Transcriptomic Identification of Long Noncoding RNAs Modulating MPK3/MPK6-Centered Immune Networks in Arabidopsis
Abstract
1. Introduction
2. Results
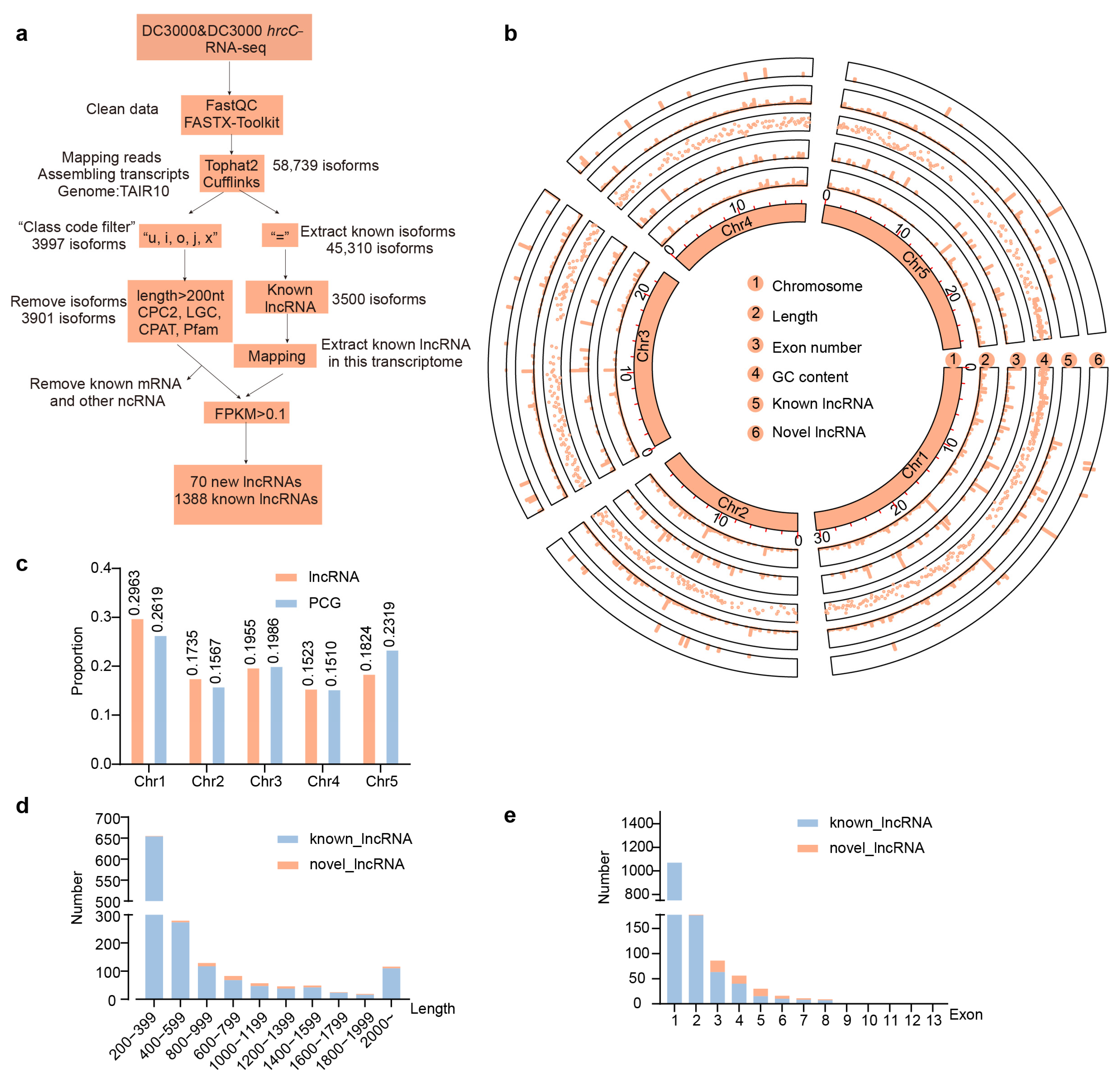
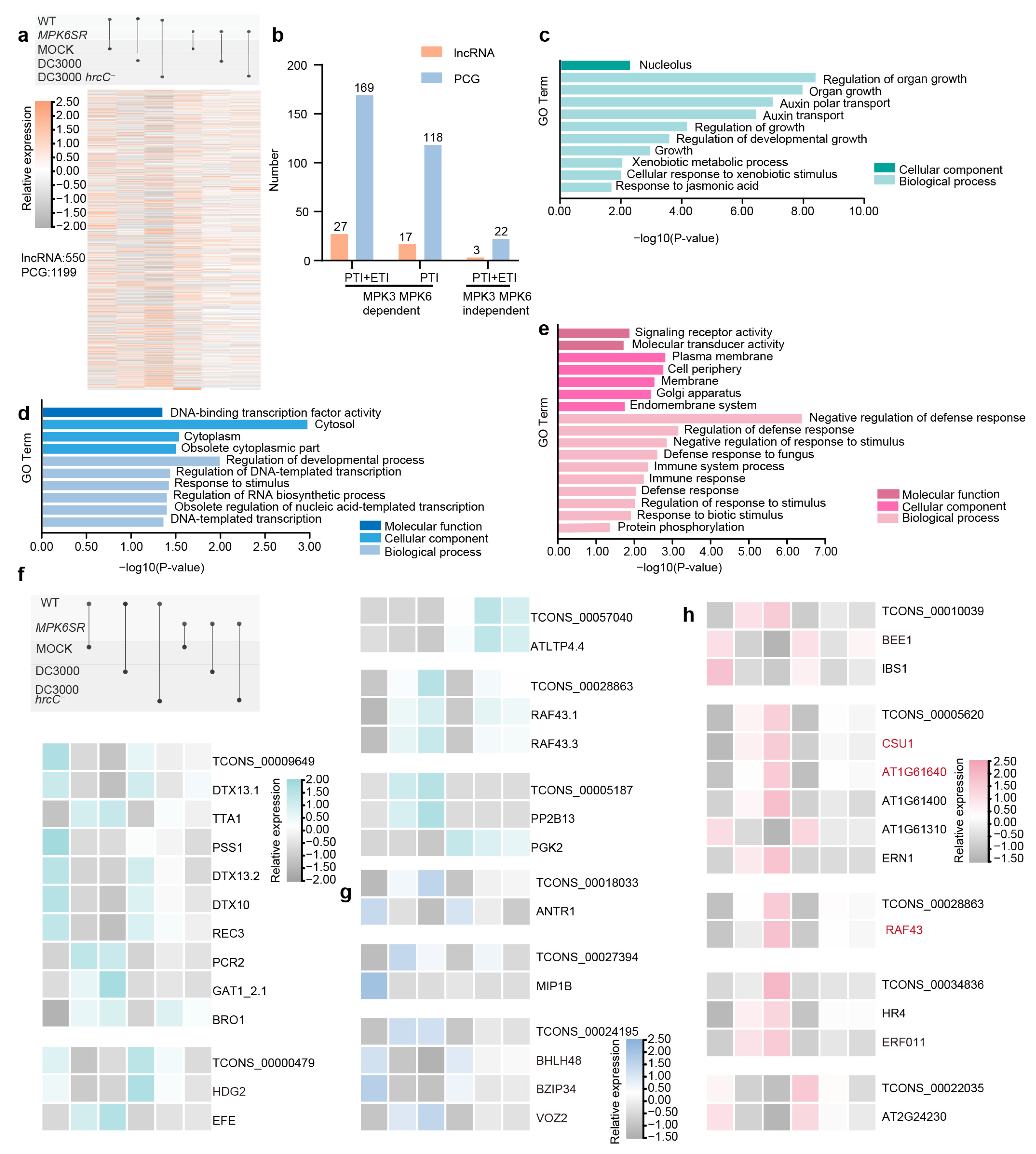
2.1. Characterization of Pathogen-Induced Immune Responses and Genome-Wide Identification of lncRNAs in Arabidopsis
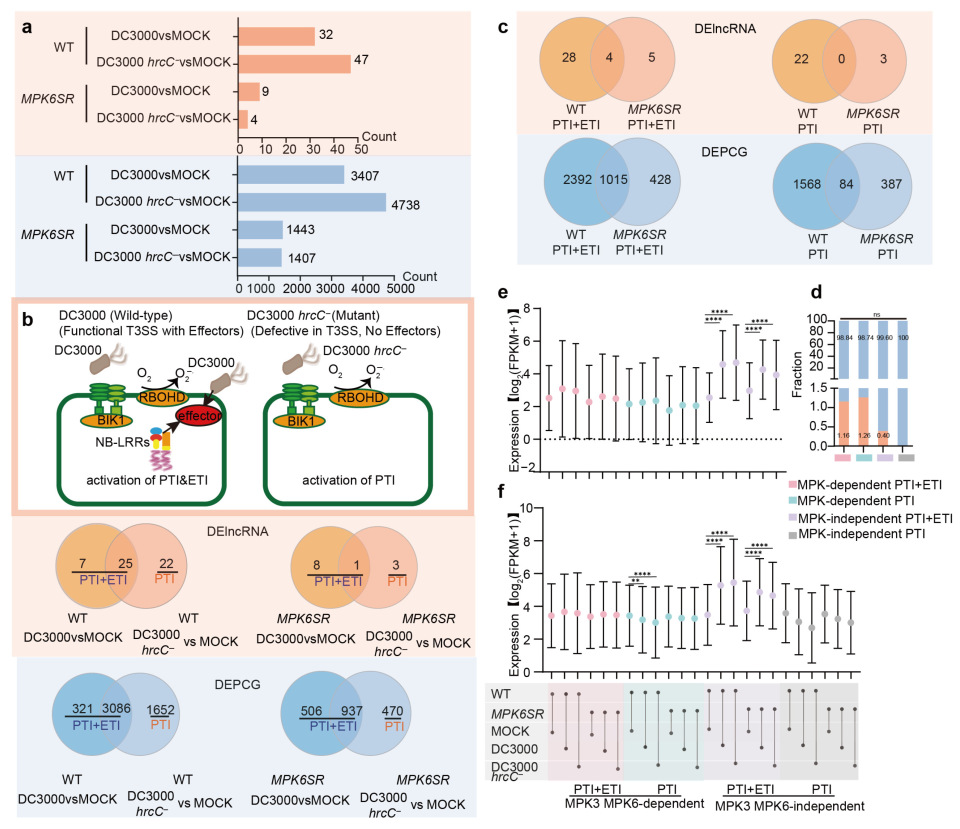
2.2. Differential Expression Analysis of lncRNAs and PCGs in Response to Pathogen Infection
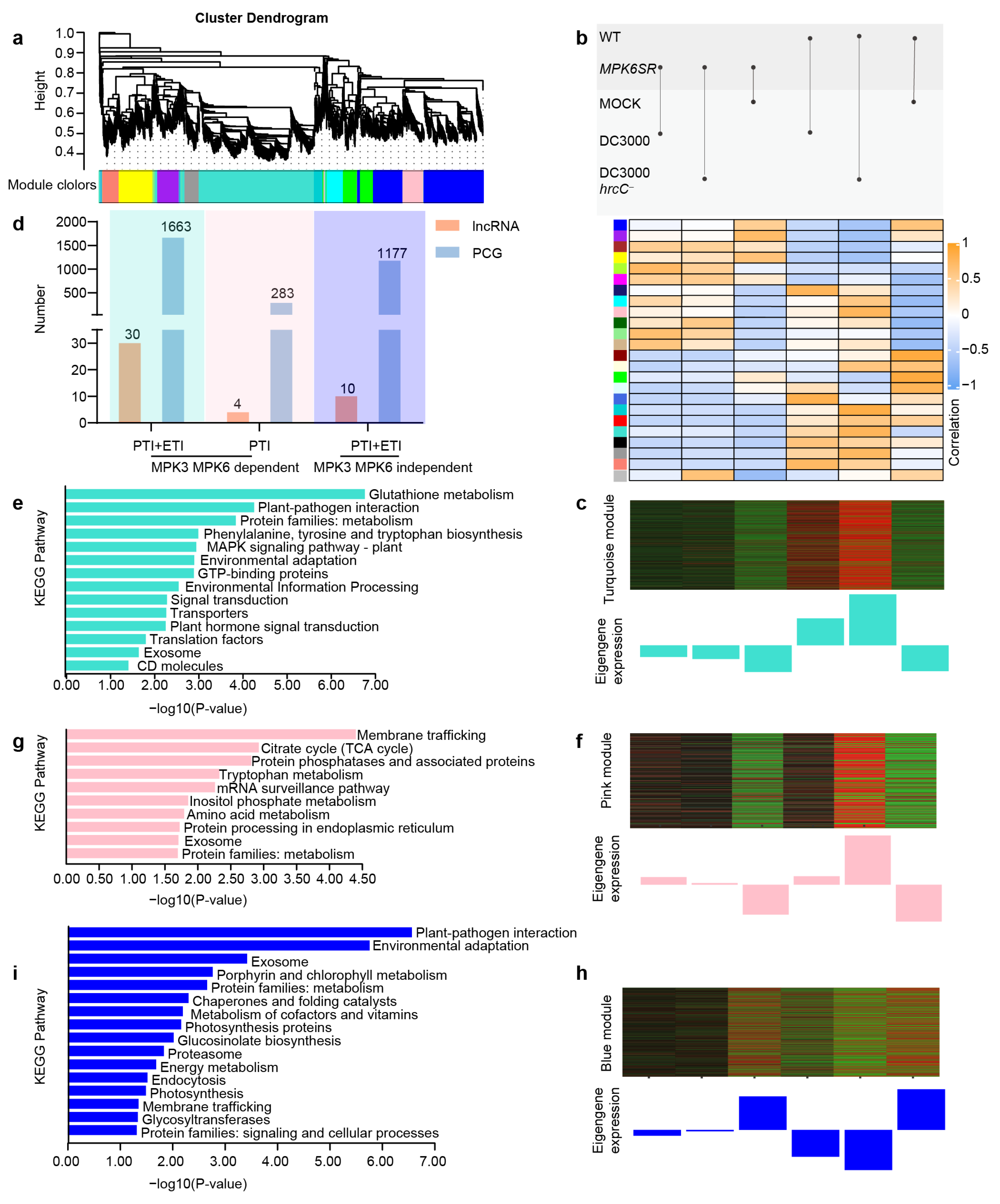
2.3. WGCNA Identification of Core Modules and Functional Analysis of lncRNA-PCG Regulatory Networks
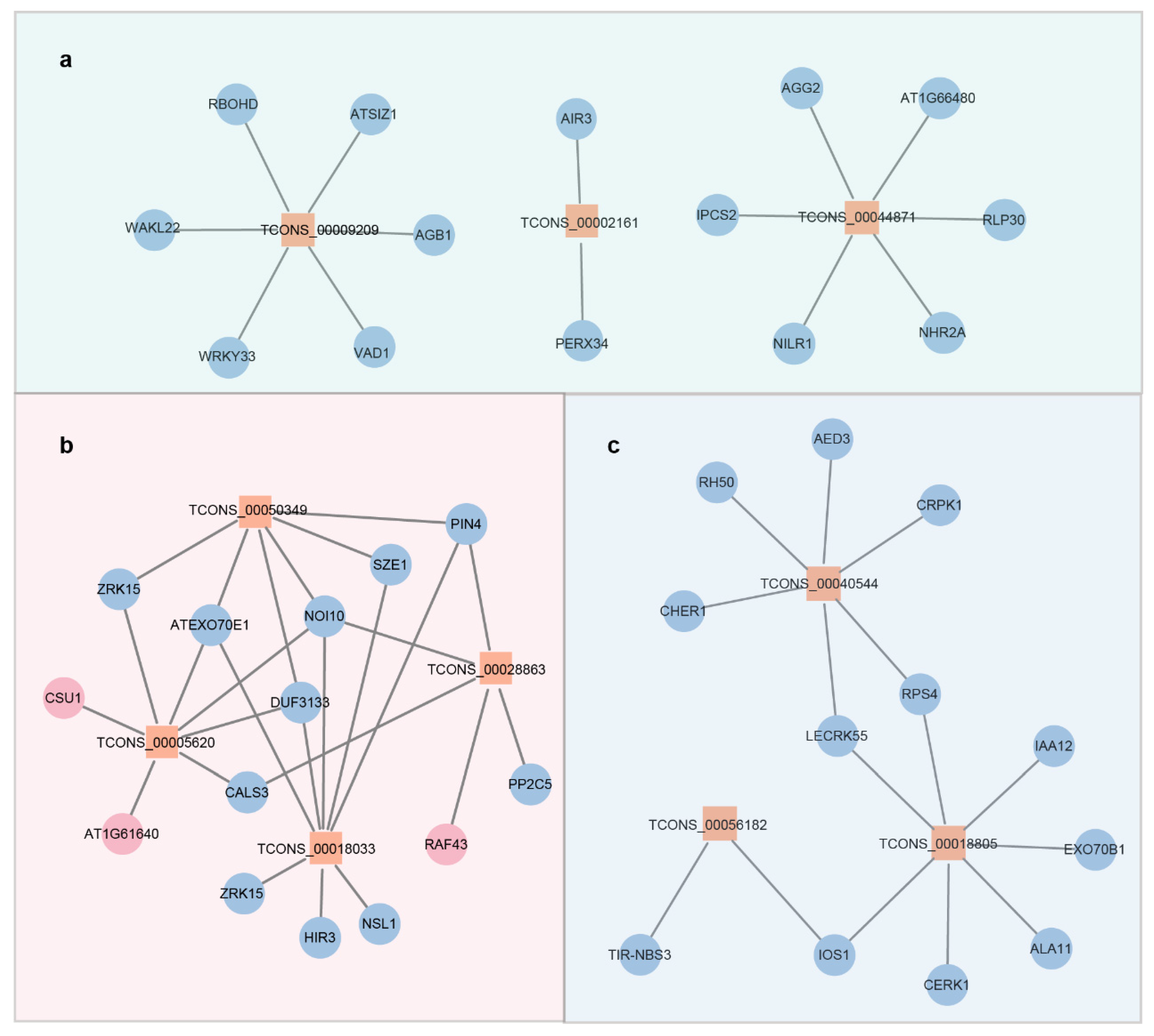
2.4. Cis-Regulatory lncRNA-PCG Pairs and Functional Classification of Proteins in Immune Pathways
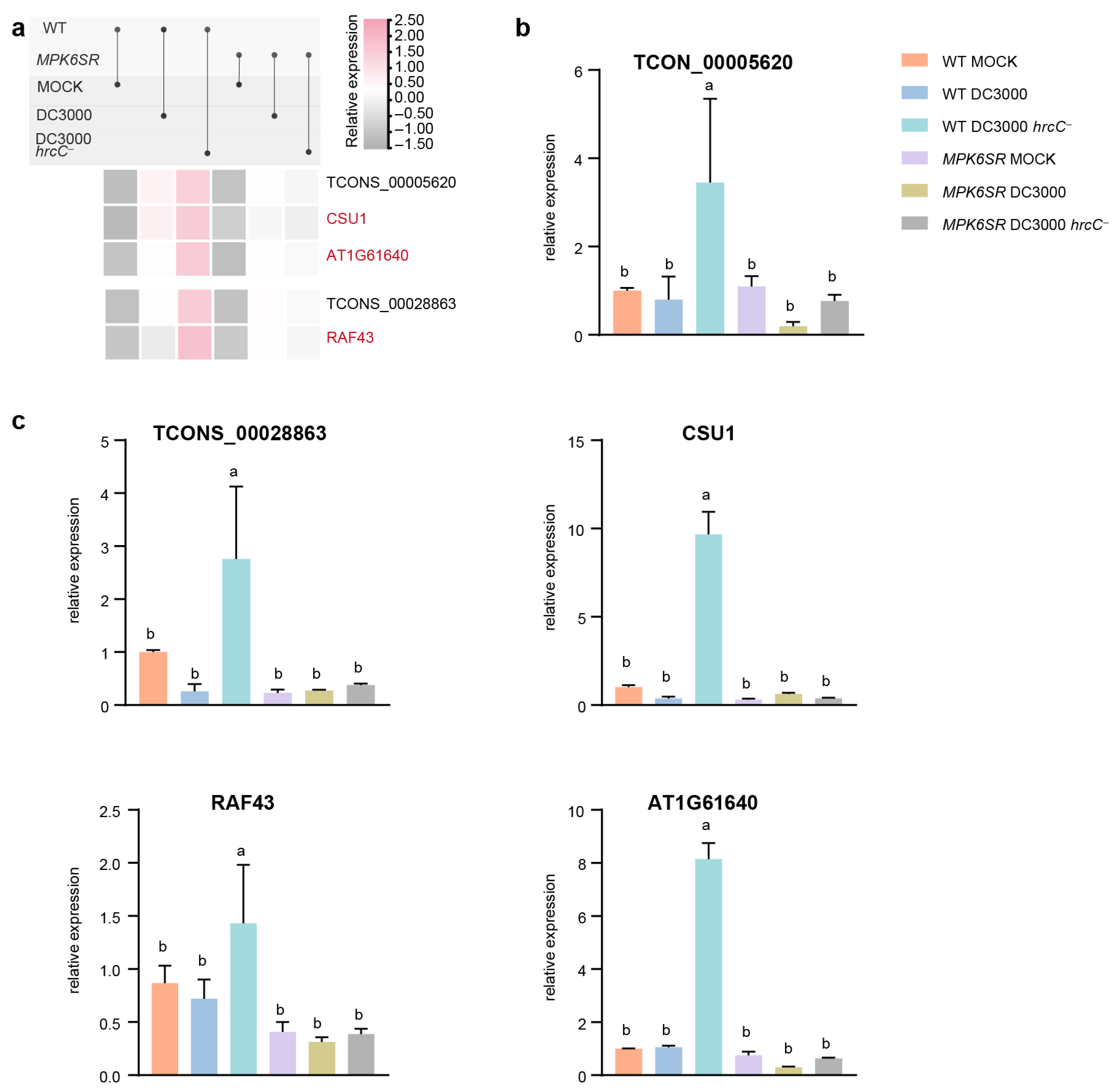
2.5. Validation of RNA-Seq Data by RT-qPCR Analysis
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Treatment
4.2. RNA-Seq and lncRNA Identification
4.3. Transcriptomic Feature Characterization and Differential Expression Analysis
4.4. Identification of lncRNA Targets via Trans-Regulatory Co-Expression Networks
4.5. Identification of Cis-Regulated Protein-Coding Gene Targets for lncRNAs
4.6. RNA Isolation and Quantitative RT-PCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Jones, J.D.G.; Dangl, J.L. The Plant Immune System. Nature 2006, 444, 323–329. [Google Scholar] [CrossRef]
- Yu, X.; Niu, H.; Liu, C.; Wang, H.; Yin, W.; Xia, X. PTI-ETI Synergistic Signal Mechanisms in Plant Immunity. Plant Biotechnol. J. 2024, 22, 2113–2128. [Google Scholar] [CrossRef]
- Zhang, M.; Zhang, S. Mitogen-activated Protein Kinase Cascades in Plant Signaling. J. Integr. Plant Biol. 2022, 64, 301–341. [Google Scholar] [CrossRef]
- Asai, T.; Tena, G.; Plotnikova, J.; Willmann, M.R.; Chiu, W.-L.; Gomez-Gomez, L.; Boller, T.; Ausubel, F.M.; Sheen, J. MAP Kinase Signalling Cascade in Arabidopsis Innate Immunity. Nature 2002, 415, 977–983. [Google Scholar] [CrossRef]
- Guan, R.; Su, J.; Meng, X.; Li, S.; Liu, Y.; Xu, J.; Zhang, S. Multilayered Regulation of Ethylene Induction Plays a Positive Role in Arabidopsis Resistance against Pseudomonas syringae. Plant Physiol. 2015, 169, 299–312. [Google Scholar] [CrossRef]
