New Insights into the Role of Secondary Metabolic Pathways in Resistance of Potato to Dickeya solani
Abstract
1. Introduction
2. Results
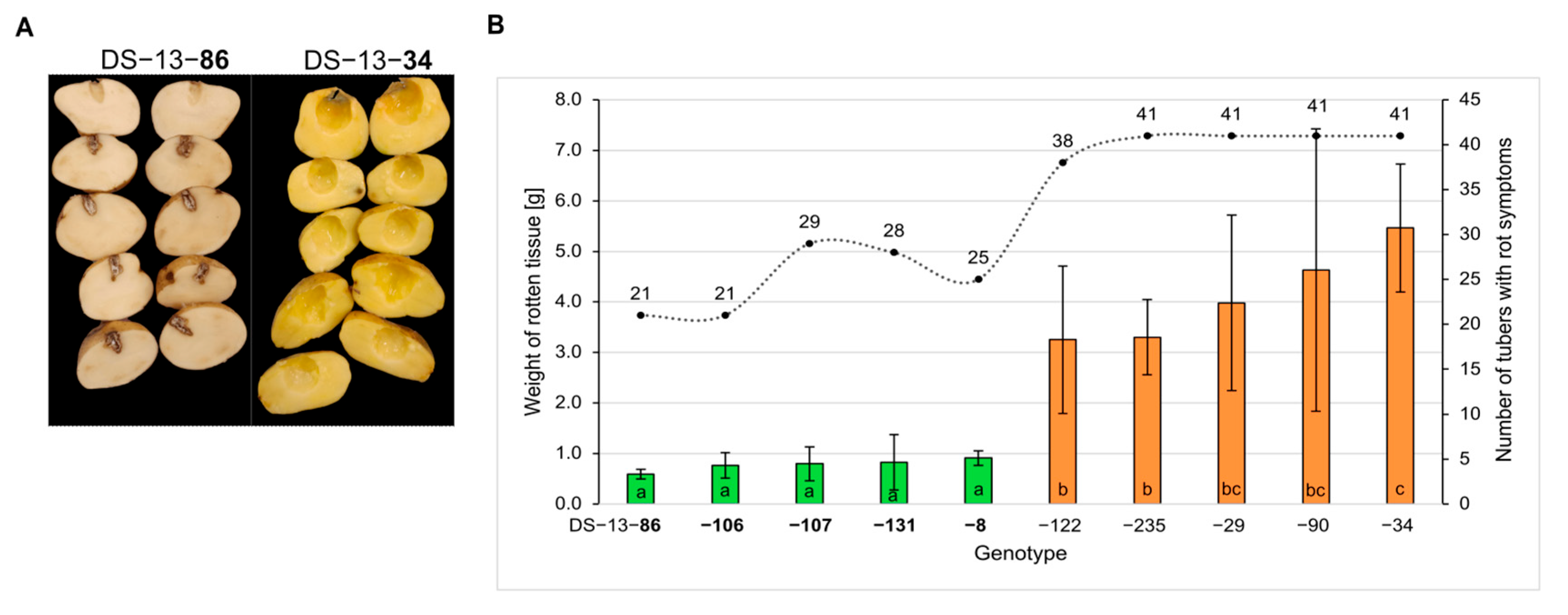
2.1. Phenotype of Selected Genotypes
2.2. Sequencing Data
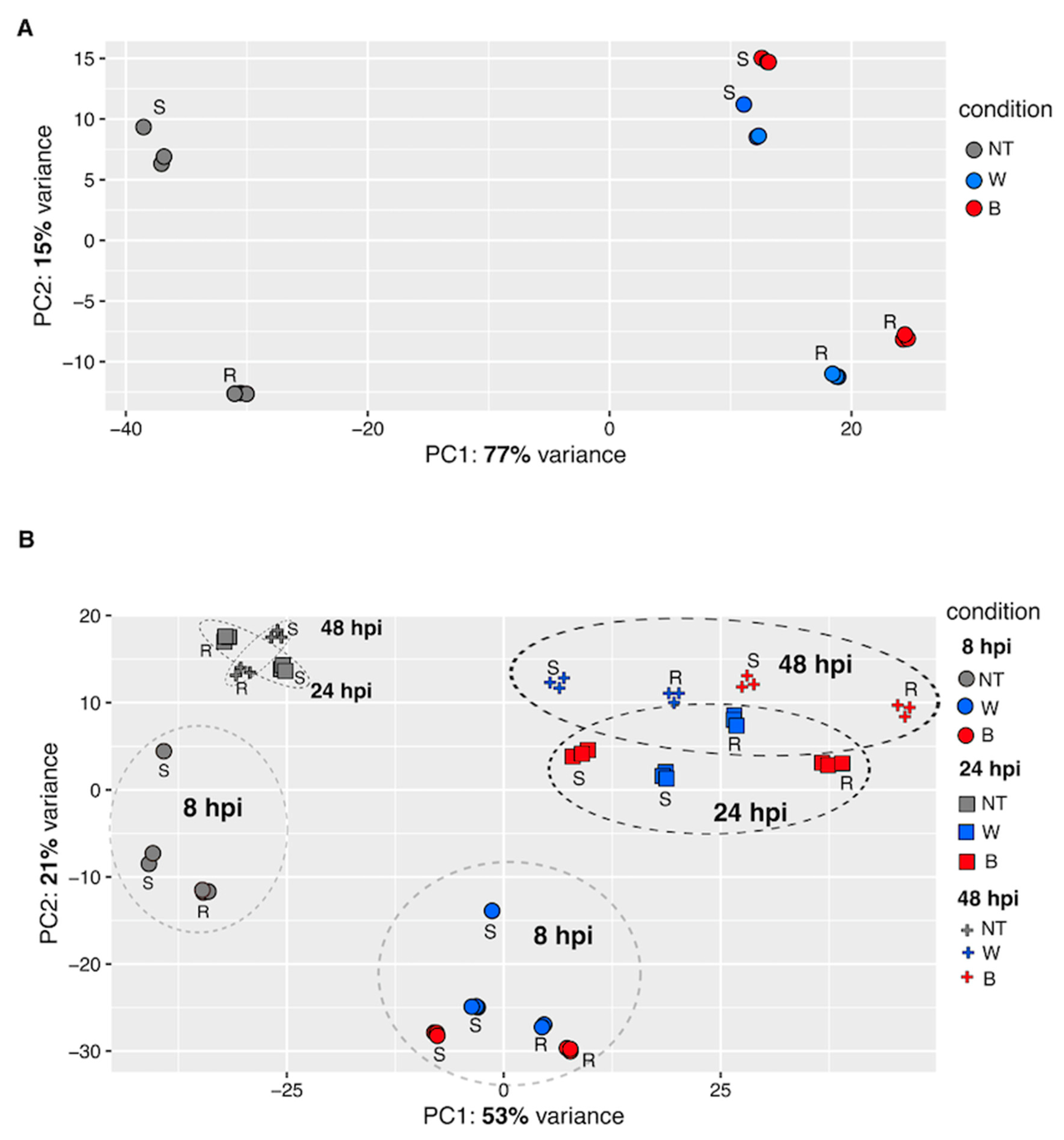
2.3. Principal Component Analysis (PCA)
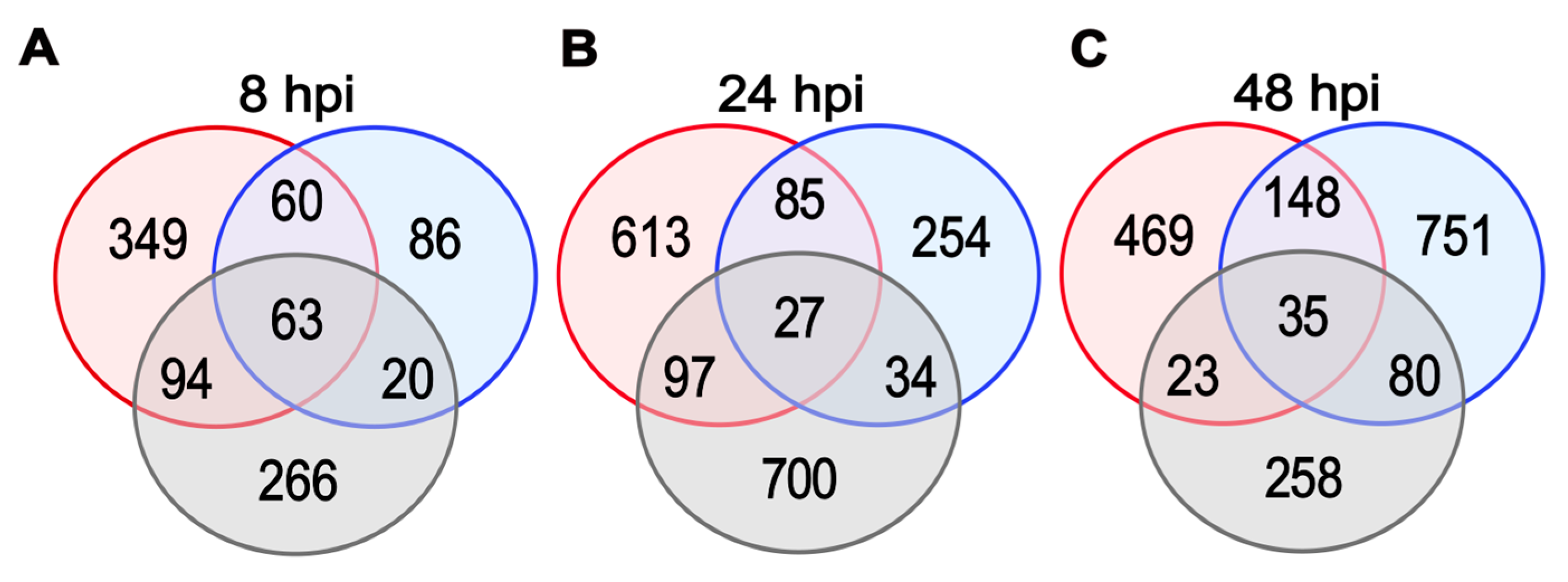
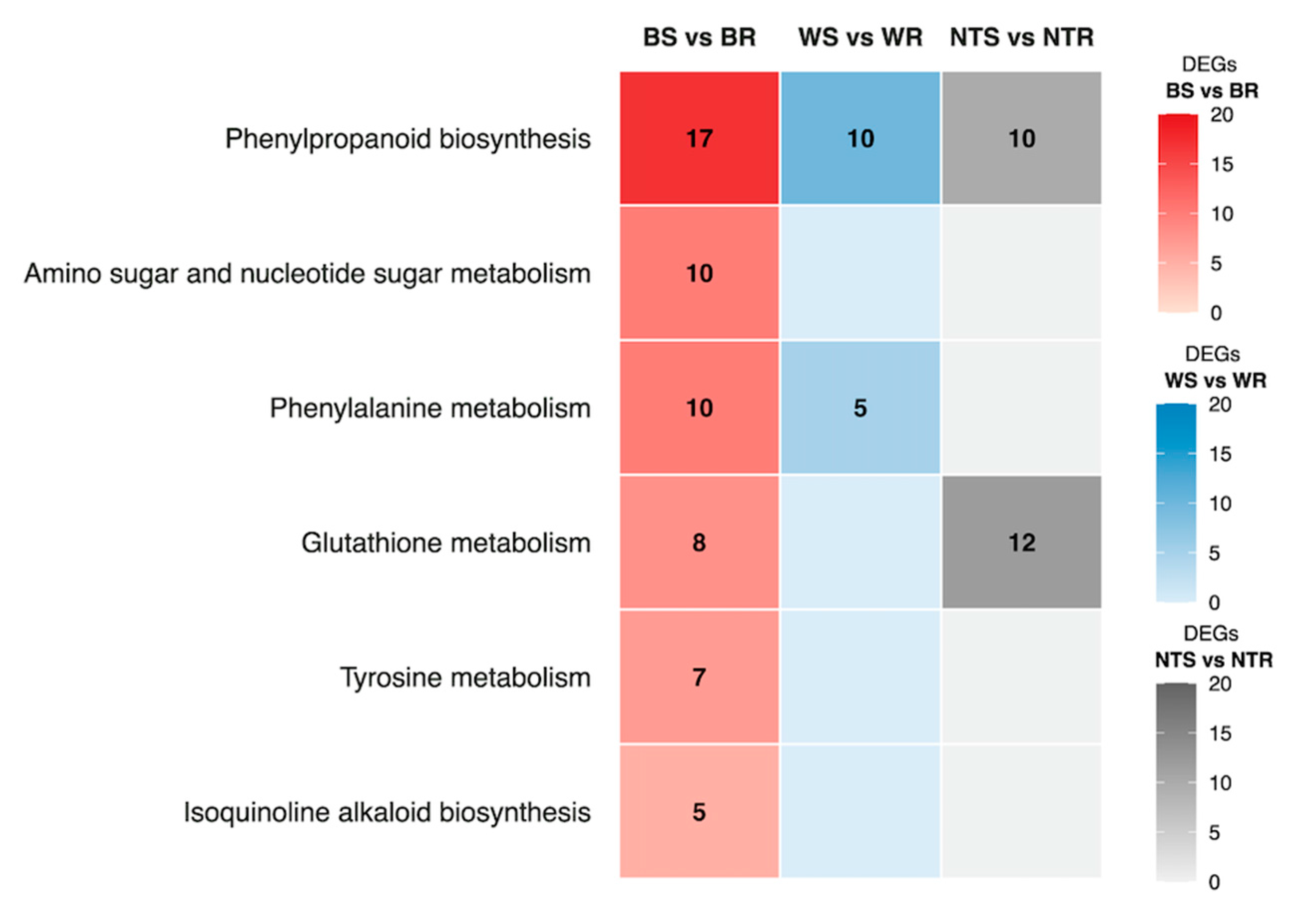
2.4. Detection of DEGs
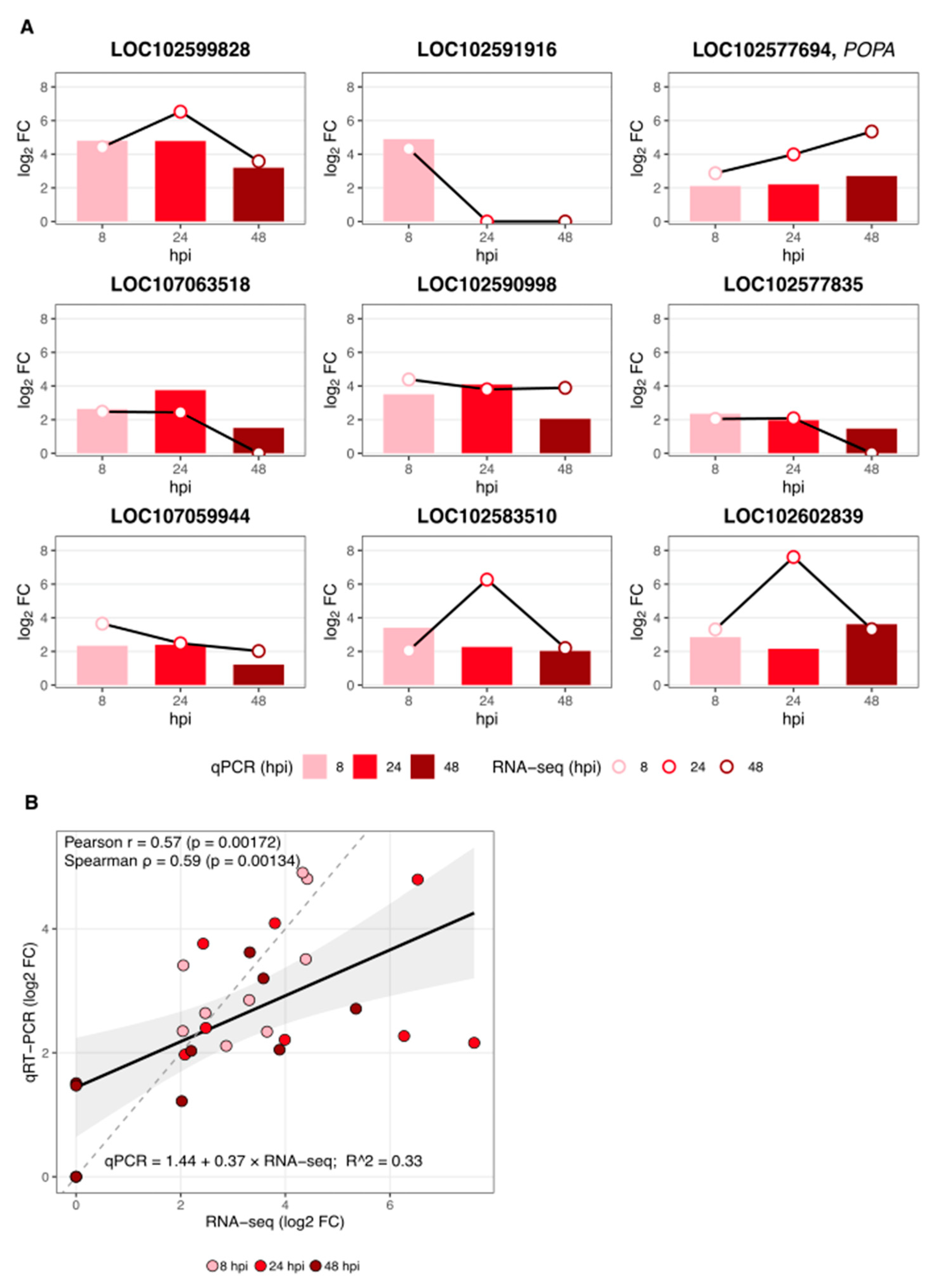
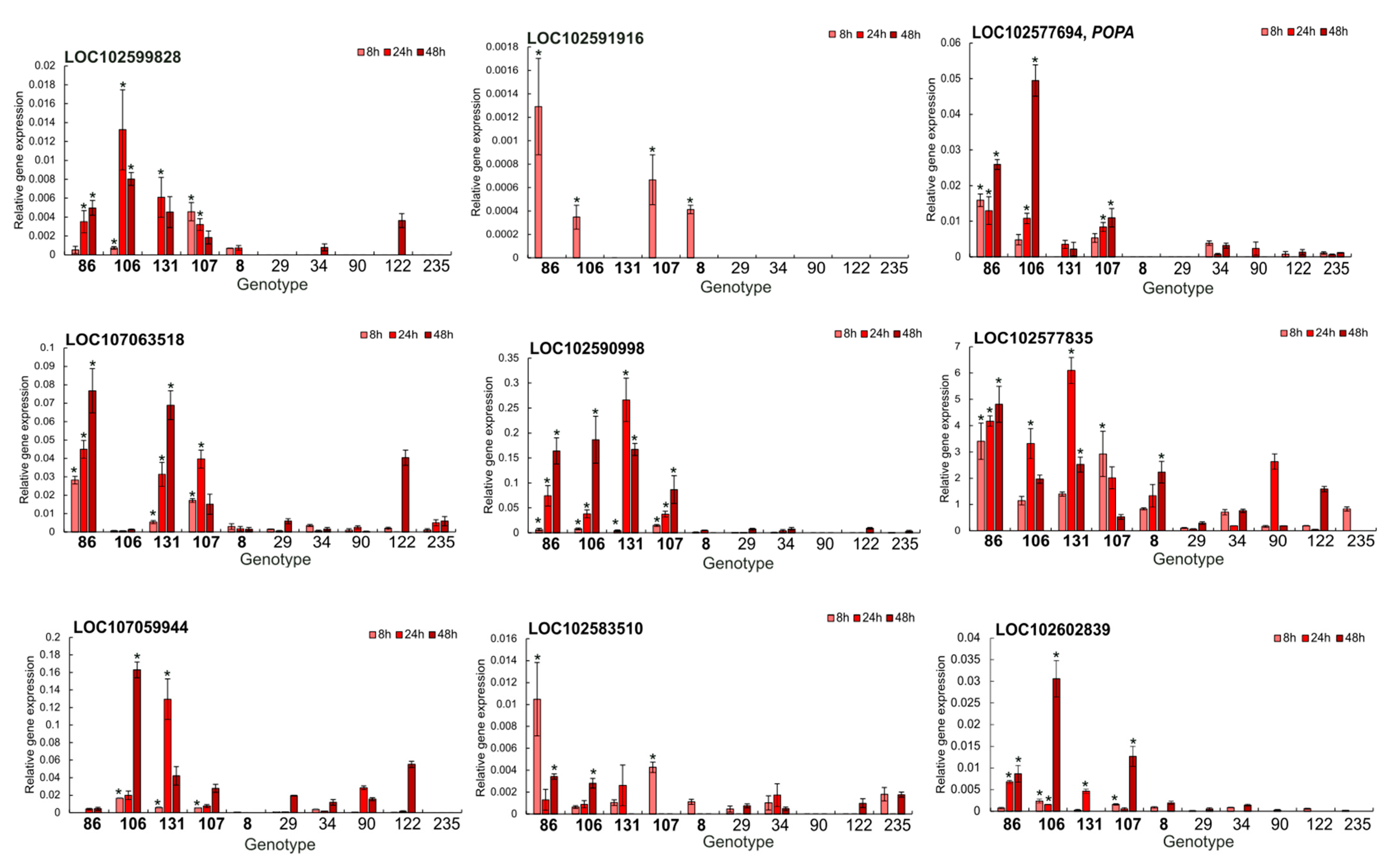
2.5. RT-qPCR Validation
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Bacterial Inoculum
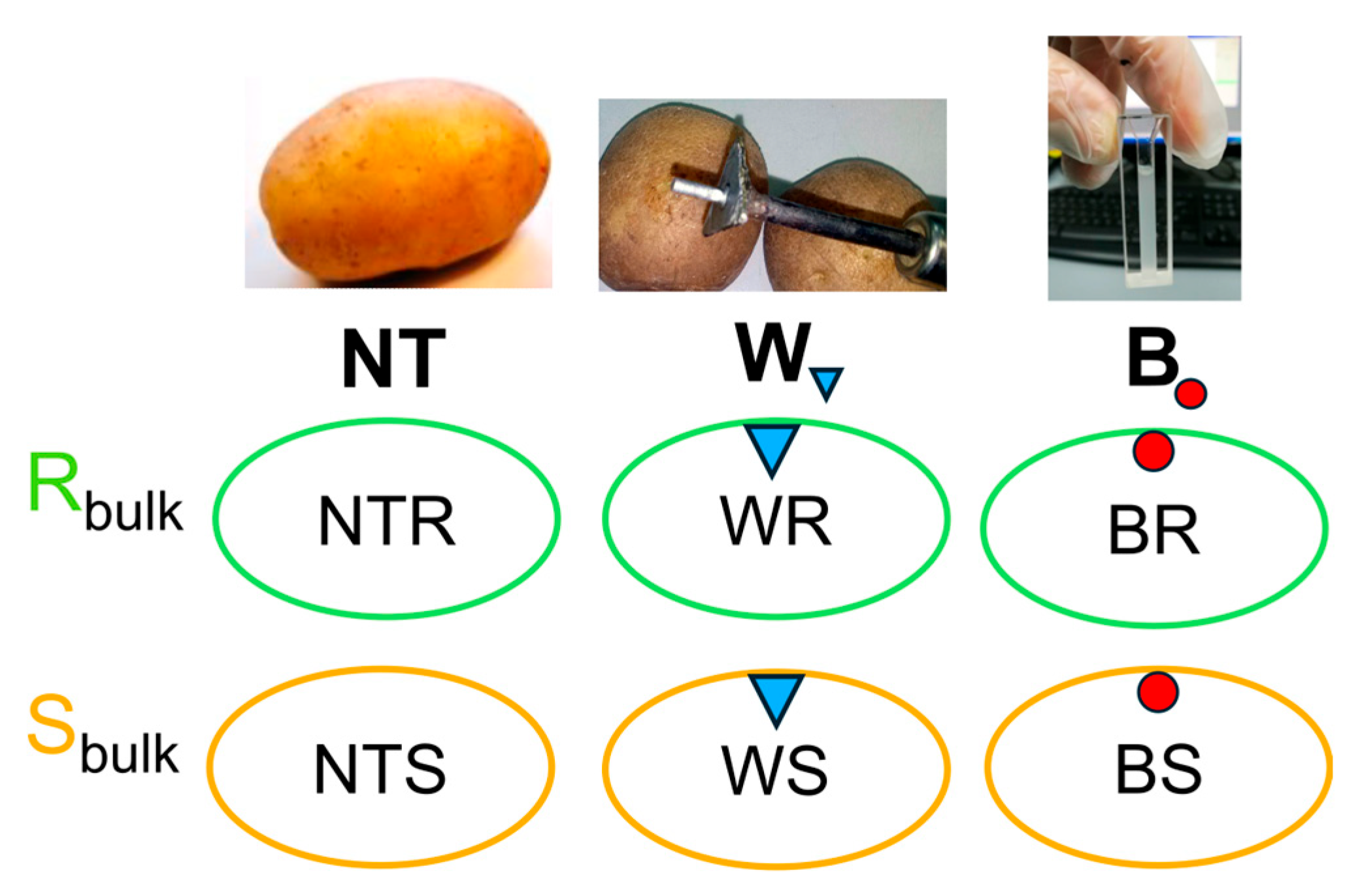
4.3. Experimental Procedure
4.4. RNA Isolation and Preparation of Sequencing Samples
4.5. RNA Sequencing
4.6. Bioinformatics Analysis
4.7. Reverse Transcription and cDNA Synthesis
4.8. PCR Amplification
4.9. Quantitative PCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Birch, P.R.J.; Bryan, G.; Fenton, B.; Gilroy, E.M.; Hein, I.; Jones, J.T.; Prashar, A.; Taylor, M.A.; Torrance, L.; Toth, I.K. Crops That Feed the World 8: Potato: Are the Trends of Increased Global Production Sustainable? Food Sec. 2012, 4, 477–508. [Google Scholar] [CrossRef]
- OECD; Food and Agriculture Organization of the United Nations. OECD-FAO Agricultural Outlook 2024–2033; OECD-FAO Agricultural Outlook; OECD: Paris, France, 2024; ISBN 978-92-64-72259-0. [Google Scholar]
- Mansfield, J.; Genin, S.; Magori, S.; Citovsky, V.; Sriariyanum, M.; Ronald, P.; Dow, M.; Verdier, V.; Beer, S.V.; Machado, M.A.; et al. Top 10 Plant Pathogenic Bacteria in Molecular Plant Pathology. Mol. Plant Pathol. 2012, 13, 614–629. [Google Scholar] [CrossRef]
- Czajkowski, R.; Pérombelon, M.C.M.; Van Veen, J.A.; Van Der Wolf, J.M. Control of Blackleg and Tuber Soft Rot of Potato Caused by Pectobacterium and Dickeya Species: A Review. Plant Pathol. 2011, 60, 999–1013. [Google Scholar] [CrossRef]
- Toth, I.K.; Barny, M.; Brurberg, M.B.; Condemine, G.; Czajkowski, R.; Elphinstone, J.G.; Helias, V.; Johnson, S.B.; Moleleki, L.N.; Pirhonen, M.; et al. Pectobacterium and Dickeya: Environment to Disease Development. In Plant Diseases Caused by Dickeya and Pectobacterium Species; Van Gijsegem, F., Van Der Wolf, J.M., Toth, I.K., Eds.; Springer International Publishing: Cham, Switzerland, 2021; pp. 39–84. ISBN 978-3-030-61458-4. [Google Scholar]
