Pan-Genome Analysis of Cannabis sativa: Insights on Genomic Diversity, Evolution, and Environment Adaption
Abstract
1. Introduction
2. Results
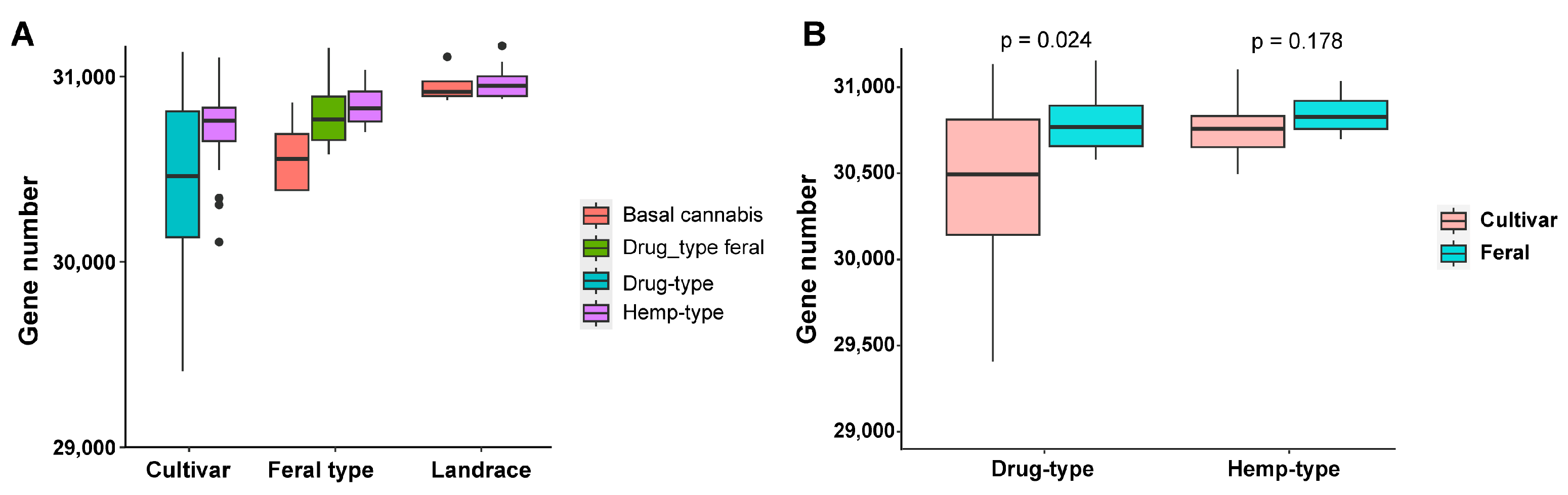
2.1. Assembly of Cannabis Pan-Genome and Gene Prediction
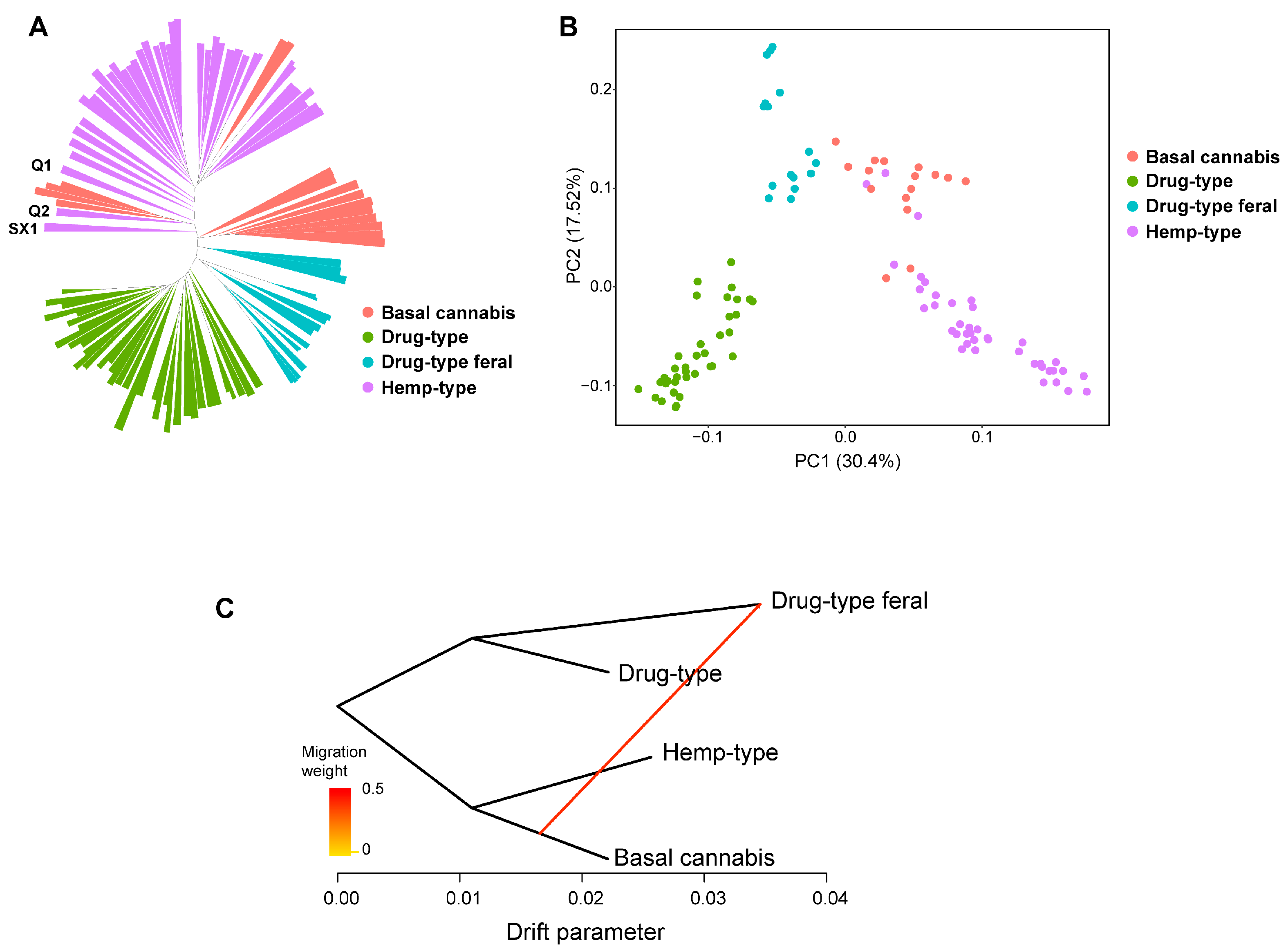
2.2. Population Genetic Analysis Based on Pan-Genome
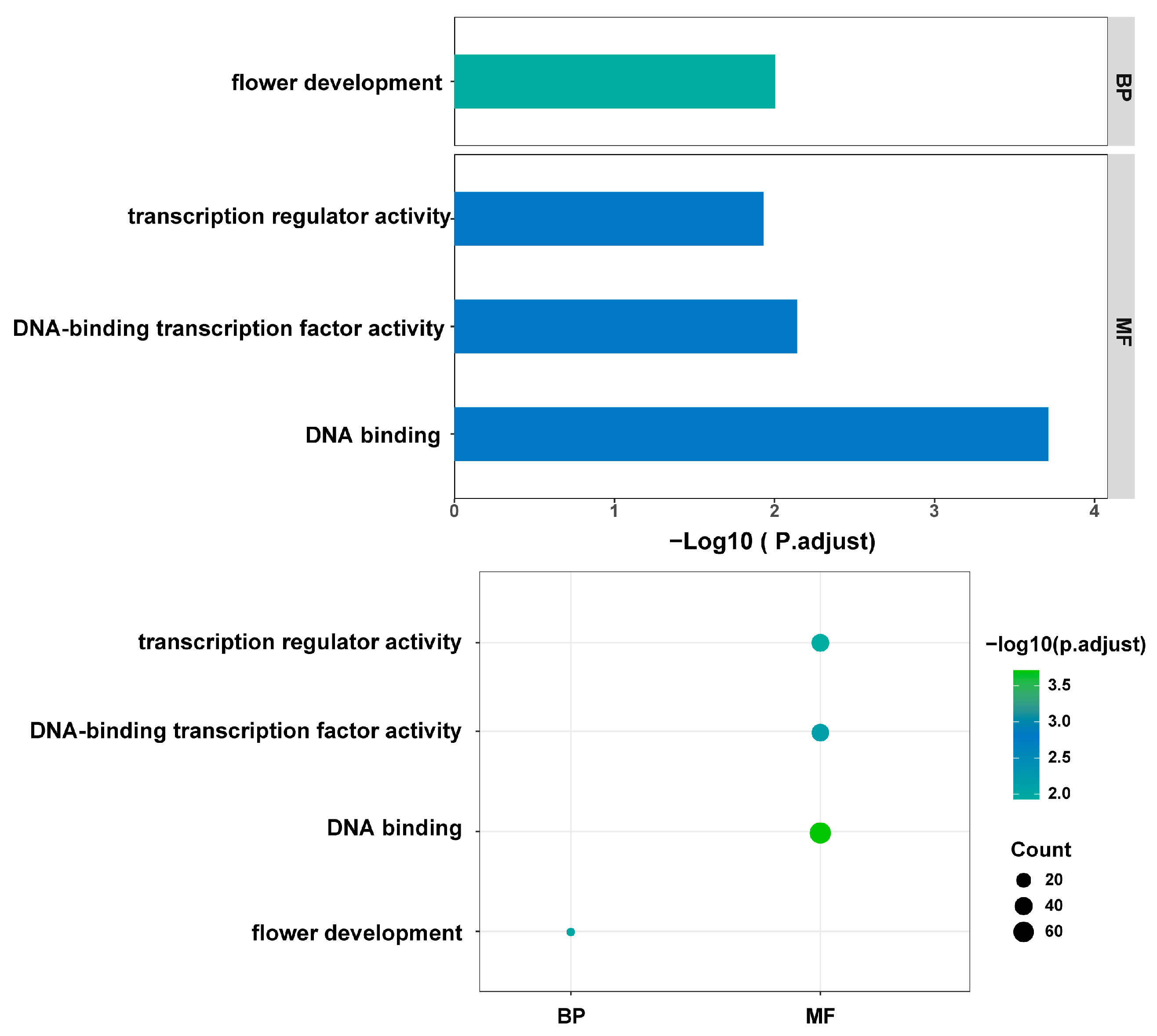
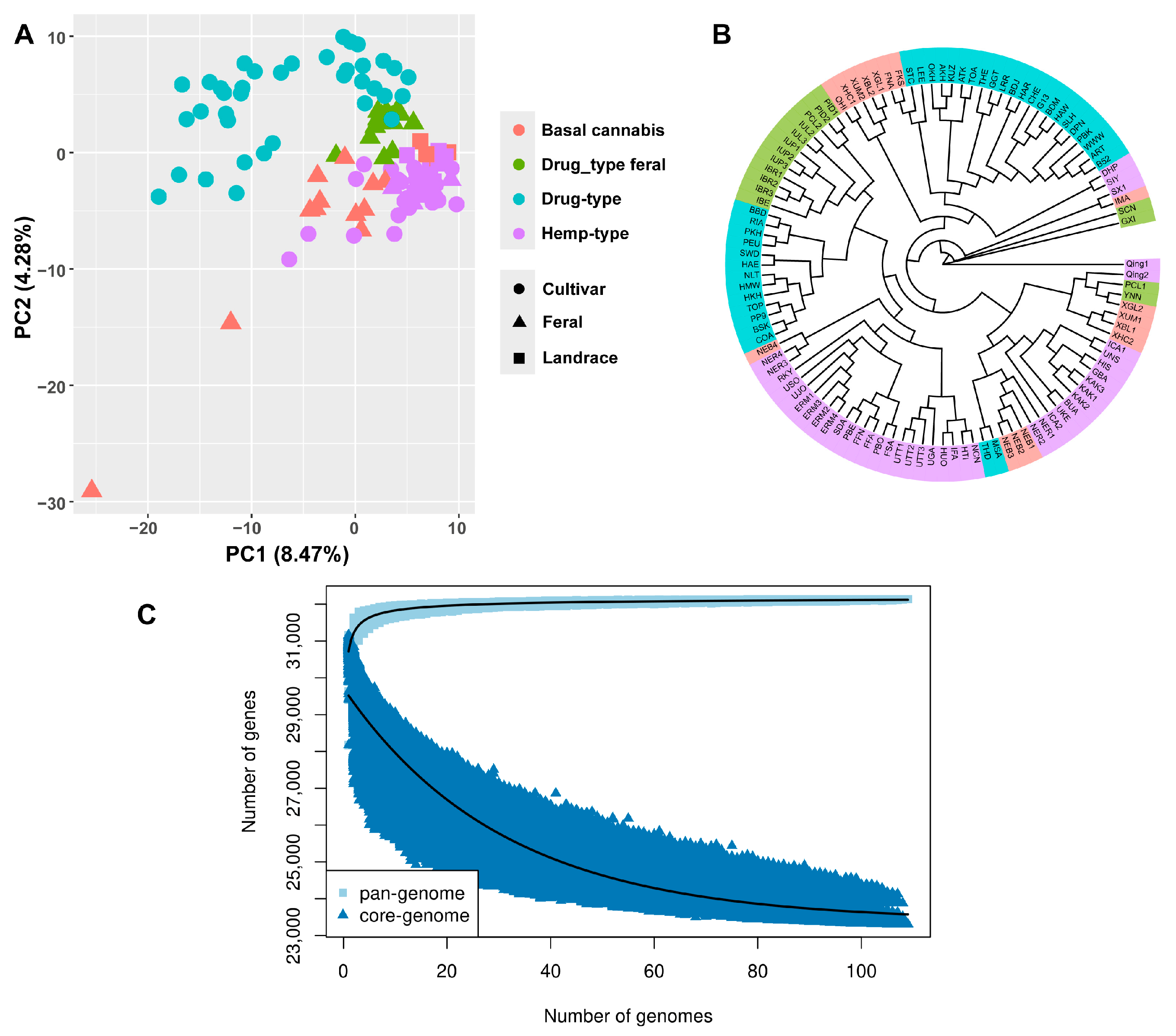
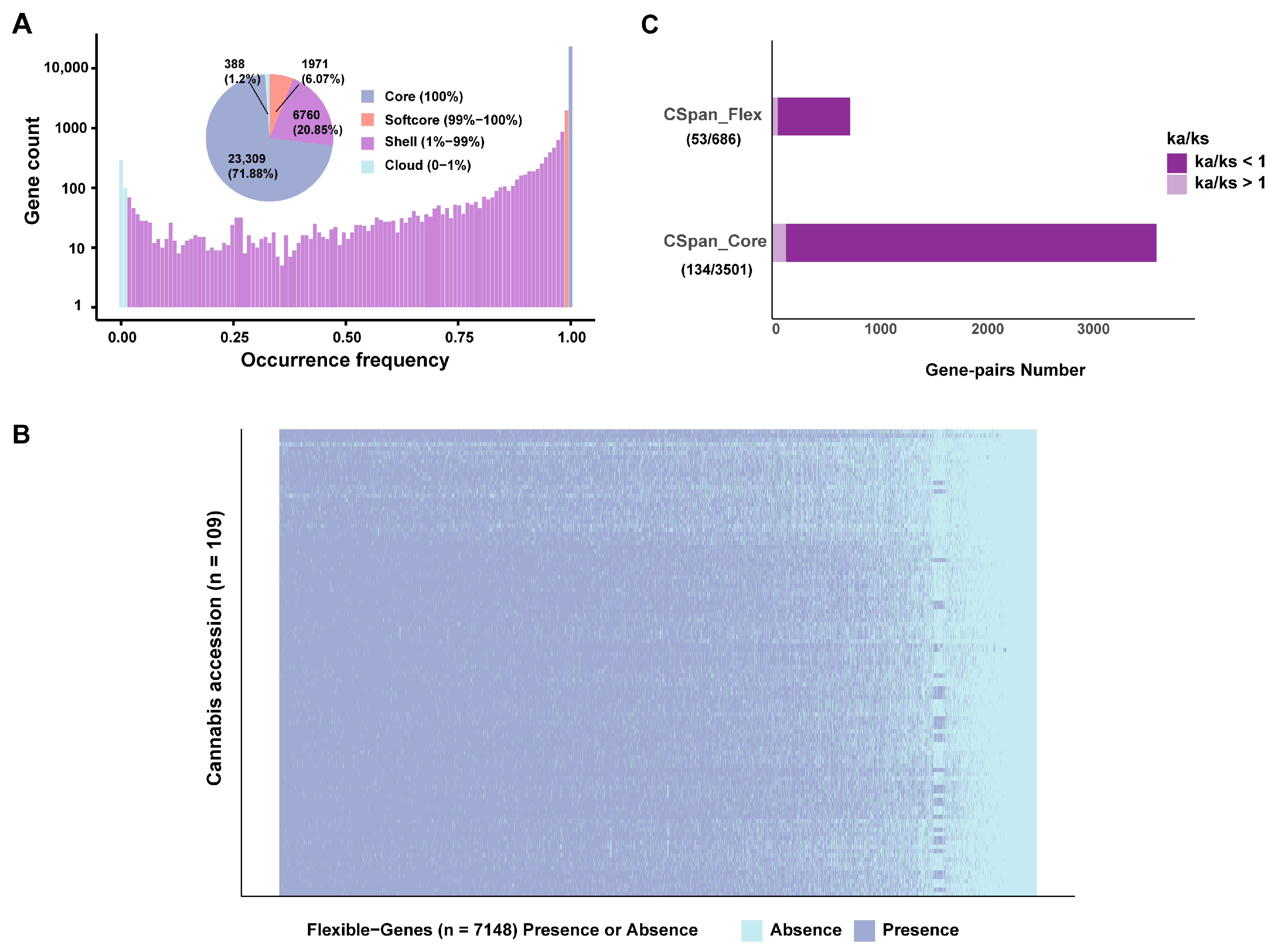
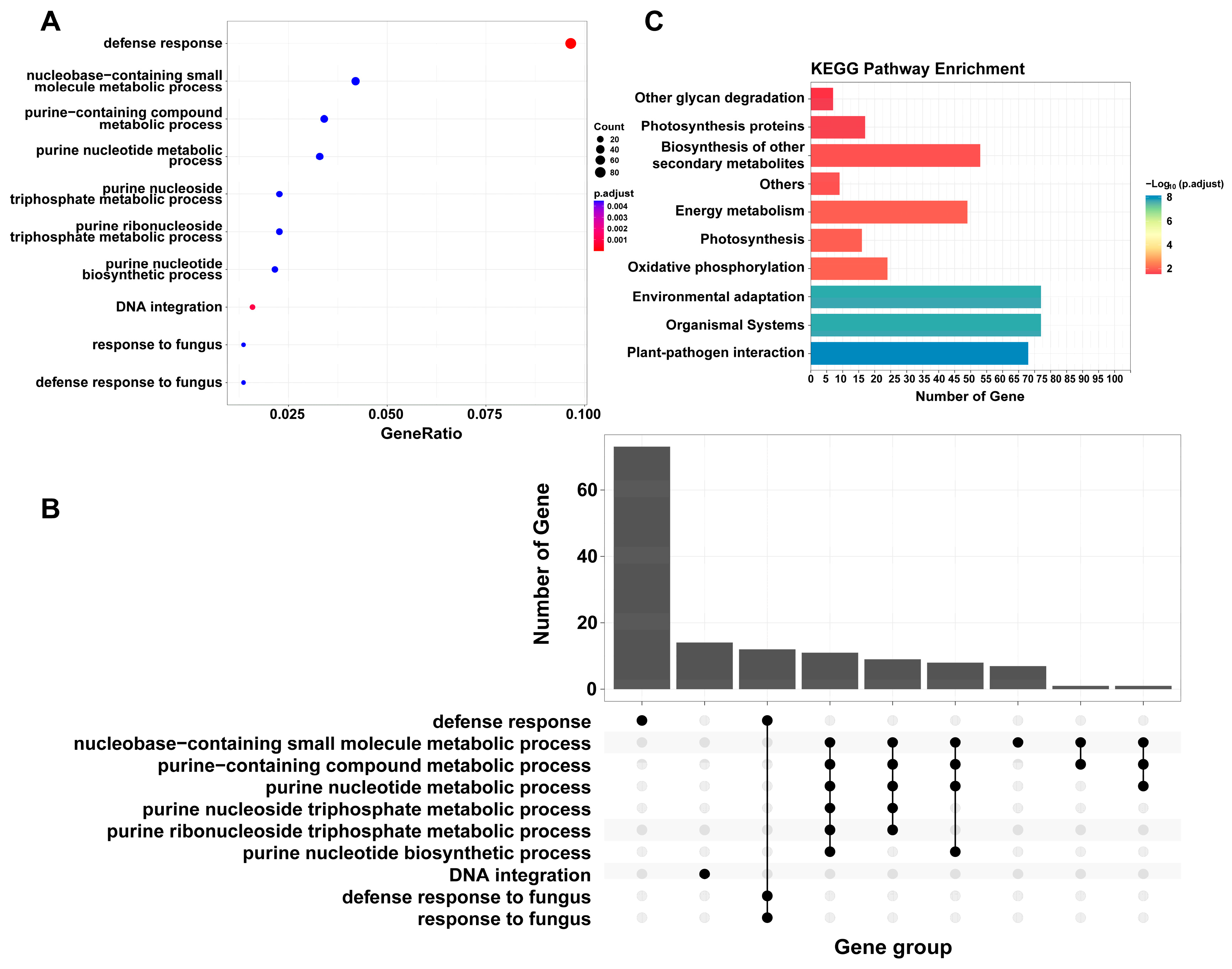
2.3. PAV Analysis of Cannabis Pan-Genome
3. Discussion
4. Materials and Methods
4.1. Construction of Cannabis Pan-Genome
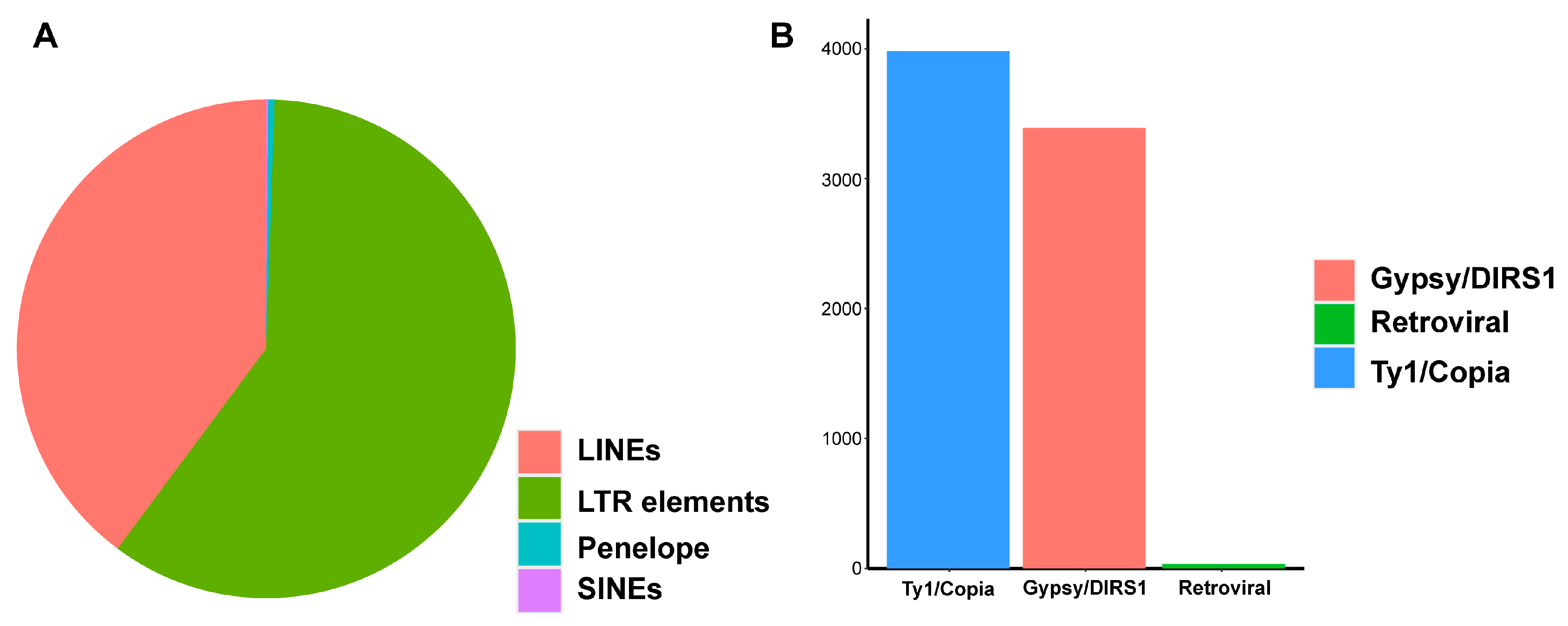
4.2. Repetitive Sequence Masking of Non-Reference Sequences
4.3. Gene Prediction and Annotation of Cannabis Pan-Genome
4.4. Population Genetic Analysis Based on Cannabis Pan-Genome
4.5. PAV Analysis Based on Cannabis Pan-Genome
4.6. Assessment of the Pan-Genome of Cannabis
4.7. Statistics on the Number and Frequency of Genes in the Cannabis Pan-Genome
4.8. Principal Component Analysis and Phylogenetic Analysis of Cannabis Pan-Genome PAVs
4.9. Analysis of Flexible Genes and Core Genes
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Salentijn, E.M.J.; Zhang, Q.; Amaducci, S.; Yang, M.; Trindade, L.M. New developments in fiber hemp (Cannabis sativa L.) breeding. Ind. Crops Prod. 2015, 68, 32–41. [Google Scholar] [CrossRef]
- Li, H.-L. The origin and use of cannabis in eastern asia linguistic-cultural implications. Econ. Bot. 1974, 28, 293–301. [Google Scholar] [CrossRef]
- Ren, M.; Tang, Z.; Wu, X.; Spengler, R.; Jiang, H.; Yang, Y.; Boivin, N. The origins of cannabis smoking: Chemical residue evidence from the first millennium BCE in the Pamirs. Sci. Adv. 2019, 5, eaaw1391. [Google Scholar] [CrossRef] [PubMed]
- Hofmann-Aßmus, M. THC/CBD wirkt besser als THC allein. MMW—Fortschritte Med. 2020, 162, 37. [Google Scholar] [CrossRef]