- Tsuda, K.; Mine, A.; Bethke, G.; Igarashi, D.; Botanga, C.J.; Tsuda, Y.; Glazebrook, J.; Sato, M.; Katagiri, F. Dual Regulation of Gene Expression Mediated by Extended MAPK Activation and Salicylic Acid Contributes to Robust Innate Immunity in Arabidopsis Thaliana. PLoS Genet. 2013, 9, e1004015. [Google Scholar] [CrossRef] [PubMed]
- Chekanova, J.A.; Wang, H.V. Long Noncoding RNAs in Plants. In Advances in Experimental Medicine and Biology; Springer: Singapore, 2017; pp. 133–154. ISBN 978-981-10-5202-6. [Google Scholar]
- Fu, X.-D. Non-Coding RNA: A New Frontier in Regulatory Biology. Natl. Sci. Rev. 2014, 1, 190–204. [Google Scholar] [CrossRef] [PubMed]
- Schlackow, M.; Nojima, T.; Gomes, T.; Dhir, A.; Carmo-Fonseca, M.; Proudfoot, N.J. Distinctive Patterns of Transcription and RNA Processing for Human lincRNAs. Mol. Cell 2017, 65, 25–38. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Cheng, S.-J.; Ren, L.-C.; Wang, Q.; Kang, Y.-J.; Ding, Y.; Hou, M.; Yang, X.-X.; Lin, Y.; Liang, N.; et al. An Expanded Landscape of Human Long Noncoding RNA. Nucleic Acids Res. 2019, 47, 7842–7856. [Google Scholar] [CrossRef]
- Melé, M.; Mattioli, K.; Mallard, W.; Shechner, D.M.; Gerhardinger, C.; Rinn, J.L. Chromatin Environment, Transcriptional Regulation, and Splicing Distinguish lincRNAs and mRNAs. Genome Res. 2017, 27, 27–37. [Google Scholar] [CrossRef]
- Yu, Y.; Zhang, Y.; Chen, X.; Chen, Y. Plant Noncoding RNAs: Hidden Players in Development and Stress Responses. Annu. Rev. Cell Dev. Biol. 2019, 35, 407–431. [Google Scholar] [CrossRef] [PubMed]
- Han, P.; Chang, C.-P. Long Non-Coding RNA and Chromatin Remodeling. RNA Biol. 2015, 12, 1094–1098. [Google Scholar] [CrossRef]
- Sebastian-delaCruz, M.; Gonzalez-Moro, I.; Olazagoitia-Garmendia, A.; Castellanos-Rubio, A.; Santin, I. The Role of lncRNAs in Gene Expression Regulation through mRNA Stabilization. Non-Coding RNA 2021, 7, 3. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wang, W.; Zhu, W.; Dong, J.; Cheng, Y.; Yin, Z.; Shen, F. Mechanisms and Functions of Long Non-Coding RNAs at Multiple Regulatory Levels. Int. J. Mol. Sci. 2019, 20, 5573. [Google Scholar] [CrossRef]
