Effects of Temperature and Salinity on Ovarian Development and Differences in Energy Metabolism Between Reproduction and Growth During Ovarian Development in the Lateolabrax maculatus
Abstract
1. Introduction
2. Results
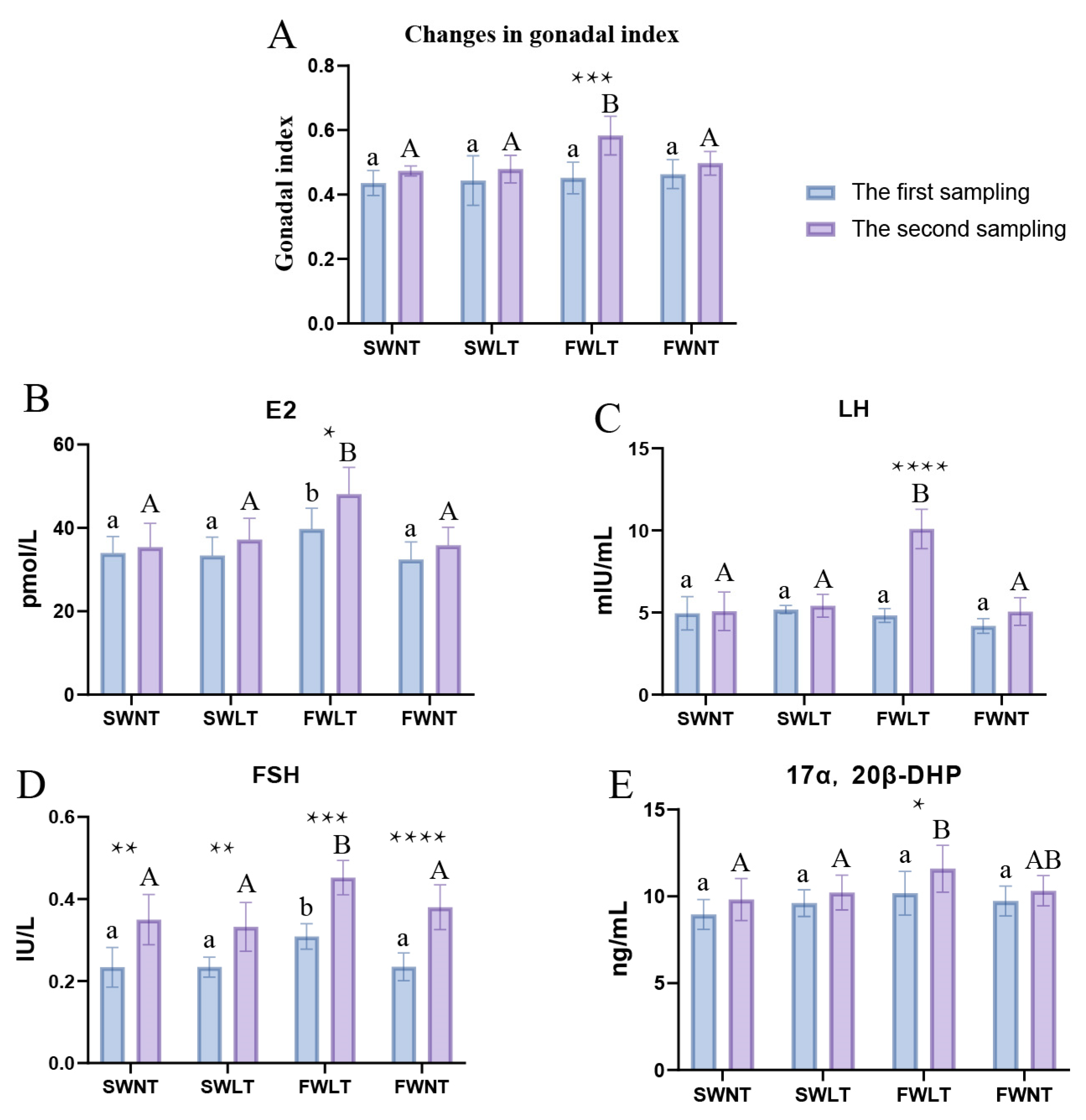
2.1. Analysis of the Effect of Temperature and Salinity on Ovarian Development
2.2. Effects of Temperature and Salinity on Reproduction-Related Hormones in Lateolabrax Maculatus
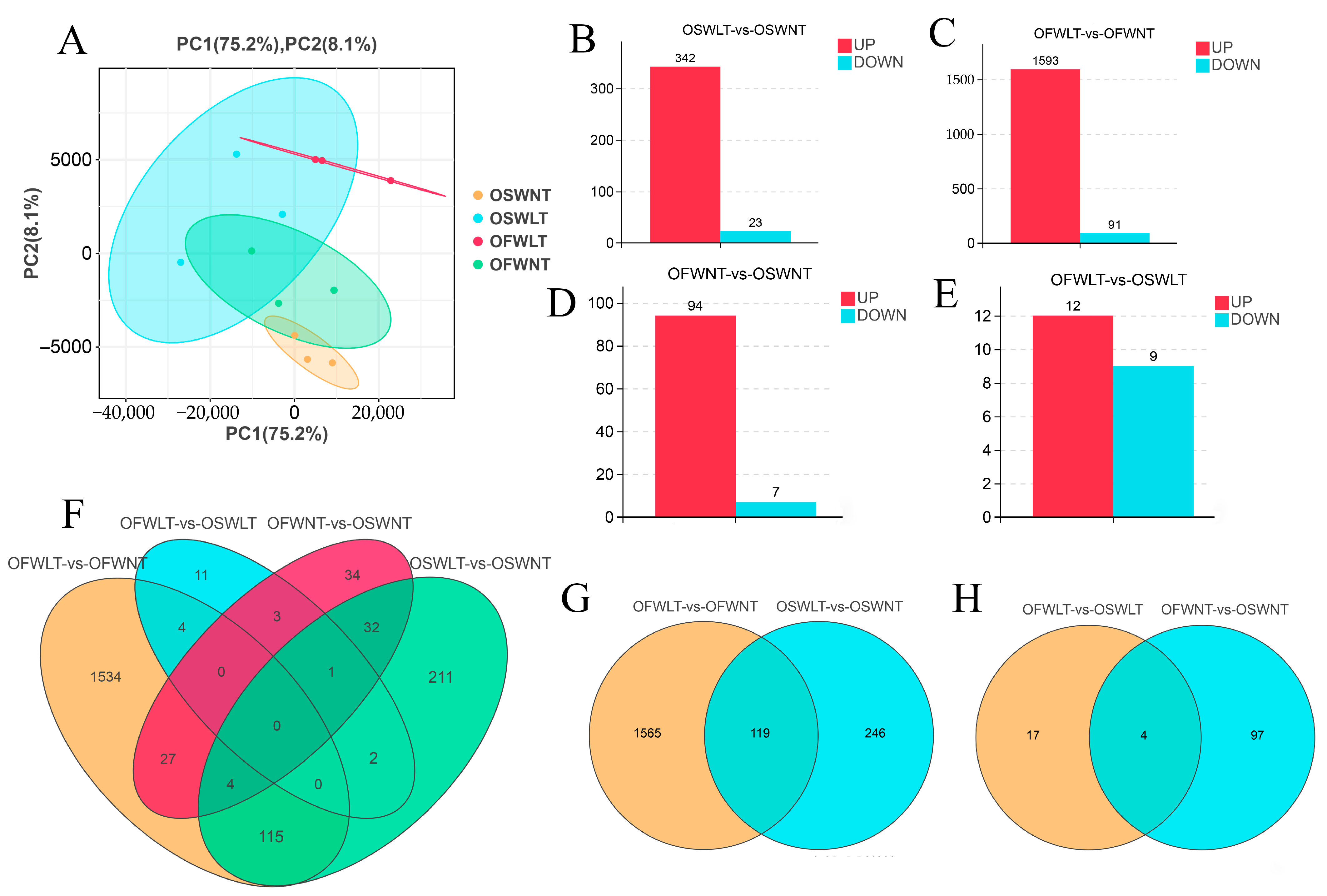
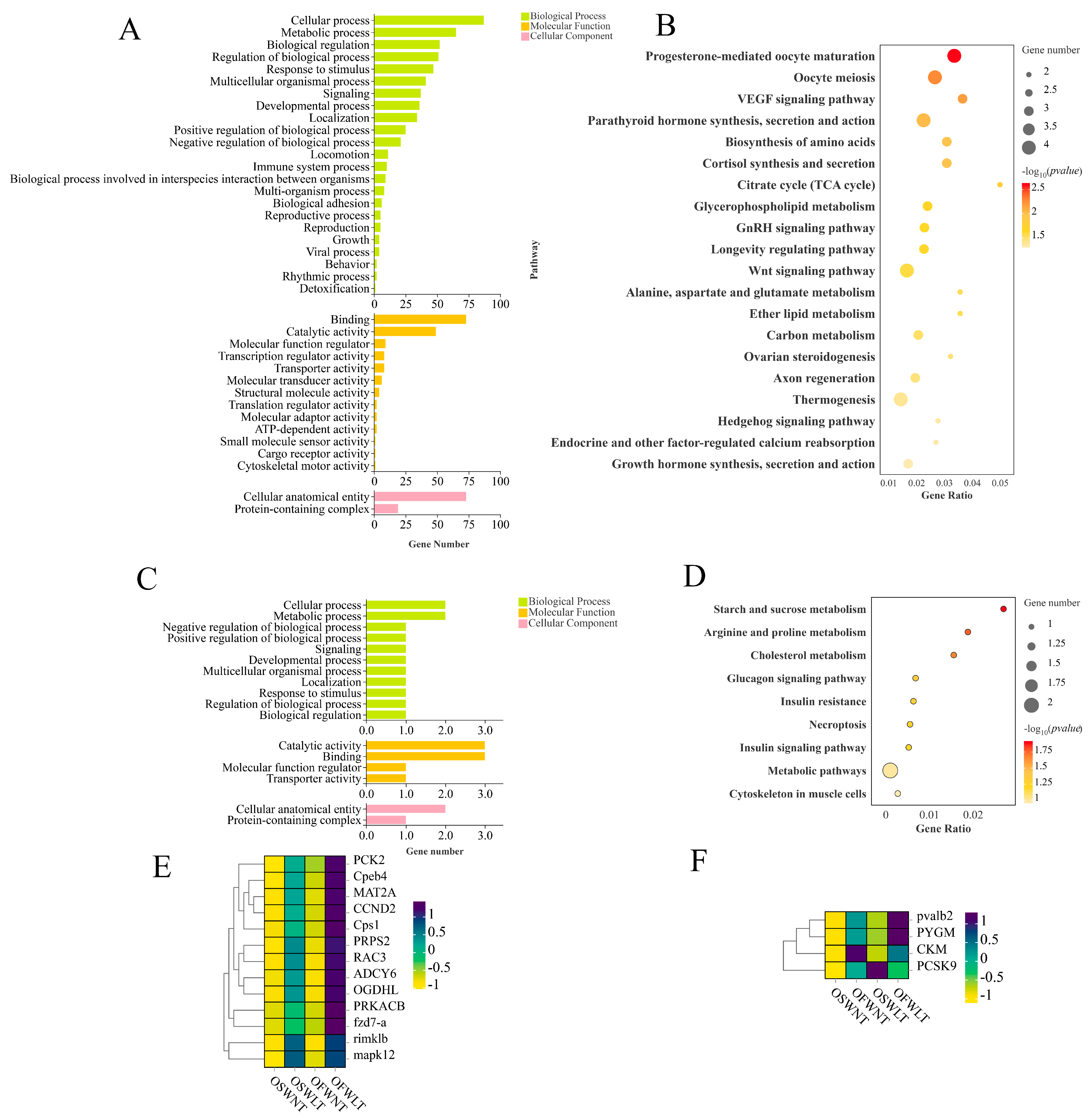
2.3. Transcriptome Analysis of Ovaries at Different Temperatures and Salinities
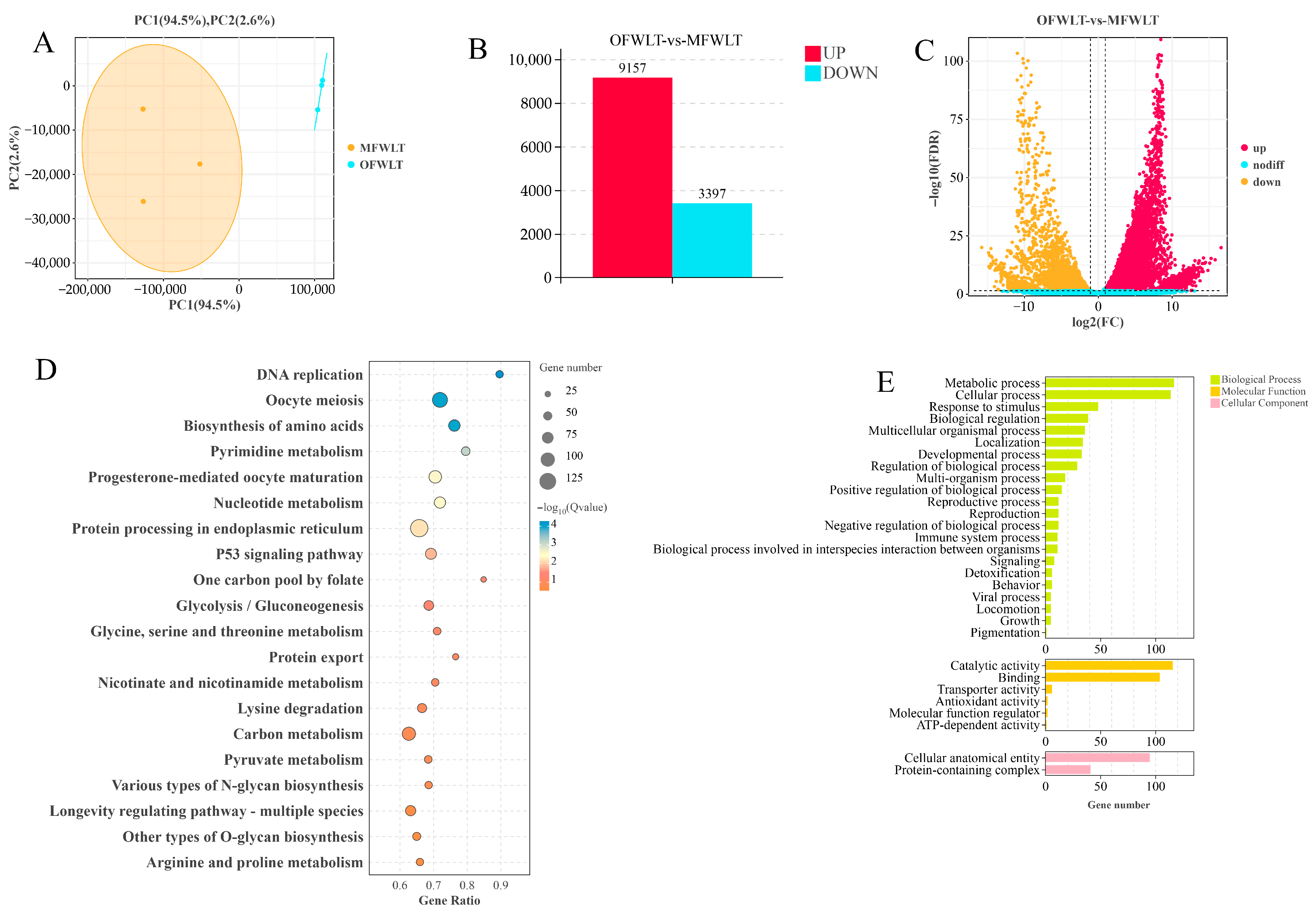
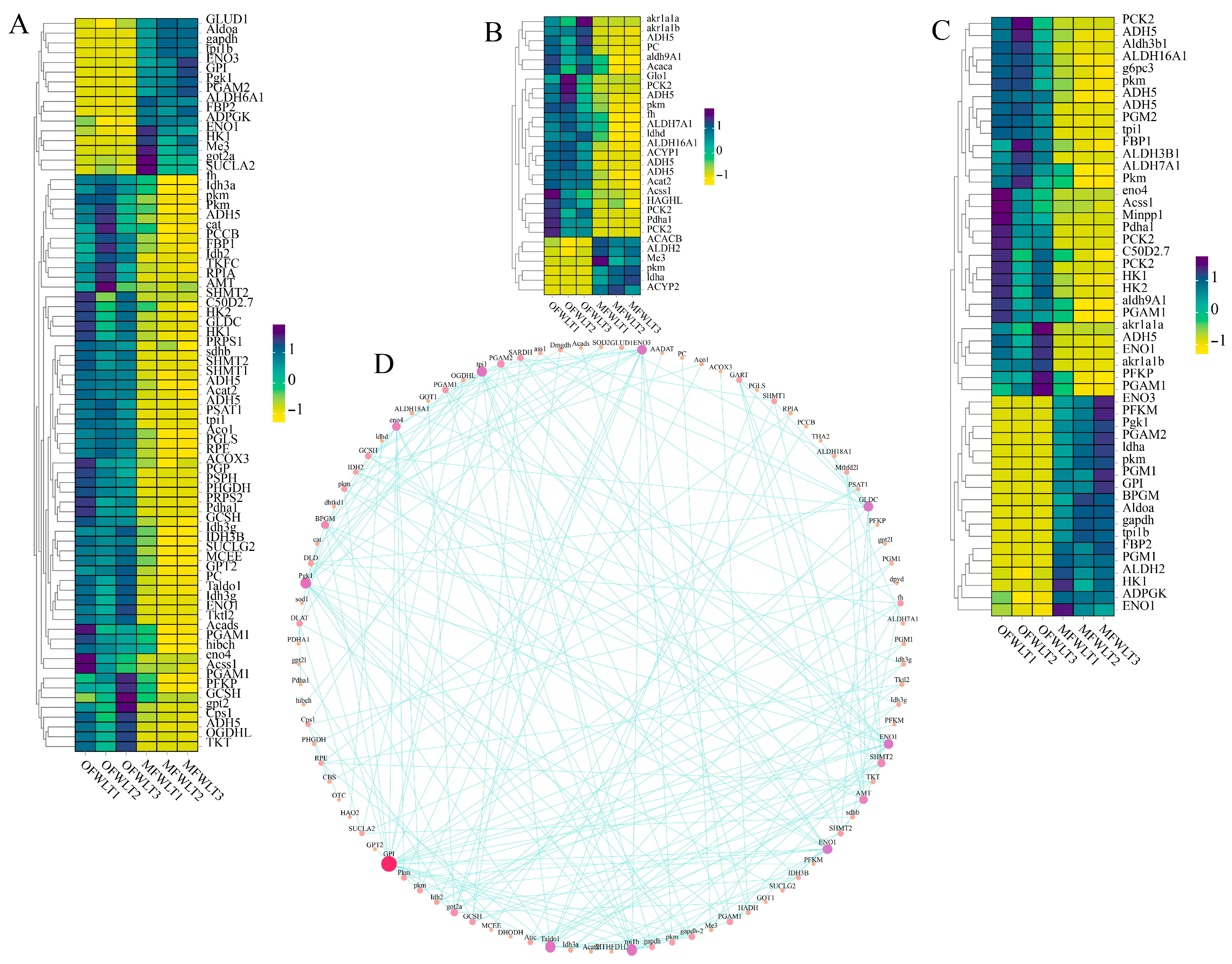
2.4. Transcriptome and Energy Metabolism Analysis of Ovarian and Muscle Tissues During Ovarian Development
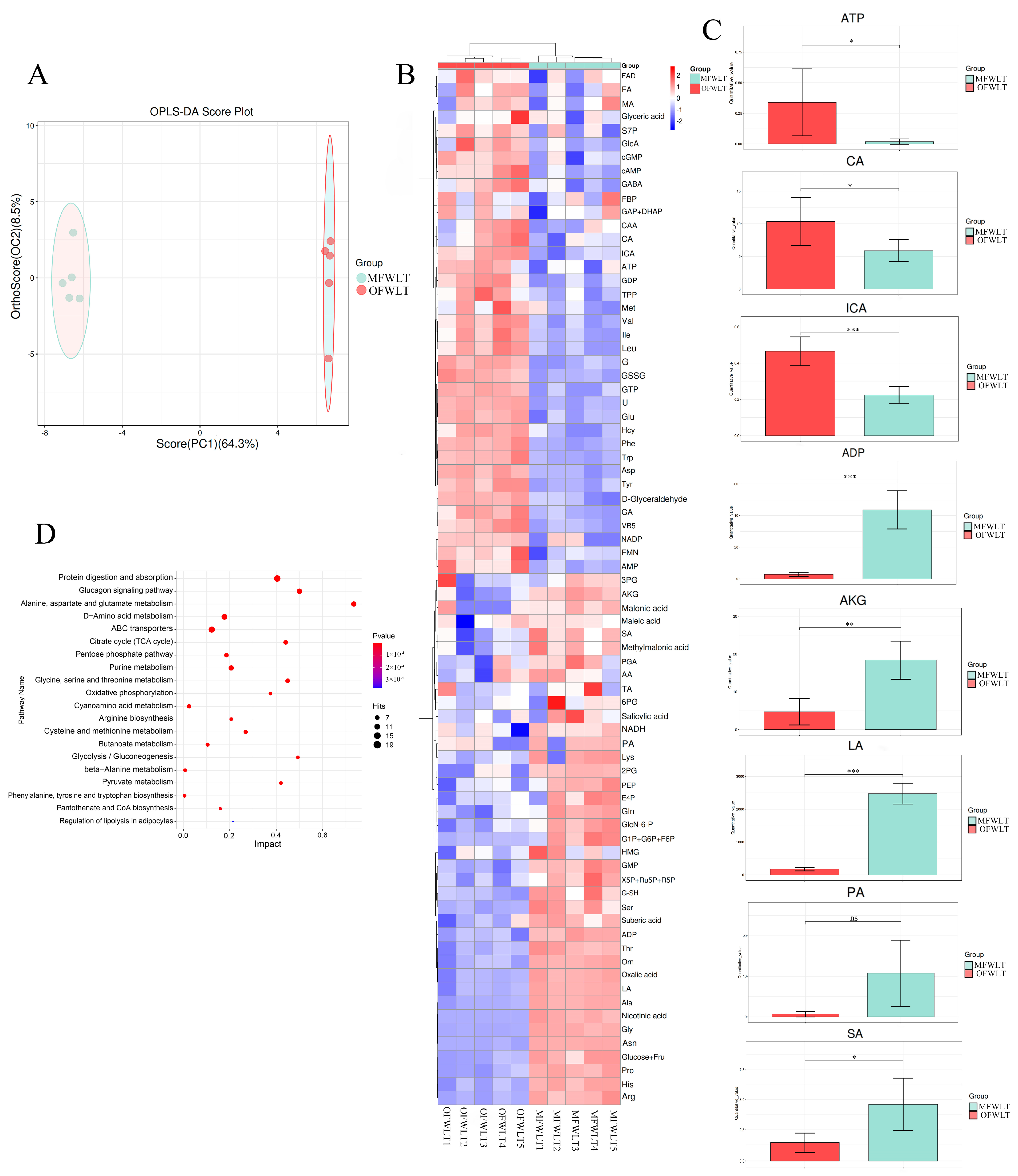
2.5. Combined Transcriptome and Energy Metabolome Analysis
3. Discussion
3.1. Analysis of the Effects of Temperature and Salinity on Ovarian Development
3.2. Analysis of the Effects of Temperature and Salinity on the Ovarian Transcriptome
3.2.1. Analysis of the Effect of Temperature on the Ovary Transcriptome
3.2.2. Effect of Salinity on Ovarian Transcriptome
3.3. Transcriptome and Energy Metabolome Analysis of Gonadal and Muscle Tissues During Gonadal Development
3.3.1. Transcriptome Analysis of Gonadal and Muscle Tissue
3.3.2. Analysis of Gonadal and Muscle Tissue-Targeted Energy Metabolism
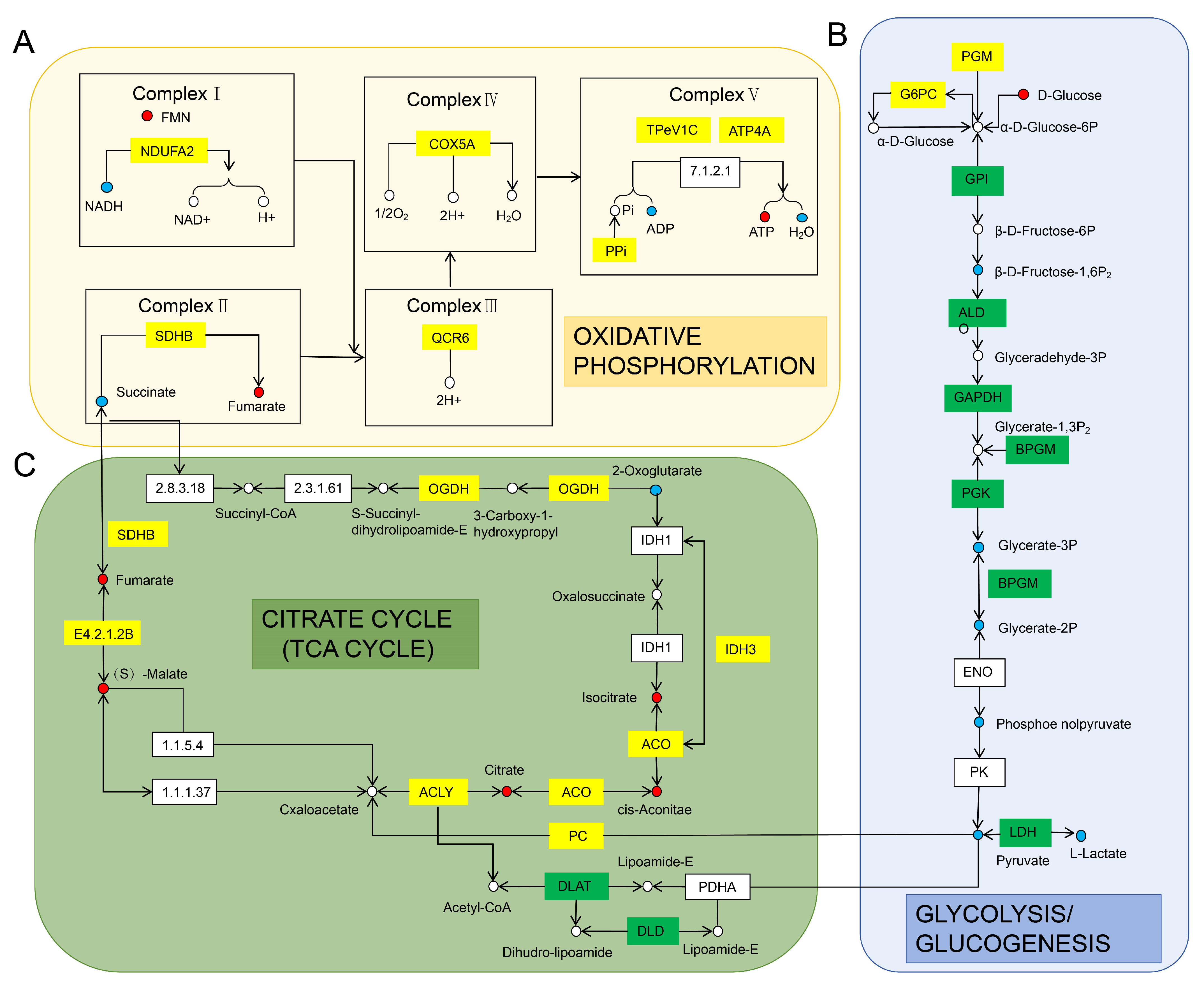
3.3.3. Transcriptome and Targeted Energy Metabolome Association Analysis of Gonadal and Muscle Tissues
4. Materials and Methods
4.1. Ethics Statement
4.2. Experimental Animals and Sample Collection
4.3. Calculation of Gonadal Index
4.4. Detection of Reproduction-Related Hormones
4.5. RNA Extraction and Transcriptome Analysis
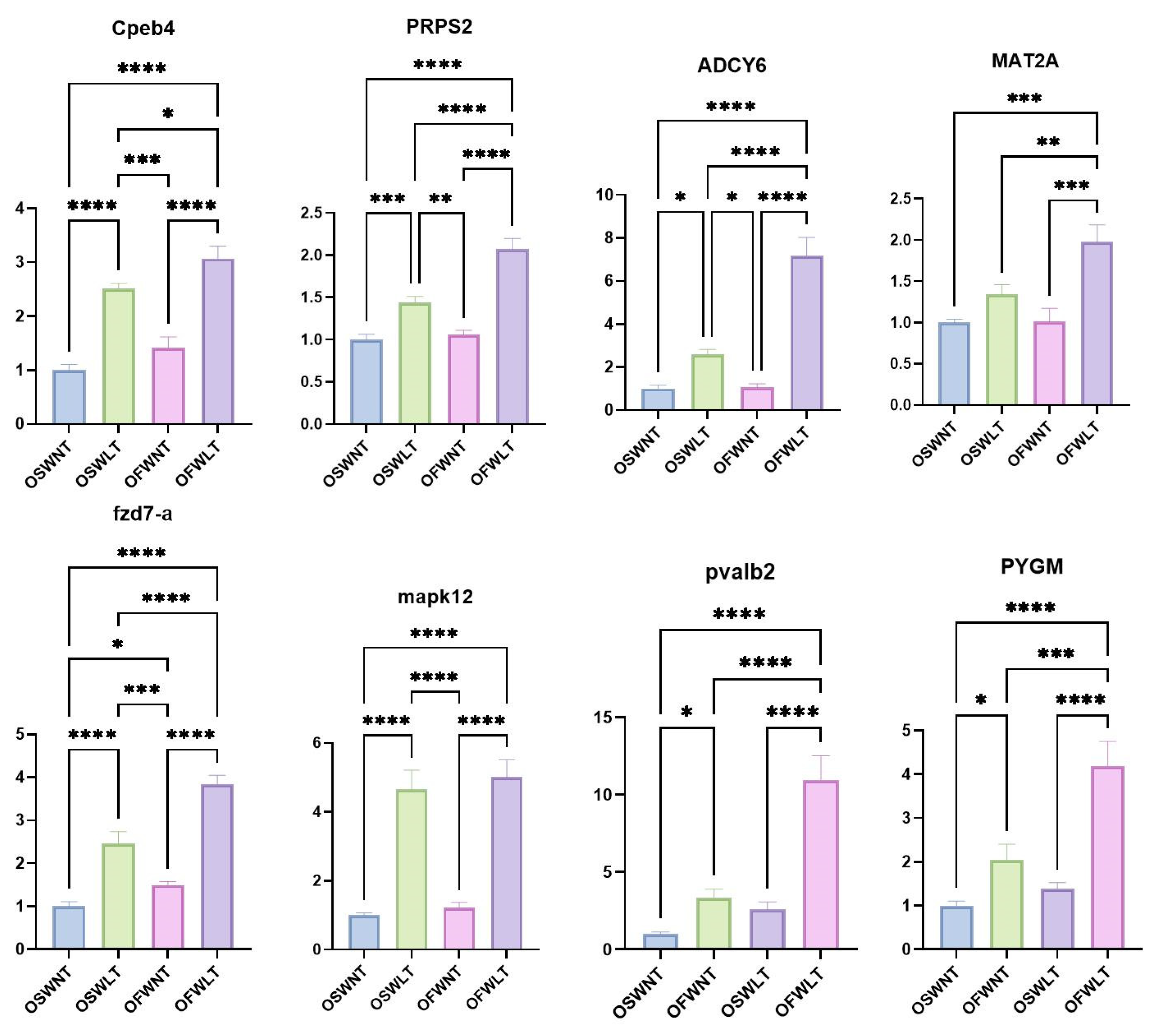
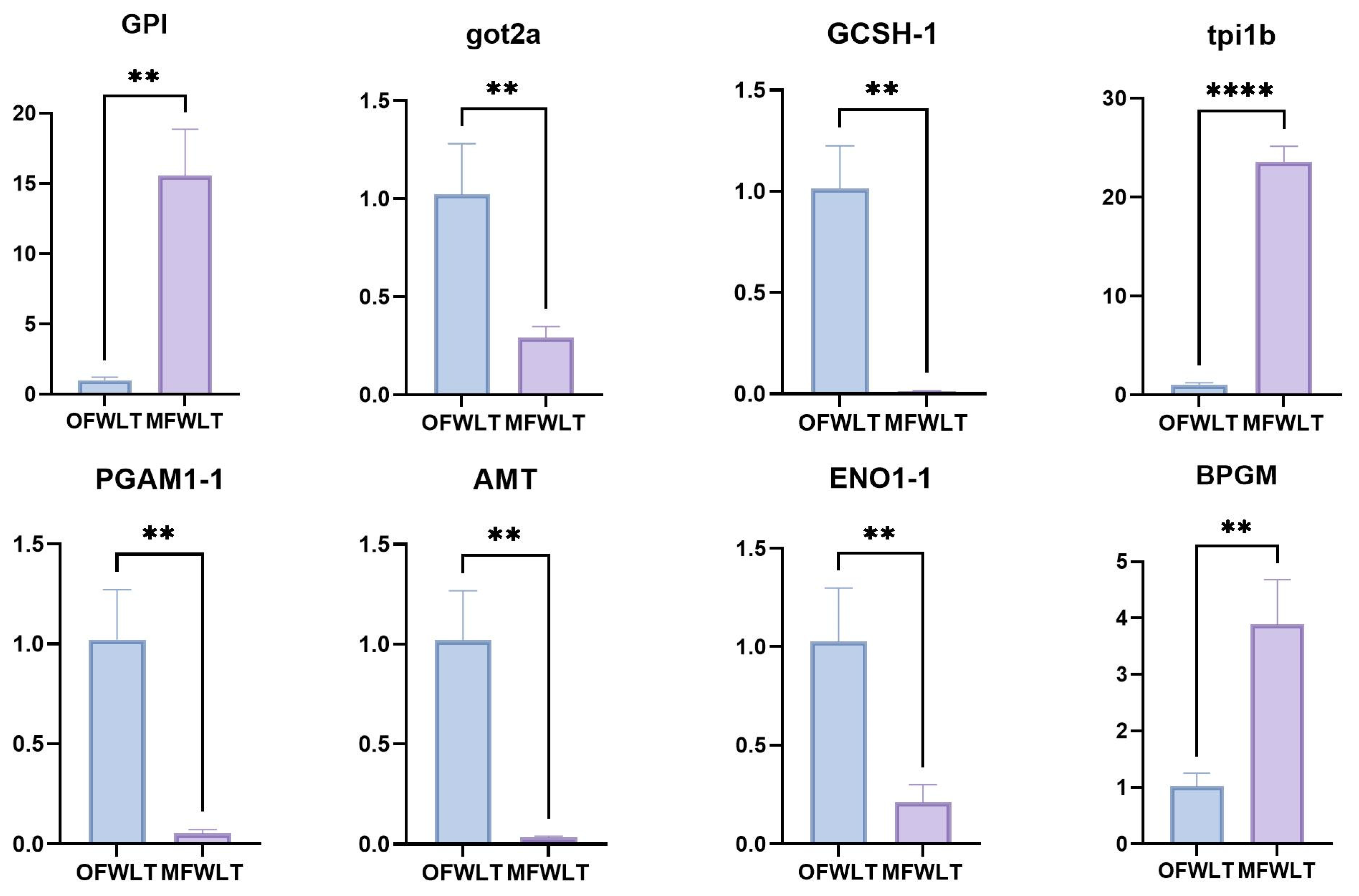
4.6. RT-qPCR Validation of Differentially Expressed Genes
4.7. Targeted Energy Metabolomics Analysis
4.8. Combined Transcriptome and Energy Metabolome Analysis
4.9. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| Lateolabrax maculatus | L. maculatus |
| Gonadosomatic index | GSI |
| Estradiol | E2 |
| Luteinizing hormone | LH |
| Follicle-stimulating hormone | FSH |
| 17α,20β-dihydroxyprogesterone | 17α,20β-DHP |
| Gonadotropin-releasing hormone | GnRH |
| Real-time quantitative reverse transcription PCR | RT-qPCR |
| Isocitrate | ICA |
| Cis aconitic acid | CAA |
| Citric acid | CA |
| Malic acid | MA |
| Fumaric acid | FA |
| Succinic acid | SA |
| Pyruvic acid | PA |
| α-Ketoglutarate | AKG |
| Flavin mononucleotide | FMN |
| Beta-D-fructose 1,6-bisphosphate | BDFP |
| Final oocyte maturation | FOM |
References
- Schulte, P.M.; Healy, T.M.; Fangue, N.A. Thermal performance curves, phenotypic plasticity, and the time scales of temperature exposure. Integr. Comp. Biol. 2011, 51, 691–702. [Google Scholar] [CrossRef]
- Pankhurst, N.W.; Thomas, P.M. Maintenance at elevated temperature delays the steroidogenic and ovulatory responsiveness of rainbow trout Oncorhynchus mykiss to luteinizing hormone releasing hormone analogue. Aquaculture 1998, 166, 163–177. [Google Scholar] [CrossRef]
- Zohar, Y.; Muñoz-Cueto, J.A.; Elizur, A.; Kah, O. Neuroendocrinology of reproduction in teleost fish. Gen. Comp. Endocrinol. 2010, 165, 438–455. [Google Scholar] [CrossRef]
- Lethimonier, C.; Madigou, T.; Muñoz-Cueto, J.A.; Lareyre, J.J.; Kah, O. Evolutionary aspects of GnRHs, GnRH neuronal systems and GnRH receptors in teleost fish. Gen. Comp. Endocrinol. 2004, 135, 1–16. [Google Scholar] [CrossRef]
- Pankhurst, N.; Munday, P. Effects of climate change on fish reproduction and early life history stages. Mar. Freshw. Res. 2011, 62, 1015–1026. [Google Scholar] [CrossRef]
- Alix, M.; Kjesbu, O.S.; Anderson, K.C. From gametogenesis to spawning: How climate-driven warming affects teleost reproductive biology. J. Fish. Biol. 2020, 97, 607–632, Erratum in J. Fish. Biol. 2021, 98, 1494. [Google Scholar] [CrossRef] [PubMed]
- Donelson, J.; Munday, P.; McCormick, M.; Pankhurst, N.; Pankhurst, P. Effects of elevated water temperature and food on the reproductive performance of a coral reef fish. Mar. Ecol. Prog. Ser. 2010, 401, 233–243. [Google Scholar] [CrossRef]
- Donelson, J.M.; McCormick, M.I.; Booth, D.J.; Munday, P.L. Reproductive Acclimation to Increased Water Temperature in a Tropical Reef Fish. PLoS ONE 2014, 9, e97223. [Google Scholar] [CrossRef] [PubMed]
- Gillet, C. Egg production in an Arctic charr (Salvelinus alpinus L.) brood stock: Effects of temperature on the timing of spawning and the quality of eggs. Aquat. Living Resour. 1991, 4, 109–116. [Google Scholar] [CrossRef]