- Zimnoch-Guzowska, E.; Marczewski, W.; Lebecka, R.; Flis, B.; Schäfer-Pregl, R.; Salamini, F.; Gebhardt, C. QTL Analysis of New Sources of Resistance to Erwinia Carotovora Ssp. Atroseptica in Potato Done by AFLP, RFLP, and Resistance-Gene-Like Markers. Crop Sci. 2000, 40, 1156–1167. [Google Scholar] [CrossRef]
- Lebecka, R.; Zimnoch-Guzowska, E.; Kaczmarek, Z. Resistance to Soft Rot (Erwinia Carotovora Subsp.Atroseptica) in Tetraploid Potato Families Obtained from 4x-2x Crosses. Am. J. Potato Res. 2005, 82, 203–210. [Google Scholar] [CrossRef]
- Lebecka, R.; Wasilewicz-Flis, I.; Mańkowski, D. Diploid Potato Germplasm with Resistance to Dickeya Solani. Potato Res. 2021, 64, 375–385. [Google Scholar] [CrossRef]
- Lebecka, R.; Śliwka, J.; Grupa-Urbańska, A.; Szajko, K.; Marczewski, W. QTLs for Potato Tuber Resistance to Dickeya Solani Are Located on Chromosomes II and IV. Plant Pathol. 2021, 70, 1745–1756. [Google Scholar] [CrossRef]
- Lebecka, R.; Kistowski, M.; Dębski, J.; Szajko, K.; Murawska, Z.; Marczewski, W. Quantitative Proteomic Analysis of Differentially Expressed Proteins in Tubers of Potato Plants Differing in Resistance to Dickeya Solani. Plant Soil 2019, 441, 317–329. [Google Scholar] [CrossRef]
- Yang, W.-L.; Bernards, M.A. Wound-Induced Metabolism in Potato (Solanum tuberosum) Tubers: Biosynthesis of Aliphatic Domain Monomers. Plant Signal. Behav. 2006, 1, 59–66. [Google Scholar] [CrossRef]
- Woolfson, K.N.; Zhurov, V.; Wu, T.; Kaberi, K.M.; Wu, S.; Bernards, M.A. Transcriptomic Analysis of Wound-Healing in Solanum Tuberosum (Potato) Tubers: Evidence for a Stepwise Induction of Suberin-Associated Genes. Phytochemistry 2023, 206, 113529. [Google Scholar] [CrossRef]
- Kraepiel, Y.; Barny, M. Gram-negative Phytopathogenic Bacteria, All Hemibiotrophs after All? Mol. Plant Pathol. 2016, 17, 313–316. [Google Scholar] [CrossRef] [PubMed]
- Charkowski, A.O. The Changing Face of Bacterial Soft-Rot Diseases. Annu. Rev. Phytopathol. 2018, 56, 269–288. [Google Scholar] [CrossRef]
- Gonzalez, J.M.; Aranda, B. Microbial Growth under Limiting Conditions-Future Perspectives. Microorganisms 2023, 11, 1641. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Ren, Y.; Guo, S.; Zhang, H.; Gong, G.; Du, Y.; Xu, Y. Application of Comparative Genomics in Developing Markers Tightly Linked to the Pm-2F Gene for Powdery Mildew Resistance in Melon (Cucumis melo L.). Euphytica 2013, 190, 157–168. [Google Scholar] [CrossRef]