- Mattick, J.S.; Rinn, J.L. Discovery and Annotation of Long Noncoding RNAs. Nat. Struct. Mol. Biol. 2015, 22, 5–7. [Google Scholar] [CrossRef]
- Seo, J.S.; Sun, H.-X.; Park, B.S.; Huang, C.-H.; Yeh, S.-D.; Jung, C.; Chua, N.-H. ELF18-INDUCED LONG-NONCODING RNA Associates with Mediator to Enhance Expression of Innate Immune Response Genes in Arabidopsis. Plant Cell 2017, 29, 1024–1038. [Google Scholar] [CrossRef]
- Zhu, Q.; Stephen, S.; Taylor, J.; Helliwell, C.A.; Wang, M. Long Noncoding RNAs Responsive to Fusarium oxysporum Infection in Arabidopsis thaliana. New Phytol. 2014, 201, 574–584. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Luan, Y.; Jiang, N.; Bao, H.; Meng, J. Comparative Transcriptome Analysis between Resistant and Susceptible Tomato Allows the Identification of LncRNA16397 Conferring Resistance to Phytophthora infestans by Co-expressing Glutaredoxin. Plant J. 2017, 89, 577–589. [Google Scholar] [CrossRef]
- Wang, J.; Yu, W.; Yang, Y.; Li, X.; Chen, T.; Liu, T.; Ma, N.; Yang, X.; Liu, R.; Zhang, B. Genome-Wide Analysis of Tomato Long Non-Coding RNAs and Identification as Endogenous Target Mimic for microRNA in Response to TYLCV Infection. Sci. Rep. 2015, 5, 16946. [Google Scholar] [CrossRef]
- Li, R.; Jin, J.; Xu, J.; Wang, L.; Li, J.; Lou, Y.; Baldwin, I.T. Long Non-coding RNAs Associate with Jasmonate-mediated Plant Defence against Herbivores. Plant Cell Environ. 2021, 44, 982–994. [Google Scholar] [CrossRef]
- Lei, M.-Q.; He, R.-R.; Zhou, Y.-F.; Yang, L.; Zhang, Z.-F.; Yuan, C.; Zhao, W.-L.; Cheng, Y.; Lian, J.-P.; Zhang, Y.-C.; et al. The Long Noncoding RNA ALEX1 Confers a Functional Phase State of ARF3 to Enhance Rice Resistance to Bacterial Pathogens. Mol. Plant 2025, 18, 114–129. [Google Scholar] [CrossRef]
- Kwenda, S.; Birch, P.R.J.; Moleleki, L.N. Genome-Wide Identification of Potato Long Intergenic Noncoding RNAs Responsive to Pectobacterium Carotovorum Subspecies Brasiliense Infection. BMC Genom. 2016, 17, 614. [Google Scholar] [CrossRef]
- Zhang, H.; Hu, W.; Hao, J.; Lv, S.; Wang, C.; Tong, W.; Wang, Y.; Wang, Y.; Liu, X.; Ji, W. Genome-Wide Identification and Functional Prediction of Novel and Fungi-Responsive lincRNAs in Triticum Aestivum. BMC Genom. 2016, 17, 238. [Google Scholar] [CrossRef] [PubMed]