- Pankhurst, N.W.; Purser, G.J.; Van Der Kraak, G.; Thomas, P.M.; Forteath, G.N.R. Effect of holding temperature on ovulation, egg fertility, plasma levels of reproductive hormones and in vitro ovarian steroidogenesis in the rainbow trout Oncorhynchus mykiss. Aquaculture 1996, 146, 277–290. [Google Scholar] [CrossRef]
- Taranger, G.L.; Hansen, T. Ovulation and egg survival following exposure of Atlantic salmon, Salmo salar L., broodstock to different water temperatures. Aquac. Res. 1993, 24, 151–156. [Google Scholar] [CrossRef]
- Lema, S.C.; Chow, M.I.; Dittman, A.H.; May, D.; Housh, M.J. Accustomed to the heat: Temperature and thyroid hormone influences on oogenesis and gonadal steroidogenesis pathways vary among populations of Amargosa pupfish (Cyprinodon nevadensis amargosae). Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2022, 272, 111280. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.K.; Yang, S.W. Relationship between ovarian development and serum levels of gonadal steroid hormones, and induction of oocyte maturation and ovulation in the cultured female Korean spotted sea bass Lateolabrax maculatus (Jeom-nong-eo). Aquaculture 2002, 207, 169–183. [Google Scholar] [CrossRef]
- Watts, M.; Pankhurst, N.W.; King, H.R. Maintenance of Atlantic salmon (Salmo salar) at elevated temperature inhibits cytochrome P450 aromatase activity in isolated ovarian follicles. Gen. Comp. Endocrinol. 2004, 135, 381–390. [Google Scholar] [CrossRef] [PubMed]
- Lema, S.C.; Luckenbach, J.A.; Yamamoto, Y.; Housh, M.J. Fish reproduction in a warming world: Vulnerable points in hormone regulation from sex determination to spawning. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2024, 379, 20220516. [Google Scholar] [CrossRef]
- Lubzens, E.; Young, G.; Bobe, J.; Cerdà, J. Oogenesis in teleosts: How eggs are formed. Gen. Comp. Endocrinol. 2010, 165, 367–389. [Google Scholar] [CrossRef]
- Bock, S.L.; Chow, M.I.; Forsgren, K.L.; Lema, S.C. Widespread alterations to hypothalamic-pituitary-gonadal (HPG) axis signaling underlie high temperature reproductive inhibition in the eurythermal sheepshead minnow (Cyprinodon variegatus). Mol. Cell. Endocrinol. 2021, 537, 111447. [Google Scholar] [CrossRef]
- Lahnsteiner, F.; Kletzl, M. The effect of water temperature on gamete maturation and gamete quality in the European grayling (Thymalus thymallus) based on experimental data and on data from wild populations. Fish Physiol. Biochem. 2012, 38, 455–467. [Google Scholar] [CrossRef]
- Beuvard, C.; Imsland, A.K.D.; Thorarensen, H. The effect of temperature on growth performance and aerobic metabolic scope in Arctic charr, Salvelinus alpinus (L.). J. Therm. Biol. 2022, 104, 103117. [Google Scholar] [CrossRef]
- Cruz Vieira, A.B.; Weber, A.A.; Ribeiro, Y.M.; Luz, R.K.; Bazzoli, N.; Rizzo, E. Influence of salinity on spermatogenesis in adult Nile tilapia (Oreochromis niloticus) testis. Theriogenology 2019, 131, 1–8. [Google Scholar] [CrossRef]
- Xue, Y.; Jiang, G.; Shu, H.; Wang, W.; Huang, X. Effects of Temperature and Salinity on the Growth, Reproduction, and Carotenoid Accumulation in Artemia sinica and Transcriptome Analysis. Fishes 2024, 9, 437. [Google Scholar] [CrossRef]
- Pham, H.Q.; Kjørsvik, E.; Nguyen, A.T.; Nguyen, M.D.; Arukwe, A. Reproductive cycle in female Waigieu seaperch (Psammoperca waigiensis) reared under different salinity levels and the effects of dopamine antagonist on steroid hormone levels. J. Exp. Mar. Biol. Ecol. 2010, 383, 137–145. [Google Scholar] [CrossRef]
- Haffray, P.; Fostier, A.; Normant, Y.; Fauré, A.; Loir, M.; Jalabert, B.; Maisse, G.; Le Gac, F. Influence du maintien en mer ou de la période du transfert en eau douce des reproducteurs de saumon Atlantique (Salmo salar) sur la maturation sexuelle et la qualité des gamètes. Aquat. Living Resour. 1995, 8, 135–145. [Google Scholar] [CrossRef]
- Sower, S.A.; Schreck, C.B.; Donaldson, E.M. Hormone-induced Ovulation of Coho Salmon (Oncorhynchus kisutch) Held in Seawater and Fresh Water. Can. J. Fish. Aquat. Sci. 1982, 39, 627–632. [Google Scholar] [CrossRef]
- Haddy, J.A.; Pankhurst, N.W. The effects of salinity on reproductive development, plasma steroid levels, fertilisation and egg survival in black bream Acanthopagrus butcheri. Aquaculture 2000, 188, 115–131. [Google Scholar] [CrossRef]
- Chu, Y.T.; Yeh, S.L.; Hseu, J.R.; Tin, Y.Y. Influence of salin- ity on gonadal development of black porgy, Acanthopagrus schlegeli, during final maturation. J. Taiwan Fish. Res. 2004, 12, 33–38. [Google Scholar]
- Ladisa, C.; Ma, Y.; Habibi, H.R. Metabolic Changes During Growth and Reproductive Phases in the Liver of Female Goldfish (Carassius auratus). Front. Cell Dev. Biol. 2022, 10, 834688. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Li, X.; Li, J.; Li, Y. Insights Into Energy Accumulation and Allocation Strategy of Reproductive Migration of Black Amur Bream (Megalobrama terminalis) in the Pearl River Basin, China. Front. Ecol. Evol. 2022, 10, 848228. [Google Scholar] [CrossRef]