- Daris, B.; Tancer Verboten, M.; Knez, Z.; Ferk, P. Cannabinoids in cancer treatment: Therapeutic potential and legislation. Bosn. J. Basic. Med. Sci. 2019, 19, 14–23. [Google Scholar] [CrossRef]
- Kuzumi, A.; Yoshizaki-Ogawa, A.; Fukasawa, T.; Sato, S.; Yoshizaki, A. The Potential Role of Cannabidiol in Cosmetic Dermatology: A Literature Review. Am. J. Clin. Dermatol. 2024, 25, 951–966. [Google Scholar] [CrossRef]
- van Bakel, H.; Stout, J.M.; Cote, A.G.; Tallon, C.M.; Sharpe, A.G.; Hughes, T.R.; Page, J.E. The draft genome and transcriptome of Cannabis sativa. Genome Biol. 2011, 12, R102. [Google Scholar] [CrossRef]
- Grassa, C.J.; Weiblen, G.D.; Wenger, J.P.; Dabney, C.; Poplawski, S.G.; Timothy Motley, S.; Michael, T.P.; Schwartz, C.J. A new Cannabis genome assembly associates elevated cannabidiol (CBD) with hemp introgressed into marijuana. New Phytol. 2021, 230, 1665–1679. [Google Scholar] [CrossRef]
- Ren, G.; Zhang, X.; Li, Y.; Ridout, K.; Serrano-Serrano, M.L.; Yang, Y.; Liu, A.; Ravikanth, G.; Nawaz, M.A.; Mumtaz, A.S.; et al. Large-scale whole-genome resequencing unravels the domestication history of Cannabis sativa. Sci. Adv. 2021, 7, eabg2286. [Google Scholar] [CrossRef]
- Bian, P.P.; Zhang, Y.; Jiang, Y. Pan-genome: Setting a new standard for high-quality reference genomes. Yi Chuan 2021, 43, 1023–1037. [Google Scholar] [CrossRef]
- Song, J.M.; Guan, Z.; Hu, J.; Guo, C.; Yang, Z.; Wang, S.; Liu, D.; Wang, B.; Lu, S.; Zhou, R.; et al. Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat. Plants 2020, 6, 34–45. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Xue, H.; Dong, X.; Li, M.; Zheng, X.; Li, Z.; Xu, J.; Wang, W.; Wei, C. Long-read sequencing of 111 rice genomes reveals significantly larger pan-genomes. Genome Res. 2022, 32, 853–863. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Mauleon, R.; Hu, Z.; Chebotarov, D.; Tai, S.; Wu, Z.; Li, M.; Zheng, T.; Fuentes, R.R.; Zhang, F.; et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 2018, 557, 43–49. [Google Scholar] [CrossRef]