- Jin, X.; Wang, Z.; Li, X.; Ai, Q.; Wong, D.C.J.; Zhang, F.; Yang, J.; Zhang, N.; Si, H. Current Perspectives of lncRNAs in Abiotic and Biotic Stress Tolerance in Plants. Front. Plant Sci. 2024, 14, 1334620. [Google Scholar] [CrossRef] [PubMed]
- Abdallah, H.B.; Bauer, P. Quantitative Reverse Transcription-qPCR-Based Gene Expression Analysis in Plants. In Methods in Molecular Biology; Springer: New York, NY, USA, 2016; pp. 9–24. ISBN 978-1-4939-3114-9. [Google Scholar]
- Rigo, R.; Bazin, J.; Romero-Barrios, N.; Moison, M.; Lucero, L.; Christ, A.; Benhamed, M.; Blein, T.; Huguet, S.; Charon, C.; et al. The Arabidopsis Lnc RNA ASCO Modulates the Transcriptome through Interaction with Splicing Factors. EMBO Rep. 2020, 21, e48977. [Google Scholar] [CrossRef]
- Kim, J.Y.; Lee, J.; Kang, M.H.; Trang, T.T.M.; Lee, J.; Lee, H.; Jeong, H.; Lim, P.O. Dynamic Landscape of Long Noncoding RNAs during Leaf Aging in Arabidopsis. Front. Plant Sci. 2022, 13, 1068163. [Google Scholar] [CrossRef]
- Liu, N.; Xu, Y.; Li, Q.; Cao, Y.; Yang, D.; Liu, S.; Wang, X.; Mi, Y.; Liu, Y.; Ding, C.; et al. A lncRNA Fine-Tunes Salicylic Acid Biosynthesis to Balance Plant Immunity and Growth. Cell Host Microbe 2022, 30, 1124–1138.e8. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Li, X.; Yao, X.; Ma, J.; Lu, K.; An, Y.; Sun, Z.; Wang, Q.; Zhou, M.; Qin, L.; et al. MYB44 Regulates PTI by Promoting the Expression of EIN2 and MPK3/6 in Arabidopsis. Plant Commun. 2023, 4, 100628. [Google Scholar] [CrossRef]
- Bi, G.; Zhou, Z.; Wang, W.; Li, L.; Rao, S.; Wu, Y.; Zhang, X.; Menke, F.L.H.; Chen, S.; Zhou, J.-M. Receptor-Like Cytoplasmic Kinases Directly Link Diverse Pattern Recognition Receptors to the Activation of Mitogen-Activated Protein Kinase Cascades in Arabidopsis. Plant Cell 2018, 30, 1543–1561. [Google Scholar] [CrossRef]
- Lang, J.; Genot, B.; Bigeard, J.; Colcombet, J. MPK3 and MPK6 Control Salicylic Acid Signaling by Up-Regulating NLR Receptors during Pattern- and Effector-Triggered Immunity. J. Exp. Bot. 2022, 73, 2190–2205. [Google Scholar] [CrossRef]