- Doucett, R.R.; Booth, R.K.; Power, G.; McKinley, R.S. Effects of the spawning migration on the nutritional status of anadromous Atlantic salmon (Salmo salar): Insights from stable-isotope analysis. Can. J. Fish. Aquat. Sci. 1999, 56, 2172–2180. [Google Scholar] [CrossRef]
- Kiessling, A.; Lindahl-Kiessling, K.; Kiessling, K.-H. Energy utilization and metabolism in spawning migrating Early Stuart sockeye salmon (Oncorhynchus nerka): The migratory paradox. Can. J. Fish. Aquat. Sci. 2004, 61, 452–465. [Google Scholar] [CrossRef]
- Li, Y.; Li, C.; Fu, Y.; Wang, R.; Yang, Y.; Zhang, M.; Zhang, Y.; Wang, X.; Wang, G.; Jiang, H.; et al. Insulin-like growth factor 1 promotes the gonadal development of Pampus argenteus by regulating energy metabolism. Biol. Reprod. 2023, 109, 227–237. [Google Scholar] [CrossRef]
- Kronberg, J.; Byrne, J.J.; Jansen, J.; Antczak, P.; Hines, A.; Bignell, J.; Katsiadaki, I.; Viant, M.R.; Falciani, F. Modeling the metabolic profile of Mytilus edulis reveals molecular signatures linked to gonadal development, sex and environmental site. Sci. Rep. 2021, 11, 12882. [Google Scholar] [CrossRef]
- Roff, D.A. An Allocation Model of Growth and Reproduction in Fish. Can. J. Fish. Aquat. Sci. 1983, 40, 1395–1404. [Google Scholar] [CrossRef]
- Saborido-Rey, F.; Garabana, D.; Stransky, C.; Melnikov, S.; Shibanov, V. Review of the population structure and ecology of S. mentella in the Irminger sea and adjacent waters. Rev. Fish Biol. Fish. 2004, 14, 455–479. [Google Scholar] [CrossRef]
- Tyler, C.R.; Sumpter, J.P.; Witthames, P.R. The dynamics of oocyte growth during vitellogenesis in the rainbow trout (Oncorhynchus mykiss). Biol. Reprod. 1990, 43, 202–209. [Google Scholar] [CrossRef]
- Varsamos, S.; Nebel, C.; Charmantier, G. Ontogeny of osmoregulation in postembryonic fish: A review. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 2005, 141, 401–429. [Google Scholar] [CrossRef]
- Xiong, Y.; Wang, X.; Dong, S.; Wang, F.; Yang, J.; Zhou, Y. Comparisons of Salinity Adaptation in Terms of Growth, Body Composition, and Energy Budget in Juveniles of Rainbow and Steelhead Trouts (Oncorhynchus mykiss). J. Ocean. Univ. China 2019, 18, 509–518. [Google Scholar] [CrossRef]
- Kang, D.Y.; Kim, H.C.; Im, J.H. Reproduction and Maturation of Sea Bass, Lateolabrax japonicus, after Transportation from Net-Cages to Indoor Tanks. Dev. Reprod. 2021, 25, 157–171. [Google Scholar] [CrossRef]
- Planas, J.V.; Swanson, P. Maturation-associated changes in the response of the salmon testis to the steroidogenic actions of gonadotropins (GTH I and GTH II) in vitro. Biol. Reprod. 1995, 52, 697–704. [Google Scholar] [CrossRef]
- Kawauchi, H.; Suzuki, K.; Itoh, H.; Swanson, P.; Naito, N.; Nagahama, Y.; Nozaki, M.; Nakai, Y.; Itoh, S. The duality of teleost gonadotropins. Fish Physiol. Biochem. 1989, 7, 29–38. [Google Scholar] [CrossRef]
- Swanson, P.; Suzuki, K.; Kawauchi, H.; Dickhoff, W.W. Isolation and characterization of two coho salmon gonadotropins, GTH I and GTH II. Biol. Reprod. 1991, 44, 29–38. [Google Scholar] [CrossRef]
- Kusakabe, M.; Nakamura, I.; Evans, J.; Swanson, P.; Young, G. Changes in mRNAs encoding steroidogenic acute regulatory protein, steroidogenic enzymes and receptors for gonadotropins during spermatogenesis in rainbow trout testes. J. Endocrinol. 2006, 189, 541–554. [Google Scholar] [CrossRef] [PubMed]
- Pan, B.; Chai, J.; Fei, K.; Zheng, T.; Jiang, Y. Dynamic changes in the transcriptome and metabolome of pig ovaries across developmental stages and gestation. BMC Genom. 2024, 25, 1193. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Liu, D.; Shaner, Z.C.; Chen, S.; Hong, W.; Stellwag, E.J. Nuclear progestin receptor (pgr) knockouts in zebrafish demonstrate role for pgr in ovulation but not in rapid non-genomic steroid mediated meiosis resumption. Front. Endocrinol. 2015, 6, 37. [Google Scholar] [CrossRef] [PubMed]
- Tokumoto, T.; Yamaguchi, T.; Ii, S.; Tokumoto, M. In vivo induction of oocyte maturation and ovulation in zebrafish. PLoS ONE 2011, 6, e25206. [Google Scholar] [CrossRef]
- Mathur, P.D.; Zou, J.; Neiswanger, G.; Zhu, D.; Wang, Y.; Almishaal, A.A.; Vashist, D.; Hammond, H.K.; Park, A.H.; Yang, J. Adenylyl cyclase 6 plays a minor role in the mouse inner ear and retina. Sci. Rep. 2023, 13, 7075. [Google Scholar] [CrossRef]
- Nagahama, Y.; Yamashita, M. Regulation of oocyte maturation in fish. Dev. Growth Differ. 2008, 50 (Suppl. 1), S195–S219. [Google Scholar] [CrossRef]