- Zou, C.; Wang, P.; Xu, Y. Bulked Sample Analysis in Genetics, Genomics and Crop Improvement. Plant Biotechnol. J. 2016, 14, 1941–1955. [Google Scholar] [CrossRef]
- Hardigan, M.A.; Crisovan, E.; Hamilton, J.P.; Kim, J.; Laimbeer, P.; Leisner, C.P.; Manrique-Carpintero, N.C.; Newton, L.; Pham, G.M.; Vaillancourt, B.; et al. Genome Reduction Uncovers a Large Dispensable Genome and Adaptive Role for Copy Number Variation in Asexually Propagated Solanum Tuberosum. Plant Cell 2016, 28, 388–405. [Google Scholar] [CrossRef]
- Liu, H.; Coulthurst, S.J.; Pritchard, L.; Hedley, P.E.; Ravensdale, M.; Humphris, S.; Burr, T.; Takle, G.; Brurberg, M.-B.; Birch, P.R.J.; et al. Quorum Sensing Coordinates Brute Force and Stealth Modes of Infection in the Plant Pathogen Pectobacterium Atrosepticum. PLoS Pathog. 2008, 4, e1000093. [Google Scholar] [CrossRef]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A Revolutionary Tool for Transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Everaert, C.; Luypaert, M.; Maag, J.L.V.; Cheng, Q.X.; Dinger, M.E.; Hellemans, J.; Mestdagh, P. Benchmarking of RNA-Sequencing Analysis Workflows Using Whole-Transcriptome RT-qPCR Expression Data. Sci. Rep. 2017, 7, 1559. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE Guidelines: Minimum Information for Publication of Quantitative Real-Time PCR Experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef]
- Lee, D.-K.; Ahn, S.; Cho, H.Y.; Yun, H.Y.; Park, J.H.; Lim, J.; Lee, J.; Kwon, S.W. Metabolic Response Induced by Parasitic Plant-Fungus Interactions Hinder Amino Sugar and Nucleotide Sugar Metabolism in the Host. Sci. Rep. 2016, 6, 37434. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, J. Transcriptomic and Proteomic Effects of Gene Deletion Are Not Evolutionarily Conserved. Genome Res. 2025, 35, 512–521. [Google Scholar] [CrossRef]
- Gullner, G.; Komives, T.; Király, L.; Schröder, P. Glutathione S-Transferase Enzymes in Plant-Pathogen Interactions. Front. Plant Sci. 2018, 9, 1836. [Google Scholar] [CrossRef] [PubMed]
- Kuźniak, E.; SkŁodowska, M. Differential Implication of Glutathione, Glutathione-Metabolizing Enzymes and Ascorbate in Tomato Resistance to Pseudomonas Syringae. J. Phytopathol. 2004, 152, 529–536. [Google Scholar] [CrossRef]