- Su, C.; Li, X.; Dong, Y.; Daniel, B.; Liu, C.; Xing, Y.; Ma, D. Identification and Functional Analysis of Wheat lincRNAs in Response to Fusarium Graminearum Infection. Plant Physiol. Biochem. 2025, 224, 109898. [Google Scholar] [CrossRef]
- Zhang, L.; Lin, T.; Zhu, G.; Wu, B.; Zhang, C.; Zhu, H. LncRNAs Exert Indispensable Roles in Orchestrating the Interaction among Diverse Noncoding RNAs and Enrich the Regulatory Network of Plant Growth and Its Adaptive Environmental Stress Response. Hortic. Res. 2023, 10, uhad234. [Google Scholar] [CrossRef]
- Zhu, F.; Zhang, Q.; Che, Y.; Zhu, P.; Zhang, Q.; Ji, Z. Glutathione Contributes to Resistance Responses to TMV through a Differential Modulation of Salicylic Acid and Reactive Oxygen Species. Mol. Plant Pathol. 2021, 22, 1668–1687. [Google Scholar] [CrossRef]
- Levine, A.; Tenhaken, R.; Dixon, R.; Lamb, C. H2O2 from the Oxidative Burst Orchestrates the Plant Hypersensitive Disease Resistance Response. Cell 1994, 79, 583–593. [Google Scholar] [CrossRef]
- Boro, P.; Chattopadhyay, S. Crosstalk between MAPKs and GSH under Stress: A Critical Review. J. Biosci. 2022, 47, 71. [Google Scholar] [CrossRef]
- Dalton, T.P.; Shertzer, H.G.; Puga, A. Regulation of gene expression by reactive oxygen. Annu. Rev. Pharmacol. Toxicol. 1999, 39, 67–101. [Google Scholar] [CrossRef]
- Noctor, G.; Queval, G.; Mhamdi, A.; Chaouch, S.; Foyer, C.H. Glutathione. Arab. Book 2011, 9, 1–32. [Google Scholar] [CrossRef] [PubMed]
- Mao, G.; Meng, X.; Liu, Y.; Zheng, Z.; Chen, Z.; Zhang, S. Phosphorylation of a WRKY Transcription Factor by Two Pathogen-Responsive MAPKs Drives Phytoalexin Biosynthesis in Arabidopsis. Plant Cell 2011, 23, 1639–1653. [Google Scholar] [CrossRef]
- Nguyen, N.H.; Trotel-Aziz, P.; Clément, C.; Jeandet, P.; Baillieul, F.; Aziz, A. Camalexin Accumulation as a Component of Plant Immunity during Interactions with Pathogens and Beneficial Microbes. Planta 2022, 255, 116. [Google Scholar] [CrossRef] [PubMed]
- Kamata, N.; Okada, H.; Komeda, Y.; Takahashi, T. Mutations in Epidermis-specific HD-ZIP IV Genes Affect Floral Organ Identity in ARabidopsis thaliana. Plant J. 2013, 75, 430–440. [Google Scholar] [CrossRef]