- He, M.; Zhang, T.; Yang, Y.; Wang, C. Mechanisms of Oocyte Maturation and Related Epigenetic Regulation. Front. Cell Dev. Biol. 2021, 9, 654028. [Google Scholar] [CrossRef]
- Pierre, K.J.; Tremblay, J.J. Differential Response of Transcription Factors to Activated Kinases in Steroidogenic and Non-Steroidogenic Cells. Int. J. Mol. Sci. 2022, 23, 13153. [Google Scholar] [CrossRef]
- Igea, A.; Méndez, R. Meiosis requires a translational positive loop where CPEB1 ensues its replacement by CPEB4. Embo J. 2010, 29, 2182–2193. [Google Scholar] [CrossRef]
- Chen, L.; Jiang, J.C.; Dai, X.X.; Fan, H.Y. Function and molecular mechanism of mitogen-activated protein kinase (MAPK) in regulating oocyte meiotic maturation and ovulation. Sheng Li Xue Bao 2020, 72, 48–62. [Google Scholar]
- Yuan, J.; Xu, B.Z.; Qi, S.T.; Tong, J.S.; Wei, L.; Li, M.; Ouyang, Y.C.; Hou, Y.; Schatten, H.; Sun, Q.Y. MAPK-activated protein kinase 2 is required for mouse meiotic spindle assembly and kinetochore-microtubule attachment. PLoS ONE 2010, 5, e11247. [Google Scholar] [CrossRef]
- Yang, Y.; Workman, S.; Wilson, M. The molecular pathways underlying early gonadal development. J. Mol. Endocrinol. 2018, 62, R47–R64. [Google Scholar] [CrossRef] [PubMed]
- Qian, X.; Li, X.; Tan, L.; Lee, J.H.; Xia, Y.; Cai, Q.; Zheng, Y.; Wang, H.; Lorenzi, P.L.; Lu, Z. Conversion of PRPS Hexamer to Monomer by AMPK-Mediated Phosphorylation Inhibits Nucleotide Synthesis in Response to Energy Stress. Cancer Discov. 2018, 8, 94–107. [Google Scholar] [CrossRef]
- Qiao, H.; Tan, X.; Lv, D.J.; Xing, R.W.; Shu, F.P.; Zhong, C.F.; Li, C.; Zou, Y.G.; Mao, X.M. Phosphoribosyl pyrophosphate synthetases 2 knockdown inhibits prostate cancer progression by suppressing cell cycle and inducing cell apoptosis. J. Cancer 2020, 11, 1027–1037. [Google Scholar] [CrossRef]
- Li, C.; Gui, G.; Zhang, L.; Qin, A.; Zhou, C.; Zha, X. Overview of Methionine Adenosyltransferase 2A (MAT2A) as an Anticancer Target: Structure, Function, and Inhibitors. J. Med. Chem. 2022, 65, 9531–9547. [Google Scholar] [CrossRef] [PubMed]
- de Cima, S.; Polo, L.M.; Díez-Fernández, C.; Martínez, A.I.; Cervera, J.; Fita, I.; Rubio, V. Structure of human carbamoyl phosphate synthetase: Deciphering the on/off switch of human ureagenesis. Sci. Rep. 2015, 5, 16950. [Google Scholar] [CrossRef]
- Guo, D.; He, H.; Meng, Y.; Luo, S.; Lu, Z. Determiners of cell fates: The tricarboxylic acid cycle versus the citrate-malate shuttle. Protein Cell 2023, 14, 162–164. [Google Scholar] [CrossRef] [PubMed]
- Fernie, A.R.; Carrari, F.; Sweetlove, L.J. Respiratory metabolism: Glycolysis, the TCA cycle and mitochondrial electron transport. Curr. Opin. Plant Biol. 2004, 7, 254–261. [Google Scholar] [CrossRef]
- Inigo, M.; Deja, S.; Burgess, S.C. Ins and Outs of the TCA Cycle: The Central Role of Anaplerosis. Annu. Rev. Nutr. 2021, 41, 19–47. [Google Scholar] [CrossRef]
- Bunik, V.I.; Degtyarev, D. Structure-function relationships in the 2-oxo acid dehydrogenase family: Substrate-specific signatures and functional predictions for the 2-oxoglutarate dehydrogenase-like proteins. Proteins 2008, 71, 874–890. [Google Scholar] [CrossRef]
- Arnold, P.K.; Finley, L.W.S. Regulation and function of the mammalian tricarboxylic acid cycle. J. Biol. Chem. 2023, 299, 102838. [Google Scholar] [CrossRef]
- Yu, S.; Meng, S.; Xiang, M.; Ma, H. Phosphoenolpyruvate carboxykinase in cell metabolism: Roles and mechanisms beyond gluconeogenesis. Mol. Metab. 2021, 53, 101257. [Google Scholar] [CrossRef]
- Stark, R.; Guebre-Egziabher, F.; Zhao, X.; Feriod, C.; Dong, J.; Alves, T.C.; Ioja, S.; Pongratz, R.L.; Bhanot, S.; Roden, M.; et al. A role for mitochondrial phosphoenolpyruvate carboxykinase (PEPCK-M) in the regulation of hepatic gluconeogenesis. J. Biol. Chem. 2014, 289, 7257–7263. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Zhou, Y.; Ding, N.; Fang, Y. Recent Advances in Metabolic Engineering for the Biosynthesis of Phosphoenol Pyruvate-Oxaloacetate-Pyruvate-Derived Amino Acids. Molecules 2024, 29, 2893. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Shi, X.; Shi, Y.; Wang, Z. The Signaling Pathways Involved in Ovarian Follicle Development. Front. Physiol. 2021, 12, 730196. [Google Scholar] [CrossRef]
- Ma, H.; Gao, G.; Palti, Y.; Tripathi, V.; Birkett, J.E.; Weber, G.M. Transcriptomic Response of the Ovarian Follicle Complex in Post-Vitellogenic Rainbow Trout to 17α,20β-Dihdroxy-4-pregnen-3-one In Vitro. Int. J. Mol. Sci. 2024, 25, 12683. [Google Scholar] [CrossRef] [PubMed]
- Anastas, J.N.; Moon, R.T. WNT signalling pathways as therapeutic targets in cancer. Nat. Rev. Cancer 2013, 13, 11–26. [Google Scholar] [CrossRef]
- Steinhart, Z.; Angers, S. Wnt signaling in development and tissue homeostasis. Development 2018, 145, 146589. [Google Scholar] [CrossRef]