- Satish, L.; Shamili, S.; Yolcu, S.; Lavanya, G.; Alavilli, H.; Swamy, M.K. Biosynthesis of Secondary Metabolites in Plants as Influenced by Different Factors. In Plant-Derived Bioactives; Swamy, M.K., Ed.; Springer: Singapore, 2020; pp. 61–100. ISBN 9789811517600. [Google Scholar]
- Anjali; Kumar, S.; Korra, T.; Thakur, R.; Arutselvan, R.; Kashyap, A.S.; Nehela, Y.; Chaplygin, V.; Minkina, T.; Keswani, C. Role of Plant Secondary Metabolites in Defence and Transcriptional Regulation in Response to Biotic Stress. Plant Stress 2023, 8, 100154. [Google Scholar] [CrossRef]
- Chen, Z.; Shao, S.; Zhu, X.; Zhang, Y.; Lan, Z.; Jin, H. Transcriptome Responses to Ralstonia Solanacearum Infection in Tetraploid Potato. Heliyon 2025, 11, e41903. [Google Scholar] [CrossRef]
- Jha, Y. Endophytic Bacteria Mediated Anti-Autophagy and Induced Catalase, β-1,3-Glucanases Gene in Paddy after Infection with Pathogen Pyricularia Grisea. Indian Phytopathol. 2019, 72, 99–106. [Google Scholar] [CrossRef]
- Jha, Y. Higher induction of defense enzymes and cell wall reinforcement in maize by root associated bacteria for better protection against Aspergillus niger. J. Plant Prot. Res. 2023, 59, 341–349. [Google Scholar] [CrossRef]
- Woolfson, K.N.; Esfandiari, M.; Bernards, M.A. Suberin Biosynthesis, Assembly, and Regulation. Plants 2022, 11, 555. [Google Scholar] [CrossRef]
- Lavhale, S.G.; Kalunke, R.M.; Giri, A.P. Structural, Functional and Evolutionary Diversity of 4-Coumarate-CoA Ligase in Plants. Planta 2018, 248, 1063–1078. [Google Scholar] [CrossRef]
- Bernards, M.A.; Fleming, W.D.; Llewellyn, D.B.; Priefer, R.; Yang, X.; Sabatino, A.; Plourde, G.L. Biochemical Characterization of the Suberization-Associated Anionic Peroxidase of Potato. Plant Physiol. 1999, 121, 135–146. [Google Scholar] [CrossRef]
- Du, J.; Tian, Z.; Liu, J.; Vleeshouwers, V.G.A.A.; Shi, X.; Xie, C. Functional Analysis of Potato Genes Involved in Quantitative Resistance to Phytophthora Infestans. Mol. Biol. Rep. 2013, 40, 957–967. [Google Scholar] [CrossRef]
- Yang, Y.; Jiang, R.; Wang, H.; Tian, Z.; Xie, C. StPOPA, Encoding an Anionic Peroxidase, Enhances Potato Resistance against Phytophthora Infestans. Mol. Breed. 2020, 40, 16. [Google Scholar] [CrossRef]