- Boller, T.; Felix, G. A Renaissance of Elicitors: Perception of Microbe-Associated Molecular Patterns and Danger Signals by Pattern-Recognition Receptors. Annu. Rev. Plant Biol. 2009, 60, 379–406. [Google Scholar] [CrossRef] [PubMed]
- Bigeard, J.; Colcombet, J.; Hirt, H. Signaling Mechanisms in Pattern-Triggered Immunity (PTI). Mol. Plant 2015, 8, 521–539. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Meng, X.; Shan, L.; He, P. Transcriptional Regulation of Pattern-Triggered Immunity in Plants. Cell Host Microbe 2016, 19, 641–650. [Google Scholar] [CrossRef]
- Xu, J.; Xie, J.; Yan, C.; Zou, X.; Ren, D.; Zhang, S. A Chemical Genetic Approach Demonstrates That MPK3/MPK6 Activation and NADPH Oxidase-mediated Oxidative Burst Are Two Independent Signaling Events in Plant Immunity. Plant J. 2014, 77, 222–234. [Google Scholar] [CrossRef]
- Galletti, R.; Ferrari, S.; De Lorenzo, G. Arabidopsis MPK3 and MPK6 Play Different Roles in Basal and Oligogalacturonide- or Flagellin-Induced Resistance against Botrytis cinerea. Plant Physiol. 2011, 157, 804–814. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Su, T.; Li, P.; Xin, X.; Cao, Y.; Wang, W.; Zhao, X.; Zhang, D.; Yu, Y.; Li, D.; et al. Identification of Long Noncoding RNAs Involved in Resistance to Downy Mildew in Chinese Cabbage. Hortic. Res. 2021, 8, 44. [Google Scholar] [CrossRef]
- Schweighofer, A.; Hirt, H.; Meskiene, I. Plant PP2C Phosphatases: Emerging Functions in Stress Signaling. Trends Plant Sci. 2004, 9, 236–243. [Google Scholar] [CrossRef]
- Brock, A.K.; Willmann, R.; Kolb, D.; Grefen, L.; Lajunen, H.M.; Bethke, G.; Lee, J.; Nürnberger, T.; Gust, A.A. The Arabidopsis Mitogen-Activated Protein Kinase Phosphatase PP2C5 Affects Seed Germination, Stomatal Aperture, and Abscisic Acid-Inducible Gene Expression. Plant Physiol. 2010, 153, 1098–1111. [Google Scholar] [CrossRef]
- Xu, D.; Lin, F.; Jiang, Y.; Huang, X.; Li, J.; Ling, J.; Hettiarachchi, C.; Tellgren-Roth, C.; Holm, M.; Deng, X.W. The RING-Finger E3 Ubiquitin Ligase COP1 SUPPRESSOR1 Negatively Regulates COP1 Abundance in Maintaining COP1 Homeostasis in Dark-Grown Arabidopsis Seedlings. Plant Cell 2014, 26, 1981–1991. [Google Scholar] [CrossRef]