- Hino, S.; Tanji, C.; Nakayama, K.I.; Kikuchi, A. Phosphorylation of beta-catenin by cyclic AMP-dependent protein kinase stabilizes beta-catenin through inhibition of its ubiquitination. Mol. Cell. Biol. 2005, 25, 9063–9072. [Google Scholar] [CrossRef] [PubMed]
- King, P.J.; Guasti, L.; Laufer, E. Hedgehog signalling in endocrine development and disease. J. Endocrinol. 2008, 198, 439–450. [Google Scholar] [CrossRef]
- Wijgerde, M.; Ooms, M.; Hoogerbrugge, J.W.; Grootegoed, J.A. Hedgehog signaling in mouse ovary: Indian hedgehog and desert hedgehog from granulosa cells induce target gene expression in developing theca cells. Endocrinology 2005, 146, 3558–3566. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Marin, D.; Stroman, G.C.; Fulton, C.J.; Pruitt, K. Frizzled receptors: Gatekeepers of Wnt signaling in development and disease. Front. Cell Dev. Biol. 2025, 13, 1599355. [Google Scholar] [CrossRef]
- Liu, J.; Xiao, Q.; Xiao, J.; Niu, C.; Li, Y.; Zhang, X.; Zhou, Z.; Shu, G.; Yin, G. Wnt/β-catenin signalling: Function, biological mechanisms, and therapeutic opportunities. Signal Transduct. Target. Ther. 2022, 7, 3. [Google Scholar] [CrossRef]
- Kwon, S.; Lim, H.J. Small GTPases and formins in mammalian oocyte maturation: Cytoskeletal organizers. Clin. Exp. Reprod. Med. 2011, 38, 1–5. [Google Scholar] [CrossRef]
- Gampel, A.; Parker, P.J.; Mellor, H. Regulation of epidermal growth factor receptor traffic by the small GTPase rhoB. Curr. Biol. 1999, 9, 955–958. [Google Scholar] [CrossRef] [PubMed]
- Pang, Y.; Ge, W. Epidermal growth factor and TGFalpha promote zebrafish oocyte maturation in vitro: Potential role of the ovarian activin regulatory system. Endocrinology 2002, 143, 47–54. [Google Scholar] [CrossRef]
- Van Der Kraak, G.; Lister, A.L. The inhibitory control of oocyte maturation in the zebrafish (Danio rerio): The role of the G protein-coupled estrogen receptor and epidermal growth factor. Biol. Reprod. 2011, 85, 6–8. [Google Scholar] [CrossRef]
- Habas, R.; Dawid, I.B.; He, X. Coactivation of Rac and Rho by Wnt/Frizzled signaling is required for vertebrate gastrulation. Genes. Dev. 2003, 17, 295–309. [Google Scholar] [CrossRef]
- Han, Y.; Xia, G.; Tsang, B.K. Regulation of cyclin D2 expression and degradation by follicle-stimulating hormone during rat granulosa cell proliferation in vitro. Biol. Reprod. 2013, 88, 57. [Google Scholar] [CrossRef] [PubMed]
- Sicinski, P.; Donaher, J.L.; Geng, Y.; Parker, S.B.; Gardner, H.; Park, M.Y.; Robker, R.L.; Richards, J.S.; McGinnis, L.K.; Biggers, J.D.; et al. Cyclin D2 is an FSH-responsive gene involved in gonadal cell proliferation and oncogenesis. Nature 1996, 384, 470–474. [Google Scholar] [CrossRef]
- Mill, P.; Mo, R.; Fu, H.; Grachtchouk, M.; Kim, P.C.; Dlugosz, A.A.; Hui, C.C. Sonic hedgehog-dependent activation of Gli2 is essential for embryonic hair follicle development. Genes. Dev. 2003, 17, 282–294. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Yin, Y.L.; Li, D.; Kim, S.W.; Wu, G. Amino acids and immune function. Br. J. Nutr. 2007, 98, 237–252. [Google Scholar] [CrossRef]
- Szlas, A.; Kurek, J.M.; Krejpcio, Z. The Potential of L-Arginine in Prevention and Treatment of Disturbed Carbohydrate and Lipid Metabolism-A Review. Nutrients 2022, 14, 961. [Google Scholar] [CrossRef] [PubMed]
- Rettori, V.; McCann, S.M. Role of nitric oxide and alcohol on gonadotropin release in vitro and in vivo. Ann. N. Y. Acad. Sci. 1998, 840, 185–193. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, J.; Wang, C.; Li, W.; Ge, Q.; Qin, Z.; Li, J.; Li, J. The Responses of the Ovary and Eyestalk in Exopalaemon carinicauda under Low Salinity Stress. Fishes 2022, 7, 365. [Google Scholar] [CrossRef]
- Moghadam-Kia, S.; Oddis, C.V.; Aggarwal, R. Approach to asymptomatic creatine kinase elevation. Clevel. Clin. J. Med. 2016, 83, 37–42. [Google Scholar] [CrossRef]
- McLeish, M.J.; Kenyon, G.L. Relating structure to mechanism in creatine kinase. Crit. Rev. Biochem. Mol. Biol. 2005, 40, 1–20. [Google Scholar] [CrossRef]
- Zhu, Y.; Wu, J.; Leng, X.; Du, H.; Wu, J.; He, S.; Luo, J.; Liang, X.; Liu, H.; Wei, Q.; et al. Metabolomics and gene expressions revealed the metabolic changes of lipid and amino acids and the related energetic mechanism in response to ovary development of Chinese sturgeon (Acipenser sinensis). PLoS ONE 2020, 15, e0235043. [Google Scholar] [CrossRef]
- Horton, J.D.; Cohen, J.C.; Hobbs, H.H. Molecular biology of PCSK9: Its role in LDL metabolism. Trends Biochem. Sci. 2007, 32, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Zhao, D.; Xu, L.; Guo, W.; Nie, L.; Lei, Y.; Long, Y.; Liu, M.; Wang, Y.; Zhang, X.; et al. Role of PCSK9 in lipid metabolic disorders and ovarian dysfunction in polycystic ovary syndrome. Metabolism 2019, 94, 47–58. [Google Scholar] [CrossRef] [PubMed]