- Hubner, S.; Bercovich, N.; Todesco, M.; Mandel, J.R.; Odenheimer, J.; Ziegler, E.; Lee, J.S.; Baute, G.J.; Owens, G.L.; Grassa, C.J.; et al. Sunflower pan-genome analysis shows that hybridization altered gene content and disease resistance. Nat. Plants 2019, 5, 54–62. [Google Scholar] [CrossRef]
- Liu, Y.; Du, H.; Li, P.; Shen, Y.; Peng, H.; Liu, S.; Zhou, G.A.; Zhang, H.; Liu, Z.; Shi, M.; et al. Pan-Genome of Wild and Cultivated Soybeans. Cell 2020, 182, 162–176.e13. [Google Scholar] [CrossRef]
- Cai, X.; Chang, L.; Zhang, T.; Chen, H.; Zhang, L.; Lin, R.; Liang, J.; Wu, J.; Freeling, M.; Wang, X. Impacts of allopolyploidization and structural variation on intraspecific diversification in Brassica rapa. Genome Biol. 2021, 22, 166. [Google Scholar] [CrossRef]
- Qin, P.; Lu, H.; Du, H.; Wang, H.; Chen, W.; Chen, Z.; He, Q.; Ou, S.; Zhang, H.; Li, X.; et al. Pan-genome analysis of 33 genetically diverse rice accessions reveals hidden genomic variations. Cell 2021, 184, 3542–3558. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Yuan, D.; Wang, P.; Wang, Q.; Sun, M.; Liu, Z.; Si, H.; Xu, Z.; Ma, Y.; Zhang, B.; et al. Cotton pan-genome retrieves the lost sequences and genes during domestication and selection. Genome Biol. 2021, 22, 119. [Google Scholar] [CrossRef]
- Dolatabadian, A.; Bayer, P.E.; Tirnaz, S.; Hurgobin, B.; Edwards, D.; Batley, J. Characterization of disease resistance genes in the Brassica napus pangenome reveals significant structural variation. Plant Biotechnol. J. 2020, 18, 969–982. [Google Scholar] [CrossRef]
- Benedetti, E.; Resce, G.; Brunori, P.; Molinaro, S. Cannabis Policy Changes and Adolescent Cannabis Use: Evidence from Europe. Int. J. Environ. Res. Public Health 2021, 18, 5174. [Google Scholar] [CrossRef]