- Golanowska, M.; Galardini, M.; Bazzicalupo, M.; Hugouvieux-Cotte-Pattat, N.; Mengoni, A.; Potrykus, M.; Slawiak, M.; Lojkowska, E. Draft Genome Sequence of a Highly Virulent Strain of the Plant Pathogen Dickeya Solani, IFB0099. Genome Announc. 2015, 3, e00109-15. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt Removes Adapter Sequences from High-Throughput Sequencing Reads. EMBnet. J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-Based Genome Alignment and Genotyping with HISAT2 and HISAT-Genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq—A Python Framework to Work with High-Throughput Sequencing Data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef]
- Tian, F.; Yang, D.-C.; Meng, Y.-Q.; Jin, J.; Gao, G. PlantRegMap: Charting Functional Regulatory Maps in Plants. Nucleic Acids Res. 2019, 48, D1104–D1113. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Kanehisa, M. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L.; et al. clusterProfiler 4.0: A Universal Enrichment Tool for Interpreting Omics Data. Innovation 2021, 2, 100141. [Google Scholar] [CrossRef] [PubMed]
- Nahar, N.; Westerberg, E.; Arif, U.; Huchelmann, A.; Olarte Guasca, A.; Beste, L.; Dalman, K.; Dutta, P.C.; Jonsson, L.; Sitbon, F. Transcript Profiling of Two Potato Cultivars during Glycoalkaloid-Inducing Treatments Shows Differential Expression of Genes in Sterol and Glycoalkaloid Metabolism. Sci. Rep. 2017, 7, 43268. [Google Scholar] [CrossRef] [PubMed]
- Vogel, C.; Marcotte, E.M. Insights into the Regulation of Protein Abundance from Proteomic and Transcriptomic Analyses. Nat. Rev. Genet. 2012, 13, 227–232. [Google Scholar] [CrossRef]
- Sorokan, A.V.; Burhanova, G.F.; Maksimov, I.V. Anionic Peroxidase-mediated Oxidative Burst Requirement for Jasmonic Acid-dependent Solanum tuberosum Defence against Phytophthora infestans. Plant Pathol. 2018, 67, 349–357. [Google Scholar] [CrossRef]
| Time (hpi) | Treatment | Number of Genes | ||||
|---|---|---|---|---|---|---|
| Total | Significant (p ≤ 0.01) | |||||
| Upregulated | Downregulated | |||||
| Log2 Fold Change ≥ 2 | Log2 Fold Change ≤ −2 | |||||
| 8 | BS vs. BR | 21,068 | 3134 | 566 | 3345 | 170 |
| WS vs. WR | 21,228 | 2000 | 229 | 2457 | 140 | |
| NTS vs. NTR | 20,730 | 2768 | 443 | 2937 | 231 | |