- Stocco, D.M. The role of the StAR protein in steroidogenesis: Challenges for the future. J. Endocrinol. 2000, 164, 247–253. [Google Scholar] [CrossRef]
- Rajakumar, A.; Senthilkumaran, B. Expression analysis of cyp11a1 during gonadal development, recrudescence and after hCG induction and sex steroid analog treatment in the catfish, Clarias batrachus. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2014, 176, 42–47. [Google Scholar] [CrossRef] [PubMed]
- Saltiel, A.R.; Kahn, C.R. Insulin signalling and the regulation of glucose and lipid metabolism. Nature 2001, 414, 799–806. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, C.M.; Emanuelli, B.; Kahn, C.R. Critical nodes in signalling pathways: Insights into insulin action. Nat. Rev. Mol. Cell Biol. 2006, 7, 85–96. [Google Scholar] [CrossRef]
- Das, D.; Arur, S. Conserved insulin signaling in the regulation of oocyte growth, development, and maturation. Mol. Reprod. Dev. 2017, 84, 444–459. [Google Scholar] [CrossRef]
- Duleba, A.J.; Spaczynski, R.Z.; Olive, D.L.; Behrman, H.R. Effects of insulin and insulin-like growth factors on proliferation of rat ovarian theca-interstitial cells. Biol. Reprod. 1997, 56, 891–897. [Google Scholar] [CrossRef][Green Version]
- Louhio, H.; Hovatta, O.; Sjöberg, J.; Tuuri, T. The effects of insulin, and insulin-like growth factors I and II on human ovarian follicles in long-term culture. Mol. Hum. Reprod. 2000, 6, 694–698. [Google Scholar] [CrossRef]
- Xu, J.; Bernuci, M.P.; Lawson, M.S.; Yeoman, R.R.; Fisher, T.E.; Zelinski, M.B.; Stouffer, R.L. Survival, growth, and maturation of secondary follicles from prepubertal, young, and older adult rhesus monkeys during encapsulated three-dimensional culture: Effects of gonadotropins and insulin. Reproduction 2010, 140, 685–697. [Google Scholar] [CrossRef]
- Di Mauro, S. Muscle glycogenoses: An overview. Acta Myol. 2007, 26, 35–41. [Google Scholar]
- Katz, A. The role of glycogen phosphorylase in glycogen biogenesis in skeletal muscle after exercise. Sports Med. Health Sci. 2023, 5, 29–33. [Google Scholar] [CrossRef]
- Xu, Y.X.; Zhu, Z.Y.; Lo, L.C.; Wang, C.M.; Lin, G.; Feng, F.; Yue, G.H. Characterization of two parvalbumin genes and their association with growth traits in Asian seabass (Lates calcarifer). Anim. Genet. 2006, 37, 266–268. [Google Scholar] [CrossRef]
- Heizmann, C.W.; Berchtold, M.W.; Rowlerson, A.M. Correlation of parvalbumin concentration with relaxation speed in mammalian muscles. Proc. Natl. Acad. Sci. USA 1982, 79, 7243–7247. [Google Scholar] [CrossRef]
- Wilwert, J.L.; Madhoun, N.M.; Coughlin, D.J. Parvalbumin correlates with relaxation rate in the swimming muscle of sheepshead and kingfish. J. Exp. Biol. 2006, 209 Pt 2, 227–237. [Google Scholar] [CrossRef] [PubMed]
- Wilson, D.F.; Matschinsky, F.M. Metabolic Homeostasis in Life as We Know It: Its Origin and Thermodynamic Basis. Front. Physiol. 2021, 12, 658997. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.; Gupta, N. Glycolysis and Gluconeogenesis. In Fundamentals of Bacterial Physiology and Metabolism; Springer: Singapore, 2021; pp. 267–287. [Google Scholar]
- Yetkin-Arik, B.; Vogels, I.M.C.; Nowak-Sliwinska, P.; Weiss, A.; Houtkooper, R.H.; Van Noorden, C.J.F.; Klaassen, I.; Schlingemann, R.O. The role of glycolysis and mitochondrial respiration in the formation and functioning of endothelial tip cells during angiogenesis. Sci. Rep. 2019, 9, 12608. [Google Scholar] [CrossRef]
- Zhang, X.; Yang, S.; Chen, J.; Su, Z. Unraveling the Regulation of Hepatic Gluconeogenesis. Front. Endocrinol. 2018, 9, 802. [Google Scholar] [CrossRef]
- Zangari, J.; Petrelli, F.; Maillot, B.; Martinou, J.C. The Multifaceted Pyruvate Metabolism: Role of the Mitochondrial Pyruvate Carrier. Biomolecules 2020, 10, 1068. [Google Scholar] [CrossRef]
- Payne, A.H.; Hales, D.B. Overview of steroidogenic enzymes in the pathway from cholesterol to active steroid hormones. Endocr. Rev. 2004, 25, 947–970. [Google Scholar] [CrossRef] [PubMed]
- Estévez-Herrera, J.; Domínguez, N.; Pardo, M.R.; González-Santana, A.; Westhead, E.W.; Borges, R.; Machado, J.D. ATP: The crucial component of secretory vesicles. Proc. Natl. Acad. Sci. USA 2016, 113, E4098–E4106. [Google Scholar] [CrossRef]
- Vakilian, M.; Tahamtani, Y.; Ghaedi, K. A review on insulin trafficking and exocytosis. Gene 2019, 706, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Burnstock, G. Purinergic signalling in endocrine organs. Purinergic Signal 2014, 10, 189–231. [Google Scholar] [CrossRef]
- Novak, I. ATP as a signaling molecule: The exocrine focus. News Physiol. Sci. 2003, 18, 12–17. [Google Scholar] [CrossRef]