- Startek, M.; Szafranski, P.; Gambin, T.; Campbell, I.M.; Hixson, P.; Shaw, C.A.; Stankiewicz, P.; Gambin, A. Genome-wide analyses of LINE-LINE-mediated nonallelic homologous recombination. Nucleic Acids Res. 2015, 43, 2188–2198. [Google Scholar] [CrossRef]
- Dittwald, P.; Gambin, T.; Szafranski, P.; Li, J.; Amato, S.; Divon, M.Y.; Rodriguez Rojas, L.X.; Elton, L.E.; Scott, D.A.; Schaaf, C.P.; et al. NAHR-mediated copy-number variants in a clinical population: Mechanistic insights into both genomic disorders and Mendelizing traits. Genome Res. 2013, 23, 1395–1409. [Google Scholar] [CrossRef]
- Stankiewicz, P.; Lupski, J.R. Genome architecture, rearrangements and genomic disorders. Trends Genet. 2002, 18, 74–82. [Google Scholar] [CrossRef]
- Sun, X.; Jiao, C.; Schwaninger, H.; Chao, C.T.; Ma, Y.; Duan, N.; Khan, A.; Ban, S.; Xu, K.; Cheng, L.; et al. Phased diploid genome assemblies and pan-genomes provide insights into the genetic history of apple domestication. Nat. Genet. 2020, 52, 1423–1432. [Google Scholar] [CrossRef]
- Sun, X.; Zhang, Z.; Wu, J.; Cui, X.; Feng, D.; Wang, K.; Xu, M.; Zhou, L.; Han, X.; Gu, X.; et al. The Oryza sativa Regulator HDR1 Associates with the Kinase OsK4 to Control Photoperiodic Flowering. PLoS Genet. 2016, 12, e1005927. [Google Scholar] [CrossRef] [PubMed]
- Sun, A.; Yu, B.; Zhang, Q.; Peng, Y.; Yang, J.; Sun, Y.; Qin, P.; Jia, T.; Smeekens, S.; Teng, S. MYC2-Activated TRICHOME BIREFRINGENCE-LIKE37 Acetylates Cell Walls and Enhances Herbivore Resistance. Plant Physiol. 2020, 184, 1083–1096. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Chen, S.; Zhou, Y.; Shen, Y.; Qin, Z.; Wu, L. VERNALIZATION1 represses FLOWERING PROMOTING FACTOR1-LIKE1 in leaves for timely flowering in Brachypodium distachyon. Plant Cell 2023, 35, 3697–3711. [Google Scholar] [CrossRef] [PubMed]
