholar] [CrossRef]
- Zhang, Z.; Kukita, C.; Humayun, M.Z.; Saier, M.H. Environment-Directed Activation of the Escherichia coli flhDC Operon by Transposons. Microbiology 2017, 163, 554–569. [Google Scholar] [CrossRef]
- Barker, C.S.; Prüss, B.M.; Matsumura, P. Increased Motility of Escherichia coli by Insertion Sequence Element Integration into the Regulatory Region of the FlhD Operon. J. Bacteriol. 2004, 186, 7529–7537. [Google Scholar] [CrossRef]
- Rosner, J.L.; Guyer, M.S. Transposition of IS1-LambdaBIO-IS1 from a Bacteriophage Lambda Derivative Carrying the IS1-Cat-IS1 Transposon (Tn9). Mol. Gen. Genet. 1980, 178, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Couchoud, C.; Bertrand, X.; Valot, B.; Hocquet, D. Deciphering the Role of Insertion Sequences in the Evolution of Bacterial Epidemic Pathogens with PanISa Software. Microb. Genom. 2020, 6, e000356. [Google Scholar] [CrossRef] [PubMed]
- Datsenko, K.A.; Wanner, B.L. One-Step Inactivation of Chromosomal Genes in Escherichia coli K-12 Using PCR Products. Proc. Natl. Acad. Sci. USA 2000, 97, 6640–6645. [Google Scholar] [CrossRef] [PubMed]
- Thomason, L.C.; Costantino, N.; Court, D.L.E. Coli Genome Manipulation by P1 Transduction. Curr. Protoc. Mol. Biol. 2007, 79, 1–17. [Google Scholar] [CrossRef]
- Zhang, Z.; Huo, J.; Velo, J.; Zhou, H.; Flaherty, A.; Saier, M.H., Jr. Comprehensive Characterization of FucAO Operon Activation in Escherichia coli. Int. J. Mol. Sci. 2024, 25, 3946. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Saier, M.H., Jr. A Mechanism of Transposon-Mediated Directed Mutation. Mol. Microbiol. 2009, 74, 29–43. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zhou, K.; Tran, D.; Saier, M. Insertion Sequence (IS) Element-Mediated Activating Mutations of the Cryptic Aromatic β-Glucoside Utilization (BglGFB) Operon Are Promoted by the Anti-Terminator Protein (BglG) in Escherichia coli. Int. J. Mol. Sci. 2022, 23, 1505. [Google Scholar] [CrossRef] [PubMed]
- Cairns, J.; Foster, P.L. Adaptive Reversion of a Frameshift Mutation in Escherichia coli. Genetics 1991, 128, 695–701. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Smith, S.; Zhang, Z.; Ho, A.; Karnani, T.; Ord, J.; Saier, M.H., Jr. Examining the Roles of Genomic Context and Endogenous Regulatory Elements on IS1 Transposition Within the Escherichia coli Genome. Int. J. Mol. Sci. 2025, 26, 8375. https://doi.org/10.3390/ijms26178375
Smith S, Zhang Z, Ho A, Karnani T, Ord J, Saier MH Jr. Examining the Roles of Genomic Context and Endogenous Regulatory Elements on IS1 Transposition Within the Escherichia coli Genome. International Journal of Molecular Sciences. 2025; 26(17):8375. https://doi.org/10.3390/ijms26178375
Chicago/Turabian StyleSmith, Sofia, Zhongge Zhang, Allyson Ho, Tusha Karnani, Jack Ord, and Milton H. Saier, Jr. 2025. "Examining the Roles of Genomic Context and Endogenous Regulatory Elements on IS1 Transposition Within the Escherichia coli Genome" International Journal of Molecular Sciences 26, no. 17: 8375. https://doi.org/10.3390/ijms26178375
APA StyleSmith, S., Zhang, Z., Ho, A., Karnani, T., Ord, J., & Saier, M. H., Jr. (2025). Examining the Roles of Genomic Context and Endogenous Regulatory Elements on IS1 Transposition Within the Escherichia coli Genome. International Journal of Molecular Sciences, 26(17), 8375. https://doi.org/10.3390/ijms26178375