- Hoque, S.A.M.; Kawai, T.; Zhu, Z.; Shimada, M. Mitochondrial Protein Turnover Is Critical for Granulosa Cell Proliferation and Differentiation in Antral Follicles. J. Endocr. Soc. 2019, 3, 324–339. [Google Scholar] [CrossRef]
- Hayashi, Y.; Matsui, Y. Metabolic Control of Germline Formation and Differentiation in Mammals. Sex. Dev. 2022, 16, 388–403. [Google Scholar] [CrossRef]
- Chen, W.; Zhang, Z.; Chang, C.; Yang, Z.; Wang, P.; Fu, H.; Wei, X.; Chen, E.; Tan, S.; Huang, W.; et al. A bioenergetic shift is required for spermatogonial differentiation. Cell Discov. 2020, 6, 56. [Google Scholar] [CrossRef]
- Tantama, M.; Martínez-François, J.R.; Mongeon, R.; Yellen, G. Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP ratio. Nat. Commun. 2013, 4, 2550. [Google Scholar] [CrossRef] [PubMed]
- Hamidoghli, A.; Jinho, B.; Seonghun, W.; Seunghan, L.; Dae-Jung, K.; Bai, S.C. A Review on Japanese Eel (Anguilla japonica) Aquaculture, With Special Emphasis on Nutrition. Rev. Fish. Sci. Aquac. 2019, 27, 226–241. [Google Scholar] [CrossRef]
- Granchi, D.; Baldini, N.; Ulivieri, F.M.; Caudarella, R. Role of Citrate in Pathophysiology and Medical Management of Bone Diseases. Nutrients 2019, 11, 2576. [Google Scholar] [CrossRef]
- Kynshi, B.L.; Sachu, M.; Syiem, M.B. Modulation in isocitrate dehydrogenase activity under citrate enrichment affects carbon and nitrogen fixations in the cyanobacterium Nostoc muscorum Meg 1. Biochimie 2021, 186, 94–104. [Google Scholar] [CrossRef]
- Kumari, A. Chapter 2—Citric Acid Cycle. In Sweet Biochemistry; Kumari, A., Ed.; Academic Press: Cambridge, MA, USA, 2018; pp. 7–11. [Google Scholar]
- Patil, N.K.; Bohannon, J.K.; Hernandez, A.; Patil, T.K.; Sherwood, E.R. Regulation of leukocyte function by citric acid cycle intermediates. J. Leukoc. Biol. 2019, 106, 105–117. [Google Scholar] [CrossRef]
- Biswas, S.; Ghosh, S.; Maitra, S. Role of insulin-like growth factor 1 (IGF1) in the regulation of mitochondrial bioenergetics in zebrafish oocytes: Lessons from in vivo and in vitro investigations. Front. Cell Dev. Biol. 2023, 11, 1202693. [Google Scholar] [CrossRef]
- Johnston, I.A. A comparative study of glycolysis in red and white muscles of the trout (Salmo gairdneri) and mirror carp (Cyprinus carpio). J. Fish Biol. 1977, 11, 575–588. [Google Scholar] [CrossRef]
- Kam, J.C.; Milligan, C.L. Fuel use during glycogenesis in rainbow trout (Oncorhynchus mykiss Walbaum) white muscle studied in vitro. J. Exp. Biol. 2006, 209 Pt 5, 871–880. [Google Scholar] [CrossRef]
- Soengas, J.L.; Sanmartín, B.; Barciela, P.; Aldegunde, M.; Rozas, G. Changes in carbohydrate metabolism in domesticated rainbow trout (Oncorhynchus mykiss) related to spermatogenesis. Comp. Biochem. Physiol. Part B Comp. Biochem. 1993, 105, 665–671. [Google Scholar] [CrossRef]
- Kiessling, A.; Larsson, L.; Kiessling, K.H.; Lutes, P.B.; Storebakken, T.; Hung, S.S.S. Spawning induces a shift in energy metabolism from glucose to lipid in rainbow trout white muscle. Fish Physiol. Biochem. 1995, 14, 439–448. [Google Scholar] [CrossRef] [PubMed]
- Ahongo, Y.D.; Le Cam, A.; Montfort, J.; Bugeon, J.; Lefèvre, F.; Rescan, P.Y. Gene expression profiling of trout muscle during flesh quality recovery following spawning. BMC Genom. 2022, 23, 9. [Google Scholar] [CrossRef]
- Martínez-Reyes, I.; Chandel, N.S. Mitochondrial TCA cycle metabolites control physiology and disease. Nat. Commun. 2020, 11, 102. [Google Scholar] [CrossRef] [PubMed]
- Jitrapakdee, S.; St Maurice, M.; Rayment, I.; Cleland, W.W.; Wallace, J.C.; Attwood, P.V. Structure, mechanism and regulation of pyruvate carboxylase. Biochem. J. 2008, 413, 369–387. [Google Scholar] [CrossRef] [PubMed]
- Wellen, K.E.; Thompson, C.B. A two-way street: Reciprocal regulation of metabolism and signalling. Nat. Rev. Mol. Cell Biol. 2012, 13, 270–276. [Google Scholar] [CrossRef]
- Zaidi, N.; Swinnen, J.V.; Smans, K. ATP-citrate lyase: A key player in cancer metabolism. Cancer Res. 2012, 72, 3709–3714. [Google Scholar] [CrossRef]
- Lunt, S.Y.; Vander Heiden, M.G. Aerobic glycolysis: Meeting the metabolic requirements of cell proliferation. Annu. Rev. Cell Dev. Biol. 2011, 27, 441–464. [Google Scholar] [CrossRef] [PubMed]
- Ying, W. NAD+/NADH and NADP+/NADPH in cellular functions and cell death: Regulation and biological consequences. Antioxid. Redox Signal 2008, 10, 179–206. [Google Scholar] [CrossRef] [PubMed]
- TeSlaa, T.; Ralser, M.; Fan, J.; Rabinowitz, J.D. The pentose phosphate pathway in health and disease. Nat. Metab. 2023, 5, 1275–1289. [Google Scholar] [CrossRef] [PubMed]
- Sutton-McDowall, M.L.; Gilchrist, R.B.; Thompson, J.G. The pivotal role of glucose metabolism in determining oocyte developmental competence. Reproduction 2010, 139, 685–695. [Google Scholar] [CrossRef]