- Taurino, M.; Costantini, S.; De Domenico, S.; Stefanelli, F.; Ruano, G.; Delgadillo, M.O.; Sanchez-Serrano, J.J.; Sanmartin, M.; Santino, A.; Rojo, E. SEIPIN Proteins Mediate Lipid Droplet Biogenesis to Promote Pollen Transmission and Reduce Seed Dormancy. Plant Physiol. 2018, 176, 1531–1546. [Google Scholar] [CrossRef]
- Wang, X.; Chai, X.; Gao, B.; Deng, C.; Gunther, C.S.; Wu, T.; Zhang, X.; Xu, X.; Han, Z.; Wang, Y. Multi-omics analysis reveals the mechanism of bHLH130 responding to low-nitrogen stress of apple rootstock. Plant Physiol. 2023, 191, 1305–1323. [Google Scholar] [CrossRef]
- Weingartner, M.; Subert, C.; Sauer, N. LATE, a C(2)H(2) zinc-finger protein that acts as floral repressor. Plant J. 2011, 68, 681–692. [Google Scholar] [CrossRef]
- Cheng, Q.; Tong, Y.; Wang, Z.; Su, P.; Gao, W.; Huang, L. Molecular cloning and functional identification of a cDNA encoding 4-hydroxy-3-methylbut-2-enyl diphosphate reductase from Tripterygium wilfordii. Acta Pharm. Sin. B 2017, 7, 208–214. [Google Scholar] [CrossRef]
- Huang, X.; Huang, S.; Han, B.; Li, J. The integrated genomics of crop domestication and breeding. Cell 2022, 185, 2828–2839. [Google Scholar] [CrossRef]
- Ingvardsen, C.R.; Brinch-Pedersen, H. Challenges and potentials of new breeding techniques in Cannabis sativa. Front. Plant Sci. 2023, 14, 1154332. [Google Scholar] [CrossRef]
- Barcaccia, G.; Palumbo, F.; Scariolo, F.; Vannozzi, A.; Borin, M.; Bona, S. Potentials and Challenges of Genomics for Breeding Cannabis Cultivars. Front. Plant Sci. 2020, 11, 573299. [Google Scholar] [CrossRef]
- Li, Y.H.; Zhou, G.; Ma, J.; Jiang, W.; Jin, L.G.; Zhang, Z.; Guo, Y.; Zhang, J.; Sui, Y.; Zheng, L.; et al. De novo assembly of soybean wild relatives for pan-genome analysis of diversity and agronomic traits. Nat. Biotechnol. 2014, 32, 1045–1052. [Google Scholar] [CrossRef]