- Lohscheider, J.N.; Friso, G.; Van Wijk, K.J. Phosphorylation of Plastoglobular Proteins in Arabidopsis thaliana. J. Exp. Bot. 2016, 67, 3975–3984. [Google Scholar] [CrossRef]
- Virk, N.; Li, D.; Tian, L.; Huang, L.; Hong, Y.; Li, X.; Zhang, Y.; Liu, B.; Zhang, H.; Song, F. Arabidopsis Raf-Like Mitogen-Activated Protein Kinase Kinase Kinase Gene Raf43 Is Required for Tolerance to Multiple Abiotic Stresses. PLoS ONE 2015, 10, e0133975. [Google Scholar] [CrossRef] [PubMed]
- Deng, P.; Liu, S.; Nie, X.; Weining, S.; Wu, L. Conservation Analysis of Long Non-Coding RNAs in Plants. Sci. China Life Sci. 2018, 61, 190–198. [Google Scholar] [CrossRef]
- Li, Y.; Liu, K.; Tong, G.; Xi, C.; Liu, J.; Zhao, H.; Wang, Y.; Ren, D.; Han, S. MPK3/MPK6-Mediated Phosphorylation of ERF72 Positively Regulates Resistance to Botrytis cinerea through Directly and Indirectly Activating the Transcription of Camalexin Biosynthesis Enzymes. J. Exp. Bot. 2022, 73, 413–428. [Google Scholar] [CrossRef]
- Chen, T.; Chen, X.; Zhang, S.; Zhu, J.; Tang, B.; Wang, A.; Dong, L.; Zhang, Z.; Yu, C.; Sun, Y.; et al. The Genome Sequence Archive Family: Toward Explosive Data Growth and Diverse Data Types. Genom. Proteom. Bioinform. 2021, 19, 578–583. [Google Scholar] [CrossRef]
- CNCB-NGDC Members and Partners; Bao, Y.; Bai, X.; Bu, C.; Chen, H.; Chen, H.; Chen, K.; Chen, M.; Chen, M.; Chen, M.; et al. Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2025. Nucleic Acids Res. 2025, 53, D30–D44. [Google Scholar] [CrossRef]
- Pang, Y.; Zheng, K.; Min, Q.; Wang, Y.; Xue, X.; Li, W.; Zhao, H.; Qiao, F.; Han, S. Long Noncoding RNAs in Response to Hyperosmolarity Stress, but Not Salt Stress, Were Mainly Enriched in the Rice Roots. Int. J. Mol. Sci. 2024, 25, 6226. [Google Scholar] [CrossRef]
- Zheng, K.; Wu, X.; Xue, X.; Li, W.; Wang, Z.; Chen, J.; Zhang, Y.; Qiao, F.; Zhao, H.; Zhang, F.; et al. Transcriptome Screening of Long Noncoding RNAs and Their Target Protein-Coding Genes Unmasks a Dynamic Portrait of Seed Coat Coloration Associated with Anthocyanins in Tibetan Hulless Barley. Int. J. Mol. Sci. 2023, 24, 10587. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Zheng, K.; Long, W.; Zhao, L.; Li, W.; Xue, X.; Han, S. Cytosolic and Nucleosolic Calcium-Regulated Long Non-Coding RNAs and Their Target Protein-Coding Genes in Response to Hyperosmolarity and Salt Stresses in Arabidopsis Thaliana. Int. J. Mol. Sci. 2025, 26, 2086. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Osak, M.; Bogu, G.K.; Stanton, L.W.; Johnson, R.; Lipovich, L. Genome-Wide Computational Identification and Manual Annotation of Human Long Noncoding RNA Genes. RNA 2010, 16, 1478–1487. [Google Scholar] [CrossRef]
- Zhou, Z.; Windhorst, A.; Schenke, D.; Cai, D. RNAseq-Based Working Model for Transcriptional Regulation of Crosstalk between Simultaneous Abiotic UV-B and Biotic Stresses in Plants. Genes 2023, 14, 240. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Wilcox, R.R. Two-way ANOVA: Inferences about Interactions Based on Robust Measures of Effect Size. Br. J. Math. Stat. Psychol. 2022, 75, 46–58. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, T.; Zheng, K.; Min, Q.; Li, Y.; Xue, X.; Li, W.; Han, S. Transcriptomic Identification of Long Noncoding RNAs Modulating MPK3/MPK6-Centered Immune Networks in Arabidopsis. Int. J. Mol. Sci. 2025, 26, 8331. https://doi.org/10.3390/ijms26178331
Wang T, Zheng K, Min Q, Li Y, Xue X, Li W, Han S. Transcriptomic Identification of Long Noncoding RNAs Modulating MPK3/MPK6-Centered Immune Networks in Arabidopsis. International Journal of Molecular Sciences. 2025; 26(17):8331. https://doi.org/10.3390/ijms26178331
Chicago/Turabian StyleWang, Tianjiao, Kaifeng Zheng, Qinyue Min, Yihao Li, Xiuhua Xue, Wanjie Li, and Shengcheng Han. 2025. "Transcriptomic Identification of Long Noncoding RNAs Modulating MPK3/MPK6-Centered Immune Networks in Arabidopsis" International Journal of Molecular Sciences 26, no. 17: 8331. https://doi.org/10.3390/ijms26178331
APA StyleWang, T., Zheng, K., Min, Q., Li, Y., Xue, X., Li, W., & Han, S. (2025). Transcriptomic Identification of Long Noncoding RNAs Modulating MPK3/MPK6-Centered Immune Networks in Arabidopsis. International Journal of Molecular Sciences, 26(17), 8331. https://doi.org/10.3390/ijms26178331