| Sample | Raw Data | Clean Data (%) | BF_Q30 (%) | BF_GC (%) |
|---|---|---|---|---|
| OSWNT1 | 8,444,406,300 | 832,965,228 (98.64%) | 7,787,987,434 (92.23%) | 4,307,011,897 (51.00%) |
| OSWNT2 | 7,264,622,400 | 719,390,209 (99.03%) | 6,689,383,501 (92.08%) | 3,687,134,980 (50.75%) |
| OSWNT3 | 7,129,401,000 | 705,316,141 (98.93%) | 6,580,108,021(92.30%) | 3,638,707,983 (51.04%) |
| OSWLT1 | 7,944,971,100 | 778,703,561 (98.01%) | 7,413,924,316 (93.32%) | 3,894,264,316 (49.02%) |
| OSWLT2 | 7,986,083,700 | 783,286,290 (98.08%) | 7,441,185,395 (93.18%) | 4,081,109,505 (51.10%) |
| OSWLT3 | 8,026,868,700 | 786,990,444 (98.04%) | 7,487,352,558(93.28%) | 4,026,903,542 (50.17%) |
| OFWLT1 | 7,953,954,300 | 786,547,287 (98.89%) | 7,343,776,104 (92.33%) | 3,783,132,177 (47.56%) |
| OFWLT2 | 7,734,025,200 | 762,521,171 (98.59%) | 7,132,487,673 (92.22%) | 3,891,404,658 (50.32%) |
| OFWLT3 | 8,354,551,200 | 825,821,772 (98.85%) | 7,735,196,526 (92.59%) | 4,032,074,641 (48.26%) |
| OFWNT1 | 8,932,483,500 | 882,958,414 (98.85%) | 8,264,811,490(92.53%) | 4,508,584,371 (50.47%) |
| OFWNT2 | 7,383,731,700 | 727,708,069 (98.56%) | 6,809,242,523(92.22%) | 3,755,759,027 (50.87%) |
| OFWNT3 | 7,595,829,600 | 7,367,262,628 (96.99%) | 6,995,732,246 (92.10%) | 3,874,770,036 (51.01%) |
| Primer Name | Sequence (5′→3′) |
|---|---|
| Cpeb4-F | TATGAGCCGCAGCGACAG |
| Cpeb4-R | AGAGGCCCATCTCCACCCT |
| PRPS2-F | GGACATGCCTAACATCGTGC |
| PRPS2-R | CTCCTGGTTGCTGAACTTCTTG |
| ADCY6-F | TTACAATCTGTGAGCGGAAACT |
| ADCY6-R | CCATGTGCCAAATACAACCAT |
| MAT2A-F | GCAGCAGTCTCCTGACATCG |
| MAT2A-R | CATCAGTGGCATATCCAAACA |
| fzd7-a-F | ACTCCCTTTACTCCCACTTGA |
| fzd7-a-R | CCTACTGGTGTTTCCCTGATT |
| mapk12-F | TCAAAGCCATGCCGTAAGC |
| mapk12-R | GTCACCGAAAGCCACTCCC |
| pvalb2-F | GCCGACTCTTTTCAAGCACAA |
| pvalb2-R | CCTCCTCAATGAAGCCACTC |
| PYGM-F | GCCAGGAGAAAGTCAATGCT |
| PYGM-R | TACTCGGTAATGGGTGCGATC |
| GPI-F | ACCGCTTCACTGCCTACTTC |
| GPI-F | GTGGTCTTGCCCGTCATC |
| got2aR | CAAGCCTGAGGAGTGGAAGG |
| got2aR | CAAGCCTGAGGAGTGGAAGG |
| GCSH-1-F | CCGCTCCCTGTCTTCCAACT |
| GCSH-1-F | GTTCCCACCTCTGGCAGTCC |
| tpi1b-F | CAGCCAGTCCACGCCTTTA |
| tpi1b-R | CCTTCCCACCCTTCTACCC |
| PGAM1-1-F | GCCAAGAAGTAAAGCAACAGAA |
| PGAM1-1-F | AACATCCCAGTCGTTCACCA |
| AMT-R | GTCACATCGCCTCGCACAT |
| AMT-R | GGTCATCAATAATCCCTCCTTTCTC |
| ENO1-F | CCCACGCAGGCAACAAG |
| ENO1-R | AAGCCGCCCTCATCTCC |
| BPGM-F | AGGCTATGGGTCAGGAGTGG |
| BPGM-R | CCAGGAAGTAAGGGTGGGAT |
| β-actin-F | CAACTGGGGATGACATGGGGAGAAG |
| β-actin-R | TTGGCTTTGGGGGGTTCAGG |
| Sample | Raw Data | Clean Data (%) | BF_Q30 (%) | BF_GC (%) |
|---|---|---|---|---|
| OFWLT1 | 7,953,954,300 | 7,865,472,876 (98.89%) | 7,003,646,337 (93.09%) | 3,811,150,486 (50.66%) |
| OFWLT2 | 7,734,025,200 | 7,625,211,711 (98.59%) | 5,747,382,251 (92.94%) | 3,232,272,528 (52.27%) |
| OFWLT3 | 8,354,551,200 | 8,258,217,726 (98.85%) | 7,072,551,701 (92.49%) | 3,993,848,545 (52.23%) |
| MFWLT1 | 7,523,556,600 | 7,454,735,551 (99.09%) | 7,343,776,104 (92.33%) | 3,783,132,177 (47.56%) |
| MFWLT2 | 6,184,004,400 | 6,118,434,737 (98.94%) | 7,132,487,673 (92.22%) | 3,891,404,658 (50.32%) |
| MFWLT3 | 7,647,044,700 | 7,576,414,880 (99.08%) | 7,735,196,526 (92.59%) | 4,032,074,641 (48.26%) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Peng, Y.; Yan, L.; Zhao, C.; Zhang, B.; Zhang, B.; Qiu, L. Effects of Temperature and Salinity on Ovarian Development and Differences in Energy Metabolism Between Reproduction and Growth During Ovarian Development in the Lateolabrax maculatus. Int. J. Mol. Sci. 2025, 26, 8295. https://doi.org/10.3390/ijms26178295
Peng Y, Yan L, Zhao C, Zhang B, Zhang B, Qiu L. Effects of Temperature and Salinity on Ovarian Development and Differences in Energy Metabolism Between Reproduction and Growth During Ovarian Development in the Lateolabrax maculatus. International Journal of Molecular Sciences. 2025; 26(17):8295. https://doi.org/10.3390/ijms26178295
Chicago/Turabian StylePeng, Yangtao, Lulu Yan, Chao Zhao, Bo Zhang, Bo Zhang, and Lihua Qiu. 2025. "Effects of Temperature and Salinity on Ovarian Development and Differences in Energy Metabolism Between Reproduction and Growth During Ovarian Development in the Lateolabrax maculatus" International Journal of Molecular Sciences 26, no. 17: 8295. https://doi.org/10.3390/ijms26178295
APA StylePeng, Y., Yan, L., Zhao, C., Zhang, B., Zhang, B., & Qiu, L. (2025). Effects of Temperature and Salinity on Ovarian Development and Differences in Energy Metabolism Between Reproduction and Growth During Ovarian Development in the Lateolabrax maculatus. International Journal of Molecular Sciences, 26(17), 8295. https://doi.org/10.3390/ijms26178295