- Gao, L.; Gonda, I.; Sun, H.; Ma, Q.; Bao, K.; Tieman, D.M.; Burzynski-Chang, E.A.; Fish, T.L.; Stromberg, K.A.; Sacks, G.L.; et al. The tomato pan-genome uncovers new genes and a rare allele regulating fruit flavor. Nat. Genet. 2019, 51, 1044–1051. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Li, H. New strategies to improve minimap2 alignment accuracy. Bioinformatics 2021, 37, 4572–4574. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Li, D.; Liu, C.M.; Luo, R.; Sadakane, K.; Lam, T.W. MEGAHIT: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 2015, 31, 1674–1676. [Google Scholar] [CrossRef]
- Steinegger, M.; Soding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 2017, 35, 1026–1028. [Google Scholar] [CrossRef]
- Quinlan, A.R.; Hall, I.M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef]
- Vergara, D.; White, K.H.; Keepers, K.G.; Kane, N.C. The complete chloroplast genomes of Cannabis sativa and Humulus lupulus. Mitochondrial DNA A DNA Mapp. Seq. Anal. 2016, 27, 3793–3794. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Madan, A. CAP3: A DNA sequence assembly program. Genome Res. 1999, 9, 868–877. [Google Scholar] [CrossRef]
- Flynn, J.M.; Hubley, R.; Goubert, C.; Rosen, J.; Clark, A.G.; Feschotte, C.; Smit, A.F. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA 2020, 117, 9451–9457. [Google Scholar] [CrossRef]
- Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinform. 2004, 5, 4.10.1–4.10.14. [Google Scholar] [CrossRef] [PubMed]
- Cantarel, B.L.; Korf, I.; Robb, S.M.; Parra, G.; Ross, E.; Moore, B.; Holt, C.; Sanchez Alvarado, A.; Yandell, M. MAKER: An easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 2008, 18, 188–196. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef]
- Aramaki, T.; Blanc-Mathieu, R.; Endo, H.; Ohkubo, K.; Kanehisa, M.; Goto, S.; Ogata, H. KofamKOALA: KEGG Ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics 2020, 36, 2251–2252. [Google Scholar] [CrossRef]