| 24 | BS vs. BR | 20,695 | 3339 | 822 | 3164 | 485 |
| WS vs. WR | 21,196 | 3391 | 400 | 3606 | 544 | |
| NTS vs. NTR | 20,063 | 3857 | 858 | 3737 | 819 | |
| 48 | BS vs. BR | 22,337 | 2879 | 675 | 2693 | 348 |
| WS vs. WR | 22,105 | 2787 | 1014 | 2811 | 204 | |
| NTS vs. NTR | 21,520 | 3034 | 396 | 3253 | 861 | |
| Gene ID | Log2 Fold Change | Functional Annotation | |||||
|---|---|---|---|---|---|---|---|
| WS vs. WR hpi | BS vs. BR hpi | ||||||
| 8 | 24 | 48 | 8 | 24 | 48 | ||
| Phenylpropanoid biosynthesis | |||||||
| 102596330 | 4.65 | - | 7.41 | 4.62 | - | 6.88 | Acetyl-CoA-benzyl alcohol acetyltransferase-like |
| 102578320 | - | - | - | 4.49 | - | - | acetyl-CoA-benzyl alcohol acetyltransferase-like |
| 102599828 | - | - | - | 4.42 | 6.53 | 3.58 | Lignin-forming anionic peroxidase-like |
| 102598379 | - | - | - | 4.34 | - | - | Salutaridinol 7-O-acetyltransferase-like |
| 102591916 | 4.18 | - | - | 4.33 | - | - | Peroxidase 43-like |
| 102580092 | 3.76 | 2.03 | - | 4.21 | - | - | Peroxidase P7-like |
| 102605292 | 3.86 | - | - | 2.5 | - | 2.74 | Suberisation-associated anionic peroxidase 2-like |
| 102577694 | - | - | - | 2.87 | 3.99 | 5.35 | Suberisation-associated anionic peroxidase, POPA |
| 107063518 | - | - | 3.88 | 2.47 | 2.43 | - | Peroxidase 21 |
| 102601606 | 2.02 | - | - | 2.64 | - | 5.69 | Cationic peroxidase 1-like |
| 102589793 | - | - | - | 2.43 | 3.72 | - | 4-coumarate-CoA ligase-like 6 |
| 102584603 | - | - | - | 2.39 | 3.56 | 2.95 | 4-coumarate-CoA ligase 2 |
| 102588483 | - | - | - | 2.06 | 2.61 | - | 4-coumarate-CoA ligase-like 9 |
| 102588050 | - | - | 2.12 | 4.96 | - | - | Lignin-forming anionic peroxidase-like |
| 102580211 | - | - | - | 2.29 | 3.77 | - | Cinnamoyl-CoA reductase 1 |
| 102586332 | 2.93 | - | - | 3.61 | 6.03 | - | Caffeoyl-CoA O-methyltransferase-like |
| Phenylpropanoid biosynthesis; Phenylalanine metabolism | |||||||
| 1025826182 | 2.06 | 2.35 | - | 4.05 | 4.36 | 4.04 | Phenylalanine ammonia-lyase-like |
| Phenylalanine metabolism | |||||||
| 102591703 | 2.12 | - | 4.15 | 3.6 | 6.85 | 4.07 | Histidine decarboxylase-like |
| 102581954 | 3.38 | - | - | 3.34 | 5.27 | 2.00 | |
| 102592044 | 2.06 | - | 2.43 | 3.31 | 3.3 | 3.84 | |
| 102581292 | - | - | - | 3.29 | 3.45 | ||
| 102580376 | - | - | 2.09 | 2.32 | 4.02 | - | |
| 102579686 | - | - | - | 2.07 | 2.29 | - | Putative amidase |