- Tarasov, A.; Vilella, A.J.; Cuppen, E.; Nijman, I.J.; Prins, P. Sambamba: Fast processing of NGS alignment formats. Bioinformatics 2015, 31, 2032–2034. [Google Scholar] [CrossRef] [PubMed]
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve years of SAMtools and BCFtools. Gigascience 2021, 10, 1–4. [Google Scholar] [CrossRef]
- Lefort, V.; Desper, R.; Gascuel, O. FastME 2.0: A Comprehensive, Accurate, and Fast Distance-Based Phylogeny Inference Program. Mol. Biol. Evol. 2015, 32, 2798–2800. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
- Pickrell, J.K.; Pritchard, J.K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 2012, 8, e1002967. [Google Scholar] [CrossRef]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Duan, Z.; Qiao, Y.; Lu, J.; Lu, H.; Zhang, W.; Yan, F.; Sun, C.; Hu, Z.; Zhang, Z.; Li, G.; et al. HUPAN: A pan-genome analysis pipeline for human genomes. Genome Biol. 2019, 20, 149. [Google Scholar] [CrossRef]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef] [PubMed]
- Zwaenepoel, A.; Van de Peer, Y. wgd-simple command line tools for the analysis of ancient whole-genome duplications. Bioinformatics 2019, 35, 2153–2155. [Google Scholar] [CrossRef] [PubMed]
- Emms, D.M.; Kelly, S. OrthoFinder: Phylogenetic orthology inference for comparative genomics. Genome Biol. 2019, 20, 238. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 2012, 16, 284–287. [Google Scholar] [CrossRef] [PubMed]
| Number | Length (bp) | Proportion | |
|---|---|---|---|
| Retroelement | 12,772 | 8,080,389 | 32.74% |
| DNA transposon | 2138 | 783,571 | 3.18% |
| Rolling circles | 272 | 81,427 | 0.33% |
| Unclassified | 14,822 | 3,105,612 | 12.58% |
| Small RNA | 73 | 28,829 | 0.12% |
| Satellites | 86 | 16,393 | 0.07% |
| Simple repeats | 8912 | 383,968 | 1.56% |
| Low complexity | 2248 | 120,709 | 0.49% |
| Reference Genome | Non-Reference Sequences | Non-Reference Sequences (The Length of the Encoded Protein > 100 aa) | |
|---|---|---|---|
| Gene number | 31,170 | 1919 | 1313 |
| mRNA number | 33,639 | 1919 | 1313 |
| Exon number | 234,131 | 3619 | 2858 |
| CDS number | 190,424 | 3592 | 2840 |
| CDS length (bp) | 46,210,977 | 1,143,501 | 1,006,458 |
| The Number of Sequences Annotated | Proportion of All Sequences | |
|---|---|---|
| Swiss-Prot | 26,868 | 75.49% |
| Trembl | 35,168 | 98.81% |
| Uniref50 | 35,275 | 99.11% |
| NCBI_nr | 35,198 | 98.89% |
| Interproscan | 33,345 | 93.68% |
| GO | 20,953 | 58.87% |
| KEGG | 13,865 | 38.95% |
| Uniprot and GOA | 33,402 | 93.84% |
| NCBI_nr and Uniprot | 35,294 | 99.16% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, S.; Zhong, X.; Cheng, Y.; Yu, Y.; Wan, J.; Liu, Q.; Shu, Y.; Wu, X.; Li, Y. Pan-Genome Analysis of Cannabis sativa: Insights on Genomic Diversity, Evolution, and Environment Adaption. Int. J. Mol. Sci. 2025, 26, 8354. https://doi.org/10.3390/ijms26178354
Wang S, Zhong X, Cheng Y, Yu Y, Wan J, Liu Q, Shu Y, Wu X, Li Y. Pan-Genome Analysis of Cannabis sativa: Insights on Genomic Diversity, Evolution, and Environment Adaption. International Journal of Molecular Sciences. 2025; 26(17):8354. https://doi.org/10.3390/ijms26178354
Chicago/Turabian StyleWang, Shuyu, Xue Zhong, Yuhui Cheng, Ying Yu, Jifeng Wan, Qingqing Liu, Yongjun Shu, Xiuju Wu, and Yong Li. 2025. "Pan-Genome Analysis of Cannabis sativa: Insights on Genomic Diversity, Evolution, and Environment Adaption" International Journal of Molecular Sciences 26, no. 17: 8354. https://doi.org/10.3390/ijms26178354
APA StyleWang, S., Zhong, X., Cheng, Y., Yu, Y., Wan, J., Liu, Q., Shu, Y., Wu, X., & Li, Y. (2025). Pan-Genome Analysis of Cannabis sativa: Insights on Genomic Diversity, Evolution, and Environment Adaption. International Journal of Molecular Sciences, 26(17), 8354. https://doi.org/10.3390/ijms26178354