| Phenylalanine metabolism; Tyrosine metabolism; Isoquinoline alkaloid biosynthesis | |||||||
| 1025838023 | - | - | 3.38 | 4.71 | 5.02 | - | Primary amine oxidase-like |
| 1026004133 | - | - | 4.16 | 4.03 | 2.52 | ||
| 1025818353 | 2.07 | - | - | 2.43 | 2.16 | 2.07 | |
| Tyrosine metabolism; Isoquinoline alkaloid biosynthesis | |||||||
| 1026043802 | - | 2.67 | - | 3.14 | 2.89 | Polyphenol oxidase D, chloroplastic | |
| 1025777272 | 2.7 | 2.49 | 0.49 | 3.1 | - | 2.67 | PPO, Catechol oxidase B |
| 102597889 | 3.22 | - | - | 3.35 | 8.54 | Alcohol dehydrogenase 1 | |
| 102599478 | 2.33 | - | 2.52 | 2.48 | - | 2.58 | |
| Amino sugar and nucleotide sugar metabolism | |||||||
| 102590097 | - | 3.09 | - | 5.59 | 5.18 | 2.39 | Endochitinase 3-like |
| 102590998 | - | 2.95 | - | 4.39 | 3.8 | 3.89 | Basic endochitinase |
| 102592970 | - | - | - | 2.64 | Solanum tuberosum endochitinase 4-like | ||
| 102583127 | - | - | - | 2.14 | 4.63 | 2.34 | Endochitinase EP3 |
| 102577835 | - | 2.06 | 2.30 | 2.04 | 2.08 | Acidic endochitinase pcht28 | |
| 102584052 | 3.22 | - | - | 2.48 | 1.47 | - | Alpha-1,4-glucan-protein synthase [UDP-forming] 2-like |
| 102600109 | - | - | 1.7 | 2.25 | - | 2.22 | Acidic mammalian chitinase-like |
| 107059944 | - | - | - | 3.65 | 2.48 | 2.02 | Chitotriosidase-1-like |
| 102599450 | - | - | 2.07 | 3.33 | 2.96 | 2.3 | |
| 102594281 | - | 2.16 | - | 2.17 | 2.8 | Mannose-1-phosphate Guanylyltransferase 1-like | |
| Glutathione metabolism | |||||||
| 102599376 | - | 5.31 | 3.93 | 4.63 | 5.18 | 2.70 | Probable glutathione S-transferase |
| 102600356 | 2.49 | - | 2.03 | 2.61 | 4.96 | 3.15 | |
| 102599054 | - | - | 3.43 | 2.6 | 5.01 | 3.56 | |
| 102602051 | - | - | 4.48 | 2.41 | 2.23 | 5.17 | |
| 102604161 | 3.75 | 7.39 | 5.21 | 2.41 | 5.54 | 5.59 | Glutathione S-transferase L1-like |
| Gene ID | Log2 Fold Change | Physical Position on the Chromosome a | Functional Annotation | |
|---|---|---|---|---|
| Beginning | End | |||
| ChrII | ||||
| LOC102589748 | 5.26 | 22771975 | 22769355 | Probable N-acetyltransferase HLS1 |
| LOC102590998 | 4.39 | 19537268 | 19539189 | Basic endochitinase [Solanum tuberosum (potato)] |
| LOC102580833 | 4.21 | 33215044 | 33210712 | NAC domain-containing protein 73 [Solanum tuberosum (potato)] |
| LOC102604927 | 4.04 | 41059989 | 41058531 | ST1 homolog |
| LOC102594583 | 3.88 | 29338193 | 29336454 | Tetrahydrocannabinolic acid synthase-like |
| LOC102594665 | 3.54 | 36280755 | 36276180 | Protein DETOXIFICATION 40-like |
| LOC102605761 | 3.02 | 17250771 | 17248588 | WRKY transcription factor 6-like |
| LOC102577694 | 2.87 | 35172274 | 35169042 | Suberisation-associated anionic, POPA |
| LOC102582119 | 2.73 | 47475919 | 47479431 | NAC domain-containing protein 8 |
| LOC102579839 | 2.16 | 27067654 | 27064953 | 2-oxoglutarate-dependent dioxygenase |
| LOC102005497 | 2.13 | 33319425 | 33320828 | Protein E6 |
| LOC102579526 | 2.06 | 20712931 | 20711738 | GATA transcription factor 5-like |
| LOC102583510 | 2.05 | 26944018 | 26946758 | UDP-glycosyltransferase 87A2-like |
| LOC102578310 | 2.03 | 35301108 | 35305753 | G-type lectin S-receptor-like Serine/threonine-protein kinase At1g11300 |
| ChrIV | ||||
| LOC102603519 | 4.40 | 49272319 | 49274102 | Premnaspirodiene oxygenase-like |
| LOC102602839 | 3.31 | 49258137 | 49260195 | Cytochrome P450 71D7 |
| LOC102582168 | 2.55 | 52289066 | 52291659 | L-ascorbate oxidase-like |
| LOC102601591 | 2.19 | 52050946 | 52058860 | General transcription factor IIH subunit 2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grupa-Urbańska, A.; Szajko, K.; Marczewski, W.; Lebecka, R. New Insights into the Role of Secondary Metabolic Pathways in Resistance of Potato to Dickeya solani. Int. J. Mol. Sci. 2025, 26, 8370. https://doi.org/10.3390/ijms26178370
Grupa-Urbańska A, Szajko K, Marczewski W, Lebecka R. New Insights into the Role of Secondary Metabolic Pathways in Resistance of Potato to Dickeya solani. International Journal of Molecular Sciences. 2025; 26(17):8370. https://doi.org/10.3390/ijms26178370
Chicago/Turabian StyleGrupa-Urbańska, Anna, Katarzyna Szajko, Waldemar Marczewski, and Renata Lebecka. 2025. "New Insights into the Role of Secondary Metabolic Pathways in Resistance of Potato to Dickeya solani" International Journal of Molecular Sciences 26, no. 17: 8370. https://doi.org/10.3390/ijms26178370
APA StyleGrupa-Urbańska, A., Szajko, K., Marczewski, W., & Lebecka, R. (2025). New Insights into the Role of Secondary Metabolic Pathways in Resistance of Potato to Dickeya solani. International Journal of Molecular Sciences, 26(17), 8370. https://doi.org/10.3390/ijms26178370






